Abstract
Background
Age-related macular degeneration (AMD) is a major cause of irreversible blindness among elderly people in developed countries. Many studies suggested that age-related maculopathy susceptibility 2 (ARMS2) is the second major susceptibility gene for AMD. Increasing evidence was found recently that inflammatory processes and oxidative stress may contribute to the pathogenesis of AMD. Meanwhile, the mechanisms underlying the contributions of ARMS2 to the pathogenesis of AMD remain unclear. The purpose of the current study was to elucidate the relationship between the ARMS2 gene and proinflammatory mediators, for further assessment of the associated biologic effects.
Methods
siRNA was used to knock down ARMS2 mRNA, and Western blotting and reverse real-time PCR were used to detect the effect of siRNA on the expression of ARMS2 in ARPE-19 cells. The expressions of C3, C5, IL-6, IL-8, and TNF-α after si-RNA knockdown were evaluated by SYBR Green I real-time PCR and ELISA.
Results
Transcription accumulative indexes (TAI = 2−delta delta CT) of ARMS2 by real-time PCR revealed that the transfection rate in the positive control group was 72.0 ± 2.07 % (P < 0.01). The ratio of absorbance values (by Western blotting) of AMRS2 to β-actin was 0.85 ± 0.122, 0.87 ± 0.143, and 0.61 ± 0.240 in the blank control group, scrambled ARMS2-siRNA group, and ARMS2-siRNA group respectively (F = 42.5, P < 0.01). The secreted protein levels of C3, C5, IL-6, IL-8, and TNF-α were found by ELISA to be reduced by 34.24 ± 1.81 %, 37.15 ± 2.02 %, 35.11 ± 1.75 %, 30.11 ± 2.19 %, and 34.33 ± 2.18 % respectively, in the siRNA-ARMS2 group (P < 0.05). Compared with the blank control group, reduced TAI of C3, C5, IL-6, IL-8, and TNF-α were detected by real-time PCR in the ARMS2-siRNA group.
Conclusion
This study produced evidence supporting the notion that the ARMS2 risk allele for AMD is linked directly or indirectly to proinflammatory mediators. More importantly, our data indicate that the change in ARMS2 may affect C3, C5, IL-6, IL-8, and TNF-α levels, and this may be one of the mechanisms of AMD development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Age-related macular degeneration (AMD), a major cause of irreversible blindness among elderly people in developed countries, is a multifactorial disease with heterogeneous clinical manifestations [1]. In Asian populations, most vision-threatening cases of AMD are of the exudative type, while the atrophic type is more common in Caucasian populations [2, 3]. The cause of AMD is complicated because multiple genetic and environmental factors are involved in its pathogenesis. Epidemiological studies have indicated that factors such as age, smoking, gender, obesity, hypertension, and genetic background are associated with AMD [2, 4]. Increasing evidence has been found recently that inflammatory processes and oxidative stress may contribute to the pathogenesis of AMD [5–7].
Investigations on genome-wide and targeted genetic association have identified various polymorphisms that are important for AMD susceptibility, among which polymorphisms of complement factor H (CFH; Y402H) and age-related maculopathy susceptibility 2 (ARMS2; A69S) might have the strongest influence on the pathogenesis of AMD in the Caucasian and Asian populations respectively [8–12]. The complement pathway is interfered with AMD by inflammatory process, but how ARMS2 influences the disease is still in dispute [6–8, 13–17]. Yasuma et al. [15] reported that people over 60 years carried AMD risk alleles in the ARMS2 locus; the serum high sensitivity C-reaction protein levels were elevated. They proposed that ARMS2 may heighten the risk of developing AMD by chronic systemic inflammation. The purpose of the current study was to elucidate the relationship between the ARMS2 gene and those proinflammatory mediators, so as to further assess the associated biologic effects.
Materials and methods
Cell cultures
Human ARPE-19 (ATCC, catalog No. CRL-2302, Rockefeller, Maryland, USA) cells were cultured in Dulbecco’s modified Eagle medium/nutrient mixture F12 (DMEM/F12; Invitrogen, Carlsbad, CA, USA) (1:1) containing 10 % fetal bovine serum (FBS, Invitrogen), 100 IU/mL penicillin G (InvitrogenUSA), and 100 μg/mL streptomycin (Invitrogen). The cells were incubated at 37°C in a humidified atmosphere of 5 % CO2 and 95 % air. The culture medium was changed every 3 days. After 90 % confluence was reached, the cells were routinely passaged by dissociation in 0.05 % trypsin/0.02 % ethylene diamine tetraacetic acid (EDTA). The cells were seeded in 96- or 6-well plates, according to the different requirements.
Transfection of siRNA
The transfection reagent (FlexTube siRNA Premix) was purchased from Qiagen (Hilden, Germany), including a short-interfering RNA (siRNA) designed for the human ARMS2 gene (cat. no. SI00507990, target sequence CTCCATGATCCCAGCTGCTAA) and a control siRNA, which contained a scrambled sequence (cat. no. SI03650318, target sequence CAGCATCAATGTGAAGCCAAA). FlexTube siRNA Premix was resuspended before the first use according to the manufacturer’s protocol. Twenty-four hours before siRNA transfection, 250,000 cells/well were seeded in 6-well plates. At the time of transfection with siRNA, the cells were approximately 50–60 % confluent in 2 ml of complete DMEM/F12 medium, and then were transfected for 24 hrs with 42 μl of transfection reagent diluted in 2 ml of DMEM/F12 (HyClone, Thermo Fisher Scientific, Beijing, China) medium. The final concentration of the siRNAs was 25 nmol/l each. After transfection, the supernatant was removed from all wells, and some of the cells were incubated with complete DMEM/F12 medium for another 24 hrs to monitor changes in ARMS2 expression with real-time PCR, Western blotting, and ELISA.
Human ARPE-19 cells transfected with ARMS2-siRNA served as a positive control group to detect the transfection efficiency. The non-transfected ARPE-19 cells and scrambled transfected ARPE-19 cells with PBS and scrambled-ARMS2-siRNA were taken respectively as blank control and blank transfection groups. The forward and downward specific sequences for ARMS2, complement component 3 (C3), complement component (C5), interleukin (IL)-6, IL-8, tumor necrosis factor α (TNF-α), and β-actin are shown in Table 1.
Detection of ARMS2, C3, C5, IL-6, IL-8, and TNF-α by SYBR Green I Real-Time PCR
Total mRNA was extracted from ARPE-19 cells using a NucleoSpin RNAII Assay Kit (Macherey-Nagel, Düren, Germany) according to the manufacturer’s instructions. The DNA concentration was standardized for working DNA using an Eppendorff pipetting machine (Eppendorf AG, Hamburg, Germany). The first strand complementary DNA (cDNA) was synthesized from 1 μg of total RNA in a 10-μl reaction mixture using a PrimeScript 1st Strand cDNA Synthesis Kit (TaKaRa, Dalian, China). PCR reactions were performed using GoTaq Flexi DNA polymerase (Promega, Madison, WI, USA) in a 10-μl mixture containing 0.25 μmmol/L each of dNTP (Deoxyribonucleoside5′-Triphosphates), 2 μl 5 × Colorless GoTaq Flexi Buffer, 2.5 mmol/l MgCl2, 0.25 mol/l of each primer, 4 units GoTaq DNA polymerase, and 25 ng cDNA. After an initial denaturation step at 95 °C for 3 min, amplification was performed for a total of 30 cycles under the following conditions: denature at 95 °C for 30 s, annealing at 40.7 °C for 30 s, and extension at 72 °C for 30 s. Then the final step was at 72 °C for 10 min. A blank control substitution of water for the cDNA template was included in all experiments. The housekeeping gene β-actin was used as an internal control because it was believed to be continuously expressed in cells. The purity and amount of RNA was determined using an ultraviolet (UV) spectrophotometer at 260 nm and 280 nm (A260/A280 > 1.8).
SYBR green I was diluted with a ratio of 1:10 000 and added to the reaction. The primers were designed with Primer 5.0. The mRNA expression was expressed as the transcription accumulative index (TAI = 2−delta delta CT). The three groups were used to detect the expressions of ARMS2, C3, C5, IL-6, IL-8, and TNF-α.
Western blotting assay
The cells in the three groups were seeded in tissue culture dishes, and allowed to grow for 72 hrs in the medium described above, before they were incubated for another 24 hrs in the same medium but without serum. Cells were lysed with lysis buffer containing 150 mM NaCl, 1 % Igepal, 1 mM EDTA, 50 mM Tris, and 10 % protease inhibitor mixture on ice for 30 min. The supernatants were extracted by centrifugation at 4 °C with 14,000 r/min and stored in –80 °C. Protein quantification was performed with the bicinchoninic acid (BCA) method according to the manufacturer’s instructions (Thermo Scientific, Rockford, IL, USA). Thirty μg protein for each sample was subjected to 10 % SDS-PAGE, electro-transferred to nitrocellulose membrane (NC), blocked with 1 % Tris-Buffered Saline and Tween 20 (TBST)/5 % skimmed milk powder, incubated with ARMS2 (1:1000; Abcam, Cambridge, UK) primary antibody or β-actin (1: 20,000; Oncogene Research Products, San Diego, CA, USA) diluted in 1 % TBST/5 % skimmed milk powder overnight, rinsed three times in 0.1 % TBST, and incubated with Horseradish peroxidase labeled goat anti-rabbit IgG (1:5000, ZSGB-BIO, Beijing, China) for 2 hrs. After diaminobenzidin (DAB) coloration for 5–10 min, brown bands were taken as positive results. By using GAS9500 image analysis system (UVItecy Co., London, UK), the absorbance values of hybridity bands on NC membrane and the ratio to β-actin representing the protein relative quantity were determined.
Determination of C3, C5, IL-6, IL-8, and TNF-α by ELISA
The supernatants in the three groups were collected from cell cultures described above. The supernatants were tested by ELISA for C3, C5, IL-6, IL-8, and TNF-α. ELISA kits were purchased from R&D Systems, Inc. (Minneapolis, MN, USA) and were performed based on the kit instruction. Values were expressed as pg/ml of conditioned medium.
Statistical analysis
Data were expressed as X− ± s. All analyses for statistically significant differences were performed with the one-way ANOVA test. P < 0.01 was considered significant. All data were presented as means (SD) of three independent experiments, each performed in duplicate.
Results
Transfection rate of siRNA SYBR Green I Real-Time PCR
Transcription accumulative indexes (TAI = 2−delta delta CT) of ARMS2 in the blank control group, scrambled ARMS2-siRNA group, and ARMS2-siRNA group were 1.06 ± 0.043, 1.08 ± 0.016, and 0.71 ± 0.011 respectively (P < 0.01) in an ABI 7500 real-time PCR (Applied Biosystems, Foster City, California, USA). The transfection rate in the positive control group was 72.0 % (±1.07, P < 0.01).
Detection of ARMS2 protein expression by Western blotting
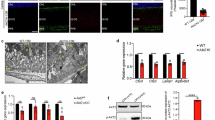
The ratio of absorbance values of AMRS2 to β-actin was 0.61 ± 0.240, 0.87 ± 0.143, and 0.85 ± 0.122 (F = 42.53, P < 0.01) (Fig. 1) in the ARMS2-siRNA group, blank control group, and scrambled ARMS2-siRNA group respectively. The ARMS2 level was decreased by 29.0 ± 1.12 % after transfection in the ARMS2-siRNA group.
ARMS2 protein expression detected by Western blotting. M: marker; 1: normal group; 2: scrambled ARMS2-siRNA transfected group; 3: ARMS2-siRNA transfected group. ARMS2 level was decreased by 29.0 ± 2.244 % after transfection in the ARMS2-siRNA group. Significant difference was found between the ARMS2-siRNA group and the blank control group
Detection of C3, C5, IL-6, IL-8, and TNF-α expressions by real-time PCR
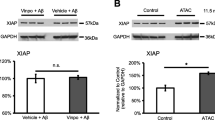
Transcription accumulative indexes (TAI = 2−delta delta CT) of C3, C5, IL-6, IL-8, and TNF-α in differently treated cells are summarized in Fig. 2. Compared with the blank control group, the expressions of C3, C5, IL-6, IL-8, and TNF-α in the ARMS2-siRNA group was significantly reduced (P < 0.05). In the untreated and scrambled siRNA-transfected groups, the expressions of C3, C5, IL-6, IL-8, and TNF-α were similar, and no significant differences were found between these two groups (P = 0.441).
Transcription accumulative indexes (TAI = 2−delta delta CT) of ARMS2 in the blank control group, scrambled ARMS2-siRNA group, and ARMS2-siRNA group were 1.06 ± 0.043, 1.08 ± 0.016, and 0.71 ± 0.011 respectively (P < 0.01), in an ABI 7500 real-time PCR (Applied Biosystems, Foster City, CA, USA). The transfection rate in the positive control group was 72.0 % (±1.07, P < 0.01). Compared with the blank control group, the TAI of C3, C5, IL-6, IL-8, and TNF-α in the ARMS2-siRNA group was reduced. In the normal group and the scrambled siRNA-transfected group, the expressions of C3, C5, IL-6, IL-8, and TNF-α were similar, and no significant differences were found in these two groups (P = 0.441). No significant difference was found between the blank control group and the scrambled ARMS2-siRNA group (*). Significant difference was found between the ARMS2-siRNA group and the blank control group (**). Error bars indicate SD. **P < 0.05
Detection of C3, C5, IL-6, IL-8, and TNF-α protein expressions by ELISA
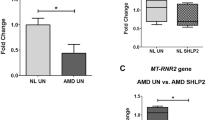
The protein expression level was assayed by ELISA to examine the effect of siRNA-ARMS2 interference on C3, C5, IL-6, IL-8, and TNF-α. The values of C3, C5, IL-6, IL-8, and TNF-α in the three groups are shown in Fig. 3. The secreted protein levels of the siRNA-ARMS2 group continued to reduce by 34.24 ± 1.812 %, 37.15 ± 2.021 %, 35.11 ± 1.751 %, 30.11 ± 2.191 %, and 34.33 ± 2.182 % respectively (P < 0.05), after transduction as compared with the contents in the medium of the blank control group. No significant differences were found in the normal group and the scrambled siRNA-transfected group (P = 0.343).
The secreted protein levels of the siRNA-ARMS2 group were reduced by 34.24 ± 1.812 %, 37.15 ± 2.021 %, 35.11 ± 1.751 %, 30.11 ± 2.191 %, and 34.33 ± 2.182 % respectively (P < 0.05), after transduction as compared with the content in the medium of blank control group. No significant difference was found between the blank control group and the scrambled ARMS2-siRNA group (*). Significant difference was found between the ARMS2-siRNA group and the blank control group (**). Error bars indicate SD. ** P < 0.05
Discussion
Although it is generally accepted that both ARMS2 and inflammation are related with AMD, the association of ARMS2 and proinflammatory mediators has not been reported. In the current study, we investigated the effect of ARMS2 interference on the expression of some inflammatory cytokines. The results showed that interference of ARMS2 can down-regulate the expression of some of the major inflammatory cytokines, including IL-6, IL-8, and TNF-α, as well as the complement factors C3 and C5.
To date, the well documented alteration of inflammatory markers in AMD patients is the elevation of the C3, C5, CFH, IL-6, IL-8, and TNF-α levels [5–7]. These proinflammatory mediators are also associated with disease progression. CFH, C3, and C5 are main components of the alternative pathway. Factor H is an inhibitor of complement activation, while C3 is a central component of all complement activation pathways. Dysfunction of the complement system may result in local inflammation, autoimmune reactions, and tissue damage, and may play an important role in the etiology of AMD. IL-6 is a proinflammatory cytokine, which amplifies immune and inflammatory responses and plays a critical role in the occurrence of autoimmune diseases. It is also the key factor for the stimulation of immune reactions, inflammatory processes, and the occurrence of autoimmune diseases. Increased systemic or local IL-6 levels have been observed in various autoimmune diseases. IL-6–deficient mice were resistant to various experimentally-induced autoimmune diseases [13–17]. It has been reported that serum IL-6 levels correlated with the progression of AMD, and high levels of serum IL-6 were associated with the geographic atrophy type of AMD [13, 14]. IL-8 is a group of peptides produced by various types of cells, which activate and recruit polymorphonuclear leukocytes in acute and chronic inflammatory processes. It is supposed that IL-6 and IL-8 are released by degenerating retinal pigment epithelial cells and associated with drusen in AMD. TNF-α is a proinflammatory cytokine produced by macrophages and T-cells, and involved in inflammation and apoptosis. Activation of caspases, TUNEL-positive staining, and observation of hypodiploid cells suggest that apoptosis occur in cells treated with AMD alone or with AMD/TNF. In the eye, TNF-α appears to play a role in the pathogenesis of inflammatory, edematous, neovascular, and neurodegenerative disorders. Our results showed that all these inflammatory cytokines had a close relationship with the expression of ARMS2. A strong association between AMD and ARMS2 has been confirmed by multiple studies, but the exact mechanism remains unclear. The proposed functions of ARMS2 are in mitochondria, extracellular matrix, or as a noncoding RNA [18–24]. Yasuma et al. [15] found that normal subjects with AMD risk alleles in the ARMS2/HTRA1 locus may be at risk of increased hypersensitive C-reaction protein levels and chronic systemic inflammation with aging. These findings indicate that the ARMS2 risk allele might increase the risk of AMD by inflammation pathway. Our study provided evidence supporting the above notion that the ARMS2 risk allele for AMD is linked directly or indirectly to chronic systemic inflammation.
HtrA1 and ARMS2 are two neighboring genes located at chromosome 10q26. This area is regarded as a major candidate region associated with the susceptibility of several types of AMD [25], including polypoidal choroidal vasculopathy [26]. There is strong linkage disequilibrium across the ARMS2/HTRA1 region, making genetic association studies alone insufficient to distinguish the two candidates. Functional investigations of these two genes showed that decrease of ARMS2 message was associated with increase in HTRA1 expression [27–29]. HTRA1 is a multifunctional serine protease that is expressed in a variety of human tissues and cells, including the retinal pigment epithelium and vascular endothelium [24, 27–29]. Preliminary evidence suggests that HtrA1 may increase the risk of AMD by inflammation pathway [24–29]. Therefore, we audaciously hypothesize that the ARMS2 gene might be involved in the inflammation mechanism through influencing HtrA1 genes and subsequently the proinflammatory mediators.
There was an obvious limitation in this study. We did not investigate the relationship between ARMS2 and the protective factors, such as CFH, toll-like receptor-3 (TLR-3), IL-10, IL-27, etc. [30–32]. TLR-3 is an essential receptor of the innate immune system and the first responder for protection against bacterial and viral pathogens. Recently, genetic variants of both TLR3 and the key complement regulatory protein CFH were found to confer protection against AMD [32]. Hasegawa et al. [30] suggested that IL-10 had anti-angiogenic properties on choroidal neovascularization formation, and IL-27, a member of the IL-12 cytokine family, had an angiostatic effect and regulated the development of laser-induced experimental choroidal neovascularization in mice. IL-27 did not affect macrophage migration, but inhibited its VEGF production. However, our data indicated that ARMS2 was associated with the expression levels of C3, C5, IL-6, IL-8, and TNF-α. Our findings can further support the hypothesis that ARMS2 might be involved in the pathogenesis of AMD, at least partially, through inflammatory and immune-related mechanisms. However, the exact mechanism requires further investigations.
References
Jager RD, Mieler WF, Miller JW (2008) Age-related macular degeneration. N Engl J Med 358:2606–2617
Kikuchi M, Nakamura M, Ishikawa K, Suzuki T, Nishihara H, Yamakoshi T, Nishio K, Taki K, Niwa T, Hamajima N, Terasaki H (2007) Elevated C-reactive protein levels in patients with polypoidal choroidal vasculopathy and patients with neovascular age-related macular degeneration. Ophthalmology 114:1722–1727
Oshima Y, Ishibashi T, Murata T, Tahara Y, Kiyohara Y, Kubota T (2001) Prevalence of age related maculopathy in a representative Japanese population: the Hisayama study. Br J Ophthalmol 85:1153–1157
Klein R, Peto T, Bird A, Vannewkirk MR (2004) The epidemiology of age-related macular degeneration. Am J Ophthalmol 137:486–495
Ambati J, Ambati BK, Yoo SH, Ianchulev S, Adamis AP (2003) Age related macular degeneration: etiology, pathogenesis, and therapeutic strategies. Surv Ophthalmol 48:257–293
Donoso LA, Kim D, Frost A, Callahan A, Hageman G (2006) The role of inflammation in the pathogenesis of age-related macular degeneration. Surv Ophthalmol 51:137–152
Hollyfield JG, Bonilha VL, Rayborn ME, Yang X, Shadrach KG, Lu L, Ufret RL, Salomon RG, Perez VL (2008) Oxidative damage-induced inflammation initiates age-related macular degeneration. Nat Med 14:194–198
Rivera A, Fisher SA, Fritsche LG, Keilhauer CN, Lichtner P, Meitinger T, Weber BH (2005) Hypothetical LOC387715 is a second major susceptibility gene for age-related macular degeneration, contributing independently of complement factor H to disease risk. Hum Mol Genet 14:3227–3236
Jakobsdottir J, Conley YP, Weeks DE, Mah TS, Ferrell RE, Gorin MB (2005) Susceptibility genes for age-related maculopathy on chromosome 10q26. Am J Hum Genet 77:389–407
Klein RJ, Zeiss C, Chew EY, Tsai JY, Sackler RS, Haynes C, Henning AK, SanGiovanni JP, Mane SM, Mayne ST, Bracken MB, Ferris FL, Ott J, Barnstable C, Hoh J (2005) Complement factor H polymorphism in age-related macular degeneration. Science 308:385–389
Edwards AO, Ritter R 3rd, Abel KJ, Manning A, Panhuysen C, Farrer LA (2005) Complement factor H polymorphism and age related macular degeneration. Science 308:421–424
Haines JL, Hauser MA, Schmidt S, Scott WK, Olson LM, Gallins P, Spencer KL, Kwan SY, Noureddine M, Gilbert JR, Schnetz-Boutaud N, Agarwal A, Postel EA, Pericak-Vance MA (2005) Complement factor H variant increases the risk of age related macular degeneration. Science 308:419–421
Yates JR, Sepp T, Matharu BK, Khan JC, Thurlby DA, Shahid H, Clayton DG, Hayward C, Morgan J, Wright AF, Armbrecht AM, Dhillon B, Deary IJ, Redmond E, Bird AC, Moore AT (2007) Genetic Factors in AMD Study Group. Complement C3 variant and the risk of age-related macular degeneration. N Engl J Med 357:553–561
Gold B, Merriam JE, Zernant J, Hancox LS, Taiber AJ, Gehrs K, Cramer K, Neel J, Bergeron J, Barile GR, Smith RT, AMD Genetics Clinical Study Group, Hageman GS, Dean M, Allikmets R (2007) Variation in factor B (BF) and complement component 2 (C2) genes is associated with age-related macular degeneration. Nat Genet 38:458–462
Yasuma TR, Nakamura M, Nishiguchi KM, Kikuchi M, Kaneko H, Niwa T, Hamajima N, Terasaki H (2010) Elevated C-reactive protein levels and ARMS2/HTRA1 gene variants in subjects without age-related macular degeneration. Mol Vis 16:2923–2930
Seddon JM, George S, Rosner B, Rifai N (2005) Prospective assessment of C-reactive protein, interleukin 6, and other cardiovascular biomarkers. Arch Ophthalmol 123:774–782
Baas DC, Ho L, Ennis S, Merriam JE, Tanck MW, Uitterlinden AG, de Jong PT, Cree AJ, Griffiths HL, Rivadeneira F, Hofman A, van Duijn C, Smith RT, Barile GR, Gorgels TG, Vingerling JR, Klaver CC, Lotery AJ, Allikmets R, Bergen AA (2010) The complement component 5 gene and age-related macular degeneration. Ophthalmology 117:500–511
Bessho H, Honda S, Kondo N, Negi A (2011) The association of age-related maculopathy susceptibility 2 polymorphisms with phenotype in typical neovascular age-related macular degeneration and polypoidal choroidal vasculopathy. Mol Vis 17:977–982
Fritsche LG, Loenhardt T, Janssen A, Fisher SA, Rivera A, Keilhauer CN, Weber BH (2008) Age-related macular degeneration is associated with an unstable ARMS2 (LOC387715) mRNA. Nat Genet 40:892–896
Xu YT, Wang Y, Chen P, Xu HF (2012) Age-related maculopathy susceptibility 2 participates in the phagocytosis functions of the retinal pigment epithelium. Int J Ophthalmol 5:125–132
Kanda A, Chen W, Othman M, Branham KE, Brooks M, Khanna R, He S, Lyons R, Abecasis GR, Swaroop A (2007) A variant of mitochondrial protein LOC387715/ARMS2, not HTRA1, is strongly associated with age-related macular degeneration. Proc Natl Acad Sci USA 104:16227–16232
Wang G, Spencer KL, Scott WK, Whitehead P, Court BL, Ayala-Haedo J, Mayo P, Schwartz SG, Kovach JL, Gallins P, Polk M, Agarwal A, Postel EA, Haines JL, Pericak-Vance MA (2010) Analysis of the indel at the ARMS2 3’UTR in age-related macular degeneration. Hum Genet 127:595–602
Wang G, Spencer KL, Court BL, Olson LM, Scott WK, Haines JL, Pericak-Vance MA (2009) Localization of age-related macular degeneration-associated ARMS2 in cytosol, not mitochondria. Invest Ophthalmol Vis Sci 50:3084–3090
Kanda A, Stambolian D, Chen W, Curcio CA, Abecasis GR, Swaroop A (2010) Age-related macular degeneration-associated variants at chromosome 10q26 do not significantly alter ARMS2 and HTRA1 transcript levels in the human retina. Mol Vis 16:1317–1323
Yang Z, Tong Z, Chen Y, Zeng J, Lu F, Sun X, Zhao C, Wang K, Davey L, Chen H, London N, Muramatsu D, Salasar F, Carmona R, Kasuga D, Wang X, Bedell M, Dixie M, Zhao P, Yang R, Gibbs D, Liu X, Li Y, Li C, Campochiaro B, Constantine R, Zack DJ, Campochiaro P, Fu Y, Li DY, Katsanis N, Zhang K (2010) Genetic and functional dissection of HTRA1 and LOC387715 in age-related macular degeneration. PLoS Genet 6:e1000836
Gotoh N, Yamada R, Nakanishi H, Saito M, Iida T, Matsuda F, Yoshimura N (2008) Correlation between CFH Y402H and HTRA1 rs11200638 genotype to typical exudative age-related macular degeneration and polypoidal choroidal vasculopathy phenotype in the Japanese population. Clin Exp Ophthalmol 36:437–442
Dewan A, Liu M, Hartman S, Zhang SS, Liu DT, Zhao C, Tam PO, Chan WM, Lam DS, Snyder M, Barnstable C, Pang CP, Hoh J (2006) HTRA1 promoter polymorphism in wet age-related macular degeneration. Science 314:989–992
Yang Z, Camp NJ, Sun H, Tong Z, Gibbs D, Cameron DJ, Chen H, Zhao Y, Pearson E, Li X, Chien J, Dewan A, Harmon J, Bernstein PS, Shridhar V, Zabriskie NA, Hoh J, Howes K, Zhang K (2006) A variant of the HTRA1 gene increases susceptibility to age-related macular degeneration. Science 314:992–993
Chen H, Yang Z, Gibbs D, Yang X, Hau V, Zhao P, Ma X, Zeng J, Luo L, Pearson E, Constantine R, Kaminoh Y, Harmon J, Tong Z, Stratton CA, Cameron DJ, Tang S, Zhang K (2008) Association of HTRA1 polymorphism and bilaterality in advanced age-related macular degeneration. Vision Res 48:690–694
Hasegawa E, Oshima Y, Takeda A, Saeki K, Yoshida H, Sonoda KH, Ishibashi T (2012) IL-27 inhibits pathophysiological intraocular neovascularization due to laser burn. J Leukoc Biol 91:267–273
Matsumura N, Kamei M, Tsujikawa M, Suzuki M, Xie P, Nishida K (2012) Low-dose lipopolysaccharide pretreatment suppresses choroidal neovascularization via IL-10 induction. PLoS One 7:e39890
Patel AK, Hackam AS (2012) Toll-like receptor 3 (TLR3) protects retinal pigmented epithelium (RPE) cells from oxidative stress through a STAT3-dependent mechanism. Mol Immunol 54:122–131
Acknowledgments
The authors thank Ms Ping Lin for her editorial assistance.
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zeng, F., Zhang, M., Xu, Y. et al. ARMS2 interference leads to decrease of proinflammatory mediators. Graefes Arch Clin Exp Ophthalmol 251, 2539–2544 (2013). https://doi.org/10.1007/s00417-013-2442-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00417-013-2442-0