Abstract
Histone methylation is one of the main epigenetic mechanisms by which methyl groups are dynamically added to the lysine and arginine residues of histone tails in nucleosomes. This process is catalyzed by specific histone methyltransferase enzymes. Methylation of these residues promotes gene expression regulation through chromatin remodeling. Functional analysis and knockout studies have revealed that the histone lysine methyltransferases SETD1B, SETDB1, SETD2, and CFP1 play key roles in establishing the methylation marks required for proper oocyte maturation and follicle development. As oocyte quality and follicle numbers progressively decrease with advancing maternal age, investigating their expression patterns in the ovaries at different reproductive periods may elucidate the fertility loss occurring during ovarian aging. The aim of our study was to determine the spatiotemporal distributions and relative expression levels of the Setd1b, Setdb1, Setd2, and Cxxc1 (encoding the CFP1 protein) genes in the postnatal mouse ovaries from prepuberty to late aged periods. For this purpose, five groups based on their reproductive periods and histological structures were created: prepuberty (3 weeks old; n = 6), puberty (7 weeks old; n = 7), postpuberty (18 weeks old; n = 7), early aged (52 weeks old; n = 7), and late aged (60 weeks old; n = 7). We found that Setd1b, Setdb1, Setd2, and Cxxc1 mRNA levels showed significant changes among postnatal ovary groups (P < 0.05). Furthermore, SETD1B, SETDB1, SETD2, and CFP1 proteins exhibited different subcellular localizations in the ovarian cells, including oocytes, granulosa cells, stromal and germinal epithelial cells. In general, their levels in the follicles, oocytes, and granulosa cells as well as in the germinal epithelial and stromal cells significantly decreased in the aged groups when compared the other groups (P < 0.05). These decreases were concordant with the reduced numbers of the follicles at different stages and the luteal structures in the aged groups (P < 0.05). In conclusion, these findings suggest that altered expression of the histone methyltransferase genes in the ovarian cells may be associated with female fertility loss in advancing maternal age.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
There has been a worldwide trend in recent decades toward postponement of first births (delayed parenting) or having more children. The main drivers of these decisions are career planning, economic situation/problems, and lack of supportive family policies (Mills et al. 2011). Consistent with this trend, fertility rates in many European countries, such as Norway, Spain, and the UK have gradually decreased in parallel with advancing maternal age (Molina-Garcia et al. 2019). This decreased fertility substantially results from reduced oocyte quality and follicle numbers in the ovaries during reproductive aging (Ford 2013; Li et al. 2021).
Epigenetic alterations occurring during reproductive aging are one of the main factors influencing the quality of oocytes, early embryos, and granulosa cells (Chamani and Keefe 2019). Genome-wide DNA methylation profiles and associated enzyme levels were found to significantly decrease in the oocytes and early embryos of old mice (Yue et al. 2012). Reproductive aging also alters histone acetylation levels in mouse germinal vesicle (GV) and metaphase II (MII) oocytes (Manosalva and Gonzalez 2009) as well as in human metaphase I (MI) and MII oocytes (van den Berg et al. 2011). Shao et al. (2015) demonstrated that aging also affects global histone methylation profiles in mouse GV (Manosalva and Gonzalez 2010) and MII (Shao et al. 2015) oocytes. These studies indicate that epigenetic modifications undergo remarkable changes in association with reproductive aging, possibly due to alterated expression of related genes. As histone methylation plays crucial roles in regulating gene expression during oocyte maturation and follicular development (reviewed in He et al. 2021; Bilmez et al. 2022), analyzing histone methylation-related gene expression in ovaries during reproductive aging seems to be an important step towards deciphering the factors underlying fertility loss at later periods of life.
Histone lysine methyltransferases (KMTs) participate in the regulation of various cellular events, including gene expression, DNA replication, cell division, and chromatin reorganization through dynamically methylating target lysine residues (Pan et al. 2012; Cui et al. 2019). A highly conserved KMT protein SET domain containing 1B (SETD1B) performs histone 3 lysine 4 (H3K4) methylation (H3K4me) to modulate gene expression during oocyte maturation in mice (Sha et al. 2020b). Conditional loss of SETD1B causes growth retardation from embryonic day (E) 7.5, with embryos dying up to approximately E11.5 (Bledau et al. 2014). On the other hand, maternal lack of Setd1b gene leads to sterility in female mice, decreased levels of such key transcription factors as OBOXs, MEIS2, and SALL4 in the oocytes, increased follicular loss with age, and meiotic spindle and zona pellucida defects in the MII oocytes (Brici et al. 2017). Importantly, there is no embryonic development beyond 4-cell stage in the knockout group.
Another KMT, SET domain bifurcated 1 (SETDB1), modulates global H3K9 dimethylation (H3K9me2) levels in growing oocytes (Eymery et al. 2016; Kim et al. 2016). Kim et al. (2016) and Eymery et al. (2016) reported that conditional knockout (cKO) of Setdb1 gene in mouse oocytes leads to an arrest at GV or MI stage, and impaired meiotic resumption. Moreover, Setdb1 cKO oocytes show disrupted gene expression, derepression of retrotransposons, and increased DNA damage (Eymery et al. 2016; Kim et al. 2016). The SET domain containing 2 (SETD2) implicates in establishing oocyte epigenome via H3K36 methylation (Xu et al. 2019). Maternal deficiency of SETD2 causes reduced H3K36me3 levels, aberrant oocyte maturation, decreased number of MII oocytes, and female sterility (Xu et al. 2019). Abnormal DNA methylation in the oocytes, embryonic development arrest at the 1-cell stage, failures in DNA replication, and embryonic genome activation (EGA) also occur in the absence of SETD2.
The CpG binding protein, CXXC finger protein 1 (CFP1, encoded by Cxxc1 gene), is an important component of the SET1 complex with SETD1B, both of which work together to perform H3K4 tri-methylation (H3K4me3) in oocytes (Lee and Skalnik 2005). This complex recognizes unmethylated DNA regions and preformed H3K4me3 domains through the CFP1 protein and thus promotes transcriptional activity (Brown et al. 2017). As expected, deletion of the Cxxc1 gene in the developing mouse oocytes results in decreased H3K4me3 levels in the GV and MII oocytes (Yu et al. 2017). CFP1 deficiency also leads to altered timing of maternal mRNAs clearance, disrupted EGA, and an arrest at the 1- or 2-cell embryo stage. A subsequent study by Sha et al. (2018) demonstrated that while oocyte-specific null of Cxxc1 causes maturation arrest and chromosome misalignment in the MI oocytes, formation of the meiotic spindle is impaired in both MI and MII oocytes upon its overexpression (Sha et al. 2018).
Reproductive aging in females is a natural phenomenon that results in menopause. Fertility features gradually decline with ovarian aging, most likely due to decreased oocyte quality and follicle numbers (Ottolenghi et al. 2004; Rambags et al. 2014; Shirasuna and Iwata 2017). A limited number of studies have examined the potential effects of aging on histone methylation marks in mammalian oocytes at different stages. One of these studies (Manosalva and Gonzalez 2010) demonstrated that levels of the histone lysine methylation marks, including H3K36me2, H3K79me2, H4K20me2, and H3K9me3 were predominantly reduced in the GV and MII oocytes obtained from aged mice. Shao et al. (2015) found that H3K4me2 and H3K4me3 methylation profiles were altered in the GV and MII oocytes of aged mice. In in vitro preovulatory aging, a significant decrease in the H3K9me3 level in the MII oocytes was found, but this effect was not observed in the in vivo preovulatory aged oocytes (Demond et al. 2016). On the other hand, in vitro postovulatory aging leads to increased H3K9me3 level (Petri et al. 2020), H3K4me2 and H3K4me3 intensities (Xing et al. 2021) in the MII oocytes. Further, H3K4me2 and H3K27me2 levels were found to be enhanced in the MII oocytes cultured for a long time (Nie et al. 2018). Overall, different aging states, including biological, in vitro, or in vivo, are capable of changing histone lysine methylation profiles in oocytes, some of which are methylated by the SETD1B, SETDB1, SETD2, and CFP1 proteins. It is most likely that changed methylation marks in histone variants during aging arise from altered expression of these KMT genes. Thus, the aim of our study was to determine spatiotemporal distributions and relative expression levels of the Setd1b, Setdb1, Setd2, and Cxxc1 genes in the postnatal mouse ovaries from prepuberty to the late aged periods.
Materials and methods
Animal and sample collection
In the present study, we used the mouse as an experimental model due to its high rate of biological similarities with human (Emes et al. 2003) and because many epigenetic studies have been performed on mice (Blewitt and Whitelaw 2013), thus providing an opportunity to compare newly obtained knowledge. We created five groups of BALB/c female mice, as follows: prepuberty (3 weeks old; n = 6), puberty (7 weeks old; n = 7), postpuberty (18 weeks old; n = 7), early aged (52 weeks old; n = 7), and late aged (60 weeks old; n = 7). As described in our previous studies (Ozturk et al. 2015; Kosebent et al. 2020; Uysal and Ozturk 2020), these groups were formed based on the numbers and types of ovarian follicles, stromal content, and availability of corpus luteum and corpus albicans structures. All mice were provided by Akdeniz University Experimental Animals Application and Research Center, and they were kept under a 12/12-h light/dark cycle without water and food restrictions. All experimental protocols were carried out in accordance with relevant guidelines and regulations approved by the Akdeniz University Institutional Animal Care and Use Committee (Protocol no 1037/2020.02.011). Following cervical dislocation, which was performed immediately after ether inhalation, we dissected ovaries from each mouse under sterile conditions using a stereomicroscope (Zeiss, Oberkochen, Germany). One of the two ovaries was used in the quantitative real-time PCR (qRT-PCR) analysis, and the other was for hematoxylin and eosin (HE) and immunohistochemistry staining after paraffin embedding.
Gene expression analysis
The Setd1b, Setdb1, Setd2, and Cxxc1 gene expression analysis in the postnatal ovaries was performed as described in previous studies (Ozturk et al. 2015; Kosebent and Ozturk 2021b). We extracted total RNA from mouse ovaries using TRIzol® Reagent according to the manufacturer’s instructions (Life Technologies GmbH, Darmstadt, Germany). Concentration and absorbance values of the isolated RNA were measured using the Epoch microplate spectrophotometer (BioTek, Winooski, VT, USA). A 5-µg sample of total RNA was treated with DNase I (Ambion, Austin, TX, USA) to eliminate any genomic DNA contamination, following which complementary DNA (cDNA) was generated from 2 µg of the treated RNA using the RETROscript™ Kit (Ambion), based on the instructions provided by the manufacturer.
The relative mRNA levels of the Setd1b, Setdb1, Setd2, and Cxxc1 genes were determined in the postnatal mouse ovaries by qRT-PCR in a total reaction volume of 25 μl containing 12.5 μl of 2× SYBR Green Supermix (Bio-Rad Laboratories, Inc., Hercules, CA, USA), 10 μM of primers, and 1 μl of cDNA. PCR cycling was performed in a Rotor-Gene instrument (Corbett Research, Sydney, Australia). The primer sequences and product sizes are given in Table 1. qRT-PCR assays were carried out in triplicate for each group. Melting curve analysis was used to confirm specificity of the PCR products. Beta-actin (β-actin) gene expression was used as an internal control to normalize the relative mRNA levels of the target genes. The 2−ΔΔCt (where Ct stands for cycle threshold) formula was used to calculate the fold changes. Using this protocol, we were able to detect and determine the relative mRNA levels of the Setd1b, Setdb1, Setd2, and Cxxc1 genes in the postnatal ovaries from prepuberty to late aged groups.
Paraffin embedding
The postnatal mouse ovaries were prepared using a routine protocol for Bouin’s-fixed paraffin-embedded tissue processing (Ozturk et al. 2012). Ovaries were immersed in Bouin’s solution at +4 °C for 12 h, then dehydrated in increasing concentrations of ethanol, cleared in xylene, and subsequently embedded in paraffin. The paraffin blocks were cut into 5-μm-thick serial sections using a rotary microtome (Leica, Nussloch, Germany) and the sections were mounted on Superfrost Plus glass slides (Thermo Scientific, Rockford, IL, USA) for subsequent analysis.
Histological analysis
The sections were treated with HE stain not only to enable evaluation of the histological structures of the postnatal ovaries but also to determine the relative abundance of the follicles at different stages, from the primordial to antral stages, and atretic follicles as well as the luteal structures, corpus luteum and corpus albicans. Briefly, sections were incubated in an oven at 60 °C for 1 h before HE staining. The sections were then treated twice with xylene for 10 min each time and rehydrated in a series of decreasing ethanol concentrations for 5 min at each concentration, following which the sections were washed under tap water and then stained, first with hematoxylin, subsequent with eosin. Histological structures of postnatal ovaries were evaluated under a bright-field microscope (Carl Zeiss Inc., Thornwood, NY, USA).
The follicles from primordial to antral stages, and atretic follicles as well as the corpus luteum and corpus albicans structures were counted on randomly selected and HE-stained ovarian sections. We classified the follicles based on generally accepted criteria (Myers et al. 2004; Mehlmann et al. 2004). In brief, primordial follicles were composed of an oocyte surrounded by a layer of squamous follicular cells (also known as pre-granulosa cells). We described primary follicles as having an oocyte enclosed by a layer of cuboidal follicular cells, and secondary follicles as having two or more layers of cuboidal granulosa cells surrounding oocyte, but with no visible space among granulosa cells. In comparison, follicles with small spaces among multi-layered granulosa cells and having a centrally localized oocyte were defined as preantral follicles (also called as early antral follicles), and antral follicles were distinguished by having a large antrum filled with follicular fluid and eccentrically localized oocyte. Atretic follicles were defined as those with a structurally disrupted oocyte and clusters of granulosa and theca cells. Corpus luteum structures which appear following ovulation possess granulosa and theca lutein cells, both of show eosinophilic staining. The corpus albicans is the regressed form of the corpus luteum and is largely composed of connective tissue. Based on these distinguishing morphological criteria, we counted follicles, corpus luteum, and corpus albicans structures per square micron (µm2) in each ovarian section of the postnatal ovaries.
Immunohistochemistry
The spatiotemporal distributions of the SETD1B, SETDB1, SETD2, and CFP1 proteins were evaluated in the postnatal ovaries from the prepuberty to late aged groups using immunohistochemistry, as described in our previous studies (Uysal and Ozturk 2020; Kosebent and Ozturk 2021a). The sections obtained from paraffin-embedded blocks were deparaffinized in fresh xylene after incubation in an oven at 60 °C for 1 h and then rehydrated in a series of decreasing ethanol concentrations. The sections were then boiled for 5 min in Tris–EDTA solution (including 10 mM Tris base and 1 mM EDTA) for antigen retrieval in a microwave set at 665 W. Subsequently, endogenous peroxidase activity was blocked in 3% H2O2 solution (prepared in methanol) for 25 min at room temperature (RT). The sections were then washed with 1× Tris-buffered saline with Tween 20 (TBS-T) and blocked using an Ultra V Block solution (Thermo Fisher Scientific, Waltham, MA, USA) at RT for 7 min (for the SETD2 and CFP1 proteins staining) or 20 min (for the SETD1B and SETDB1 proteins staining) to prevent non-specific binding. We then incubated the sections with the SETD1B (diluted 1:1000; catalog no. 55005–1-AP; Proteintech, Rosemont, IL, USA), SETDB1 (diluted 1:500; catalog no. bs-11670R; Bioss, Woburn, MA, USA), SETD2 (diluted 1:500; catalog no. LS-C81092; LifeSpan Biosciences, Seattle, WA, USA), and CFP1 (diluted 1:1750; catalog no. ab198977; Abcam, Cambridge, UK) primary antibodies at +4 °C overnight. It should be noted that isotype IgG antibody (catalog no. I5006; Sigma-Aldrich, St. Louis, MO, USA) was used in negative control sections to determine specificity of the primary antibodies. Following incubation with the primary antibody, the sections were washed in TBS-T three times for 15 min each, and then incubated with biotinylated secondary antibody (diluted 1:750; catalog no. BA-1000; Vector Laboratories, Stuttgart, Germany) for 1 h at RT, following which the sections were treated with streptavidin–horseradish peroxidase (HRP) complex (catalog no. TS-125-HR; Thermo Fisher Scientific) for 30 min at RT. The immunoreaction was visualized using 3, 3’-diaminobenzidine (DAB) substrate (catalog no. D4168; Sigma-Aldrich) under a bright-field light microscope (Olympus, Tokyo, Japan). After washing sections in running tap water, we counterstained them with Mayer’s hematoxylin. Total, follicular, and cellular levels of the SETD1B, SETDB1, SETD2, and CFP1 proteins were evaluated using ImageJ software (National Institutes of Health, Bethesda, MD, USA) in the postnatal ovaries from prepuberty to late aged groups. The micrographs were captured under a bright-field microscope equipped with an Axiocam 105 color camera (Zeiss, Jena, Germany) at × 200 and × 400 original magnifications. The integrated mean values measured by the ImageJ program were divided into area to determine unit expression levels. At least six images from each group were analyzed.
Statistical analysis
The obtained data were analyzed using one-way analysis of variance (one-way ANOVA) on ranks, followed by a suitable post hoc test, i.e., Tukey’s, Newman-Keuls or Bonferroni. We conducted statistical calculations using GraphPad Prism 5 software (GraphPad Software Inc., San Diego, CA, USA). P < 0.05 was considered statistically significant.
Results
Histological analysis
We first evaluated histological structures of the postnatal mouse ovaries from the prepuberty to late aged groups through analyzing HE staining (Fig. 1a). The prepubertal ovaries had a small stromal area but, except for the antral ones, there were a number of follicles occupying a large area. In the puberty and postpuberty groups, the stromal area was enlarged, and all follicular stages were observed in addition to corpus luteum and corpus albicans structures. However, dramatic changes in the aged groups were observed, with follicular counts predominantly decreasing in both the early and aged groups. As in the puberty and postpuberty groups, we monitored the corpus luteum and corpus albicans structures in the aged groups (Fig. 1a).
a Representative micrographs of the postnatal mouse ovaries. Histological structures of hematoxylin and eosin (HE)-stained ovarian tissue sections obtained from prepuberty (PreP), puberty (P), postpuberty (PostP), early aged (EA), and late aged (LA) groups were assessed. All ovaries exhibited normal histological features. The micrographs were captured at ×40 original magnification. Scale bars: 100 µm. b Numbers of ovarian follicles and luteal structures in each group. Follicles were counted from primordial to antral stages and atretic follicles as well as corpus luteum and corpus albicans per µm2. Numbers of primordial, secondary, and preantral follicles significantly decreased from the prepuberty to aged groups (P < 0.05), but there were no predominant changes in the primary, antral, and atretic follicles, and corpus luteum counts among groups. Notably, the aged groups had lower numbers of corpus albicans structures when compared to the postpuberty group. Data on the primordial, primary, secondary, antral, and atretic follicles, and on the corpus luteum were evaluated using one-way analysis of variance (one-way ANOVA) and Tukey’s post hoc test. Bonferroni’s post hoc test was used for the analysis on the preantral follicles and corpus albicans. Data are presented as the mean ± standard error of mean (SEM). Asterisks indicate a significant difference at *P < 0.05, **P < 0.01, and ***P < 0.001. PF Primordial follicle, PrF primary follicle, SF secondary follicle, PAF preantral follicle, AF antral follicle, AtF atretic follicle, CL corpus luteum, CA corpus albicans
Counting normally appearing and atretic follicles as well as the luteal structures revealed that significant changes had occurred from prepuberty to aged groups (Fig. 1b). The numbers of ovarian follicles at the primordial, secondary, or preantral stages decreased from the prepuberty to early aged groups, and further decreased in the late aged group (P < 0.05); however, no significant changes were noted in the primary, antral, and atretic follicles among groups. Notably, although there were no significant changes in the corpus luteum counts between groups, the early and late aged groups had lower numbers of corpus albicans structures compared to the postpuberty group (P < 0.05; Fig. 1b).
Setd1b, Setdb1, Setd2, and Cxxc1 gene expression in the postnatal ovaries
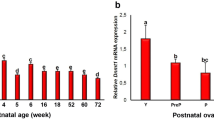
The levels of Setd1b, Setdb1, Setd2, and Cxxc1 mRNAs were analyzed in the postnatal ovaries from the prepuberty to late aged groups (Fig. 2). Setd1b gene expression gradually increased from the prepuberty to postpuberty groups (P < 0.001), and then progressively decreased towards the late aged group (P < 0.001; Fig. 2a). It is worth noting that the late aged group had lower Setd1b mRNA levels than the prepuberty (P < 0.05), puberty (P < 0.001), and postpuberty (P < 0.001) groups, and that the early aged group had lower Setd1b mRNA levels than the puberty (P < 0.001) and postpuberty (P < 0.001) groups. The level of Setdb1 mRNA was lowest in the prepuberty group, progressively increased toward the early aged group (P < 0.001), and then slightly decreased in the late aged group (Fig. 2b). Interestingly, the early aged group had a higher Setdb1 mRNA level when compared to the prepuberty (P < 0.001), puberty (P < 0.05), and postpuberty (P < 0.05) groups, and the late aged group had a higher Setdb1 mRNA level than the prepuberty group (P < 0.001; Fig. 2b).
Setd1b, Setdb1, Setd2, and Cxxc1 gene expression in the postnatal mouse ovaries. Expression levels of these genes were determined in the prepuberty (PreP, n = 6), puberty (P, n = 7), postpuberty (PostP, n = 7), early aged (EA, n = 7), and late aged (LA, n = 7) groups by quantitative real-time PCR (qRT-PCR). Notably, Beta-actin (β-Actin) was used as a housekeeping gene for normalization of target gene expression. a Relative Setd1b mRNA levels in the postnatal ovary groups, showing a significantly increase from the prepuberty to postpuberty groups, and then a decrease toward the late aged group (P < 0.001). Setd1b gene expression in the postpuberty group was set to 1. b Relative Setdb1 mRNA levels in the postnatal ovary groups, showing a gradual increase from the prepuberty to early aged groups (P < 0.001) and a minimal decrease in the late aged group. Setdb1 gene expression in the early aged group was set to 1. c Relative Setd2 mRNA levels in the postnatal ovary groups, showing fluctuations in Setd2 expression from the prepuberty to late aged groups (P < 0.05). Setd2 gene expression in the puberty group was set to 1. d Relative Cxxc1 mRNA levels in the postnatal ovary groups, showing significantly higher Cxxc1 gene expression in the early aged group when compared to the other groups (P < 0.01). Cxxc1 gene expression in the early aged group was set to 1. Data for Setd1b and Cxxc1 gene expression were evaluated using one-way ANOVA and Tukey’s post hoc test and Newman-Keuls post hoc test for Setdb1 and Setd2 gene expressions, and are presented as the mean ± standard deviation. Asterisks indicate a significant difference at *P < 0.05, **P < 0.01, and ***P < 0.001
Expression of the Setd2 gene fluctuated from the prepuberty to late aged groups (Fig. 2c). The prepuberty group showed the lowest expression level of Setd2, with a prominent increase in the puberty group (P < 0.01), followed by a significant decrease in the postpuberty group (P < 0.05), then a partial increase in the early aged group and finally a reduction again in the late aged group (P < 0.05; Fig. 2c). It should be noted that the early and late aged groups showed lower Setd2 expression than the puberty group (P < 0.01; Fig. 2c). When Cxxc1 gene expression was analyzed among these groups (Fig. 2d), we observed low levels of Setd2 mRNA in the prepuberty and puberty groups, with an enhancement in the postpuberty group (P < 0.05), attainment of the highest level in the early aged group (P < 0.01), and then a dramatic reduction in the late aged group (P < 0.001).
Spatiotemporal distributions and relative levels of the histone methyltransferases in the postnatal ovaries
The spatiotemporal distributions of the SETD1B, SETDB1, SETD2, and CFP1 proteins were comprehensively evaluated in the ovaries from the prepuberty to late aged groups. The relative levels of these proteins in the follicles, oocytes, granulosa cells, theca cells, and other ovarian cells were also analyzed.
Cellular distribution and relative level of SETD1B protein
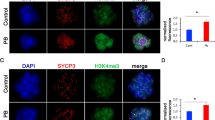
We detected SETD1B protein in the follicles, stromal area, luteal structures, and germinal epithelium (Fig. 3a). All follicles from the primordial to antral stages included SETD1B either in the granulosa cells or oocytes. Although SETD1B was intensively localized to nuclei of granulosa cells, oocytes showed weak staining intensities in the cytoplasm and nucleus. Most cells in the stromal area, corpus luteum, corpus albicans, germinal epithelium, and theca layers showed strong nuclear SETD1B immunoexpression when compared to the other ovarian somatic cells (Fig. 3a).
Cellular distribution and relative level of SETD1B protein in the postnatal mouse ovaries. a Representative micrographs of SETD1B immunoexpression in the prepuberty, puberty, postpuberty, early aged, and late aged groups. Strong nuclear localization of SETD1B was observed in the ovarian cells, including granulosa cells, germinal epithelium, stromal cells, theca, and luteal cells, whereas weak nuclear and cytoplasmic localizations were detected in the oocytes. Asterisks indicate antral space, black arrow indicates germinal epithelium. The micrographs and their inserts were captured at ×200 and ×400 original magnifications, respectively. Scale bars: 50 µm. b Relative SETD1B protein levels in the prepuberty (PreP, n = 6), puberty (P, n = 7), postpuberty (PostP, n = 7), early aged (EA, n = 7), and late aged (LA, n = 7) groups. In terms of total area, there was a gradual decrease in the SETD1B protein levels from prepuberty to the early/late aged groups (P < 0.05). c–h Relative SETD1B protein levels in the primordial (c), primary (d), secondary (e), preantral (f), antral (g), and atretic follicles (h), and in their respective oocytes [cytoplasm (C) and nucleus (N)], granulosa and theca cells. No significant changes were observed in these analyses (P > 0.05), although there was a decreasing tendency towards aged groups. i Relative SETD1B protein levels in the ovarian cells localized in the germinal epithelium, stroma, and corpus luteum. Despite minor decreases from the prepuberty to aged groups, no significant change between groups (P > 0.05) was detected. Data were analyzed using one-way ANOVA and Tukey’s post hoc test, and are presented as the mean ± standard deviation (SD). Asterisks (b) indicate a significant difference at *P < 0.05, **P < 0.01, and ***P < 0.001. A antrum, O oocyte, TL theca layer, S stroma; see Fig. 1 caption for other abbreviations
The relative levels of SETD1B protein in the total area, in follicles, and in each cell type were evaluated. In terms of total area analysis, SETD1B levels gradually decreased from the prepuberty to early/late aged groups (P < 0.05; Fig. 3b). Notably, the early and late aged groups had lower SETD1B levels compared to the puberty group (P < 0.05). In follicles from the primordial to antral stages as well as in atretic follicles, and in their oocytes, granulosa cells, and theca cells, although there was a decreasing tendency in some analyses toward the aged groups (Fig. 3c−i), no significant differences were observed among groups (Fig. 3c−h). Similarly, there were no remarkable changes in the germinal epithelial, stromal, theca, and granulosa lutein cells (Fig. 3i).
Cellular distribution and relative level of SETDB1 protein
The oocytes and granulosa cells of the follicles from the primordial to antral stages, as well as atretic follicles, showed SETDB1 immunoexpression, either in the cytoplasm or nucleus, in all groups from prepuberty to late aged (Fig. 4a). The prepuberty group showed stronger SETDB1 intensity in the granulosa cells and oocytes of the follicles from primordial to preantral stages when compared to the other groups. We also found that most ovarian cells, including stromal, germinal epithelial, and luteal cells, exhibited strong nuclear and weak cytoplasmic staining. It is important to note that stromal cells of the puberty, postpuberty, early and late aged groups showed intense nuclear SETDB1 immunoexpression, as opposed to the weak nuclear SETDB1 immunoexpression seen in the prepuberty group (Fig. 4a).
Cellular distribution and relative level of SETDB1 protein in the postnatal mouse ovaries. a Representative micrographs of SETDB1 immunoexpression in the prepuberty, puberty, postpuberty, early aged, and late aged groups. Strong nuclear localization was observed in the stromal cells, whereas weak nuclear and cytoplasmic immunostaining were detected in the oocytes and granulosa cells of the follicles at distinct stages. Asterisks indicate antral space, black arrow indicates germinal epithelium. The micrographs and their inserts were captured at ×200 and ×400 original magnifications, respectively. Scale bars: 50 µm. b Relative SETDB1 protein levels in the prepuberty (PreP, n = 6), puberty (P, n = 7), postpuberty (PostP, n = 7), early aged (EA, n = 7), and late aged (LA, n = 7) groups. In the total area analysis, there was a gradual decrease in SETDB1 protein levels from prepuberty to early aged groups (P < 0.01), followed by a sharp increase in the late aged group (P < 0.01). c–h Relative SETDB1 protein levels in the primordial (c), primary (d), secondary (e), preantral (f), antral (g), and atretic follicles (h), and in their respective oocytes [cytoplasm (C) and nucleus (N)], granulosa and theca cells. Significant decreases in the primordial, primary, secondary, preantral, and antral follicles were noted towards the aged groups (P < 0.05). i Relative SETDB1 protein levels in the ovarian cells located in the germinal epithelium, stroma, and corpus luteum. No significant alteration between groups (P > 0.05) was found. One-way ANOVA and Tukey’s post hoc test were used to analyze the data of total area, primordial, primary, and antral follicles, and ovarian cells, Newman-Keuls post hoc test was used for the secondary follicles, and the Bonferroni post hoc test was used for the preantral follicles. Data are presented as the mean ± SD. Asterisks (b–g) indicate a significant difference at *P < 0.05, **P < 0.01, and ***P < 0.001. A antrum, O oocyte, TL theca layer, S stroma; see captions to Figs. 1 and 2 for other abbreviations
Relative SETDB1 levels in the total area (Fig. 4b) gradually decreased from the prepuberty to early aged groups (P < 0.01), and sharply increased in the late aged group (P < 0.001). Lower SETDB1 levels were detected in the early and late aged groups than in the prepuberty, puberty, and postpuberty groups (P < 0.05). The analyses of the follicles from the primordial to antral stages, atretic follicles, and ovarian cells (Fig. 4c) revealed significant changes in some comparisons. In the primordial follicles, cytoplasmic SETDB1 levels in the oocytes progressively decreased from the prepuberty to puberty groups (P < 0.05). The level of cytoplasmic SETDB1 was highest in the oocytes of primary follicles when compared to the other groups (P < 0.001), but lowest in the early and late aged groups (Fig. 4d).
In the secondary follicles and their theca cells, the highest level of SETDB1 was detected in the prepuberty group (P < 0.001; Fig. 4e). It progressively decreased in the cytoplasm of oocytes and granulosa cells from prepuberty to early/late aged groups (P < 0.05). SEDTB1 levels in the preantral follicles and their granulosa cells in the prepuberty and puberty groups were significantly higher than those in the other groups, and the theca cells in the prepuberty group had the highest SETDB1 level (P < 0.01; Fig. 4f). The antral follicles, cytoplasm of the oocytes, and granulosa cells in the early and late aged groups possessed predominantly lower SETDB1 levels when compared to the puberty and postpuberty groups (P < 0.05; Fig. 4g). However, no statistical significance was found in the atretic follicles (Fig. 4h) and other ovarian cells (Fig. 4i).
Cellular distribution and relative level of SETD2 protein
In the postnatal ovaries from the prepuberty to late aged groups, SETD2 was intensively localized in the follicles from the primordial to antral stages (Fig. 5a). While granulosa cells of the follicles showed weak nuclear and strong cytoplasmic immunoexpression intensities, SETD2 immunoexpression was weak in both the cytoplasm and nucleus of the oocytes. The ovarian cells in the germinal epithelium, stroma, and theca layer, as well as in the granulosa and theca lutein cells showed cytoplasmic and nuclear SETD2 localization (Fig. 5a).
Cellular distribution and relative level of the SETD2 protein in the postnatal mouse ovaries. a Representative micrographs of SETD2 immunolocalization in the prepuberty, puberty, postpuberty, early aged, and late aged groups. SETD2 protein exhibited strong cytoplasmic and weak nuclear localization in the follicles from the primordial to antral stages. Asterisks indicate antral space, black arrow indicates germinal epithelium. The micrographs and their inserts were captured at ×200 and ×400 original magnifications, respectively. Scale bars: 50 µm. b Relative SETD2 protein levels in the prepuberty (PreP, n = 6), puberty (P, n = 7), postpuberty (PostP, n = 7), early aged (EA, n = 7), and late aged (LA, n = 7) groups. In the total area analysis, the highest and lowest SETD2 levels were detected in the puberty and late aged groups, respectively (P < 0.05). c–h Relative SETD2 protein levels in the primordial (c), primary (d), secondary (e), preantral (f), antral (g), and atretic follicles (h), and in their respective oocytes [cytoplasm (C) and nucleus (N)], granulosa and theca cells. Significant decreases towards the aged groups were noted in the follicles, granulosa and theca lutein cells (P < 0.05). i Relative SETD2 levels in the ovarian cells of germinal epithelium, stroma, and corpus luteum. Theca lutein cells displayed fluctuating changes from prepuberty to late aged groups (P < 0.01). One-way ANOVA and Tukey’s post hoc test were used to analyze the data for total area, primordial, primary, secondary, antral and atretic follicles, and other ovarian cells, and Newman-Keuls post hoc test was used for the preantral follicles. Data are presented as the mean ± SD. Asterisks (b–i) indicate a significant difference at *P < 0.05, **P < 0.01, and ***P < 0.001. A antrum, O oocyte, TL theca layer, S stroma; see captions to Figs. 1 and 2 for other abbreviations
We also analyzed the relative SETD2 protein levels in the total area, follicles, and each ovarian cell type among groups. SETD2 protein in the total area slightly increased from the prepuberty to puberty groups, and then gradually decreased toward the late aged group (P < 0.05; Fig. 5b). Notably, the late aged group showed the lowest SETD2 level when compared to the other groups (P < 0.01). While the granulosa cells of the primordial follicles in the puberty group had higher SETD2 levels than those in the postpuberty group (P < 0.05; Fig. 5c), no significant changes were found in the secondary (Fig. 5e) and antral (Fig. 5g) follicles among groups.
SETD2 protein levels in the granulosa cells of the primary follicles increased from the prepuberty to puberty groups and then progressively decreased toward the late aged group (P < 0.05; Fig. 5d). In the preantral follicles, SETD2 protein level was lowest in the prepuberty group, increased markedly in the puberty group, and subsequently decreased toward the late aged group (P < 0.01; Fig. 5f). Moreover, the granulosa cells of the puberty group and the theca cells of the postpuberty group showed higher SETD2 levels than those of the prepuberty group (P < 0.05). Interestingly, there was a gradual increase in SETD2 levels from the prepuberty to late aged groups in both the atretic follicles and their granulosa cells (P < 0.05; Fig. 5h). In the theca lutein cells of the corpus luteum, SETD2 protein level was highest in the puberty group, then dramatically decreased in the postpuberty group, increased in the early aged group (P < 0.001), and finally decreased again in the late aged group (P < 0.01; Fig. 5i).
Cellular distribution and relative level of CFP1 protein
CFP1 immunostaining was detected in the follicles and other ovarian cells of the postnatal ovaries from the prepuberty to late aged groups (Fig. 6a). Follicles from the primordial to antral stages showed strong nuclear and cytoplasmic immunoexpression in the granulosa cells, and their oocytes had strong nuclear but weak cytoplasmic CFP1 intensity. The ovarian cells showed intense nuclear and cytoplasmic immunoexpression in most cells located in the stroma, theca layer, corpus luteum, and germinal epithelium (Fig. 6a).
Cellular distribution and relative level of the CFP1 protein in the postnatal mouse ovaries. a Representative micrographs of CFP1 immunolocalization in the prepuberty, puberty, postpuberty, early aged, and late aged groups. CFP1 protein exhibited strong nuclear intensity in the granulosa cells and other ovarian cells, including stromal cells, luteal cells, and theca cells, whereas there was weak nuclear and cytoplasmic immunoreaction in the oocytes. Asterisks indicate antral space. The micrographs and their inserts were captured at ×200 and ×400 original magnifications, respectively. Scale bars: 50 µm. b. Relative CFP1 protein levels in the prepuberty (PreP, n = 6), puberty (P, n = 7), postpuberty (PostP, n = 7), early aged (EA, n = 7), and late aged (LA, n = 7) groups. In the total area analysis, there was a gradual decrease in CFP1 protein levels from the prepuberty to aged groups (P < 0.01). c–h Relative CFP1 protein levels in the primordial (c), primary (d), secondary (e), preantral (f), antral (g), and atretic follicles (h), and in their respective oocytes [cytoplasm (C) and nucleus (N)], granulosa and theca cells. In these analyses, there were significant changes in the primary follicles, their oocytes and granulosa cells as well as in the granulosa and theca cells of the secondary follicles (P < 0.05). i Relative CFP1 levels in the ovarian cells localized in the germinal epithelium, stroma, and corpus luteum. No significant change was noted among groups. One-way ANOVA and Tukey’s post hoc test were used to analyze the data of all follicle types and other ovarian cells. Data are presented as the mean ± SD. Asterisks (b, d, f, h, i) indicated significant differences at *P < 0.05, **P < 0.01, and ***P < 0.001. A antrum, O oocyte, TL theca layer, S stroma; see captions to Figs. 1 and 2 for other abbreviations
The relative CFP1 levels were also analyzed for total area, follicles, and other ovarian cells. Total area analysis showed a gradual decrease in CFP1 levels from the prepuberty to late aged groups (P < 0.01; Fig. 6b), whereas there was no predominant change between the early and late aged groups. Despite fluctuations in the CFP1 protein levels, no significant differences were found in the primordial (Fig. 6c), preantral (Fig. 6f), antral (Fig. 6g), atretic follicles (Fig. 6h), and ovarian cells (Fig. 6i). The primary follicles of the early and late aged groups had lower CFP1 levels than those of the puberty group (P < 0.05), and the granulosa cells and oocyte nuclei of the primary follicles showed highest levels in the prepuberty group (P < 0.05; Fig. 6d). In the secondary follicles, there were lower CFP1 levels in the granulosa cells of the early and late aged groups than in those of the prepuberty group (P < 0.05), and higher levels in the theca cells of the prepuberty group compared to the postpuberty group (P < 0.05; Fig. 6e).
Discussion
Reproductive aging mainly arises from disrupted biological processes in the reproduction-related germ and somatic cells (Yureneva et al. 2021). These disruptions may give rise to various abnormalities, including genetic defects, morphological changes, and mitochondrial dysfunction in the aged oocytes and granulosa cells (Nelson et al. 2013; Wilding 2015). Thus, in terms of extending reproductive lifespan, uncovering these biological processes is becoming increasingly important. In this study, we found significant changes in levels of the histone lysine methyltransferases in association with aging.
To our knowledge, this is the first study aimed at elucidating the relative levels of Setd1b, Setdb1, Setd2, and Cxxc1 gene expression and the spatiotemporal distribution of their encoded proteins in the postnatal mouse ovaries. We found significant changes in the expression of these genes in the postnatal ovary groups analyzed, from the prepuberty to late aged groups. Although there were fluctuations in the expression of the Setd1b, Setdb1, Setd2, and Cxxc1 genes among groups, the trend was toward a decrease with ageing, i.e. toward the late aged group. The fluctuations in the expression of these genes may result from changes in cellular contents, such as oocytes, granulosa cells, granulosa and theca lutein cells. As revealed in the counting results, the numbers of follicles from the primordial to antral stages and the luteal structures exhibited remarkable changes among groups, with a trend towards a decrease in the aged groups. As demonstrated in previous studies (de Bruin et al. 2004; Broekmans et al. 2009; Ozturk et al. 2015; Uysal et al. 2021), biological aging leads to a gradual decrease in the quality and numbers of ovarian follicles and corpus luteum in mice and humans. In the light of these findings, we suggest that changed gene expression among groups results from altered numbers of the ovarian contents, follicles and luteal structures, among groups. As is well-known, there can be a converse relationship between protein and mRNA levels (Maier et al. 2009; Liu et al. 2016); consequently, decreased Setd1b, Setdb1, Setd2, and Cxxc1 gene expression may arise from increased protein levels in the corresponding groups.
Localization of the SETD1B, SETDB1, SETD2, and CFP1 proteins in the follicles at different stages of development and stromal and germinal epithelial cells of the postnatal ovaries indicate that they contribute to establishing certain histone methylation processes to regulate main cellular events, such as gene expression and chromatin remodeling (Bilmez et al. 2022) in these cells. To understand their exact functions in each ovarian cell type, further work, especially in vitro studies, is warranted to elucidate their functional importance from the early to aged periods. No study has yet analyzed spatiotemporal expression patterns of these methyltransferases in the ovaries from prepuberty to aged periods, these findings would help researchers choose which histone methyltransferases to study in more detailed molecular biological investigations.
In our study, we found that SETD1B levels gradually decreased in the ovaries from the prepuberty to aged groups. As the growing follicles from the primary to antral stages exhibited strong SETD1B immunoexpression in all groups, decreased SETD1B levels in the aged groups possibly resulted from having lower numbers of these follicles, as shown in the follicle count analysis. A previous study reported that bsence of the Setd1b gene leads to various defects in the mouse oocytes, including formation of short spindle, impaired transition from a non-surrounded nucleolus (NSN)-type to surrounded nucleolus (SN)-type, cytoplasmic granularities due to abnormal organelle distribution, structural disorders in the zona pellucida, and disrupted nuclear and cytoplasmic maturation (Brici et al. 2017). These defects most likely result from changes in the levels of H3K4me3, which is catalyzed by SETD1B during oocyte maturation to stimulate timely expression of the genes encoding required proteins (Hanna et al. 2022). Importantly, the cellular defects occurring due to SETD1B deficiency largely resemble those in the aged oocytes (Miao et al. 2009; Cimadomo et al. 2018). Clearly detectable changes in H3K4me3 levels in the GV and MII oocytes undergoing biological aging (Shao et al. 2015), in vitro postovulatory aging (Xing et al. 2021), or in vitro long culturing (Nie et al. 2018) suggest that decreased SETD1B levels in the early and late aged groups may be associated with the reduced oocyte quality that accompanies aging.
Maternal loss of SETDB1 in the mouse oocytes causes reduced H3K9me2 and H3K9me3 levels, meiotic arrest at the GV or MI stage, production of fewer mature oocytes, and defective early embryo development (Kim et al. 2016). Upon oocyte-specific knockdown of Sedb1 gene expression, the chromatin defects, increased DNA damage, derepression of retrotransposons, gene dysregulation, and spindle and chromosome abnormalities appear (Eymery et al. 2016). Previous studies also revealed that biological aging (Manosalva and Gonzalez 2010) and postovulatory in vitro aging (Demond et al. 2016) could change H3K9me3 levels in the GV or MII oocytes. In light of these findings, it is possible that reduced SETDB1 levels in the oocytes, granulosa and theca cells of the follicles in the aged groups may contribute to decreased oocyte quality during aging through changes in the H3K9me2 and H3K9me3 levels.
In our study, SETD2 displayed an intense immunolocalization in the ovarian follicles, corpus luteum, and other ovarian cells, and its total level was significantly reduced in the late aged group. The reduced numbers of the follicles from primordial to preantral stages may underlie the decrease in the late aged group. In addition, predominant decreases in SETD2 levels in the granulosa cells of the primary follicles and preantral follicles, and theca lutein cells may be associated with the reduced SETD2 level in the late aged group. These decreases may influence the basic roles of SETD2 protein in these cells during aging.
SETD2 catalyzes H3K36me3 to promote transcriptional activity, which is in turn required for encoding meiotic maturation-related proteins in mouse oocytes (Li et al. 2018). In one study, knockdown of Setd2 gene expression in the mouse oocytes using small interfering RNA (siRNA) technology led to impaired oocyte maturation, abnormal spindle formation and kinetochore-microtubule attachments, chromosome misalignment, and reduced polar body extrusion rate (Li et al. 2018). Furthermore, Xu et al. (2019) reported that maternal SETD2 deficiency causes decreased H3K36me3 profiles in the mouse oocytes, reduced MII oocyte numbers, and abnormal DNA methylome establishment (Xu et al. 2019). Occurrence of many cellular abnormalities inoocytes upon knockdown or absence of SETD2 suggests that decreased SETD2 levels in the aged ovaries may be associated with a gradual loss of oocyte quality during aging. Consistent with this hypothesis, similar abnormalities have been observed in the aged oocytes, such as irregular organelle distribution, disorganized spindles, chromosome misalignment (Miao et al. 2009), and changed transcriptional activity (Llonch et al. 2021; Ntostis et al. 2021).
With regard to SETD2 immunoexpression, Diao et al. (2018) reported that it strongly localized in the nucleus of NSN-type porcine oocytes and also in the cytoplasm of the SN, MI, and MII oocytes, but not in their nuclei (Diao et al. 2018). We detected SETD2 immunoexpression either in the cytoplasm or nucleus of the oocytes in the follicles from primordial to antral stages. As growing follicles contain NSN-type oocytes until the surge in luteinizing hormone (LH) (De La Fuente and Eppig 2001; Xiao et al. 2015), there may be a species-dependent difference related to intracellular SETD2 distribution.
The CFP1 protein is an important component of the SET1 complex and contributes to regulating H3K4me3 establishment in growing oocytes (Yu et al. 2017; Sha et al. 2018). Its maternal deficiency results in a predominant decrease in the H3K4me3 levels, changed maternal mRNA clearance, and impaired gene expression related to DNA (de)methylation process in the mouse oocyte (Yu et al. 2017). Also, transcriptional activity is largely reduced in the CFP1-deficient oocytes. Recent studies have reported that oocyte-specific CFP1 deficiency leads to disrupted spindle formation, chromosome misalignment, meiotic arrest at MI stage, impaired H3K4me3 accumulation, and changed chromatin configurations and DNA methylation levels in the mouse oocyte (Sha et al. 2018, 2021). In the present study, we observed a significant decrease of CFP1 levels in the aged groups, which most likely arises from reduced CFP1 levels in the oocytes, granulosa or theca cells of the primary or secondary follicles. As loss of CFP1 causes various cellular defects in oocytes, decreasing CFP1 levels in the ovarian cells, including oocytes and granulosa cells, during aging may adversely influence oocyte quality.
In mice, CFP1 protein has been found to be localized in the cytoplasm and strongly in the nucleus of oocytes (Sha et al. 2018, 2020a) and in the nucleus of granulosa cells of the growing follicles (Sha et al. 2020a). Consistent with this result, we detected CFP1 immunoexpression in the cytoplasm of the oocytes and granulosa cells at a weak intensity and in the nucleus of these cells at a strong intensity. This expression distribution is compatible with the function of CFP1 in methylating H3K4 to modulate transcriptional activity. On the other hand, previous studies revealed that H3K4me3 levels dramatically decrease in the GV and MII oocytes obtained from aged mice (Shao et al. 2015) or MII oocytes that underwent in vitro postovulatory aging (Xing et al. 2021). Thus, the decrease in CFP1 levels during aging may be a potent cause of reduced H3K4me3 levels in the aged oocytes.
Although spatiotemporal distribution of the histone lysine methyltransferases (SETD1B, SETDB1, SETD2, and CFP1) were comprehensively evaluated in the postnatal ovaries from prepuberty to late aged periods, there are some limitations of this study. An important limitation is that we were unable to analyze the enzymatic activity of these methyltransferases and their target methylation marks in the ovarian cells, such as oocytes and granulosa cells.
Conclusion
Expression of the Setd1b, Setdb1, Setd2, and Cxxc1 genes exhibits significant changes in the mouse ovaries from prepuberty to aged periods, and their protein levels show a remarkable decrease in the aged ovaries. These changes may be inked with the fertility loss that is associated with biological aging in women. Further studies on determining the factors which drive these changes in histone lysine methyltransferase levels would contribute to an improved understanding of the potential reasons causing decreased fertility at a later age. New findings may help in creation of distinct strategies for preventing or reducing fertility loss during aging.
References
Bilmez Y, Talibova G, Ozturk S (2022) Dynamic changes of histone methylation in mammalian oocytes and early embryos. Histochem Cell Biol 157(1):7–25. https://doi.org/10.1007/s00418-021-02036-2
Binda O (2013) On your histone mark, SET, methylate! Epigenetics 8(5):457–463. https://doi.org/10.4161/epi.24451
Bledau AS, Schmidt K, Neumann K, Hill U, Ciotta G, Gupta A, Torres DC, Fu J, Kranz A, Stewart AF, Anastassiadis K (2014) The H3K4 methyltransferase Setd1a is first required at the epiblast stage, whereas Setd1b becomes essential after gastrulation. Development 141(5):1022–1035. https://doi.org/10.1242/dev.098152
Blewitt M, Whitelaw E (2013) The use of mouse models to study epigenetics. Cold Spring Harb Perspect Biol 5(11):a017939. https://doi.org/10.1101/cshperspect.a017939
Brici D, Zhang Q, Reinhardt S, Dahl A, Hartmann H, Schmidt K, Goveas N, Huang J, Gahurova L, Kelsey G, Anastassiadis K, Stewart AF, Kranz A (2017) Setd1b, encoding a histone 3 lysine 4 methyltransferase, is a maternal effect gene required for the oogenic gene expression program. Development 144(14):2606–2617. https://doi.org/10.1242/dev.143347
Broekmans FJ, Soules MR, Fauser BC (2009) Ovarian aging: mechanisms and clinical consequences. Endocr Rev 30(5):465–493. https://doi.org/10.1210/er.2009-0006
Brown DA, Di Cerbo V, Feldmann A, Ahn J, Ito S, Blackledge NP, Nakayama M, McClellan M, Dimitrova E, Turberfield AH, Long HK, King HW, Kriaucionis S, Schermelleh L, Kutateladze TG, Koseki H, Klose RJ (2017) The SET1 complex selects actively transcribed target genes via multivalent interaction with CpG Island Chromatin. Cell Rep 20(10):2313–2327. https://doi.org/10.1016/j.celrep.2017.08.030
Chamani IJ, Keefe DL (2019) Epigenetics and female reproductive aging. Front Endocrinol (Lausanne) 10:473. https://doi.org/10.3389/fendo.2019.00473
Cimadomo D, Fabozzi G, Vaiarelli A, Ubaldi N, Ubaldi FM, Rienzi L (2018) Impact of maternal age on oocyte and embryo competence. Front Endocrinol (Lausanne) 9:327. https://doi.org/10.3389/fendo.2018.00327
Cui JY, Fu ZD, Dempsey J (2019) Toxicoepigenetics. In: McCullough Shaun D, Dolinoy Dana C (eds). The role of histone methylation and methyltransferases in gene regulation. United States, Academic press, Cambridge, Massachusetts
Cuomo D, Porreca I, Ceccarelli M, Threadgill DW, Barrington WT, Petriella A, D’Angelo F, Cobellis G, De Stefano F, D’Agostino MN, De Felice M, Mallardo M, Ambrosino C (2018) Transcriptional landscape of mouse-aged ovaries reveals a unique set of non-coding RNAs associated with physiological and environmental ovarian dysfunctions. Cell Death Discov 4:112. https://doi.org/10.1038/s41420-018-0121-y
de Bruin JP, Dorland M, Spek ER, Posthuma G, van Haaften M, Looman CW, te Velde ER (2004) Age-related changes in the ultrastructure of the resting follicle pool in human ovaries. Biol Reprod 70(2):419–424. https://doi.org/10.1095/biolreprod.103.015784
De La Fuente R, Eppig JJ (2001) Transcriptional activity of the mouse oocyte genome: companion granulosa cells modulate transcription and chromatin remodeling. Dev Biol 229(1):224–236. https://doi.org/10.1006/dbio.2000.9947
Demond H, Trapphoff T, Dankert D, Heiligentag M, Grummer R, Horsthemke B, Eichenlaub-Ritter U (2016) Preovulatory aging in vivo and in vitro affects maturation rates, abundance of selected proteins, histone methylation pattern and spindle integrity in murine oocytes. PLoS ONE 11(9):e0162722. https://doi.org/10.1371/journal.pone.0162722
Diao YF, Lin T, Li X, Oqani RK, Lee JE, Kim SY, Jin DI (2018) Dynamic changes of SETD2, a histone H3K36 methyltransferase, in porcine oocytes, IVF and SCNT embryos. PLoS ONE 13(2):e0191816. https://doi.org/10.1371/journal.pone.0191816
Emes RD, Goodstadt L, Winter EE, Ponting CP (2003) Comparison of the genomes of human and mouse lays the foundation of genome zoology. Hum Mol Genet 12(7):701–709. https://doi.org/10.1093/hmg/ddg078
Eymery A, Liu Z, Ozonov EA, Stadler MB, Peters AH (2016) The methyltransferase Setdb1 is essential for meiosis and mitosis in mouse oocytes and early embryos. Development 143(15):2767–2779. https://doi.org/10.1242/dev.132746
Ford JH (2013) Reduced quality and accelerated follicle loss with female reproductive aging—does decline in theca dehydroepiandrosterone (DHEA) underlie the problem? J Biomed Sci 20:93. https://doi.org/10.1186/1423-0127-20-93
Greer EL, Shi Y (2012) Histone methylation: a dynamic mark in health, disease and inheritance. Nat Rev Genet 13(5):343–357. https://doi.org/10.1038/nrg3173
Hanna CW, Huang J, Belton C, Reinhardt S, Dahl A, Andrews S, Stewart AF, Kranz A, Kelsey G (2022) Loss of histone methyltransferase SETD1B in oogenesis results in the redistribution of genomic histone 3 lysine 4 trimethylation. Nucleic Acids Res. https://doi.org/10.1093/nar/gkac051
He M, Zhang T, Yang Y, Wang C (2021) Mechanisms of oocyte maturation and related epigenetic regulation. Front Cell Dev Biol 9:654028. https://doi.org/10.3389/fcell.2021.654028
Kim J, Zhao H, Dan J, Kim S, Hardikar S, Hollowell D, Lin K, Lu Y, Takata Y, Shen J, Chen T (2016) Maternal setdb1 is required for meiotic progression and preimplantation development in mouse. PLoS Genet 12(4):e1005970. https://doi.org/10.1371/journal.pgen.1005970
Kosebent EG, Ozturk S (2021a) The spatiotemporal expression of TERT and telomere repeat binding proteins in the postnatal mouse testes. Andrologia 53(3):e13976. https://doi.org/10.1111/and.13976
Kosebent EG, Ozturk S (2021b) Telomere associated gene expression as well as TERT protein level and telomerase activity are altered in the ovarian follicles of aged mice. Sci Rep 11(1):15569. https://doi.org/10.1038/s41598-021-95239-5
Kosebent EG, Uysal F, Ozturk S (2020) The altered expression of telomerase components and telomere-linked proteins may associate with ovarian aging in mouse. Exp Gerontol 138:110975. https://doi.org/10.1016/j.exger.2020.110975
Lee JH, Skalnik DG (2005) CpG-binding protein (CXXC finger protein 1) is a component of the mammalian Set1 histone H3-Lys4 methyltransferase complex, the analogue of the yeast Set1/COMPASS complex. J Biol Chem 280(50):41725–41731. https://doi.org/10.1074/jbc.M508312200
Li C, Diao F, Qiu D, Jiang M, Li X, Han L, Li L, Hou X, Ge J, Ou X, Liu J, Wang Q (2018) Histone methyltransferase SETD2 is required for meiotic maturation in mouse oocyte. J Cell Physiol 234(1):661–668. https://doi.org/10.1002/jcp.26836
Li CJ, Lin LT, Tsai HW, Chern CU, Wen ZH, Wang PH, Tsui KH (2021) The molecular regulation in the pathophysiology in ovarian aging. Aging Dis 12(3):934–949. https://doi.org/10.14336/AD.2020.1113
Liu Y, Beyer A, Aebersold R (2016) On the dependency of cellular protein levels on mRNA abundance. Cell 165(3):535–550. https://doi.org/10.1016/j.cell.2016.03.014
Llonch S, Barragan M, Nieto P, Mallol A, Elosua-Bayes M, Lorden P, Ruiz S, Zambelli F, Heyn H, Vassena R, Payer B (2021) Single human oocyte transcriptome analysis reveals distinct maturation stage-dependent pathways impacted by age. Aging Cell 20(5):e13360. https://doi.org/10.1111/acel.13360
Maier T, Guell M, Serrano L (2009) Correlation of mRNA and protein in complex biological samples. FEBS Lett 583(24):3966–3973. https://doi.org/10.1016/j.febslet.2009.10.036
Manosalva I, Gonzalez A (2009) Aging alters histone H4 acetylation and CDC2A in mouse germinal vesicle stage oocytes. Biol Reprod 81(6):1164–1171. https://doi.org/10.1095/biolreprod.109.078386
Manosalva I, Gonzalez A (2010) Aging changes the chromatin configuration and histone methylation of mouse oocytes at germinal vesicle stage. Theriogenology 74(9):1539–1547. https://doi.org/10.1016/j.theriogenology.2010.06.024
Mehlmann LM, Saeki Y, Tanaka S, Brennan TJ, Evsikov AV, Pendola FL, Knowles BB, Eppig JJ, Jaffe LA (2004) The Gs-linked receptor GPR3 maintains meiotic arrest in mammalian oocytes. Science 306(5703):1947–1950. https://doi.org/10.1126/science.1103974
Miao YL, Kikuchi K, Sun QY, Schatten H (2009) Oocyte aging: cellular and molecular changes, developmental potential and reversal possibility. Hum Reprod Update 15(5):573–585. https://doi.org/10.1093/humupd/dmp014
Mills M, Rindfuss RR, McDonald P, Te Velde E, Reproduction E, Society Task F (2011) Why do people postpone parenthood? Reasons and social policy incentives. Hum Reprod Update 17(6):848–860. https://doi.org/10.1093/humupd/dmr026
Molina-Garcia L, Hidalgo-Ruiz M, Cocera-Ruiz EM, Conde-Puertas E, Delgado-Rodriguez M, Martinez-Galiano JM (2019) The delay of motherhood: Reasons, determinants, time used to achieve pregnancy, and maternal anxiety level. PLoS ONE 14(12):e0227063. https://doi.org/10.1371/journal.pone.0227063
Myers M, Britt KL, Wreford NG, Ebling FJ, Kerr JB (2004) Methods for quantifying follicular numbers within the mouse ovary. Reproduction 127(5):569–580. https://doi.org/10.1530/rep.1.00095
Nelson SM, Telfer EE, Anderson RA (2013) The ageing ovary and uterus: new biological insights. Hum Reprod Update 19(1):67–83. https://doi.org/10.1093/humupd/dms043
Nie J, Xiao P, Wang X, Yang X, Xu H, Lu K, Lu S, Liang X (2018) Melatonin prevents deterioration in quality by preserving epigenetic modifications of porcine oocytes after prolonged culture. Aging (Albany NY) 10(12):3897–3909. https://doi.org/10.18632/aging.101680
Ntostis P, Iles D, Kokkali G, Vaxevanoglou T, Kanavakis E, Pantou A, Huntriss J, Pantos K, Picton HM (2021) The impact of maternal age on gene expression during the GV to MII transition in euploid human oocytes. Hum Reprod 37(1):80–92. https://doi.org/10.1093/humrep/deab226
Ottolenghi C, Uda M, Hamatani T, Crisponi L, Garcia JE, Ko M, Pilia G, Sforza C, Schlessinger D, Forabosco A (2004) Aging of oocyte, ovary, and human reproduction. Ann N Y Acad Sci 1034:117–131. https://doi.org/10.1196/annals.1335.015
Ozturk S, Guzeloglu-Kayisli O, Demir N, Sozen B, Ilbay O, Lalioti MD, Seli E (2012) Epab and Pabpc1 are differentially expressed during male germ cell development. Reprod Sci 19(9):911–922. https://doi.org/10.1177/1933719112446086
Ozturk S, Sozen B, Demir N (2015) Epab and Pabpc1 are differentially expressed in the postnatal mouse ovaries. J Assist Reprod Genet 32(1):137–146. https://doi.org/10.1007/s10815-014-0362-5
Pan Z, Zhang J, Li Q, Li Y, Shi F, Xie Z, Liu H (2012) Current advances in epigenetic modification and alteration during mammalian ovarian folliculogenesis. J Genet Genomics 39(3):111–123. https://doi.org/10.1016/j.jgg.2012.02.004
Petri T, Dankert D, Demond H, Wennemuth G, Horsthemke B, Grummer R (2020) In vitro postovulatory oocyte aging affects H3K9 trimethylation in two-cell embryos after IVF. Ann Anat 227:151424. https://doi.org/10.1016/j.aanat.2019.151424
Rambags BP, van Boxtel DC, Tharasanit T, Lenstra JA, Colenbrander B, Stout TA (2014) Advancing maternal age predisposes to mitochondrial damage and loss during maturation of equine oocytes in vitro. Theriogenology 81(7):959–965. https://doi.org/10.1016/j.theriogenology.2014.01.020
Sha QQ, Dai XX, Jiang JC, Yu C, Jiang Y, Liu J, Ou XH, Zhang SY, Fan HY (2018) CFP1 coordinates histone H3 lysine-4 trimethylation and meiotic cell cycle progression in mouse oocytes. Nat Commun 9(1):3477. https://doi.org/10.1038/s41467-018-05930-x
Sha QQ, Jiang Y, Yu C, Xiang Y, Dai XX, Jiang JC, Ou XH, Fan HY (2020a) CFP1-dependent histone H3K4 trimethylation in murine oocytes facilitates ovarian follicle recruitment and ovulation in a cell-nonautonomous manner. Cell Mol Life Sci 77(15):2997–3012. https://doi.org/10.1007/s00018-019-03322-y
Sha QQ, Zhang J, Fan HY (2020b) Function and regulation of histone H3 lysine-4 methylation during oocyte meiosis and maternal-to-zygotic transition. Front Cell Dev Biol 8:597498. https://doi.org/10.3389/fcell.2020.597498
Sha QQ, Zhu YZ, Xiang Y, Yu JL, Fan XY, Li YC, Wu YW, Shen L, Fan HY (2021) Role of CxxC-finger protein 1 in establishing mouse oocyte epigenetic landscapes. Nucleic Acids Res 49(5):2569–2582. https://doi.org/10.1093/nar/gkab107
Shao GB, Wang J, Zhang LP, Wu CY, Jin J, Sang JR, Lu HY, Gong AH, Du FY, Peng WX (2015) Aging alters histone H3 lysine 4 methylation in mouse germinal vesicle stage oocytes. Reprod Fertil Dev 27(2):419–426. https://doi.org/10.1071/RD13293
Shirasuna K, Iwata H (2017) Effect of aging on the female reproductive function. Contracept Reprod Med 2:23. https://doi.org/10.1186/s40834-017-0050-9
Uysal F, Ozturk S (2020) The loss of global DNA methylation due to decreased DNMT expression in the postnatal mouse ovaries may associate with infertility emerging during ovarian aging. Histochem Cell Biol 154(3):301–314. https://doi.org/10.1007/s00418-020-01890-w
Uysal F, Kosebent EG, Toru HS, Ozturk S (2021) Decreased expression of TERT and telomeric proteins as human ovaries age may cause telomere shortening. J Assist Reprod Genet 38(2):429–441. https://doi.org/10.1007/s10815-020-01932-1
van den Berg IM, Eleveld C, van der Hoeven M, Birnie E, Steegers EA, Galjaard RJ, Laven JS, van Doorninck JH (2011) Defective deacetylation of histone 4 K12 in human oocytes is associated with advanced maternal age and chromosome misalignment. Hum Reprod 26(5):1181–1190. https://doi.org/10.1093/humrep/der030
Wang S, Zheng Y, Li J, Yu Y, Zhang W, Song M, Liu Z, Min Z, Hu H, Jing Y, He X, Sun L, Ma L, Esteban CR, Chan P, Qiao J, Zhou Q, Izpisua Belmonte JC, Qu J, Tang F, Liu GH (2020) Single-cell transcriptomic atlas of primate ovarian aging. Cell 180(3):585-600.e519. https://doi.org/10.1016/j.cell.2020.01.009
Wilding M (2015) Potential long-term risks associated with maternal aging (the role of the mitochondria). Fertil Steril 103(6):1397–1401. https://doi.org/10.1016/j.fertnstert.2015.03.034
Xiao S, Duncan FE, Bai L, Nguyen CT, Shea LD, Woodruff TK (2015) Size-specific follicle selection improves mouse oocyte reproductive outcomes. Reproduction 150(3):183–192. https://doi.org/10.1530/REP-15-0175
Xing X, Zhang J, Wu T, Zhang J, Wang Y, Su J, Zhang Y (2021) SIRT1 reduces epigenetic and non-epigenetic changes to maintain the quality of postovulatory aged oocytes in mice. Exp Cell Res 399(2):112421. https://doi.org/10.1016/j.yexcr.2020.112421
Xu Q, Xiang Y, Wang Q, Wang L, Brind’Amour J, Bogutz AB, Zhang Y, Zhang B, Yu G, Xia W, Du Z, Huang C, Ma J, Zheng H, Li Y, Liu C, Walker CL, Jonasch E, Lefebvre L, Wu M, Lorincz MC, Li W, Li L, Xie W (2019) SETD2 regulates the maternal epigenome, genomic imprinting and embryonic development. Nat Genet 51(5):844–856. https://doi.org/10.1038/s41588-019-0398-7
Yu C, Fan X, Sha QQ, Wang HH, Li BT, Dai XX, Shen L, Liu J, Wang L, Liu K, Tang F, Fan HY (2017) CFP1 regulates histone H3K4 trimethylation and developmental potential in mouse oocytes. Cell Rep 20(5):1161–1172. https://doi.org/10.1016/j.celrep.2017.07.011
Yue MX, Fu XW, Zhou GB, Hou YP, Du M, Wang L, Zhu SE (2012) Abnormal DNA methylation in oocytes could be associated with a decrease in reproductive potential in old mice. J Assist Reprod Genet 29(7):643–650. https://doi.org/10.1007/s10815-012-9780-4
Yureneva S, Averkova V, Silachev D, Donnikov A, Gavisova A, Serov V, Sukhikh G (2021) Searching for female reproductive aging and longevity biomarkers. Aging (Albany NY) 13(12):16873–16894. https://doi.org/10.18632/aging.203206
Funding
This study was supported by Akdeniz University Research Fund (Grant no. TYL-2020-5358).
Author information
Authors and Affiliations
Contributions
YB and GT performed RT-PCR and immunohistochemistry experiments and counted follicles and luteal structures. YB analyzed all data and wrote the manuscript. GT read the manuscript. SO and YB designed the study. SO evaluated the experimental results, managed the investigation, and critically read and revised the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Ethical approval
All procedures performed in studies involving animals were in accordance with the ethical standards of international and national, and/or institutional guidelines for the care and use of animals.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Bilmez, Y., Talibova, G. & Ozturk, S. Expression of the histone lysine methyltransferases SETD1B, SETDB1, SETD2, and CFP1 exhibits significant changes in the oocytes and granulosa cells of aged mouse ovaries. Histochem Cell Biol 158, 79–95 (2022). https://doi.org/10.1007/s00418-022-02102-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00418-022-02102-3