Abstract
We detail the expression of the Arabidopsis thaliana (L.) Heynh. atExt1 extensin gene. atExt1 is normally expressed in roots and inflorescences, and is induced by wounding, exogenously supplied salicylic acid, methyl jasmonate, auxins and brassinosteroids. Northern assays and histochemical analysis of transgenics expressing an atExt1::gus fusion show that this gene is also induced by the brassica pathogen Xanthomonas campestris pv. campestris and that this induction is restricted to tissues close to the site of infection. Expression at regions of abscission and senescence also implicates atExt1 in these important developmental processes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Extensins constitute one of the classes of plant structural cell wall proteins (Showalter 1993). Their name originates from Lamport's initial assumption that they play a crucial role in cell wall extension (Lamport 1963). Since evidence for this function has not yet materialised, "extensin" is now taken to refer to the extended rod-like appearance these proteins possess due to their polyproline Ⅱ helical conformation (Cooper et al. 1987). It is now becoming clear that these enigmatic proteins play a central role in the development of the plant form. The characteristic motif for extensins in dicotyledonous species is the highly repeated pentapeptide Ser(Hyp)4. The hydroxyproline residues incorporated within the Ser(Hyp)4 motif derive from the hydroxylation of proline residues through the action of prolyl hydroxylase (Guzman et al. 1990). Following hydroxylation the hydroxyproline residues are arabinosylated with one to four arabinose units while the serine residues are mono-galactosylated. Hydrogen bonding between the oligo-arabinosides and the hydroxyprolines is believed to confer stability to the molecule and maintain the characteristic rod-like appearance (Stafstrom and Staehelin 1986). The multiple Ser(Hyp)4 motifs are separated by short oligopeptides, the spacer sequences (Tierney and Varner 1987). These are rich in valine, threonine, lysine and tyrosine and are believed to comprise functional sites that enable the extensin molecules to interact uniquely with other cell wall components (Tierney and Varner 1987).
Extensins are secreted into the plant cell wall where they become insolubilised through the formation of intra and inter-molecular cross-links. Tyrosines within the Tyr-X-Tyr-Lys (where X may be Tyr or Lys) motif have been suggested to be the sites for the proposed intramolecular cross-links (Epstein and Lamport 1984; Kieliszewski and Lamport 1994). The nature of extensin inter-molecular cross-linkages, however, still excites controversial discussion. The Val-Tyr-Lys motif and the tyrosine tetramer di-isodityrosine have both been suggested to be involved in inter-molecular cross-linking (Schnabelrauch et al. 1996; Brady et al. 1996; respectively).
Several extensin genes have been isolated and characterised from a variety of plant species (for reviews, see José and Puigdomenech 1993; Showalter 1993; Sommer-Knudsen et al. 1998). Extensins occur in multigene families and it is thought that each member is expressed in a tissue-specific manner and probably fulfils a different function (Varner and Lin 1989). Northern analysis has demonstrated that many extensin genes are expressed in the roots in many plant species, including the DC5A1 extensin gene in carrot (Chen and Varner 1985), the extA gene in oilseed rape (Shirsat et al. 1991), the aExt gene in almond (Garcia-Mas et al. 1992), the Ext 26G gene in pea (Arsenijevic-Maksimovic et al. 1997), the SbHRGP3 gene in soybean (Ahn et al. 1996), the tobacco extensin genes npExt (De Loose et al. 1991; Tiré et al. 1994), Ext1.2 (Parmentier et al. 1995) and Ext 1.4 (Hirsinger et al. 1997, 1999), and the Arabidopsis atExt1 gene (Merkouropoulos et al. 1999). Extensin gene expression in root tissues has been proposed by many workers to be associated with cell wall reinforcement in order that the cells in this region withstand the considerable pressures the root experiences as it pushes its way into the soil or when the lateral root breaks its way through the tissues of the parental root (Keller and Lamb 1989). However, there is no direct evidence so far to support this assumption.
Using the β-glucuronidase (GUS) reporter gene system, extensin gene promoter–gus fusion expression was detected in the cortical cells that surround the vascular bundles of the newly formed leaf in the nodal regions of the stem in transgenic tobacco plants (Tiré et al. 1994; Wycoff et al. 1995). Extensin gene expression in such regions was considered to be the result of the tensile stress exerted by the weight of the developing organ. In further experiments, Shirsat and his colleagues showed that expression of a Brassica napus extA extensin–gus fusion could be induced in transgenic tobacco plants by applying weights (Shirsat et al. 1996a) showing, for the first time that the extA gene promoter could be transcriptionally activated by applied external load stresses. Elliott and Shirsat (1998) further identified regions of the extA gene promoter that regulated this expression.
Extensin expression in response to applied stress conditions, such as wounding (Ludevid et al. 1990; De Loose et al. 1991; Adams et al. 1992; Bown et al. 1993; Parmentier et al. 1995; Wycoff et al. 1995; Ahn et al. 1996; Hirsinger et al. 1997, 1999), pathogen infection (Corbin et al. 1987; Memelink et al. 1993; Niebel et al. 1993; Tiré et al. 1994; Hirsinger et al. 1997) and treatment with compounds known to be involved in various plant defense responses (Ecker and Davis 1987; Tagu et al. 1992; Memelink et al. 1993; Shirsat et al. 1996b; Ahn et al. 1996; Hirsinger et al. 1999; Merkouropoulos et al. 1999) supports a role for extensins in plant defense. Increased extensin deposition and extensin cross-linking has been proposed to assist in wound healing and also in the formation of a physical barrier against invading pathogens, thereby preventing the entry of pathogens into the vascular system and therefore limiting systemic pathogen spread (Showalter 1993).
We have isolated an Arabidopsis atExt1 extensin gene with a novel and unusual coding region (Merkouropoulos et al. 1999). The amino acid sequence of the atExt1 gene is extremely repetitive with Ser(Pro)4 motifs regularly alternating with Ser(Pro)3 motifs. Lamport in his recent review considers that the alternation of Ser(Pro)4 and Ser(Pro)3 motifs in this unusual gene probably reflects exquisite control of molecular flexibility (Lamport 2001). The spacer sequences, interspersed between the Ser(Pro)n regions, are relatively short in length (three to four amino acids) and are always a combination of valine, histidine, lysine and tyrosine residues. A signal peptide of 23 amino acids is present at the N-terminus of the protein. Sequence analysis has identified a putative 95-bp intron sequence in the 3′ transcribed but untranslated region of the gene (Merkouropoulos et al. 1999). Northern analysis showed atExt1 transcript accumulation in the root tissues, in the rosette (2–4 weeks after germination) and at lower levels in the flower (Merkouropoulos et al. 1999). Wounding dramatically changed this pattern of expression, with atExt1 mRNA levels in the roots declining rapidly (expression of atExt1 was undetectable within 24 h), while expression in the leaves appeared 12 h after wounding, reaching a peak at 18 h (Merkouropoulos et al. 1999).
Extremely high levels of atExt1 gene transcripts were recently detected in a mutant (mpk4) identified by hybridisation to a microarray of 7,864 cDNAs expressed throughout Arabidopsis development (Petersen et al. 2000). This mpk4 mutant is an Arabidopsis dwarf, and has a transposon-inactivated Arabidopsis mitogen-activated protein (MAP) kinase 4 gene. It exhibits a constitutive systemic acquired resistance (SAR), and as a result, it has been proposed that the MPK4 protein is a negative regulator for the SAR (Petersen et al. 2000). Detection of high levels of atExt1 gene transcripts in the mpk4 mutant supports the idea that the atExt1 gene plays an important role in plant defence. Recently, the identification of a knockout mutant in an Arabidopsis extensin gene, which is very similar to atExt1, has shown that it is required for maintaining normal cell shape and expansion, for the correct positioning of the cell plate during cytokinesis and for normal embryo development (Hall and Cannon 2002).
In the present paper we describe the construction of the Arabidopsis atExt1::gus fusion gene and present the developmental and stress-induced expression patterns of this construct. Expression patterns in response to mechanical wounding, pathogen infection, senescence and at abscission zones are described. We have also investigated the time course of atExt1 gene activation after treatment with compounds (salicylic acid, methyl jasmonate) that are known to be involved in the response of the plant to pest and pathogen attack.
Materials and methods
Plant material and plant treatments
Plants of Arabidopsis thaliana (L.) Heynh. were grown either in soil as described in Merkouropoulos et al. (1999) or hydroponically in 1× Hoagland's solution (Hoagland and Arnon 1950). Salicylic acid (SA), methyl jasmonate (MeJA), auxin (α-naphthaleneacetic acid) and epibrassinolide were applied by adding these compounds at appropriate concentrations to the hydroponic solutions. Arabidopsis stem sections and leaves were wounded using a blade and the wounded sliced tissues were incubated in phosphate buffer as described in Merkouropoulos et al. (1999). The bacterium Xanthomonas campestris pv. campestris was used to infect Arabidopsis plants either by spraying the pathogen on the leaves or by placing 2–6 μl of 106 cfu. ml−1 of X. campestris pv. campestris suspension on many points on the adaxial and abaxial side of leaves. The infected plants were covered with a clear polyethylene bag and left at room temperature for 5–10 days. Individual leaves were collected at different time points and were used for total RNA extraction and northern blotting, or examined histochemically for GUS activity.
DNA and RNA gel blot analysis
Plant genomic DNA and total RNA were extracted, purified, and run on neutral and denaturing agarose gels as described in Merkouroupoulos et al. (1999). Ten micrograms of both DNA and total RNA were electrophoresed on neutral and denaturing gels respectively; probes for both northern and Southern hybridisation were labelled by random priming and Southern and northern hybridisations performed as described in Merkouroupoulos et al. (1999). After hybridisation, Southern blots were washed to progressively higher stringency (3×SSC, 0.1% SDS, 65 °C, 60 min; 1×SSC, 0.1% SDS, 65 °C, 60 min; 0.5×SSC, 0.1% SDS, 65 °C, 60 min) and exposed to photographic film. Northern blots were washed twice with 3×SSC, 0.1% SDS at room temperature, twice with 1×SSC, 0.1% SDS at room temperature, twice with 0.1×SSC, 0.1% SDS at room temperature and finally twice with 0.1×SSC, 0.1% SDS at 65 °C. All washes were carried out for 20 min. The blots were then exposed to an X-ray-sensitive film for an appropriate length of time before being developed.
Construction of the atExt1::gus gene fusion
The 4.4-kb SalⅠ fragment, containing the atExt1 gene coding sequence together with about 3.3 kb of the upstream promoter sequence, was isolated from the Arab A genomic clone (Merkouropoulos et al. 1999) and cloned into the plasmid vector pBluescript SK+ (Stratagene Cloning Systems, La Jolla, Calif., USA) to produce clone pGM2. Clone pGM2 was then cleaved with SmaⅠ and BanⅠ and the 3.3-kb SmaⅠ/BanⅠ fragment carrying the atExt1 gene promoter was isolated. The BanⅠ cohesive end, which is located six nucleotides upstream of the atExt1 gene translation start point, was filled-in to produce a blunt-ended fragment. This fragment was cloned into the plasmid vector pBluescript SK+ to produce clones pGM3 and pGM4. The 3.3-kb promoter sequence in the 5′-3′ orientation from pGM3 was inserted upstream of the GUS gene coding sequence in the binary Agrobacterium plant transformation vector pBI101.3 (Jefferson 1987) to create a translational gene fusion pGM5.3 (Fig. 1). The 13 amino acid N-terminal extension does not display any of the common structural features found in signal peptides (von Heijne 1983, 1984) and this additional sequence therefore does not contain any targeting information.
Restriction map of the atExt1::gus fusion gene (pGM5.3). The fusion junction between the atExt1 gene promoter and the gus (Escherichia coli GUS, uidA) coding sequence is shown. The ATG initiator codon (bold and underlined) of the atExt1 gene coding sequence is in frame with the ATG initiator codon of the gus gene (bold and double underlined) coding sequence. The predicted GUS fusion polypeptide possesses a 13-amino-acid N-terminal extension, the sequence of which is shown underneath the corresponding nucleotide sequence. The first 10 bp (ATGGGGGCAC) of the nucleotide sequence originate from the 5′ end of the atExt1 gene coding sequence, followed by GGG (a relic of the subcloning procedure), while the next 26 bp is part of the pBI101.3 vector sequence, ending in the GUS translation initiator ATG)
Arabidopsis transformation
pGM5.3 was mobilised from E. coli into the Agrobacterium tumefaciens strain LBA4404 in a tri-parental mating, using the helper plasmid pRK2013. Successfully transformed A. tumefaciens strains were selected on the basis of kanamycin resistance due to the presence of the neomycin phosphotransferase Ⅱ gene (NPTⅡ), which is present on the pBI101.3 plasmid vector. These A. tumefaciens strains were used to transform Arabidopsis cv. Wassilewskija plants according to the vacuum-infiltration method (Bechtold et al. 1993), as modified by Bent et al. (1994).
GUS histochemical assays
Histochemical assays were performed as described by Jefferson (1987). Transverse hand sections taken from a variety of organs were incubated in 50 mM NaH2PO4 (pH 7.0), containing 1 mM 5-bromo-4-chloro-3-indolyl glucuronide (X-Gluc) dissolved in dimethyl formamide and 18 mM cycloheximide, at 37 °C overnight. The sections were then fixed in 3% glutaraldehyde (prepared in NaH2PO4, pH 7.0) at 4 °C overnight and finally cleared and stored in 100% ethanol. The sections were photographed on a Leica Wild M8 microscope, using Agfa Ultra 50 color film.
Results
atExt1 is expressed in the roots
We have previously shown that the atExt1 gene is expressed in 2- and 4-week-old rosette tissue, but not in 6-week-old rosettes (Merkouropolos et al. 1999). In order to examine developmental expression patterns in different organs of the mature plant, total RNA was isolated from roots, stems, and leaves of 5- to 6-week-old hydroponically grown plants, and the RNA samples were then northern-hybridised against the atExt1 gene coding sequence. The atExt1 probe was found to hybridise strongly to root RNA (Fig. 2).
Developmental expression of the atExt1 gene in 5- to 6-week-old wild-type Arabidopsis thaliana plants. Total RNA was extracted from roots (R), rosette leaves (L), caulines (C) and stems (S), and northern-hybridised against the atExt1 gene coding sequence. Equivalence of RNA loading between tracks is shown by ethidium bromide-stained ribosomal RNA bands
Transgenics carrying the atExt1::gus gene fusion contain normal, non-rearranged copies of the transgene
Genomic DNA was isolated from T1 plants, cleaved with HindⅢ/SacⅠ to excise the fusion construct (Fig. 3A), and hybridised against a 1.6-kb fragment of the atExt1 gene promoter. A single hybridising band of 5.2 kb (comprising the 3.3-kb atExt1 promoter and the 1.87-kb gus coding sequence) appeared in all lanes, indicating that the transgene had not been rearranged (Fig. 3B). The different hybridising band intensities from the different transgenics shown in Fig. 3B probably reflect differences in copy number seen in the T1 plants. For analysis of histochemical expression patterns, T3 plants derived from homozygous T2 plants originating from the line shown in Fig. 3B, lane 4, were used.
Agarose gel electrophoresis of genomic DNA from transgenic Arabidopsis plants. Genomic DNA from different individual transgenics was digested with HindⅢ and SacⅠ (A, lanes 2–9), Southern-blotted and hybridised to a 1.6-kb BanⅠ/XhoⅠ fragment from the atExt1 gene promoter. The resulting autoradiograph is shown in B. λDNA cleaved with PstⅠ was used as size marker (B, lane 1)
Expression of the atExt1::gus fusion gene through plant development
Root system
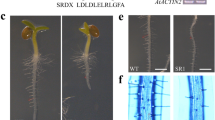
Expression of the atExt1::gus fusion gene was first seen 1–2 days after germination at the region where root hairs started to form (Fig. 4A). Expression in this region was found in the epidermis and the internal tissues of the primary root (Fig. 4B) and also in the root hairs (Fig. 4A–C). GUS staining was always heavier in those regions of the roots where root hairs were formed. In addition, GUS staining was always seen behind the root tips (in the region where new root hairs develop), both in the primary root and also in the secondary roots (Fig. 4A, C, D, E). The root tips always remained distinctively GUS-free (Fig. 4A, C, D, E). Expression of the atExt1::gus fusion gene was also observed in the hypocotyl–root transition zone where root hairs initiate. Staining in this region was always intense (Fig. 4D) and was limited to the epidermal cells of the root (data not shown). This early expression pattern of the atExt1::gus gene fusion changed later in development: expression was always seen in the root hairs while expression in the roots was variable, leaving some roots unstained (Fig. 4F).
Histochemical localisation of the atExt1::gus fusion gene expression in transgenic Arabidopsis plants during development. A Germinating seed. B Transverse section of a radicle from a 1-week-old seedling. C 4- to 5-day-old seedling. D 10-day-old seedling. E Junctions of lateral roots to the primary root. F 2-week-old seedling. G Leaf hydathodes. H Detail of leaf hydathodes. I Senescing rosette leaf. J Detail of the senescing part of a rosette leaf. K Base of a cauline leaf. L Stigma apex. M Inflorescence after the floral organs have been shed from the older pod. N Receptacle with floral organs still attached. O Floral organs after shedding. Bars = 1 mm
Rosette leaves and caulines
Upon the formation of the rosette, expression of the atExt1::gus fusion was only seen in the petioles of cotyledon leaves but not in the true leaves (Fig. 4D, F). Staining seen in the petiole of the cotyledon leaves was always limited to the junction of the cotyledon petiole with the rosette. At later developmental stages, expression of the atExt1::gus transgene was first seen at the base of the cauline leaves of both the primary stem and the bolts (Fig. 4K). After the rosette had fully expanded, expression of the atExt1::gus fusion gene was sometimes seen at the hydathodes of the rosette leaves (Fig. 4G, H).
Senescence and abscission
During leaf senescence, a blue-stained line of GUS activity was always seen separating the chlorotic-senescing part of the lamina from the healthy-green area (Fig. 4I, J). The chlorosis spread progressively to the interior of the leaf creating a wavy blue staining pattern (Fig. 4I).
In the floral tissues, expression of the atExt1::gus fusion was seen in the stigma (Fig. 4L). Expression was also detected at the abscission zone of the floral organs (sepals, petals and stamens; Fig. 4M–O). After the silique had fully developed, reached maturity and the floral parts had been shed, blue-stained scars remained on the base of the silique (Fig. 4M). The scar tissues were arranged in three whorls, each corresponding to a different floral organ.
atExt1 is expressed in response to exogenously supplied SA and MeJA
Previously, we showed that application of SA and MeJA through the roots of wild-type Arabidopsis led to accumulation of atExt1 mRNA in the leaves, while the level of atExt1 expression in the roots remains unaffected (Merkouropoulos et al. 1999). In the present work, we examined SA-induced expression of atExt1 over a time course. Figure 5A shows that atExt1 transcript accumulation in the leaves is first seen 6 h after the onset of incubation with 0.1 mM SA. atExt1 gene transcripts continue to increase up to 24 h, and decline at 48 h. In the roots, a biphasic induction of atExt1 mRNA levels in response to SA application is seen: an increase at 6 h, followed by a decrease in transcription up to 15 h, leading to a peak at 24 h followed by a small reduction 48 h after treatment (Fig. 5B). We have repeated this experiment, and always see the same slight reduction in atExt1 RNA accumulation at 48 h (data not shown).
Effect of SA and auxin supplied through the roots on atExt1 gene transcript accumulation in different organs. A, B Time-courses of atExt1 gene transcript accumulation in leaves (A) and roots (B) of wild-type Arabidopsis plants treated with 0.1 mM SA. C atExt1 transcript accumulation in leaves (L36) and roots (R36) 36 h after treatment with a 2 mg ml−1 auxin solution. L0 and R0 are control leaf and root samples not treated with auxin. In all cases, the time (hours) the RNA was extracted after the various treatments is indicated above the corresponding tracks. Equivalence of RNA loading between tracks is shown by ethidium bromide-stained ribosomal RNA bands
In order to define the exact tissues where the atExt1::gus fusion gene was expressed in response to SA and MeJA, individual leaves from transgenic Arabidopsis plants expressing the atExt1::gus fusion were incubated in 0.1 mM SA and 250 μM MeJA solutions. In mature leaves, application of SA resulted in expression of the atExt1::gus gene throughout the leaf lamina, without any obvious specificity (Fig. 6A, panel 2), while controls not incubated in SA showed no expression (Fig. 6A, panel 1). The effect of SA and MeJA on the expression of the atExt1::gus fusion gene early in development was also examined by germinating transgenic Arabidopsis seedlings on Murashige–Skoog (MS) plates containing SA and MeJA at various concentrations. Expression of the atExt1::gus gene fusion in seedlings grown on MS plates was limited to the roots and the tissues of the rosette (Fig. 6B), whereas in seedlings grown in plates containing SA or MeJA, non-specific GUS staining was seen throughout the seedlings (Fig. 6C and D, respectively).
Histochemical localisation of stress-induced atExt1::gus fusion gene expression in Arabidopsis seedlings in response to stresses and SA, MeJa and epibrassinolide application. A Control leaf (panel 1, not treated with SA), and a leaf incubated in SA solution and assayed for GUS activity (panel 2). B Transgenic seedling not treated with either SA or MeJA. C, D Transgenic seedlings grown on MS plates containing 0.1 mM SA (C) or 250 μM MeJA (D). E Transgenic seedlings incubated in 0.5 μM epibrassinolide (panel 1), and wild-type seedling not treated with epibrassinolide (panel 2). F–H, J, K GUS transgene activity in leaves that had been burnt at the right-hand section of the lamina (F), drilled (G, J), cut into small strips (H) or folded (K). I Expression of the atExt1::gus transgene at the edges of cut stem sections. L GUS activity in a leaf lamina that had been inoculated with X. campestris pv. campestris on both the left (late lesion with a circle of GUS activity surrounding a necrotic center) and right (early lesion with no visible signs of necrosis) halves. Bars = 1 mm
Expression in response to auxins and brassinosteroids
As the plant hormone auxin has been implicated as a strong positive regulator in root development and has been shown to increase both the length and number of root hairs (Bernhardt and Tierney 2000) we examined the effect of auxin application on atExt1 gene expression in root tissues by northern analysis of samples collected from wild-type Arabidopsis plants supplied with an auxin solution (2 mg ml−1) for 36 h. The northern blot (Fig. 5C) showed increased atExt1 gene transcript accumulation in the roots in response to auxin application, but there is no corresponding increase in atExt1 transcripts in leaves also treated with auxin.
Brassinosteroids are compounds known to stimulate cell elongation and division and are also involved in the differentiation of the vascular system and in stress responses. Extensin genes are expressed in dividing cells and also in rapidly growing seedlings (Cooper et al. 1994; Hirsinger et al. 1997, 1999). The effect of epibrassinolide (a brassinosteroid) on atExt1::gus fusion gene expression in young seedlings was examined by histochemical assays. Very intense, non-specific staining was observed in all tissues and organs of young seedlings exposed to brassinosteroids (Fig. 6E1) and the seedling expanded greatly, showing that expression of extensin genes in rapidly dividing cells may be a response to endogenous brassinosteroid synthesis in these regions. The wild-type seedling (Fig. 6E, panel 2) by comparison had very small cotyledon leaves, and did not show the same expansion, and in a control transgenic seedling not exposed to epibrassinolide (Fig. 6B), GUS activity was only seen in the roots and rosette tissues. This is a preliminary result, and we intend to investigate this response further by performing northern analyses and studying this response in detail.
Expression of the atExt1::gus fusion gene in response to mechanical wounding
The wound-induced expression of the atExt1::gus fusion construct was examined in stem and leaf tissues by wounding these tissues in different ways. Stems were cut into small pieces, while leaves were drilled, folded, burnt or cut into small strips. In all cases, GUS activity was seen in the immediate vicinity of the stress site (Fig. 6F–K). In the stem sections, expression of the atExt1::gus gene was limited to the wound site, leaving the region between the two sliced ends unstained (Fig. 6I). This pattern of expression occurred regardless of the length of the stem sections, since the same pattern was obtained from stem sections that were between 4 mm and 5 cm long. In leaves that had the adaxial side burnt, the damaged and destroyed parts of the lamina were separated from the non-burnt, healthy, green parts of the lamina by a blue-stained line of GUS expression (Fig. 6F). In leaves that had been folded, GUS activity was detected in straight lines along the folding crease (Fig. 6K). In leaves that had been mechanically drilled, strong expression was seen around the drilled holes on the lamina (Fig. 6G, J) and weak staining was seen in the rest of the lamina (Fig. 6G). In leaves that had been cut into strips, GUS activity was seen throughout the strips, being more intense around the cut edges (Fig. 6H). To confirm that these patterns of GUS expression were due to activation of the atExt1 gene, total RNA was isolated from wounded stems, wounded leaf strips and folded leaves of transgenic Arabidopsis plants carrying the atExt1::gus fusion gene. These RNA samples were northern-hybridised against the atExt1 coding sequence. atExt1 mRNA accumulation in response to wounding was seen in all tissues (Fig. 7), while nothing was seen in the unwounded control leaf or stem samples.
Northern blot showing transcript levels of the atExt1 gene in wounded Arabidopsis leaves carrying the atExt1::gus fusion construct. S W24 Wounded stem sections incubated in phosphate buffer for 24 h. S 0 Control unwounded stem sample. L W12–48 Wounded leaf strips incubated in phosphate buffer for 12, 24 and 48 h. L 0 Control unwounded leaf sample. Equivalence of RNA loading between tracks is shown by ethidium bromide-stained ribosomal RNA bands
Expression of the atExt1::gus fusion gene in response to pathogen infection
Extensin proteins have frequently been proposed to be involved in the response of the plant to pathogen attack—possibly by either strengthening the wall at the site of damage, or by restricting the further spread of the pathogen. In order to see if pathogen infection results in atExt1 gene expression, wild-type Arabidopsis leaves were infected in planta with X. campestris pv. campestris, which is a pathogen known to establish a compatible interaction with Arabidopsis. When necrotic lesions had appeared on the lamina (after 10 days), RNA was extracted from the leaf area that surrounded the necrotic lesion (Fig. 8, L3) and also from two adjacent sections (Fig. 8, L2, L1), which showed no signs of infection or necrosis. As a control, RNA was also isolated from healthy leaf tissue (Fig. 8, Lu). All RNA samples were then northern-hybridised against the atExt1 gene-coding region. The results show no atExt1 gene transcript accumulation in the control leaf (Fig. 8, Lu). In the infected leaf, all three samples showed a strong atExt1 transcript band, although the signal corresponding to the sample collected from the necrotic part of the lamina was stronger and was followed by visible degradation of the transcript (Fig. 8, L3). Histochemical assays were performed on transgenic Arabidopsis leaves infected with X. campestris pv. campestris. At an early stage of infection, before the development of necrotic lesions, expression of the atExt1::gus fusion gene was found in the cells around the infection site (Fig. 6L, right side of lamina). At late stages of infection (Fig. 6L, left side of lamina), a circular necrotic lesion appeared at the infected site and expression of the atExt1::gus fusion gene was restricted to the cells surrounding the necrotic lesion.
Northern blot showing expression of the atExt1 gene in Arabidopsis leaf sections adjacent to, and distal from, the site of inoculation with X. campestris pv. campestris. A representation of the leaf sections used for northern analysis is shown in the pictogram above the northern blot. Leaf section L3 was inoculated with X. campestris pv. campestris and showed a visible necrotic lesion (indicated by the circle), leaf section L2 was adjacent to L3, had not been inoculated, and showed no signs of infection. Leaf section L1 was adjacent to L2, had not been inoculated, and showed no signs of infection. The northern blot is of RNA extracted from leaf sections L1, L2, and L3 and hybridised against the atExt1 coding sequence. Root RNA (R) is included as a positive control, and RNA from uninfected leaves (L u ) as a negative control
Discussion
atExt1 is expressed in the root hairs and the elongating zone of the primary root
Extensin genes from various plant species have been found to be expressed in root tissues possibly because their synthesis in these regions acts to reinforce the cell wall in order to withstand pressures exerted on the root as it pushes its way through the soil or when the lateral root breaks its way through the tissues of the parental root. The Arabidopsis extensin gene atExt1 is no exception. Northern analysis shows that the atExt1 extensin gene is expressed to high levels in mature Arabidopsis roots but not in the leaves, caulines or stems. Histochemical analysis of expression of the fusion gene shows expression in all the tissues behind the root tip, where differentiation of the primary vascular tissue occurs. GUS activity was never detected in the root tips, suggesting that the atExt1 gene is required in cell differentiation processes rather than in the initial stages of cell ontogenesis. Histochemical GUS analysis shows that the transgene is primarily active in the trichoblasts and the epidermal atrichoblasts in the root hair bearing region. Activation of the atExt1 gene promoter in the root hairs directly implicates the involvement of the atExt1 gene in processes related to root hair function or structure and may also contribute to the reinforcement of the cell walls in order to withstand the mechanical forces the trichoblasts experience as they penetrate into the soil. It is also possible that the ATEXT1 protein may function to lock other cell wall components into position as part of the continuous wall restructuring which occurs during root hair elongation.
The atExt1::gus gene fusion expression patterns seen during the early stages of Arabidopsis development are similar (if not identical) to the expression patterns shown by the AtPRP3 Arabidopsis proline-rich cell wall protein (Fowler et al. 1999; Bernhardt and Tierney 2000). It is therefore reasonable to assume that both genes fulfil similar functions and are regulated through the same signalling pathways. Bernhardt and Tierney (2000) reported that both the number and the length of root hairs of Arabidopsis seedlings grown on plates containing the synthetic auxin α-naphthaleneacetic acid (α-NAA) increased, and that AtPRP3::gus activity in these seedlings was more than double that of the untreated control. In the present work, we show by northern analysis that when auxin is supplied to wild-type Arabidopsis plants, atExt1 transcript accumulation in the roots increases significantly. Recently, the expression of the tomato extensin-like gene LeExt1 was found to show similar patterns of expression in the roots of transgenic tomato, potato and Arabidopsis plants (Bucher et al. 2002). In particular, histochemical analysis of the LeExt1-promoter::gus fusions revealed expression of the fusion construct in the root hair-bearing zone of the growing radicle of transgenic tomato, potato and Arabidopsis plants. The root tip always remained free of GUS staining. Additionally, LeExt1 gene transcript accumulation was detected in the differentiation zone of tomato rhizodermal cells by in situ hybridisation (Bucher et al. 2002). Taken together these findings indicate that different members of the extensin family fulfil different functions through common developmental pathways.
atExt1 is expressed during senescence and abscission
GUS activity was detected at the beginning of senescence around chlorotic regions, which appeared at the margin of the senescing leaves. Chlorosis grew progressively larger and spread towards the centre of the lamina. A blue-stained wavy line of GUS activity separated the chlorotic regions from the rest of the leaf. Arabidopsis rosette leaf senescence is known to develop from the oldest leaf to the youngest and is characterised by the progressive yellowing of the leaf beginning from the periphery and spreading to the interior (Bleecker and Patterson 1997). GUS activity seen during the senescence process implies a structural and protective role for this gene and, therefore, for extensins in general. Extensins may also play a role in the plant's defence strategy by the construction of an extensin network that may form an impenetrable barrier preventing the establishment of infection foci at the site of dying cells.
Abscission is the process of organ shedding. It involves dissolution of the wall at certain points of attachment, i.e. the abscission zone (Sexton and Roberts 1982; Bleecker and Patterson 1997; Gonzáles-Carranza et al. 1998). Cells in the abscission zone are small in size and densely cytoplasmic. During abscission, a protective layer is formed immediately adjacent to the abscission layer. The cells in the protective layer become lignified or filled with gum to form an impermeable barrier that protects the exposed surface from desiccation and from the entry of parasites (Fahn 1982).
GUS activity was seen in the abscission zone of the floral organs, i.e. sepals, petals and stamens, at the site where the scar tissue forms. This finding indicates a role for the atExt1 gene during abscission and is the first time that the expression of an extensin gene has been found to be associated with the abscission process.
Expression in the rosette leaves, the caulines and the hydathodes indicates a role for the atExt1 gene in resisting mechanical stresses and pathogen infection
The rosette represents a structure where all nodal regions are very close due to the limited elongation of the internodal regions. Expression of the atExt1::gus fusion gene was detected in all tissues of the rosette, at the region where the leaves were attached to the stem. GUS staining in this region remained constant throughout plant development.
Expression of cell wall protein fusion genes in tissues experiencing mechanical stresses has been reported in several systems. The tobacco npExt::gus (Tiré et al. 1994) and Ext1.4::gus (Hirsinger et al. 1999) fusion genes are expressed in cortical cells around the leaf trace in stem nodes. Similar studies in heterologous systems, where the oilseed rape extA::gus (Shirsat et al. 1996a; Elliott and Shirsat 1998) and the bean HRGP4.1::gus fusion genes (Wycoff et al. 1995) had been transformed into tobacco, detected identical patterns of expression in the nodal regions. In these studies, expression of the extensin::gus fusion gene was proposed to be induced in response to the mechanical stresses developing in nodal regions due to the weight of the developing leaf at the petiole/stem junction. Activation of the atExt1::gus fusion gene in the region where the rosette leaves attach to the stem might therefore also be due to the mechanical loading at this region.
GUS activity was also seen at the base of caulines (where the caulines attach to the stem). It is unlikely that this expression is due to mechanical stresses because expression was never detected in the region where the branch attaches to the stem (in the same node). Expression of the atExt1::gus transgene at the cauline base may be indicative of extensin involvement in the growth of the leaf itself, since it is known that the growth of Arabidopsis leaves is a result of cell division occurring at the base of the leaves (van Lijsebettens and Clarke 1998). In this case, we might have expected to observe GUS activity in other regions of the plant where cell proliferation occurs, such as the stem and root primordia. However, no expression was ever seen in any proliferous zone.
The present work is the first to report expression of an extensin gene (atExt1) in hydathodes. Hydathodes are organised structures located on the margins, serrations and tips of the leaves. Specific expression of defence related genes in hydathodes, such as the Arabidopsis type Ⅲ acidic chitinase::gus fusion gene, has also been reported (Samac and Shah 1991). Since hydathodes represent areas without structural barriers against the entry of pathogens, expression of the atExt1::gus fusion gene in these regions may indicate that this gene product contributes (in synchrony with other mechanisms) to strengthen cells in order to withstand or avoid possible infection. It would be interesting to test this hypothesis in Arabidopsis atExt1 mutant knock out lines.
When wild-type leaves were infected with X. campestris pv. campestris and different sections were assayed for expression of the atExt1 gene using northerns, a very strong signal with considerable evidence of degradation was seen in the infected and necrotic section. It is likely that the degradation of the transcript is due to ribonucleases liberated from necrotic and infected cells. Sections adjacent to this necrotic region, which appeared to show no visible signs of infection, also expressed the atExt1 gene, albeit to lower levels. Infection of transgenic Arabidopsis plants carrying the fusion gene showed that during the early stages of infection, GUS activity was seen in a group of cells adjacent to the infection site. When infection was well advanced and necrotic lesions appeared on the lamina, GUS activity was detected around the necrotic lesion. It is interesting that while northern blots showed that leaf sections close to the infected region and which appeared to be uninfected expressed the atExt1 gene to significant levels, GUS activity driven from the atExt1 promoter was mainly restricted to tissues close to the necrotic lesion. This may be due to the different infection methods used in the experiments—in the northern experiments, the whole leaf surface was sprayed with the pathogen, while in the transgenics, the pathogen was applied to a highly localised spot on the leaf surface. As a consequence, it is possible that in the northern experiments, leaf regions that appeared not to show any infection symptoms were in fact infected and expressing the atExt1 gene prior to the development of visible lesions.
Stress-induced expression of extensin genes is usually related to the requirement for a fortified cell wall in order to protect the cell from pathogen invasion or mechanical damage (Wilson and Fry 1986). This is clearly not the case for the atExt1 gene, since expression of the atExt1 gene in the infected site did not ultimately result in the survival of the cells. atExt1 gene activation might therefore be part of the hypersensitive response (HR). The cells in the infected site rapidly collapse and die resulting in the appearance of a necrotic lesion. This may immobilize pathogens to the vicinity of the lesion and therefore block their spread. The idea that atExt1 gene expression is part of the HR is further supported by the SA-mediated induction of the atExt1 gene, since SA is believed to play a role in signalling host cell death (Dempsey et al. 1999).
SA and MeJA both activate the atExt1 gene
We have previously shown (Merkouropoulos et al. 1999) that application of SA at different concentrations to wild-type Arabidopsis roots results in atExt1 gene transcript accumulation in the stem and leaf. In the current work, application of SA at 0.1 mM through the roots led to activation of the atExt1 gene in the leaves, though the transcript levels were lower than the levels obtained from the 0.3 mM and 1.0 mM treatments (Merkouropoulos et al. 1999). These findings suggest that the level of atExt1 gene expression or the stability of the produced transcripts increases in proportion to the concentration of SA used. Histochemical analysis of transgenic Arabidopsis plants carrying the atExt1::gus fusion gene showed that incubation of leaves in SA solutions resulted in non-specific GUS activity in all the tissues of the lamina. Examination of the atExt1 gene promoter sequence found motifs highly similar to cis-acting elements known to be involved in SA-induced responses.
Activation of the atExt1 gene by both the SA-mediated and the MeJA-mediated pathways leads to the conclusion that the atExt1 extensin protein plays a role in plant stress responses. However, it is not yet clear whether the atExt1 protein functions as a specialized molecule involved in particular stress responses or whether it is synthesised under general stress conditions, playing a central role in plant defence and stress mechanisms. Only a few other genes that are inducible by both SA and MeJA/ethylene (providing evidence that cross-talk occurs between the SA- and MeJA/ethylene-mediated pathways) have been identified. Analysis of an expressed sequence tag (EST) microarray consisting of 2,375 sequences found only 10 genes that were activated upon screening with mRNA prepared from SA-, MeJA- and ethylene-treated plants (Schenk et al. 2000). This finding shows that SA and MeJA/ethylene signal interaction is limited to just a few genes (Schenk et al. 2000). Another study that shows the combinatorial interplay between the SA and MeJA/ethylene-mediated pathways used protoplasts from various Arabidopsis mutants deficient in different steps in the SA, JA and ethylene response pathways. The protoplasts were treated with the fungal toxin fumonisin B1 (FB1), which normally induces apoptosis-like programmed cell death in wild-type protoplasts (Asai et al. 2000). Programmed cell death did not occur in these mutants in response to FB1 application, indicating that SA/JA/ethylene are all involved in some aspect of the programmed cell death process. Asai et al. (2000) also identified atExt1 as one of the genes that were induced by FB1 application, implicating the atExt1 gene (and extensins in general) in programmed cell death.
Functional analysis of the various promoter sequence motifs located on the atExt1 gene (Merkouropoulos et al. 1999) will be essential to see what role (if any) they play in the control of the expression patterns described in this paper.
Abbreviations
- GUS:
-
β-glucuronidase gene, uidA
- HR:
-
hypersensitive response
- MeJA:
-
methyl jasmonate
- SA:
-
salicylic acid
- SAR:
-
systemic acquired resistance
References
Adams CA, Nelson WS, Nunberg AN, Thomas TL (1992) A wound-inducible member of the hydroxyproline-rich glycoprotein gene family in sunflower. Plant Physiol 99:775–776
Ahn JH, Choi Y, Kwon YM, Kim S-G, Choi YD, Lee JS (1996) A novel extensin gene encoding a hydroxyproline-rich glycoprotein requires sucrose for its wound-inducible expression in transgenic plants. Plant Cell 8:1477–1490
Arsenijevic-Maksimovic I, Broughton WJ, Krause A (1997) Rhizobia modulate root-hair-specific expression of extensin genes. Mol Plant Microbe Interact 10:95–101
Asai T, Stone JM, Heard JE, Kovtun Y, Yorgey P, Sheen J, Ausubel FM (2000) Fumonisin B1-induced cell death in Arabidopsis protoplasts requires jasmonate-, ethylene-, and salicylic-dependent signalling pathways. Plant Cell 12:1823–1835
Bechtold N, Ellis J, Pelletier G (1993) In planta Agrobacterium mediated gene transfer by infiltration of adult Arabidopsis thaliana plants. C R Acad Sci Paris D 316:1194–1199
Bent AF, Kunkel BN, Dahlbeck D, Brown KL, Schmidt R, Giraudat J, Leung J, Staskawicz BJ (1994) RPS2 of Arabidopsis thaliana: a leucine-rich repeat class of plant disease resistance genes. Science 265:1856–1860
Bernhardt C, Tierney ML (2000) Expression of AtPRP3, a proline-rich structural cell wall protein from Arabidopsis, is regulated by cell-type-specific developmental pathways involved in root hair formation. Plant Physiology 122:705–714
Bleecker AB, Patterson SE (1997) Last exit: senescence, abscission and meristem arrest in Arabidopsis. Plant Cell 9:1169–1179
Bown DP, Bolwell GP, Gatehouse JA (1993) Characterisation of potato (Solanum tuberosum L.) extensins: a novel extensin-like cDNA from dormant tubers. Gene 134:229–233
Brady JD, Sadler IH, Fry SC (1996) Di-isodityrosine, a novel tetrameric derivative of tyrosine in plant cell wall proteins: a new potential cross-link. Biochem J 315:323–327
Bucher M, Brunner S, Zimmermann P, Zardi GI, Amrhein N, Willmitzer L, Riesmeier JW (2002) The expression of an extensin-like protein correlates with cellular tip growth in tomato. Plant Physiol 128:911–923
Chen J, Varner JE (1985) An extracellular matrix protein in plants: characterization of a genomic clone for carrot extensin. EMBO J 4:2145–2151
Cooper JB, Chen JA, van Holst GJ, Varner JE (1987) Hydroxyproline-rich glycoproteins of plant cell walls. Trends Biochem Sci 12:24–27
Cooper JB, Heuser JE, Varner JE (1994) 3,4-Dehydroproline inhibits cell wall assembly and cell division in tobacco protoplasts. Plant Physiol 104:747–752
Corbin DR, Sauer N, Lamb CJ (1987) Differential regulation of a hydroxyproline-rich glycoprotein gene family in wounded and infected plants. Mol Cell Biol 7:4337–4344
De Loose M, Gheysen G, Tiré C, Gielen J, Villarroel R, Genetello C, Van Montagu M, Depicker A, Inzé D (1991) The extensin signal peptide allows secretion of a heterologous protein from protoplasts. Gene 99:95–100
Dempsey DA, Shah J, Klessig DF (1999) Salicylic acid and disease resistance in plants. Crit Rev Plant Sci 18:547–575
Ecker JR, Davis RW (1987) Plant defense genes are regulated by ethylene. Proc Natl Acad Sci USA 84:5202–5206
Elliott KA, Shirsat AH (1998) Promoter regions of the extA extensin gene from Brassica napus control activation in response to wounding and tensile stress. Plant Mol Biol 37:675–687
Epstein L, Lamport DTA (1984) An intramolecular linkage involving isodityrosine in extensin. Phytochemistry 23:1241–1246
Fahn A (1982) Plant anatomy, 3rd revised edn. Pergamon, Oxford
Fowler TJ, Bernhardt C, Tierney ML (1999) Characterization and expression of four proline-rich cell wall protein genes in Arabidopsis encoding two distinct subsets of multiple domain proteins. Plant Physiol 121:1081–1091
Garcia-Mas J, Messeguer R, Arús P, Puigdomènech P (1992) The extensin from Prunus amygdalus. Plant Physiol 100:1603–1604
Gonzáles-Carranza ZH, Lozoya-Gloria E, Roberts JA (1998) Recent developments in abscission: shedding light on the shedding process. Trends Plant Sci 3:10–14
Guzman NA, Fuller GC, Dixon JE (1990) Hydroxyproline-containing proteins and their hydroxylations by genetically distinct prolyl 4-hydroxylases. In: Adair WS, Mecham RP (eds) Organization and assembly of plant and animal extracellular matrix. Academic Press, San Diego, pp 301–356
Hall Q, Cannon MC (2002) The cell wall hydroxyproline-rich glycoprotein RSH is essential for normal embryo development in Arabidopsis. Plant Cell 14:1161–1172
Hirsinger C, Parmentier Y, Durr A, Fleck J, Jamet E (1997) Characterization of a tobacco extensin gene and regulation of its gene family in healthy plants and under various stress conditions. Plant Mol Biol 33:279–289
Hirsinger C, Salvá I, Marbach J, Durr A, Fleck J, Jamet E (1999) The tobacco extensin gene Ext 1.4 is expressed in cells submitted to mechanical constraints and in cells proliferating under hormone control. J Exp Bot 50:343–355
Hoagland DR, Arnon DI (1950) The water-culture method for growing plants without soil. Calif Agric Exp St Circ 347:1–32
Jefferson RA (1987) Assaying chimeric genes in plants: the GUS gene fusion system. Plant Mol Biol Rep 5:387–405
José M, Puigdomènech P (1993) Structure and expression of genes coding for structural proteins of the plant cell wall. New Phytol 125:259–282
Keller B, Lamb CJ (1989) Specific expression of a novel cell wall hydroxyproline-rich glycoprotein gene in lateral root initiation. Genes Devel 3:1639–1646
Kieliszewski MJ, Lamport DTA (1994) Extensin: repetitive motifs, functional sites, post-translational codes, and phylogeny. Plant J 5:157–172
Lamport DTA (1963) Oxygen fixation into hydroxyproline of plant cell wall protein. J Biol Chem 238:1438–1440
Lamport DTA (2001) Life behind cell walls: paradigm lost, paradigm regained. Cell Mol Life Sci 58:1363–1385
Ludevid MD, Ruiz-Avila L, Vallés MP, Stiefel V, Torrent M, Torné JM, Puigdomènech P (1990) Expression of genes for cell-wall proteins in dividing and wounded tissues of Zea mays L. Planta 180:524–529
Memelink J, Swords KMM, de Kam RJ, Schilperoort RA, Hoge JHC, Staehelin LA (1993) Structure and regulation of tobacco extensin. Plant J 4:1011–1022
Merkouropoulos G, Barnett DC, Shirsat AH (1999) The Arabidopsis extensin gene is developmentally regulated, is induced by wounding, methyl jasmonate, abscisic and salicylic acid, and codes for a protein with unusual motifs. Planta 208:212–219
Niebel A, Engler JdA, Tiré C, Engler G, Van Montagu M, Gheysen G (1993) Induction patterns of an extensin gene in tobacco upon nematode infection. Plant Cell 5:1697–1710
Parmentier Y, Durr A, Marbach J, Hirsinger C, Criqui M-C, Fleck J, Jamet E (1995) A wound-inducible extensin gene is expressed early in newly isolated protoplasts of Nicotiana sylvestris. Plant Mol Biol 29:279–292
Petersen M, Brodersen P, Naested H, Andreasson E, Lindhart U, Johansen B, Nielsen HB, Lacy M, Austin MJ, Parker JE, Sharma SB, Klessig DF, Martienssen R, Mattsson O, Jensen AB, Mundy J (2000) Arabidopsis MAP kinase 4 negatively regulates systemic acquired resistance. Cell 103:1111–1120
Samac DA, Shah DM (1991) Developmental and pathogen-induced activation of the Arabidopsis acidic chitinase promoter. Plant Cell 3:1063–1072
Schenk PM, Kazan K, Wilson I, Anderson JP, Richmond T, Somerville SC, Manners JM (2000) Coordinated plant defense responses in Arabidopsis revealed by microarray analysis. Proc Natl Acad Sci USA 97:11655–11660
Schnabelrauch LS, Kieliszewski M, Upham BL, Alizedeh H, Lamport DTA (1996) Isolation of pI 4.6 extensin peroxidase from tomato cell suspension cultures and identification of Val-Tyr-Lys as putative intermolecular cross-link site. Plant J 9:477–489
Sexton R, Roberts JA (1982) Cell biology of abscission. Annu Rev Plant Physiol 33:133–162
Shirsat AH, Wilford N, Evans IM, Gatehouse LN, Croy RRD (1991) Expression of a Brassica napus extensin gene in the vascular system of transgenic tobacco and rape plants. Plant Mol Biol 17:701–709
Shirsat AH, Bell A, Spence J, Harris JN (1996a) The Brassica napus extA extensin gene is expressed in regions of the plant subject to tensile stress. Planta 199:618–624
Shirsat AH, Wieczorek D, Kozbial P (1996b) A gene for Brassica napus extensin is differentially expressed on wounding. Plant Mol Biol 30:1291–1300
Showalter AM (1993) Structure and function of plant cell wall proteins. Plant Cell 5:9–23
Sommer-Knudsen J, Bacic A, Clarke AE (1998) Hydroxyproline-rich plant glycoproteins. Phytochemistry 47:483–497
Stafstrom JP, Staehelin LA (1986) The role of carbohydrate in maintaining extensin in an extended conformation. Plant Physiol 81:242–246
Tagu D, Walker N, Ruiz-Avila L, Burgess S, Martínez-Izquierdo JA, Leguay JJ, Netter P, Puigdomènech P (1992) Regulation of the maize HRGP gene expression by ethylene and wounding. mRNA accumulation and qualitative expression analysis of the promoter by microprojectile bombardment. Plant Mol Biol 20:529–538
Tierney ML, Varner JE (1987) The extensins. Plant Physiol 84:1–2
Tiré C, De Rycke R, De Loose M, Inzé D, Van Montagu M, Engler G (1994) Extensin gene expression is induced by mechanical stimuli leading to local cell wall strengthening in Nicotiana plumbaginifolia. Planta 195:175–181
van Lijsebettens M, Clarke J (1998) Leaf development in Arabidopsis. Plant Physiol Bioch 36:47–60
Varner JE, Lin L-S (1989) Plant cell wall architecture. Cell 56, 231–239
von Heijne G (1983) Patterns of amino acid near signal-sequence cleavage sites. Eur J Biochem 133:17–21
von Heijne G (1984) How signal sequences maintain cleavage specificity. J Mol Biol 173:243–251
Wilson LG, Fry JC (1986) Extensin, a major cell wall glycoprotein. Plant Cell Environ 9:239–260
Wycoff KL, Powell PA, Gonzales RA, Corbin DR, Lamb C, Dixon RA (1995) Stress activation of a bean hydroxyproline-rich glycoprotein promoter is superimposed on a pattern of tissue-specific developmental expression. Plant Physiol 109:41–52
Acknowledgements
G.M. was supported by the State Scholarship Foundation of Greece. We thank Daniel Price currently at the School of Biological Sciences, University of Durham, Durham UK for the data presented in Fig. 8. We also thank Prof. Mike Daniels (The Sainsbury Laboratory, John Innes Centre, Norwich, UK) for the gift of Xanthomonas campestris pv. campestris
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Merkouropoulos, G., Shirsat, A.H. The unusual Arabidopsis extensin gene atExt1 is expressed throughout plant development and is induced by a variety of biotic and abiotic stresses. Planta 217, 356–366 (2003). https://doi.org/10.1007/s00425-003-1002-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-003-1002-y