Abstract
Hereditary cerebellar ataxias (HCAs) are clinically and genetically heterogeneous neurodegenerative disorders, characterised by a cerebellar syndrome and other neurological or non-neurological signs. So far, more than 20 genes have been described in autosomal dominant HCA; in autosomal recessive HCA, even more genes are involved, in often more complex phenotypes. Because of that complexity, the genetic diagnosis of these diseases is often based on the next-generation sequencing techniques. In this review paper, we discuss the major contributions that they have made to the genetic landscape of HCAs. Numerous novel genes have been identified; still more have recently been implicated in HCAs in addition to being responsible for other diseases. The phenotypic spectrum associated with a single gene constantly gains in complexity. Novel types of mutations or transmissions in known genes are regularly being identified. All these factors make genotype–phenotype correlations particularly difficult. Some but not all of this variability can be explained by different pathophysiological consequences (loss of function, gain of function, variable levels of haploinsufficiency). This also raises the question of modifier genes. Finally, we highlight some functional pathways that increasingly appear important in HCAs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hereditary cerebellar ataxias (HCAs) are clinically and genetically heterogeneous neurodegenerative disorders. Clinically, they are characterised by a cerebellar syndrome, with dysarthria, gait or limb incoordination. Ocular movements are often altered. In many cases, other neurological or non-neurological systems are also affected as the disease progresses [1, 2]. Genetically, all modes of transmission have been described. In autosomal dominant (AD)-HCAs, called spinocerebellar ataxias (SCAs), trinucleotide CAG expansions are the most frequent mutations, followed by other non-coding expansions and conventional point mutations or rearrangements. More than 20 genes and 35 loci have been reported so far, but the genetic causes of AD-HCAs remain unknown in more than 40 % of cases [3]. The genetic landscape of autosomal recessive (AR)-HCAs is even more intricate, with numerous genes involved in complex syndromes [2], as in X-linked HCAs (ABC7 [4], SLC9A6 [5], PRPS1 [6], FMR1 [7]). This large genetic heterogeneity renders the genetic diagnosis of inherited HCAs difficult; in the last few years, much interest has, therefore, been focused on next-generation sequencing, using either targeted sequencing (TS) [8] or whole-exome sequencing (WES). The latter has considerably changed the genetic landscape of HCAs, allowing many new genes to be discovered; but it has also brought challenges in terms of genotype–phenotype correlations, with a broadening of the phenotypes linked to mutations in a single gene, and the description of novel types of mutations, or transmission modes. In this review, we summarise the recent advances in the broad panorama of HCA genetics. We also stress some of the common pathways underlining these diseases that have gained in importance recently.
A bumper harvest of novel genes
Thanks to WES, there has been an explosion in the number of novel HCA-causative genes (Table 1). In AD-HCAs, new altered functions are highlighted, such as ribosomal translation in SCA26, due to mutations in EEF2, associated with late-onset pure cerebellar ataxia [9]. Loss-of-function mutations in ELOVL5 (SCA38) were found in four independent pedigrees, and associated with low levels of serum arachidonic acid and docosahexaenoic acid [10]. Mutations in TMEM240 (SCA21) [11] and NOL3 [12] have been described in complex phenotypes. In AR-HCAs (SCAR16), STUB1 mutations were described in three kindred with early-onset ataxia [13], and one family with ataxia and hypogonadism [14]. STUB1 was subsequently implicated in later-onset ataxia, with associated pyramidal tract damage [15]. Another E3 ubiquitin ligase, RNF216, was involved in two families with AR-HCA, hypogonadism and dementia [16]. The prominence of the ubiquitin–proteasome pathway in HCAs [17] is supported by the involvement of UCHL1 [18] and, putatively, UBR4 (EA8) [19] (Table 1).
Spontaneous or knock-out mouse models have supported the pathogenicity of several new genes. Mutations were found in WWOX in two families with childhood-onset HCA, generalised tonic–clonic epilepsy and mental retardation (SCAR12); while the knock-out mice present spontaneous and audiogenic seizures as well as balance disturbances [20]. In a family with ataxia and sensorineural hearing loss, a homozygous mutation in SLC9A1 was identified; in the spontaneous swe mutant mice, degeneration of deep cerebellar, vestibular and cochlear nuclei is reported [21]. In GRID2, loss-of-function mutations have been described in early-onset ataxia phenotypes with various accompanying symptoms, concordantly with hotfoot mice [22–24]. Missense mutations were described in AD congenital to late-onset HCA [25], as previously reported in lurcher mice [26].
The preponderance of mitochondrial dysfunction in HCAs was confirmed by homozygous COX20 mutations identified in a family with congenital HCA and complex IV deficiency [27] and in a patient with dystonia–ataxia syndrome [28].
Finally, light has been shed on the vesicular compartment and, more specifically, the synaptic transmission, with mutations in VAMP1 in AD spastic ataxia (SPAX1) [29], SNAP25B in AD-HCA with myasthenia and intellectual disability [30], and SNX14 in AR-HCA [31].
The phenotypic apples and bananas of a single gene
Every geneticist knows that mutations in a single gene may be responsible for different diseases; the best-known example in SCAs is probably CACNA1A/SCA6, with mutations responsible for familial hemiplegic migraine type I, episodic ataxia type II (EA2) [32], and SCA6 [33]. The genotype–phenotype correlation is not perfect since CAG expansions are involved in EA2 [34], and, conversely, point mutations in SCA6 [35]. Recently, loss-of-function mutations in KCND3, previously implicated in cardiomyopathy (Brugada syndrome), were described in nine families with mild slowly progressive AD-HCA (SCA19 [36]/SCA22 [37]), confirming the involvement of ionotropic channels in HCAs. Along with the extensive use of WES for the genetic diagnostic came a broadening of the phenotypes linked to several genes, either through an enlargement of the clinical picture previously observed or a novel implication in a different disease. ATM mutations, responsible for ataxia–telangectasia, have been described in several families with atypical presentation, such as later age at onset, normal levels of alpha-foeto-protein, and a large range of movement disorders, including dystonia and tremor [38]. More dramatically, a TGM6 mutation previously implicated in SCA35 has been shown to segregate with acute myeloid leukaemia [39].
Several observations have identified ceroid lipofuscinosis (CLN) genes, traditionally responsible for wider neurodegenerative phenotypes, as good candidates in adult-onset AR-HCA: TPP1/CLN2 in SCAR7 [40], CLN5 in adult-onset AR-HCA with cognitive decline [41], and CTSD/CLN10 in AR-HCA with cognitive decline and retinitis pigmentosa [8].
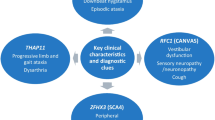
As for why the phenotype linked to a single gene may vary, the answer is not always obvious. In some cases, different types of mutations (loss of function versus gain of function) may occur. In others, the degree of haploinsufficiency might explain some of the phenotypic variability. PNPLA6, previously known to cause complex hereditary spastic paraplegia (SPG39), was recently described as a cause of Boucher-Neuhäuser and Gordon-Holmes syndromes, as well as spastic ataxia [42], exhibiting a continuous spectrum of phenotypes including features amongst spasticity, ataxia, hypogonadism and chorioretinal dystrophy, none of them being mandatory (Fig. 1). Recently, it was identified in Oliver-McFarlane syndrome, which adds trichological and pituitary abnormalities to the previous spectrum. Clinical presentation was shown to correlate with the degree of enzyme malfunction, with no activity in the latter [43].
Schematic representation of the phenotypic spectrum linked to newly described genes involved in ataxia plus hypogonadism. RNF216 mutations are responsible for a uniform phenotype (dark red) with constant clinical signs [16]. Apart from severe D-bifunctional protein deficiency, the ataxia plus hypogonadism phenotype caused by HSD17B4 mutations is also rather homogeneous (dark blue) [108–110]. In contrast, PNPLA6 (orange) and STUB1 (light blue) have been shown to induce variable phenotypes, with no or few mandatory signs [13–15, 42, 43]. This illustrates the complexity of genotype–phenotype correlations, accentuated with the next generation sequencing era
Numerous other examples of phenotypic broadening are summarised in Table 1 (DNMT1, ELOVL4/SCA34, C9orf72, UCHL1, GBA2/SPG46, HSD17B4, SRD5A3, CCDC48C/SCA40, GJB1).
Faced with this high clinical heterogeneity, many attempts have been made to identify a core phenotype linked to mutations in a given gene. ARCA3, caused by ANO10 mutations, almost always presents as cerebellar atrophy with lack of neuropathy [44]. All PLA2G6 carriers of biallelic mutations show a childhood-onset cerebellar atrophy preceding brain iron accumulation, with some correlation between the genotype and age at onset [45].
Novel types of mutations/transmission in previously described genes
On top of the description of novel genes and novel phenotypes for genes previously involved in human pathology, the new era of sequencing also reveals new types of mutations and transmission modes, for variants in previously described genes. This renders the functional interpretation of mutations particularly complex. In ITPR1/SCA15, most of the mutations reported up to 2012 were loss of function, with only one missense described [46]. Since 2012, three new missense variants, including one de novo, have been described [47, 48]. Notably, the variants described in [47] occur for congenital HCA, which is not the typical phenotype of SCA15. In SCA28, missense mutations were traditionally described in exon 15 and 16 of AFG3L2 [49]. Recently, reports mentioned a deletion of exons 14–16 in two families with classical presentation [50], a deletion encompassing the entire gene in a patient with infancy-onset global developmental delay [51], and a frameshift mutation leading to a premature stop codon associated with late-onset HCA and cognitive decline [52]. AFG3L2 was also recently involved in an AR syndrome comprised of spastic ataxia and neuropathy, SPAX5 [53]. SPTBN2, whose in-frame deletions and missense mutations account for AD-SCA5, was linked to AR ataxia and cognitive impairment, with homozygous loss-of-function mutations [54, 55].
Starting to uncover the genes underlying phenotypic heterogeneity
These elements of heterogeneity in the phenotype–genotype correlations raise the question of potential modifier genes. In SCAs with (CAG)n-repeats, the negative correlation between the size of the polyglutamine expansion and the age at onset [56] only accounts for 50–70 % of its variability. A large cohort study on 1255 affected patients allowed an assessment to be made of the influence on the age at onset of the length of the normal allele in trans (SCA1, 6, 7) and of other (CAG)n-containing genes [57]. Furthermore, intermediate-size CAG repeats in ATXN2 were recently shown to be a risk factor in other neurodegenerative diseases, such as amyotrophic lateral sclerosis (ALS), frontotemporal dementia with ALS (>29 repeats [58]), and possibly in Parkinson’s disease (>24 repeats [59]). Conventional mutations can also act as modifiers. In a family with optic atrophy plus syndrome and ataxia, a compound heterozygosity in OPA1 was established, with a non-pathogenic per se coding variant, and a deep intronic mutation responsible for aberrant splicing and a premature stop codon [60].
These specific aspects, that bring Mendelian diseases beyond their traditional fields, are still not well understood, and much effort is, therefore, being devoted to them.
Common pathways emerging in HCAs
Genetic advances have allowed the identification of novel common pathways involved in HCAs. In AD-HCAs, transcription dysregulation, protein aggregation, RNA toxicity, ion channels’ function, and mitochondrial pathways have recently been extensively reviewed [61]. Autophagy was also described as being important, in neurodegenerative diseases in general, and more specifically in SCA3 and SCA14 [61]. Recently, it has been proven to be impaired in SCA7 patients and knock-in mice [62]. The early autophagy-associated gene ATG12 was shown to be upregulated in patients’ peripheral blood mononuclear cells, correlating with the severity of disease, which might constitute a useful biomarker for therapeutic approaches. In AR-HCAs, multiple pathways have been implicated, including mitochondrial dysfunction, defects of lipoprotein assembly, deficiency of DNA repair, chaperone dysfunction [2], and ubiquitin–proteasome complex [17]. However, as new genes are described, the multiplicity of pathways involved increases. Rather than discussing all the pathways involved in detail, we shall concentrate on the ones that stand out, either because of recent advances or because of recent implication in HCAs.
Channels are not only involved in episodic ataxias
Channels function has long been an on-going topic in the field of SCAs. The first channel to be implicated in the disease was CACNA1A, encoding the alpha-1a voltage-dependent calcium channel type P/Q, whose small polyglutamine expansions [33], but also point mutations [35], are responsible for SCA6. It was long debated whether the phenotype was due to a toxic effect of the polyglutamine stretch or to an altered channel activity [63, 64]. The pacemaking activity of Purkinje cells (PC) was shown to be irregular in a mouse with spontaneous Cacna1a mutation [65]. However, the link between irregular firing and symptoms severity is not linear [66]. A second cistron encoded by CACNA1A was recently described, with a transcription factor role on genes involved in neurite outgrowth of PC, which is altered with a 33 glutamine expansion [67]. The pathophysiology of SCA6 might, therefore, involve multiple pathways.
Other genes encoding channels have recently been involved in SCA pathology, namely KCNC3/SCA13 and KCND3/SCA19/SCA22, in which the pathogenicity result from loss of function. The first human mutations in KCNC3 were associated with channel malfunction in Xenopus oocytes [68]. Recently, loss-of-function mechanisms were confirmed in mammalian cells, through reduced surface expression, shorter half-life [69], or Golgi retention and malformation for the p.R420H mutation [70]. The loss of function associated with KCND3 mutations seems to occur through impaired membrane trafficking (p.F227del [37]) or endoplasmic reticulum retention (p.T352P, p.M373I, p.S390N [36]). While coexpressed with their regulatory beta subunit Kv channel-interacting protein 2, the membrane localization was restored for all but p.S390N mutant. However, patch-clamp studies proved that these complexes were either not or less functional than with the wild-type protein.
The product of FGF14, whose mutations cause SCA27 [71], was shown to interact with the pore-forming subunits of Na+ channels [72]. In hippocampal neurons, the p.F145S mutant exerts a dominant negative effect on the wild-type protein interaction with the channel, decreasing the neurons’ excitability [72], reminiscent of the attenuated spontaneous firing of Fgf14−/− PC [73]. In granule cells, both FGF14 knock-down and overexpression of the mutant protein were recently shown to reduce Ca2+ currents, alter vesicular recycling, and, consequently, lower excitatory post-synaptic currents to the PC [74].
In the family of ligand-gated ion channels, only ITPR1/SCA15 has so far been implicated in humans [61]. Of interest is the recent involvement of GRID2 biallelic loss-of-function mutations in AR-HCAs [22–24]. In mouse, the Lurcher gain-of-function point mutation in Grid2 is responsible for a spontaneous cation leak, inducing PC death [26]; the same mutation, and others nearby, were found in patients with a semi-dominant inheritance pattern of either late-onset or congenital HCA [25].
More on mitochondrial dysfunction
Mitochondrial dysfunction has long been implicated in HCAs, with involvement of mutations in mitochondrial DNA, deficiency of coenzyme Q10 production, bioenergetic alterations (POLG, C10orf2), dysregulation of mitochondrial fusion–fission balance, and apoptosis (MFN2, OPA1, SACS, PPP2R2B/SCA12) [61]. Recent developments in the field include the growing list of genes inducing mitochondrial complex deficiency and cerebellar involvement (TTC19 and complex III [75]; COX20 and complex IV [27, 28]). Also of interest is the growing importance of protein quality control, with accumulating evidence of functional effects of AFG3L2 alterations (SCA28), and cerebellar symptoms in SPG7 patients. The mitochondrial inner membrane protein encoded by AFG3L2 belongs to the family of m-AAA proteases, either in homo-oligomeric or in hetero-oligomeric complexes with paraplegin, encoded by SPG7 [76]. Knock-out of Afg3l2 induces mitochondrial network fragmentation and reduced mitochondrial uptake of Ca++ in mouse embryonic fibroblasts [77]. In PC, early abnormal mitochondrial dynamics, respiratory dysfunction and neurodegeneration are imputed to decreased mitochondrial protein synthesis [78]. Depletion in Afg3l2 is also responsible for mitochondria anterograde transport defect [79]. In patients with SPG7 mutations, whose clinical picture includes cerebellar ataxia, skeletal muscle cells showed multiple mitochondrial DNA deletions and respiratory chain deficiencies in complexes I, III and IV [80]. In three patients with heterozygous SPG7 mutations, cerebellar signs and atrophy were described in the absence of spasticity [81].
Glutamate transmission
Glutamate transmission is also of increasing importance in HCAs. Involvement of GRID2, encoding the glutamate receptor ∂2 protein (GluRD2), which belongs to the family of ionotropic glutamate receptors, has already been mentioned above. STUB1-encoded protein has been shown, in combination with Fbx2, to promote the ubiquitination and subsequent degradation of the NR2A subunit of N-methyl-d-aspartate receptors [82], an ability that was impaired in the mutant proteins [13]. SPTBN2 protein product, whose mutations account for SCA5, stabilises the glutamate transporter EAAT4 at the cell membrane [83]; in SCA5 cerebellum extracts, EAAT4 levels were decreased, as was previously reported in SCA23 [84], and in SCA1 transgenic mouse [85]. GluRD2 membrane levels were also lower than in controls [83]. Recently, metabotropic glutamate receptor 1α (mGluR1α) was shown to have reduced localization at dendritic spines and decreased function in a SCA5 mouse model [86]. GluRD2, mGluR1 and PKCγ, whose mutations are responsible for SCA14, were all shown to interact in regulating synaptic transmission in PC [87].
Lipids biosynthesis as a growing pathway in HCAs
Lipids biosynthesis is a newly recognised pathway of importance in HCA pathology. It has previously been implicated in hereditary spastic paraplegia, with the description of mutations in CYP7B1/SPG5A, FA2H/SPG35, DDHD2/SPG54, DDHD1/SPG28 [88], CYP2U1/SPG56 [89], B4GALNT1/SPG26 [90], and GBA2/SPG46 [91]. Nonsense GBA2 mutations were shown responsible for a spastic ataxia phenotype, arguing for a loss-of-function mechanism, inducing glucosylceramide accumulation in the ER in brain and testis of Gba2-KO mice [92]. PNPLA6, which encodes a bifunctional enzyme with a role in fatty acids and glycerophosphocholine synthesis, and in 2-arachidonoyl lysophosphatidylinositol catalyzation, has been mentioned above [42, 93]. Mutations in PLA2G6, encoding phospholipase A2, account for a large phenotypic spectrum, but childhood-onset cerebellar ataxia is a core element [45]. Finally, ELOVL4 and ELOVL5, which code for elongases of very long chain fatty acids were, respectively, implicated in SCA34 and SCA38 [10, 94].
Aggregation processes strike again
Protein aggregation is a long-known hallmark of multiple neurodegenerative diseases, though its role in degeneration is still debated. In polyglutamine expansions, aggregates are mostly found in the nucleus [61]. Alongside these well-described inclusions, amyloidogenic processes, which are traditionally associated with Alzheimer’s disease, are increasingly being implicated in cerebellar malfunction. Stop mutations in ITM2B have long been recognised to account for British and Danish familial dementias, through the accumulation of a newly synthesised amyloidogenic protein [95, 96]. The phenotype, however, is not circumscribed to CA, but also includes dementia and spastic paraplegia. More strikingly, PKCγ was recently established as an amyloidogenic protein. SCA14-linked mutations seem to accelerate the amyloid-like fibril formation [97].
Implications in clinical practice
As previously mentioned, the extreme intricacy in HCAs genetics makes it practically impossible to Sanger sequence all implicated genes one by one. Nowadays, the prominent technique for molecular diagnosis would either be TS [8] or WES [98], which allow massive parallel sequencing of many or all of the HCA-implicated genes. TS provides optimised coverage of targeted regions, and is still more cost-effective than WES. On the other hand, as new genes are described on a constant basis, the list of sequenced ones becomes obsolete even before the sequencing run is launched. Whole-genome sequencing technically overcomes pitfalls of both techniques, with more harmonious coverage, higher rate of mutation detection, and non-exonic DNA sequencing, allowing molecular diagnosis in previously unsolved case [99]; however, costs in sequencing and bioinformatics processing still postpone its wide-scale use in diagnosis. In all cases, as the phenotypic spectrum associated with a specific gene broadens, and new types of mutations or transmission modes are described, an open mind should be kept in deciphering the pathogenicity of newly found mutations. However, caution should be taken as not all mutations are disease-causing. Converging evidences of the mutation causality, including database frequency, conservation or pathogenicity scores, and segregation within the family, are necessary, but not always sufficient, to establish a diagnosis. Functional testing of every variant is not yet conceivable in clinical practice. As to overcome the lack of convincing clues, efforts might be invested to set up publicly available locus-specific databases [100, 101]. Even if challenged, genotype–phenotype correlations are still a major element in the causality sentence for a given variant.
Conclusion
The heterogeneous genetic landscape of HCAs raises challenges regarding the molecular diagnosis. Not all genes are known as yet, and new ones are constantly being discovered, either as novel genes involved in human diseases or known genes newly implicated in HCAs. Moreover, the phenotypic spectrum of known genes is widely extending along with the use of next-generation sequencing as a diagnostic tool. Novel types of mutations or transmission modes for well-described genes are discovered. The golden age of strict genotype–phenotype correlations seems to have receded with the advent of new technologies. However, caution must be exercised when seeking to interpret new variants in known genes, especially when they are associated with new phenotypes, as many are not disease-causing. Efforts should be directed towards describing the core phenotypes for each gene, but in many cases, none of the clinical signs is mandatory. Therefore, the trend for genetic diagnosis in clinical practice is to use TS or WES.
In the latter case, the interpretation of results can also be challenging. In this respect, the classification of genes amongst the common networks is crucial to identify causative genes in undiagnosed patients. However, in the absence of sufficient arguments to validate new genes (lack of other affected cases to enable an analysis of co-segregation, absence of additional families), the proof of involvement is often difficult to achieve. Conversely, the description of multiple genes in a common pathway is essential for the pathophysiological understanding of HCAs, and for the future development of common therapeutic targets. We must stress the importance of genetic followed by functional studies (cell biology, biochemistry, animal models, etc.) to validate new genes, elucidate the mechanisms underlying HCAs and decipher new treatment approaches.
References
Harding AE (1983) Classification of the hereditary ataxias and paraplegias. Lancet 1(8334):1151–1155 (S0140-6736(83)90673-6)
Anheim M, Tranchant C, Koenig M (2012) The autosomal recessive cerebellar ataxias. N Engl J Med 366(7):636–646. doi:10.1056/NEJMra1006610
Durr A (2010) Autosomal dominant cerebellar ataxias: polyglutamine expansions and beyond. Lancet Neurol 9(9):885–894. doi:10.1016/S1474-4422(10)70183-6
Allikmets R, Raskind WH, Hutchinson A, Schueck ND, Dean M, Koeller DM (1999) Mutation of a putative mitochondrial iron transporter gene (ABC7) in X-linked sideroblastic anemia and ataxia (XLSA/A). Hum Mol Genet 8(5):743–749 (ddc101 [pii])
Gilfillan GD, Selmer KK, Roxrud I, Smith R, Kyllerman M, Eiklid K, Kroken M, Mattingsdal M, Egeland T, Stenmark H, Sjoholm H, Server A, Samuelsson L, Christianson A, Tarpey P, Whibley A, Stratton MR, Futreal PA, Teague J, Edkins S, Gecz J, Turner G, Raymond FL, Schwartz C, Stevenson RE, Undlien DE, Stromme P (2008) SLC9A6 mutations cause X-linked mental retardation, microcephaly, epilepsy, and ataxia, a phenotype mimicking Angelman syndrome. Am J Hum Genet 82(4):1003–1010. doi:10.1016/j.ajhg.2008.01.013
Roessler BJ, Golovoy N, Palella TD, Heidler S, Becker MA (1991) Identification of distinct PRS1 mutations in two patients with X-linked phosphoribosylpyrophosphate synthetase superactivity. Adv Exp Med Biol 309B:125–128
Hagerman PJ, Hagerman RJ (2004) The fragile-X premutation: a maturing perspective. Am J Hum Genet 74(5):805–816. doi:10.1086/386296
Nemeth AH, Kwasniewska AC, Lise S, Parolin Schnekenberg R, Becker EB, Bera KD, Shanks ME, Gregory L, Buck D, Zameel Cader M, Talbot K, de Silva R, Fletcher N, Hastings R, Jayawant S, Morrison PJ, Worth P, Taylor M, Tolmie J, O’Regan M, Valentine R, Packham E, Evans J, Seller A, Ragoussis J (2013) Next generation sequencing for molecular diagnosis of neurological disorders using ataxias as a model. Brain 136(Pt 10):3106–3118. doi:10.1093/brain/awt236
Hekman KE, Yu GY, Brown CD, Zhu H, Du X, Gervin K, Undlien DE, Peterson A, Stevanin G, Clark HB, Pulst SM, Bird TD, White KP, Gomez CM (2012) A conserved eEF2 coding variant in SCA26 leads to loss of translational fidelity and increased susceptibility to proteostatic insult. Hum Mol Genet 21(26):5472–5483. doi:10.1093/hmg/dds392
Di Gregorio E, Borroni B, Giorgio E, Lacerenza D, Ferrero M, Lo Buono N, Ragusa N, Mancini C, Gaussen M, Calcia A, Mitro N, Hoxha E, Mura I, Coviello DA, Moon YA, Tesson C, Vaula G, Couarch P, Orsi L, Duregon E, Papotti MG, Deleuze JF, Imbert J, Costanzi C, Padovani A, Giunti P, Maillet-Vioud M, Durr A, Brice A, Tempia F, Funaro A, Boccone L, Caruso D, Stevanin G, Brusco A (2014) ELOVL5 mutations cause spinocerebellar ataxia 38. Am J Hum Genet 95(2):209–217. doi:10.1016/j.ajhg.2014.07.001
Delplanque J, Devos D, Huin V, Genet A, Sand O, Moreau C, Goizet C, Charles P, Anheim M, Monin ML, Buee L, Destee A, Grolez G, Delmaire C, Dujardin K, Dellacherie D, Brice A, Stevanin G, Strubi-Vuillaume I, Durr A, Sablonniere B (2014) TMEM240 mutations cause spinocerebellar ataxia 21 with mental retardation and severe cognitive impairment. Brain 137(Pt 10):2657–2663. doi:10.1093/brain/awu202
Russell JF, Steckley JL, Coppola G, Hahn AF, Howard MA, Kornberg Z, Huang A, Mirsattari SM, Merriman B, Klein E, Choi M, Lee HY, Kirk A, Nelson-Williams C, Gibson G, Baraban SC, Lifton RP, Geschwind DH, Fu YH, Ptacek LJ (2012) Familial cortical myoclonus with a mutation in NOL3. Ann Neurol 72(2):175–183. doi:10.1002/ana.23666
Shi Y, Wang J, Li JD, Ren H, Guan W, He M, Yan W, Zhou Y, Hu Z, Zhang J, Xiao J, Su Z, Dai M, Jiang H, Guo J, Zhang F, Li N, Du J, Xu Q, Hu Y, Pan Q, Shen L, Wang G, Xia K, Zhang Z, Tang B (2013) Identification of CHIP as a novel causative gene for autosomal recessive cerebellar ataxia. PLoS One 8(12):e81884. doi:10.1371/journal.pone.0081884
Shi CH, Schisler JC, Rubel CE, Tan S, Song B, McDonough H, Xu L, Portbury AL, Mao CY, True C, Wang RH, Wang QZ, Sun SL, Seminara SB, Patterson C, Xu YM (2014) Ataxia and hypogonadism caused by the loss of ubiquitin ligase activity of the U box protein CHIP. Hum Mol Genet 23(4):1013–1024. doi:10.1093/hmg/ddt497
Synofzik M, Schule R, Schulze M, Gburek-Augustat J, Schweizer R, Schirmacher A, Krageloh-Mann I, Gonzalez M, Young P, Zuchner S, Schols L, Bauer P (2014) Phenotype and frequency of STUB1 mutations: next-generation screenings in Caucasian ataxia and spastic paraplegia cohorts. Orphanet J Rare Dis 9:57. doi:10.1186/1750-1172-9-57
Margolin DH, Kousi M, Chan YM, Lim ET, Schmahmann JD, Hadjivassiliou M, Hall JE, Adam I, Dwyer A, Plummer L, Aldrin SV, O’Rourke J, Kirby A, Lage K, Milunsky A, Milunsky JM, Chan J, Hedley-Whyte ET, Daly MJ, Katsanis N, Seminara SB (2013) Ataxia, dementia, and hypogonadotropism caused by disordered ubiquitination. N Engl J Med 368(21):1992–2003. doi:10.1056/NEJMoa1215993
Ronnebaum SM, Patterson C, Schisler JC (2014) Emerging evidence of coding mutations in the ubiquitin-proteasome system associated with cerebellar ataxias. Hum Genome Var 1 (14018). doi:10.1038/hgv.2014.18
Bilguvar K, Tyagi NK, Ozkara C, Tuysuz B, Bakircioglu M, Choi M, Delil S, Caglayan AO, Baranoski JF, Erturk O, Yalcinkaya C, Karacorlu M, Dincer A, Johnson MH, Mane S, Chandra SS, Louvi A, Boggon TJ, Lifton RP, Horwich AL, Gunel M (2013) Recessive loss of function of the neuronal ubiquitin hydrolase UCHL1 leads to early-onset progressive neurodegeneration. Proc Natl Acad Sci USA 110(9):3489–3494. doi:10.1073/pnas.1222732110
Conroy J, McGettigan P, Murphy R, Webb D, Murphy SM, McCoy B, Albertyn C, McCreary D, McDonagh C, Walsh O, Lynch S, Ennis S (2014) A novel locus for episodic ataxia:UBR4 the likely candidate. Eur J Hum Genet 22(4):505–510. doi:10.1038/ejhg.2013.173
Mallaret M, Synofzik M, Lee J, Sagum CA, Mahajnah M, Sharkia R, Drouot N, Renaud M, Klein FA, Anheim M, Tranchant C, Mignot C, Mandel JL, Bedford M, Bauer P, Salih MA, Schule R, Schols L, Aldaz CM, Koenig M (2014) The tumour suppressor gene WWOX is mutated in autosomal recessive cerebellar ataxia with epilepsy and mental retardation. Brain 137(Pt 2):411–419. doi:10.1093/brain/awt338
Guissart C, Li X, Leheup B, Drouot N, Montaut-Verient B, Raffo E, Jonveaux P, Roux AF, Claustres M, Fliegel L, Koenig M (2014) Mutation of SLC9A1, encoding the major Na+/H+ exchanger, causes ataxia-deafness Lichtenstein-Knorr syndrome. Hum Mol Genet. doi:10.1093/hmg/ddu461
Hills LB, Masri A, Konno K, Kakegawa W, Lam AT, Lim-Melia E, Chandy N, Hill RS, Partlow JN, Al-Saffar M, Nasir R, Stoler JM, Barkovich AJ, Watanabe M, Yuzaki M, Mochida GH (2013) Deletions in GRID2 lead to a recessive syndrome of cerebellar ataxia and tonic upgaze in humans. Neurology 81(16):1378–1386. doi:10.1212/WNL.0b013e3182a841a3
Utine GE, Haliloglu G, Salanci B, Cetinkaya A, Kiper PO, Alanay Y, Aktas D, Boduroglu K, Alikasifoglu M (2013) A homozygous deletion in GRID2 causes a human phenotype with cerebellar ataxia and atrophy. J Child Neurol 28(7):926–932. doi:10.1177/0883073813484967
Van Schil K, Meire F, Karlstetter M, Bauwens M, Verdin H, Coppieters F, Scheiffert E, Van Nechel C, Langmann T, Deconinck N, De Baere E (2014) Early-onset autosomal recessive cerebellar ataxia associated with retinal dystrophy: new human hotfoot phenotype caused by homozygous GRID2 deletion. Genet Med. doi:10.1038/gim.2014.95
Coutelier M, Burglen L, Mundwiller E, Abada-Bendib M, Rodriguez D, Chantot-Bastaraud S, Rougeot C, Cournelle M-A, Milh M, Toutain A, Bacq D, Meyer V, Afenjar A, Deleuze J-F, Brice A, Héron D, Stevanin G, Durr A (2015) GRID2 mutations span from congenital to mild adult onset cerebellar ataxia. Neurology 84. doi:10.1212/WNL.0000000000001524
Zuo J, De Jager PL, Takahashi KA, Jiang W, Linden DJ, Heintz N (1997) Neurodegeneration in Lurcher mice caused by mutation in delta2 glutamate receptor gene. Nature 388(6644):769–773. doi:10.1038/42009
Szklarczyk R, Wanschers BF, Nijtmans LG, Rodenburg RJ, Zschocke J, Dikow N, van den Brand MA, Hendriks-Franssen MG, Gilissen C, Veltman JA, Nooteboom M, Koopman WJ, Willems PH, Smeitink JA, Huynen MA, van den Heuvel LP (2013) A mutation in the FAM36A gene, the human ortholog of COX20, impairs cytochrome c oxidase assembly and is associated with ataxia and muscle hypotonia. Hum Mol Genet 22(4):656–667. doi:10.1093/hmg/dds473
Doss S, Lohmann K, Seibler P, Arns B, Klopstock T, Zuhlke C, Freimann K, Winkler S, Lohnau T, Drungowski M, Nurnberg P, Wiegers K, Lohmann E, Naz S, Kasten M, Bohner G, Ramirez A, Endres M, Klein C (2014) Recessive dystonia-ataxia syndrome in a Turkish family caused by a COX20 (FAM36A) mutation. J Neurol 261(1):207–212. doi:10.1007/s00415-013-7177-7
Bourassa CV, Meijer IA, Merner ND, Grewal KK, Stefanelli MG, Hodgkinson K, Ives EJ, Pryse-Phillips W, Jog M, Boycott K, Grimes DA, Goobie S, Leckey R, Dion PA, Rouleau GA (2012) VAMP1 mutation causes dominant hereditary spastic ataxia in Newfoundland families. Am J Hum Genet 91(3):548–552. doi:10.1016/j.ajhg.2012.07.018
Shen XM, Selcen D, Brengman J, Engel AG (2014) Mutant SNAP25B causes myasthenia, cortical hyperexcitability, ataxia, and intellectual disability. Neurology. doi:10.1212/WNL.0000000000001079
Thomas A, Williams H, Seto-Salvia N, Bacchelli C, Jenkins D, O’Sullivan M, Mengrelis K, Ishida M, Ocaka L, Chanudet E, James C, Lescai F, Anderson G, Morrogh D, Ryten M, Duncan A, Jin Pai Y, Saraiva J, Ramos F, Farren B, Saunders D, Vernay B, Gissen P, Straatman-Iwanowska A, Baas F, Wood N, Hersheson J, Houlden H, Hurst J, Scott R, Bitner-Glinzicz M, Moore G, Sousa S, Stanier P (2014) Mutations in SNX14 cause a distinctive autosomal-recessive cerebellar ataxia and intellectual disability syndrome. Am J Hum Genet 95(5):611–621
Ophoff RA, Terwindt GM, Vergouwe MN, van Eijk R, Oefner PJ, Hoffman SM, Lamerdin JE, Mohrenweiser HW, Bulman DE, Ferrari M, Haan J, Lindhout D, van Ommen GJ, Hofker MH, Ferrari MD, Frants RR (1996) Familial hemiplegic migraine and episodic ataxia type-2 are caused by mutations in the Ca2+ channel gene CACNL1A4. Cell 87(3):543–552 (S0092-8674(00)81373-2 [pii])
Zhuchenko O, Bailey J, Bonnen P, Ashizawa T, Stockton DW, Amos C, Dobyns WB, Subramony SH, Zoghbi HY, Lee CC (1997) Autosomal dominant cerebellar ataxia (SCA6) associated with small polyglutamine expansions in the alpha 1A-voltage-dependent calcium channel. Nat Genet 15(1):62–69. doi:10.1038/ng0197-62
Jodice C, Mantuano E, Veneziano L, Trettel F, Sabbadini G, Calandriello L, Francia A, Spadaro M, Pierelli F, Salvi F, Ophoff RA, Frants RR, Frontali M (1997) Episodic ataxia type 2 (EA2) and spinocerebellar ataxia type 6 (SCA6) due to CAG repeat expansion in the CACNA1A gene on chromosome 19p. Hum Mol Genet 6(11):1973–1978 (dda250 [pii])
Yue Q, Jen JC, Nelson SF, Baloh RW (1997) Progressive ataxia due to a missense mutation in a calcium-channel gene. Am J Hum Genet 61(5):1078–1087. doi:10.1086/301613
Duarri A, Jezierska J, Fokkens M, Meijer M, Schelhaas HJ, den Dunnen WF, van Dijk F, Verschuuren-Bemelmans C, Hageman G, van de Vlies P, Kusters B, van de Warrenburg BP, Kremer B, Wijmenga C, Sinke RJ, Swertz MA, Kampinga HH, Boddeke E, Verbeek DS (2012) Mutations in potassium channel kcnd3 cause spinocerebellar ataxia type 19. Ann Neurol 72(6):870–880. doi:10.1002/ana.23700
Lee YC, Durr A, Majczenko K, Huang YH, Liu YC, Lien CC, Tsai PC, Ichikawa Y, Goto J, Monin ML, Li JZ, Chung MY, Mundwiller E, Shakkottai V, Liu TT, Tesson C, Lu YC, Brice A, Tsuji S, Burmeister M, Stevanin G, Soong BW (2012) Mutations in KCND3 cause spinocerebellar ataxia type 22. Ann Neurol 72(6):859–869. doi:10.1002/ana.23701
Meneret A, Ahmar-Beaugendre Y, Rieunier G, Mahlaoui N, Gaymard B, Apartis E, Tranchant C, Rivaud-Pechoux S, Degos B, Benyahia B, Suarez F, Maisonobe T, Koenig M, Durr A, Stern MH, Dubois d’Enghien C, Fischer A, Vidailhet M, Stoppa-Lyonnet D, Grabli D, Anheim M (2014) The pleiotropic movement disorders phenotype of adult ataxia-telangiectasia. Neurology 83(12):1087–1095. doi:10.1212/WNL.0000000000000794
Pan LL, Huang YM, Wang M, Zhuang XE, Luo DF, Guo SC, Zhang ZS, Huang Q, Lin SL, Wang SY (2014) Positional cloning and next-generation sequencing identified a TGM6 mutation in a large Chinese pedigree with acute myeloid leukaemia. Eur J Hum Genet. doi:10.1038/ejhg.2014.67
Sun Y, Almomani R, Breedveld GJ, Santen GW, Aten E, Lefeber DJ, Hoff JI, Brusse E, Verheijen FW, Verdijk RM, Kriek M, Oostra B, Breuning MH, Losekoot M, den Dunnen JT, van de Warrenburg BP, Maat-Kievit AJ (2013) Autosomal recessive spinocerebellar ataxia 7 (SCAR7) is caused by variants in TPP1, the gene involved in classic late-infantile neuronal ceroid lipofuscinosis 2 disease (CLN2 disease). Hum Mutat 34(5):706–713. doi:10.1002/humu.22292
Mancini C, Nassani S, Guo Y, Chen Y, Giorgio E, Brussino A, Di Gregorio E, Cavalieri S, Lo Buono N, Funaro A, Pizio NR, Nmezi B, Kyttala A, Santorelli FM, Padiath QS, Hakonarson H, Zhang H, Brusco A (2014) Adult-onset autosomal recessive ataxia associated with neuronal ceroid lipofuscinosis type 5 gene (CLN5) mutations. J Neurol. doi:10.1007/s00415-014-7553-y
Synofzik M, Gonzalez MA, Lourenco CM, Coutelier M, Haack TB, Rebelo A, Hannequin D, Strom TM, Prokisch H, Kernstock C, Durr A, Schols L, Lima-Martinez MM, Farooq A, Schule R, Stevanin G, Marques W Jr, Zuchner S (2014) PNPLA6 mutations cause Boucher-Neuhauser and Gordon Holmes syndromes as part of a broad neurodegenerative spectrum. Brain 137(Pt 1):69–77. doi:10.1093/brain/awt326
Hufnagel RB, Arno G, Hein ND, Hersheson J, Prasad M, Anderson Y, Krueger LA, Gregory LC, Stoetzel C, Jaworek TJ, Hull S, Li A, Plagnol V, Willen CM, Morgan TM, Prows CA, Hegde RS, Riazuddin S, Grabowski GA, Richardson RJ, Dieterich K, Huang T, Revesz T, Martinez-Barbera JP, Sisk RA, Jefferies C, Houlden H, Dattani MT, Fink JK, Dollfus H, Moore AT, Ahmed ZM (2014) Neuropathy target esterase impairments cause Oliver-McFarlane and Laurence-Moon syndromes. J Med Genet. doi:10.1136/jmedgenet-2014-102856
Renaud M, Anheim M, Kamsteeg EJ, Mallaret M, Mochel F, Vermeer S, Drouot N, Pouget J, Redin C, Salort-Campana E, Kremer HP, Verschuuren-Bemelmans CC, Muller J, Scheffer H, Durr A, Tranchant C, Koenig M (2014) Autosomal recessive cerebellar ataxia type 3 due to ANO10 mutations: delineation and genotype–phenotype correlation study. JAMA Neurol 71(10):1305–1310. doi:10.1001/jamaneurol.2014.193
Salih MA, Mundwiller E, Khan AO, AlDrees A, Elmalik SA, Hassan HH, Al-Owain M, Alkhalidi HM, Katona I, Kabiraj MM, Chrast R, Kentab AY, Alzaidan H, Rodenburg RJ, Bosley TM, Weis J, Koenig M, Stevanin G, Azzedine H (2013) New findings in a global approach to dissect the whole phenotype of PLA2G6 gene mutations. PLoS One 8(10):e76831. doi:10.1371/journal.pone.0076831
Hara K, Shiga A, Nozaki H, Mitsui J, Takahashi Y, Ishiguro H, Yomono H, Kurisaki H, Goto J, Ikeuchi T, Tsuji S, Nishizawa M, Onodera O (2008) Total deletion and a missense mutation of ITPR1 in Japanese SCA15 families. Neurology 71(8):547–551. doi:10.1212/01.wnl.0000311277.71046.a0
Huang L, Chardon JW, Carter MT, Friend KL, Dudding TE, Schwartzentruber J, Zou R, Schofield PW, Douglas S, Bulman DE, Boycott KM (2012) Missense mutations in ITPR1 cause autosomal dominant congenital nonprogressive spinocerebellar ataxia. Orphanet J Rare Dis 7:67. doi:10.1186/1750-1172-7-67
Ohba C, Osaka H, Iai M, Yamashita S, Suzuki Y, Aida N, Shimozawa N, Takamura A, Doi H, Tomita-Katsumoto A, Nishiyama K, Tsurusaki Y, Nakashima M, Miyake N, Eto Y, Tanaka F, Matsumoto N, Saitsu H (2013) Diagnostic utility of whole exome sequencing in patients showing cerebellar and/or vermis atrophy in childhood. Neurogenetics 14(3–4):225–232. doi:10.1007/s10048-013-0375-8
Di Bella D, Lazzaro F, Brusco A, Plumari M, Battaglia G, Pastore A, Finardi A, Cagnoli C, Tempia F, Frontali M, Veneziano L, Sacco T, Boda E, Brussino A, Bonn F, Castellotti B, Baratta S, Mariotti C, Gellera C, Fracasso V, Magri S, Langer T, Plevani P, Di Donato S, Muzi-Falconi M, Taroni F (2010) Mutations in the mitochondrial protease gene AFG3L2 cause dominant hereditary ataxia SCA28. Nat Genet 42(4):313–321. doi:10.1038/ng.544
Smets K, Deconinck T, Baets J, Sieben A, Martin JJ, Smouts I, Wang S, Taroni F, Di Bella D, Van Hecke W, Parizel PM, Jadoul C, De Potter R, Couvreur F, Rugarli E, De Jonghe P (2014) Partial deletion of AFG3L2 causing spinocerebellar ataxia type 28. Neurology 82(23):2092–2100. doi:10.1212/WNL.0000000000000491
Myers KA, Warman Chardon J, Huang L, Boycott KM (2014) Deletion of AFG3L2 associated with spinocerebellar ataxia type 28 in the context of multiple genomic anomalies. Am J Med Genet A. doi:10.1002/ajmg.a.36771
Musova Z, Kaiserova M, Kriegova E, Fillerova R, Vasovcak P, Santava A, Mensikova K, Zumrova A, Krepelova A, Sedlacek Z, Kanovsky P (2014) A novel frameshift mutation in the AFG3L2 gene in a patient with spinocerebellar ataxia. Cerebellum 13(3):331–337. doi:10.1007/s12311-013-0538-z
Pierson TM, Adams D, Bonn F, Martinelli P, Cherukuri PF, Teer JK, Hansen NF, Cruz P, Mullikin For The Nisc Comparative Sequencing Program JC, Blakesley RW, Golas G, Kwan J, Sandler A, Fuentes Fajardo K, Markello T, Tifft C, Blackstone C, Rugarli EI, Langer T, Gahl WA, Toro C (2011) Whole-exome sequencing identifies homozygous AFG3L2 mutations in a spastic ataxia-neuropathy syndrome linked to mitochondrial m-AAA proteases. PLoS Genet 7(10):e1002325
Lise S, Clarkson Y, Perkins E, Kwasniewska A, Sadighi Akha E, Schnekenberg RP, Suminaite D, Hope J, Baker I, Gregory L, Green A, Allan C, Lamble S, Jayawant S, Quaghebeur G, Cader MZ, Hughes S, Armstrong RJ, Kanapin A, Rimmer A, Lunter G, Mathieson I, Cazier JB, Buck D, Taylor JC, Bentley D, McVean G, Donnelly P, Knight SJ, Jackson M, Ragoussis J, Nemeth AH (2012) Recessive mutations in SPTBN2 implicate beta-III spectrin in both cognitive and motor development. PLoS Genet 8(12):e1003074. doi:10.1371/journal.pgen.1003074
Elsayed SM, Heller R, Thoenes M, Zaki MS, Swan D, Elsobky E, Zuhlke C, Ebermann I, Nurnberg G, Nurnberg P, Bolz HJ (2014) Autosomal dominant SCA5 and autosomal recessive infantile SCA are allelic conditions resulting from SPTBN2 mutations. Eur J Hum Genet 22(2):286–288. doi:10.1038/ejhg.2013.150
Stevanin G, Durr A, Brice A (2000) Clinical and molecular advances in autosomal dominant cerebellar ataxias: from genotype to phenotype and physiopathology. Eur J Hum Genet 8(1):4–18. doi:10.1038/sj.ejhg.5200403
Tezenas du Montcel S, Durr A, Bauer P, Figueroa KP, Ichikawa Y, Brussino A, Forlani S, Rakowicz M, Schols L, Mariotti C, van de Warrenburg BP, Orsi L, Giunti P, Filla A, Szymanski S, Klockgether T, Berciano J, Pandolfo M, Boesch S, Melegh B, Timmann D, Mandich P, Camuzat A, Goto J, Ashizawa T, Cazeneuve C, Tsuji S, Pulst SM, Brusco A, Riess O, Brice A, Stevanin G (2014) Modulation of the age at onset in spinocerebellar ataxia by CAG tracts in various genes. Brain 137(Pt 9):2444–2455. doi:10.1093/brain/awu174
Lattante S, Millecamps S, Stevanin G, Rivaud-Pechoux S, Moigneu C, Camuzat A, Da Barroca S, Mundwiller E, Couarch P, Salachas F, Hannequin D, Meininger V, Pasquier F, Seilhean D, Couratier P, Danel-Brunaud V, Bonnet AM, Tranchant C, LeGuern E, Brice A, Le Ber I, Kabashi E (2014) Contribution of ATXN2 intermediary polyQ expansions in a spectrum of neurodegenerative disorders. Neurology 83(11):990–995. doi:10.1212/WNL.0000000000000778
Yamashita C, Tomiyama H, Funayama M, Inamizu S, Ando M, Li Y, Yoshino H, Araki T, Ichikawa T, Ehara Y, Ishikawa K, Mizusawa H, Hattori N (2014) The evaluation of polyglutamine repeats in autosomal dominant Parkinson’s disease. Neurobiol Aging 35 (7):1779 e1717-1721. doi:10.1016/j.neurobiolaging.2014.01.022
Bonifert T, Karle KN, Tonagel F, Batra M, Wilhelm C, Theurer Y, Schoenfeld C, Kluba T, Kamenisch Y, Carelli V, Wolf J, Gonzalez MA, Speziani F, Schule R, Zuchner S, Schols L, Wissinger B, Synofzik M (2014) Pure and syndromic optic atrophy explained by deep intronic OPA1 mutations and an intralocus modifier. Brain 137(Pt 8):2164–2177. doi:10.1093/brain/awu165
Matilla-Duenas A, Ashizawa T, Brice A, Magri S, McFarland KN, Pandolfo M, Pulst SM, Riess O, Rubinsztein DC, Schmidt J, Schmidt T, Scoles DR, Stevanin G, Taroni F, Underwood BR, Sanchez I (2014) Consensus paper: pathological mechanisms underlying neurodegeneration in spinocerebellar ataxias. Cerebellum 13(2):269–302. doi:10.1007/s12311-013-0539-y
Alves S, Cormier-Dequaire F, Marinello M, Marais T, Muriel MP, Beaumatin F, Charbonnier-Beaupel F, Tahiri K, Seilhean D, El Hachimi K, Ruberg M, Stevanin G, Barkats M, den Dunnen W, Priault M, Brice A, Durr A, Corvol JC, Sittler A (2014) The autophagy/lysosome pathway is impaired in SCA7 patients and SCA7 knock-in mice. Acta Neuropathol 128(5):705–722. doi:10.1007/s00401-014-1289-8
Frontali M (2001) Spinocerebellar ataxia type 6: channelopathy or glutamine repeat disorder? Brain Res Bull 56(3–4):227–231 (S0361-9230(01)00574-3 [pii])
Rajakulendran S, Kaski D, Hanna MG (2012) Neuronal P/Q-type calcium channel dysfunction in inherited disorders of the CNS. Nat Rev Neurol 8(2):86–96. doi:10.1038/nrneurol.2011.228
Walter JT, Alvina K, Womack MD, Chevez C, Khodakhah K (2006) Decreases in the precision of Purkinje cell pacemaking cause cerebellar dysfunction and ataxia. Nat Neurosci 9(3):389–397. doi:10.1038/nn1648
Stahl JS, Thumser ZC (2014) Flocculus Purkinje cell signals in mouse Cacna1a calcium channel mutants of escalating severity: an investigation of the role of firing irregularity in ataxia. J Neurophysiol. doi:10.1152/jn.00129.2014
Du X, Wang J, Zhu H, Rinaldo L, Lamar KM, Palmenberg AC, Hansel C, Gomez CM (2013) Second cistron in CACNA1A gene encodes a transcription factor mediating cerebellar development and SCA6. Cell 154(1):118–133. doi:10.1016/j.cell.2013.05.059
Waters MF, Minassian NA, Stevanin G, Figueroa KP, Bannister JP, Nolte D, Mock AF, Evidente VG, Fee DB, Muller U, Durr A, Brice A, Papazian DM, Pulst SM (2006) Mutations in voltage-gated potassium channel KCNC3 cause degenerative and developmental central nervous system phenotypes. Nat Genet 38(4):447–451. doi:10.1038/ng1758
Zhao J, Zhu J, Thornhill WB (2013) Spinocerebellar ataxia-13 Kv3.3 potassium channels: arginine-to-histidine mutations affect both functional and protein expression on the cell surface. Biochem J 454(2):259–265. doi:10.1042/BJ20130034
Gallego-Iradi C, Bickford JS, Khare S, Hall A, Nick JA, Salmasinia D, Wawrowsky K, Bannykh S, Huynh DP, Rincon-Limas DE, Pulst SM, Nick HS, Fernandez-Funez P, Waters MF (2014) KCNC3(R420H), a K(+) channel mutation causative in spinocerebellar ataxia 13 displays aberrant intracellular trafficking. Neurobiol Dis 71:270–279. doi:10.1016/j.nbd.2014.08.020
van Swieten JC, Brusse E, de Graaf BM, Krieger E, van de Graaf R, de Koning I, Maat-Kievit A, Leegwater P, Dooijes D, Oostra BA, Heutink P (2003) A mutation in the fibroblast growth factor 14 gene is associated with autosomal dominant cerebellar ataxia [corrected]. Am J Hum Genet 72(1):191–199 (S0002-9297(07)60518-7 [pii])
Laezza F, Gerber BR, Lou JY, Kozel MA, Hartman H, Craig AM, Ornitz DM, Nerbonne JM (2007) The FGF14(F145S) mutation disrupts the interaction of FGF14 with voltage-gated Na+ channels and impairs neuronal excitability. J Neurosci 27(44):12033–12044. doi:10.1523/JNEUROSCI.2282-07.2007
Shakkottai VG, Xiao M, Xu L, Wong M, Nerbonne JM, Ornitz DM, Yamada KA (2009) FGF14 regulates the intrinsic excitability of cerebellar Purkinje neurons. Neurobiol Dis 33(1):81–88. doi:10.1016/j.nbd.2008.09.019
Yan H, Pablo JL, Pitt GS (2013) FGF14 regulates presynaptic Ca2+ channels and synaptic transmission. Cell Rep 4(1):66–75. doi:10.1016/j.celrep.2013.06.012
Ghezzi D, Arzuffi P, Zordan M, Da Re C, Lamperti C, Benna C, D’Adamo P, Diodato D, Costa R, Mariotti C, Uziel G, Smiderle C, Zeviani M (2011) Mutations in TTC19 cause mitochondrial complex III deficiency and neurological impairment in humans and flies. Nat Genet 43(3):259–263. doi:10.1038/ng.761
Koppen M, Langer T (2007) Protein degradation within mitochondria: versatile activities of AAA proteases and other peptidases. Crit Rev Biochem Mol Biol 42(3):221–242. doi:10.1080/10409230701380452
Maltecca F, De Stefani D, Cassina L, Consolato F, Wasilewski M, Scorrano L, Rizzuto R, Casari G (2012) Respiratory dysfunction by AFG3L2 deficiency causes decreased mitochondrial calcium uptake via organellar network fragmentation. Hum Mol Genet 21(17):3858–3870. doi:10.1093/hmg/dds214
Almajan ER, Richter R, Paeger L, Martinelli P, Barth E, Decker T, Larsson NG, Kloppenburg P, Langer T, Rugarli EI (2012) AFG3L2 supports mitochondrial protein synthesis and Purkinje cell survival. J Clin Invest 122(11):4048–4058. doi:10.1172/JCI64604
Kondadi AK, Wang S, Montagner S, Kladt N, Korwitz A, Martinelli P, Herholz D, Baker MJ, Schauss AC, Langer T, Rugarli EI (2014) Loss of the m-AAA protease subunit AFG(3)L(2) causes mitochondrial transport defects and tau hyperphosphorylation. EMBO J 33(9):1011–1026. doi:10.1002/embj.201387009
Wedding IM, Koht J, Tran GT, Misceo D, Selmer KK, Holmgren A, Frengen E, Bindoff L, Tallaksen CM, Tzoulis C (2014) Spastic paraplegia type 7 is associated with multiple mitochondrial DNA deletions. PLoS One 9(1):e86340. doi:10.1371/journal.pone.0086340
Klebe S, Depienne C, Gerber S, Challe G, Anheim M, Charles P, Fedirko E, Lejeune E, Cottineau J, Brusco A, Dollfus H, Chinnery PF, Mancini C, Ferrer X, Sole G, Destee A, Mayer JM, Fontaine B, de Seze J, Clanet M, Ollagnon E, Busson P, Cazeneuve C, Stevanin G, Kaplan J, Rozet JM, Brice A, Durr A (2012) Spastic paraplegia gene 7 in patients with spasticity and/or optic neuropathy. Brain 135(Pt 10):2980–2993. doi:10.1093/brain/aws240
Nelson RF, Glenn KA, Miller VM, Wen H, Paulson HL (2006) A novel route for F-box protein-mediated ubiquitination links CHIP to glycoprotein quality control. J Biol Chem 281(29):20242–20251. doi:10.1074/jbc.M602423200
Ikeda Y, Dick KA, Weatherspoon MR, Gincel D, Armbrust KR, Dalton JC, Stevanin G, Durr A, Zuhlke C, Burk K, Clark HB, Brice A, Rothstein JD, Schut LJ, Day JW, Ranum LP (2006) Spectrin mutations cause spinocerebellar ataxia type 5. Nat Genet 38(2):184–190. doi:10.1038/ng1728
Bakalkin G, Watanabe H, Jezierska J, Depoorter C, Verschuuren-Bemelmans C, Bazov I, Artemenko KA, Yakovleva T, Dooijes D, Van de Warrenburg BP, Zubarev RA, Kremer B, Knapp PE, Hauser KF, Wijmenga C, Nyberg F, Sinke RJ, Verbeek DS (2010) Prodynorphin mutations cause the neurodegenerative disorder spinocerebellar ataxia type 23. Am J Hum Genet 87(5):593–603. doi:10.1016/j.ajhg.2010.10.001
Lin X, Antalffy B, Kang D, Orr HT, Zoghbi HY (2000) Polyglutamine expansion down-regulates specific neuronal genes before pathologic changes in SCA1. Nat Neurosci 3(2):157–163. doi:10.1038/72101
Armbrust KR, Wang X, Hathorn TJ, Cramer SW, Chen G, Zu T, Kangas T, Zink AN, Oz G, Ebner TJ, Ranum LP (2014) Mutant beta-III spectrin causes mGluR1alpha mislocalization and functional deficits in a mouse model of spinocerebellar ataxia type 5. J Neurosci 34(30):9891–9904. doi:10.1523/JNEUROSCI.0876-14.2014
Kato AS, Knierman MD, Siuda ER, Isaac JT, Nisenbaum ES, Bredt DS (2012) Glutamate receptor delta2 associates with metabotropic glutamate receptor 1 (mGluR1), protein kinase Cgamma, and canonical transient receptor potential 3 and regulates mGluR1-mediated synaptic transmission in cerebellar Purkinje neurons. J Neurosci 32(44):15296–15308. doi:10.1523/JNEUROSCI.0705-12.2012
Schuurs-Hoeijmakers JH, Geraghty MT, Kamsteeg EJ, Ben-Salem S, de Bot ST, Nijhof B, van de Vondervoort II, van der Graaf M, Nobau AC, Otte-Holler I, Vermeer S, Smith AC, Humphreys P, Schwartzentruber J, Ali BR, Al-Yahyaee SA, Tariq S, Pramathan T, Bayoumi R, Kremer HP, van de Warrenburg BP, van den Akker WM, Gilissen C, Veltman JA, Janssen IM, Vulto-van Silfhout AT, van der Velde-Visser S, Lefeber DJ, Diekstra A, Erasmus CE, Willemsen MA, Vissers LE, Lammens M, van Bokhoven H, Brunner HG, Wevers RA, Schenck A, Al-Gazali L, de Vries BB, de Brouwer AP (2012) Mutations in DDHD2, encoding an intracellular phospholipase A(1), cause a recessive form of complex hereditary spastic paraplegia. Am J Hum Genet 91(6):1073–1081. doi:10.1016/j.ajhg.2012.10.017
Tesson C, Nawara M, Salih MA, Rossignol R, Zaki MS, Al Balwi M, Schule R, Mignot C, Obre E, Bouhouche A, Santorelli FM, Durand CM, Oteyza AC, El-Hachimi KH, Al Drees A, Bouslam N, Lamari F, Elmalik SA, Kabiraj MM, Seidahmed MZ, Esteves T, Gaussen M, Monin ML, Gyapay G, Lechner D, Gonzalez M, Depienne C, Mochel F, Lavie J, Schols L, Lacombe D, Yahyaoui M, Al Abdulkareem I, Zuchner S, Yamashita A, Benomar A, Goizet C, Durr A, Gleeson JG, Darios F, Brice A, Stevanin G (2012) Alteration of fatty-acid-metabolizing enzymes affects mitochondrial form and function in hereditary spastic paraplegia. Am J Hum Genet 91(6):1051–1064. doi:10.1016/j.ajhg.2012.11.001
Boukhris A, Schule R, Loureiro JL, Lourenco CM, Mundwiller E, Gonzalez MA, Charles P, Gauthier J, Rekik I, Acosta Lebrigio RF, Gaussen M, Speziani F, Ferbert A, Feki I, Caballero-Oteyza A, Dionne-Laporte A, Amri M, Noreau A, Forlani S, Cruz VT, Mochel F, Coutinho P, Dion P, Mhiri C, Schols L, Pouget J, Darios F, Rouleau GA, Marques W Jr, Brice A, Durr A, Zuchner S, Stevanin G (2013) Alteration of ganglioside biosynthesis responsible for complex hereditary spastic paraplegia. Am J Hum Genet 93(1):118–123. doi:10.1016/j.ajhg.2013.05.006
Martin E, Schule R, Smets K, Rastetter A, Boukhris A, Loureiro JL, Gonzalez MA, Mundwiller E, Deconinck T, Wessner M, Jornea L, Oteyza AC, Durr A, Martin JJ, Schols L, Mhiri C, Lamari F, Zuchner S, De Jonghe P, Kabashi E, Brice A, Stevanin G (2013) Loss of function of glucocerebrosidase GBA2 is responsible for motor neuron defects in hereditary spastic paraplegia. Am J Hum Genet 92(2):238–244. doi:10.1016/j.ajhg.2012.11.021
Walden CM, Sandhoff R, Chuang CC, Yildiz Y, Butters TD, Dwek RA, Platt FM, van der Spoel AC (2007) Accumulation of glucosylceramide in murine testis, caused by inhibition of beta-glucosidase 2: implications for spermatogenesis. J Biol Chem 282(45):32655–32664. doi:10.1074/jbc.M702387200
Rainier S, Bui M, Mark E, Thomas D, Tokarz D, Ming L, Delaney C, Richardson RJ, Albers JW, Matsunami N, Stevens J, Coon H, Leppert M, Fink JK (2008) Neuropathy target esterase gene mutations cause motor neuron disease. Am J Hum Genet 82(3):780–785. doi:10.1016/j.ajhg.2007.12.018
Cadieux-Dion M, Turcotte-Gauthier M, Noreau A, Martin C, Meloche C, Gravel M, Drouin CA, Rouleau GA, Nguyen DK, Cossette P (2014) Expanding the clinical phenotype associated with ELOVL4 mutation: study of a large French-Canadian family with autosomal dominant spinocerebellar ataxia and erythrokeratodermia. JAMA Neurol 71(4):470–475. doi:10.1001/jamaneurol.2013.6337
Vidal R, Frangione B, Rostagno A, Mead S, Revesz T, Plant G, Ghiso J (1999) A stop-codon mutation in the BRI gene associated with familial British dementia. Nature 399(6738):776–781. doi:10.1038/21637
Vidal R, Revesz T, Rostagno A, Kim E, Holton JL, Bek T, Bojsen-Moller M, Braendgaard H, Plant G, Ghiso J, Frangione B (2000) A decamer duplication in the 3′ region of the BRI gene originates an amyloid peptide that is associated with dementia in a Danish kindred. Proc Natl Acad Sci USA 97(9):4920–4925. doi:10.1073/pnas.080076097
Takahashi H, Adachi N, Shirafuji T, Danno S, Ueyama T, Vendruscolo M, Shuvaev AN, Sugimoto T, Seki T, Hamada D, Irie K, Hirai H, Sakai N, Saito N (2014) Identification and characterization of PKCgamma, a kinase associated with SCA14, as an amyloidogenic protein. Hum Mol Genet. doi:10.1093/hmg/ddu472
Pyle A, Smertenko T, Bargiela D, Griffin H, Duff J, Appleton M, Douroudis K, Pfeffer G, Santibanez-Koref M, Eglon G, Yu-Wai-Man P, Ramesh V, Horvath R, Chinnery PF (2015) Exome sequencing in undiagnosed inherited and sporadic ataxias. Brain 138(Pt 2):276–283. doi:10.1093/brain/awu348
Gilissen C, Hehir-Kwa JY, Thung DT, van de Vorst M, van Bon BW, Willemsen MH, Kwint M, Janssen IM, Hoischen A, Schenck A, Leach R, Klein R, Tearle R, Bo T, Pfundt R, Yntema HG, de Vries BB, Kleefstra T, Brunner HG, Vissers LE, Veltman JA (2014) Genome sequencing identifies major causes of severe intellectual disability. Nature 511(7509):344–347. doi:10.1038/nature13394
Fokkema IF, Taschner PE, Schaafsma GC, Celli J, Laros JF, den Dunnen JT (2011) LOVD v. 2.0: the next generation in gene variant databases. Hum Mutat 32(5):557–563. doi:10.1002/humu.21438
Bladen CL, Salgado D, Monges S, Foncuberta ME, Kekou K, Kosma K, Dawkins H, Lamont L, Roy AJ, Chamova T, Guergueltcheva V, Chan S, Korngut L, Campbell C, Dai Y, Wang J, Barisic N, Brabec P, Lahdetie J, Walter MC, Schreiber-Katz O, Karcagi V, Garami M, Viswanathan V, Bayat F, Buccella F, Kimura E, Koeks Z, van den Bergen JC, Rodrigues M, Roxburgh R, Lusakowska A, Kostera-Pruszczyk A, Zimowski J, Santos R, Neagu E, Artemieva S, Rasic VM, Vojinovic D, Posada M, Bloetzer C, Jeannet PY, Joncourt F, Diaz-Manera J, Gallardo E, Karaduman AA, Topaloglu H, El Sherif R, Stringer A, Shatillo AV, Martin AS, Peay HL, Bellgard MI, Kirschner J, Flanigan KM, Straub V, Bushby K, Verschuuren J, Aartsma-Rus A, Beroud C, Lochmuller H (2015) The TREAT-NMD DMD global database: analysis of more than 7000 Duchenne muscular dystrophy mutations. Hum Mutat. doi:10.1002/humu.22758
Hammer MB, Eleuch-Fayache G, Schottlaender LV, Nehdi H, Gibbs JR, Arepalli SK, Chong SB, Hernandez DG, Sailer A, Liu G, Mistry PK, Cai H, Shrader G, Sassi C, Bouhlal Y, Houlden H, Hentati F, Amouri R, Singleton AB (2013) Mutations in GBA2 cause autosomal-recessive cerebellar ataxia with spasticity. Am J Hum Genet 92(2):245–251. doi:10.1016/j.ajhg.2012.12.012
Burns R, Majczenko K, Xu J, Peng W, Yapici Z, Dowling JJ, Li JZ, Burmeister M (2014) Homozygous splice mutation in CWF19L1 in a Turkish family with recessive ataxia syndrome. Neurology. doi:10.1212/WNL.0000000000001053
Synofzik M, Haack TB, Kopajtich R, Gorza M, Rapaport D, Greiner M, Schonfeld C, Freiberg C, Schorr S, Holl RW, Gonzalez MA, Fritsche A, Fallier-Becker P, Zimmermann R, Strom TM, Meitinger T, Zuchner S, Schule R, Schols L, Prokisch H (2014) Absence of BiP co-chaperone DNAJC3 Causes diabetes mellitus and multisystemic neurodegeneration. Am J Hum Genet 95(6):689–697. doi:10.1016/j.ajhg.2014.10.013
Winkelmann J, Lin L, Schormair B, Kornum BR, Faraco J, Plazzi G, Melberg A, Cornelio F, Urban AE, Pizza F, Poli F, Grubert F, Wieland T, Graf E, Hallmayer J, Strom TM, Mignot E (2012) Mutations in DNMT1 cause autosomal dominant cerebellar ataxia, deafness and narcolepsy. Hum Mol Genet 21(10):2205–2210. doi:10.1093/hmg/dds035
Lindquist SG, Duno M, Batbayli M, Puschmann A, Braendgaard H, Mardosiene S, Svenstrup K, Pinborg LH, Vestergaard K, Hjermind LE, Stokholm J, Andersen BB, Johannsen P, Nielsen JE (2013) Corticobasal and ataxia syndromes widen the spectrum of C9ORF72 hexanucleotide expansion disease. Clin Genet 83(3):279–283. doi:10.1111/j.1399-0004.2012.01903.x
Tsoi H, Yu AC, Chen ZS, Ng NK, Chan AY, Yuen LY, Abrigo JM, Tsang SY, Tsui SK, Tong TM, Lo IF, Lam ST, Mok VC, Wong LK, Ngo JC, Lau KF, Chan TF, Chan HY (2014) A novel missense mutation in CCDC88C activates the JNK pathway and causes a dominant form of spinocerebellar ataxia. J Med Genet 51(9):590–595. doi:10.1136/jmedgenet-2014-102333
Lieber DS, Hershman SG, Slate NG, Calvo SE, Sims KB, Schmahmann JD, Mootha VK (2014) Next generation sequencing with copy number variant detection expands the phenotypic spectrum of HSD17B4-deficiency. BMC Med Genet 15:30. doi:10.1186/1471-2350-15-30
Lines MA, Jobling R, Brady L, Marshall CR, Scherer SW, Rodriguez AR, Lee L, Lang AE, Mestre TA, Wanders RJ, Ferdinandusse S, Tarnopolsky MA (2014) Peroxisomal D-bifunctional protein deficiency: three adults diagnosed by whole-exome sequencing. Neurology 82(11):963–968. doi:10.1212/WNL.0000000000000219
Pierce SB, Walsh T, Chisholm KM, Lee MK, Thornton AM, Fiumara A, Opitz JM, Levy-Lahad E, Klevit RE, King MC (2010) Mutations in the DBP-deficiency protein HSD17B4 cause ovarian dysgenesis, hearing loss, and ataxia of Perrault Syndrome. Am J Hum Genet 87(2):282–288. doi:10.1016/j.ajhg.2010.07.007
Caramins M, Colebatch JG, Bainbridge MN, Scherer SS, Abrams CK, Hackett EL, Freidin MM, Jhangiani SN, Wang M, Wu Y, Muzny DM, Lindeman R, Gibbs RA (2013) Exome sequencing identification of a GJB1 missense mutation in a kindred with X-linked spinocerebellar ataxia (SCA-X1). Hum Mol Genet 22(21):4329–4338. doi:10.1093/hmg/ddt282
Acknowledgments
This work was funded by the French “Agence Nationale de la Recherche” (to GS), the Verum Foundation (to AB and GS), the European Union (OMICs grant under the 7th Framework programme, to AB; the E-Rare programme, to GS), the Fondation Roger de Spoelberch (to AB) and the “Investissements d’avenir” programme (ANR-10-AIHU-06, to AB and GS). MC is the recipient of an “Aspirant” PhD fellowship from the Fonds de la Recherche Scientifique (F.R.S.-FNRS, Belgium).
Conflicts of interest
The authors report no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Coutelier, M., Stevanin, G. & Brice, A. Genetic landscape remodelling in spinocerebellar ataxias: the influence of next-generation sequencing. J Neurol 262, 2382–2395 (2015). https://doi.org/10.1007/s00415-015-7725-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00415-015-7725-4