Abstract
When fingermarks are left on a surface, bacteria originating from the donor’s skin are also deposited. The skin microbiome is believed to be extremely diverse between individuals, allowing for potential matching between the bacterial communities and touched objects, known as “bacterial profiling”. This study stepped further and investigated how the bacterial profile could be used as an indicator of donor characteristics of potential forensic intelligence interest. Forty-five participants were asked to touch DNA-free playing cards with their dominant and non-dominant hands. Cards were swabbed and bacterial communities determined through 16S rRNA sequencing. Diversity and abundance of bacteria were compared to donor characteristics of gender, age, ethnicity, handedness, home location, sample location, occupation, diet type, use of moisturisers, use of hand sanitisers and use of public transport. Correlations between the bacterial profile with gender, ethnicity, diet type and hand sanitiser use were found. Specifically, the absence of Lactococcus indicated a primarily Chinese diet, while the absence of Alloiococcus indicated female gender, Asian ethnicity and hand sanitiser use. Testing of the prediction models demonstrated highest accuracy for gender estimation, while the prediction of other characteristics showed lower success. This study showed a correlation between the presence of certain bacterial species on donor’s hands and personal characteristics of potential forensic relevance, thus demonstrating a novel application of microbiome genotyping in forensic science.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Any information that can be obtained from a crime scene or an item collected from the scene is important. The value of fingerprints for forensic identifications is well documented and has been a cornerstone of forensic science for over 100 years [1, 2]. However, the detection and even deposition of fingermarks rely upon a number of factors, such as environmental conditions, surface composition and the chemistry of the fingermark itself [1]. Hence, not all fingermarks are of sufficient quality to be utilised for identification, while many of them remain undetected. This limitation led to a rationale of exploring additional traces within fingermarks for recovering valuable pieces of evidential importance.
Recent publication by van Dam et al. reviewed various analytical techniques that have the potential for determination donor gender, age, blood group, diet, habits and health status by examination of chemical and biochemical traces left in the fingermark [3]. However, as this review focused on chemical and biochemical traces, it ignored that along with these traces, the donor’s epidermal microbiota are also readily deposited and may provide additional intelligence information.
The human skin hosts a diverse bacterial community, whose composition is believed to be extremely variable between individuals, with only 13% of the bacterial species being shared between palms of any two individuals [4,5,6]. The microbiome is known to vary, depending on gender [7,8,9], age, [9] donor population/location [10, 11], hygiene [7] and interactions between household members and pets [9, 12]. Examination of the microbiome is usually carried out through sequencing of the 16S rRNA gene, which is sequence specific to species level. A bacterial profile can then be determined by examination of the types of bacteria within the specimen and their relative abundance.
Previous studies have shown that the significant variation of the microbiome, its ready deposition and relatively easy recovery after prolonged time had the potential to be exploited as a tool in forensic identification [12,13,14]. In these studies, bacterial traces on computer mouse devices or keyboards and door handles were compared to the microbiome from donor hands. Using just the microbiome, researchers were able to identify the donor to the microbiome trace left on the object.
To date, studies on the use of the microbiome for forensic applications have focused on donor identification [15,16,17,18,19]. Interestingly, these and other studies on bacterial traces have also shown that the traces left on mobile phones can distinguish donor geographical location and gender [15].
Thus, the current study proposed to further investigate the utilisation of bacterial traces for investigative purposes by envisaging donor personal characteristics such as gender, ethnicity, diet and movement. Furthermore, this study aims to determine if there are specific bacterial species that can be used as biomarkers for inferring aspects of a donor’s characteristics or lifestyle and if a microbiome sample obtained from a single touch would be sufficient for such analysis.
Materials and methods
Sample collection, DNA extraction and sequencing
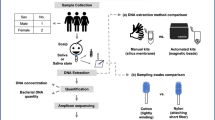
Forty-five individuals have participated in this study and samples were collected over the course of two weeks. All the participants were volunteers and were recruited randomly to gain a broad representation of the general population. In order to determine factors which could be used for donor profiling, participants were asked to fill out a questionnaire regarding intrinsic and extrinsic factors affecting the skin microbiome (Supplemental Table S1). Intrinsic factors examined were gender, age, ethnicity and handedness; while extrinsic factors were home location, sample location, occupation, diet, diet type, use of moisturisers, use of hand sanitisers and use of public transport.
All subjects were made aware of the nature of the study and gave written consent before any information gathering or sampling occurred. This study has been approved by the University of Technology Sydney Human Research Ethics committee (UTS HREC approval #2015000296).
To determine if bacterial traces can be used to gain insight into the last activities of a subject, participants were not asked to wash their hands prior to participating in the experiment. To mimic traces left at crime scenes, specimens were not taken directly from the subjects’ fingers. Instead, subjects were asked to hold a new, autoclave-sterilised playing card (Queen’s Slipper Product code 144120) for a total of 10 s in each hand. Two specimens (left and right hand) were obtained from each subject. Playing cards were placed in individual autoclave–sterilised paper bag and left for at least 2 h before being swabbed. Sterile swabs (Copan, Brescia, Italy) were moistened with sterile saline solution (0.15 M NaCl) and the entire surface of the playing card swabbed in a back-and-forth motion along one surface of the card, the card rotated 90 degrees, swabbed back-and-forth along the same surface, then turned over and the process repeated on the opposite surface. Swabs were immediately stored at − 20 °C until DNA extraction. Blank controls of unhandled sterile playing cards were processed in parallel to check for residual background bacterial DNA.
DNA was extracted from the swabs as previously described [15] using the Sigma Extract-N-Amp™ plant PCR kit (Sigma-Aldrich) with modifications to the manufacturer’s protocol. Briefly, 100 μL of extraction solution were added to sterile 1.5-ml microfuge tubes and the swab tip broken directly into the tube. Tubes were incubated at 95 °C for 10 min, followed by centrifugation for 5 min at 2500g and 100 μL of dilution solution added. Samples were stored at 4 °C. Controls for blank swabs and saline solution were also processed in parallel.
Extracted DNA was purified by ethanol precipitation, resuspended in 25 μl sterile water and quantified using the Qubit fluorometer and the HS® dsDNA kit (Invitrogen, CA, USA). Samples were subsequently processed at the Ramaciotti Centre for Genomics (University of New South Wales, Sydney, Australia) for amplification and sequencing of the V4 region of the 16S rRNA (515F-806R). Briefly, barcoding PCR for bacterial and archaeal 16S rRNA genes was carried out using a mix of 10 μL of HotMasterMix (5 PRIME), 0.2 μM of each primer and 1 μL of DNA template. Barcoded PCR primers based on 515F (5′-GTGCCAGCMGCCGCGGTAA-3′) [20] and 806R (5′-GGACTACHVGGGTWTCTAAT-3′) [21] were used. Thermal cycling conditions were as follows: initial denaturation at 94 °C for 3 min, followed by 35 cycles of denaturation at 94 °C for 45 s, annealing at 50 °C for 1 min and elongation at 72 °C for 1 min 30 s, ending with a final elongation at 72 °C for 10 min. All PCRs were carried out in 25 μL volumes. PCR products were normalised and pooled using SequalPrepT Normalization Plate Kit (ThermoFisher Scientific) according to the manufacturer’s instructions. The library was purified using AxygenR AxyPrepT Mag PCR Clean-Up Kit (Fisher Biotec) as per the manufacturer’s instructions. Concentration and quality of the pooled library were checked with Qubit Fluorometer (ThermoFisher Scientific) and the library size on an Agilent 2200 TapeStation instrument. The Agencourt AMPure XP Bead Clean-up purification was used to reduce the presence of primer dimers, as recommended by the manufacturer. The library pool was sequenced on the MiSeq platform (Illumina™) using a MiSeq Reagent Kit v2 with a 2 × 250-bp run format, using default run parameters, including adaptor trimming. For these runs, custom primers were added to the reagent cartridge for Read1, Index and Read2.
Sequence and bioinformatics analysis
The Qiime pipeline (v.1.9.1) [21] was used to process and sort the raw sequences. Briefly, paired end reads were joined using PandaSeq, while retaining and multiplexing sequences with a minimum length of 250 bp, quality score greater than 25 and 99% base accuracy. Sequences were subsequently compared to the Greengenes core set (4feb2011) using Qiime USEARCH clustering algorithm, while chimeric sequences were detected and removed. Sequences were grouped into operational taxonomic units (OTUs) at 97% sequence identity using UCLUST de novo clustering and taxonomy assigned using the RDP classifier and the Greengenes core set. Following the identification of assumed Mycoplasma contamination (abundance of 4–42% across all samples), any Mycoplasma OTU was removed from the data set. This contamination had been observed in a previous similar study using the same extraction kit [15] and its presence in the reagents was verified by the manufacturer (Sigma-Aldrich) in a personal communication. All samples were rarefied to 27,000 sequences per sample to achieve approximately equal sampling depth. The 27,000 rarefaction figure was used as it was the sequence count of the sample with the smallest acceptable number of sequences.
Rarefaction curves for alpha diversity analysis was performed using the Qiime scripts “multiple_rarefactions.py” and “alpha_diversity.py”. Arguments used in the multiple_rarefactions.py script were maximum OTU number of 27,000, sampling increments increased by intervals of 500 and sampling repeated 10 times. Metrics used in the “alpha_diversity.py” script were observed OTUs, Chao1, Shannon and phylogenetic distance. Alpha diversity metrics were summarised using “collate_alpha.py” and visualised using “make_rarefaction_plots.py”.
Beta diversity analysis was performed using the “beta_diversity.py” Qiime script with the output from the “multiple_rarefactions.py” as input. This script measures beta diversity based on weighted and unweighted UniFrac distances [22]. ANOSIM tests were performed using the “compare_categories.py” script with 999 permutations.
Biomarker identification was performed using the linear discriminant analysis effect size (LefSe) method available at https://huttenhower.sph.harvard.edu/galaxy/ and as described by Segata et al. [23]. Input used for LefSe analysis originates from the “summarize_taxa.py” Qiime script. Taxa were tested for significant differential abundance using the non-parametric factorial Kruskal-Wallis sum test. Biological significance of the detected feature was determined using a set of pairwise tests between subclasses using unpaired Wilcoxon rank-sum test. Linear discriminant analysis was used to estimate the effect size of each detected feature. Random resampling was not applied in the analysis.
SPSS statistics and SPSS Modeler were used to determine association between factors and to perform leave-one-out cross validation (LOOCV) analysis on the accuracy of the feature predictions.
Results
The sequencing of 90 samples resulted in an average of 57,230 counts per sample, which clustered into 1634 bacterial OTUs. However, rarefaction curves for each sample did not reach a plateau, implying a greater level of diversity than observed in this study. It is possible though that the true absence/abundance of “rare” OTUs may be skewed in samples with lower initial DNA concentrations, as the amount of DNA in each sample was not normalised before the initial amplification step. Our initial hypothesis was that the number of OTUs would decrease in parallel with decreasing DNA concentration, as observed by Multinu et al. 2018 [24]. To test this, we then ranked the samples according to their initial DNA concentration and compared it to the number of OTUs found. Following the application of the Spearman Rho test, we found that the association between DNA concentration and OTU number was not statistically significant (R = 0.04473, p = 0.67546). Finally, Mycoplasma contamination of varying degrees was observed in all samples. This contamination originated in the extraction reagents, provided by the supplier. While it was removed during post-sequencing analysis, we note that the abundance of Mycoplasma template may have skewed the amplification and thus abundance of other templates in the touch samples.
Beta diversity, the difference in microbiota diversity between factors, was examined using weighted (quantitative assessment based on abundance of OTUs) and unweighted (qualitative assessment based on absence/presence of OTUs) UniFrac distances [22]. Gender, ethnicity, diet type and hand sanitiser use were identified as factors influencing the clustering of bacterial fingerprint traces (Table 1). It should however be noted that the ANOSIM values, while statistically significant, are close to zero, implying minimal differences. This is likely due to the small cohort examined, while with a larger cohort these differences may be decreased or strengthened.
Based on ANOSIM values, clustering due to donor gender was likely caused by differences in OTU abundance as well as OTU absence/presence. Interestingly, differences in OTU abundance appeared to have the greater influence in clustering as evidenced by the higher weighted R value compared to the unweighted R value. This is in line with previous studies [7] which showed significant differences in microbial group abundance between male and female participants. Similarly, clustering due to ethnicity is more heavily influenced by differences in OTU abundance between ethnic groups as OTU presence/absence does not appear to be significant. In contrast, clustering defined by diet type and hand sanitiser use was influenced by the absence/presence of OTUs with differences in OTU abundance having no significant effect.
Based on the ANOSIM analysis, gender, ethnicity, use of hand sanitisers and diet type of a bacterial fingerprint donor could potentially be determined. In order to ascertain whether there are specific indicator bacteria or biomarkers for these donor characteristics, the LefSe pipeline was utilised.
As subjects were not asked to wash their hands before sampling, it is unsurprising that many of the bacteria identified as potential biomarkers were from the environment (Table 2). However, a previous study on the hand microbiome had also identified a number of these bacteria, such as Oceanospirillales and Pseudomonas [25]. Cosseau et al. also hypothesised that organisms traditionally associated with the environment may have skin-specific ecotypes, which may be the OTUs detected in this study. Interestingly, strict anaerobes such as Fusobacterium and Clostridium (Table 2 and Supplementary Table 1) were also identified. The presence of these bacteria was also likely due to subjects not washing their hands before sampling. Bacteria such as Fusobacterium, Neisseria, and Veillonella exist in the oral cavity, and frequent/recent hand-to-mouth contact would introduce these bacteria into the hand environment. Similarly, along with Clostridium, a number of gut bacteria were identified in this study (Supplementary Table S1) as potential biomarkers. It is possible that some subjects did not wash their hands after using the bathroom, resulting in the presence of these bacteria on the subjects’ hands. Indeed, Bobulsky et al. found Clostridium contamination on multiple skin sites, including the hands, of patients and carers, of individuals with Clostridium difficile–associated disease [26]. As such, a number of bacteria identified in this study may be from the transient microbiome rather than the core microbiome.
The LefSe method identified 19 bacterial groups (Table 2, Supplementary Table S1) as being linked to donor gender. Examination of the relative abundance of these bacteria between genders (Fig. 1a) showed a higher relative abundance of Corynebacterium and Staphylococcus in the male samples, consistent with Fierer et al.’s study [7]. However, examination of the number of samples containing these bacteria (Fig. 1b) showed that both genera were present in all male and female samples. As such, while the relative numbers Corynebacterium and Staphylococcus are indicative of the donor being male, their presence alone cannot be used as a distinctive marker.
Examination of the presence of other identified bacteria showed that the majority were present in at least 75% of both male and female samples (Fig. 1b). However, Alloiococcus species was found to be present in only 42% of the female samples compared to 77% presence in male samples. The prediction accuracy of the model has been evaluated using the LOOCV method and showed a 64% accuracy rate using Alloiococcus as a marker for gender. This result indicated that Alloiococcus could be a potential biomarker to indicate a male donor.
Genetic association with ethnicity identified significant correlation with 33 bacterial groups (Table 2, Supplementary Table S1). Analyses of bacterial presence in samples showed the presence of the majority of bacterial groups in at least 60% of samples from each ethnic group (Fig. 2b). Comparatively, Pediococcus was found to be completely absent in the bacterial traces from Asian donors. However, Pediococcus was also found to be rare in bacterial traces from donors of mixed and Caucasian ethnicity, appearing in 38% and 12% of samples, respectively. As such, the rarity of Pediococcus would make it an unreliable reporter for ethnicity.
Bacterial groups associated with donor ethnicity. a Relative abundance of bacterial groups as divided by ethnicity. Error bars are standard deviation. b Percentage of samples containing bacterial groups as divided by ethnicity. The dashed line indicates where 50% of samples contain a bacterial group
Interestingly, Alloiococcus, which was found to be a potential marker for gender, was also found to be a potential marker for ethnicity. In this case, Alloiococcus was present in only 38% of the bacterial traces from Asian donors, compared to 75% and 88% in bacterial traces from donors of Caucasian and mixed ethnicity respectively. Validation of the predictive model for ethnicity estimation using Alloiococcus as a marker had a 56% accuracy rate.
Interestingly, only 2 bacterial clusters, Fusobacteriales and Lactococcus, were found to be linked to the diet type, following the LefSe analysis (Table 2). The results demonstrated that the presence of Lactococcus in 25% of bacterial traces is from donors consuming a primarily Chinese diet, compared to 61% and 70% of bacterial traces from donors consuming a varied or primarily western diet, respectively (Fig. 3b). LOOCV analysis has shown that using Lactococcus as a marker for diet type had a 48% prediction accuracy rate. Based on these results, while it is not strongly discriminative, the presence of Lactococcus in the sample may indicate a donor who consumes a primarily Chinese diet.
An association analysis of the use of hand sanitisers has identified 19 bacterial groups (Table 2, Supplementary Table S1). The majority of bacterial groups were present in over 75% of the samples (Fig. 4b). However, Alloiococcus was present in only 43% of the bacterial traces from donors who used hand sanitisers compared to being present in 72% of the bacterial traces from donors who did not use hand sanitisers. The hand sanitiser usage prediction using Alloiococcus as a marker has demonstrated a 51% accuracy rate. As such, Alloiococcus has the potential to be used as an indicator for the use hand sanitisers, although a larger sample size is required to validate this model.
Bacterial groups linked to donor use of hand sanitisers. a Relative abundance of bacterial groups as divided by use of hand sanitiser. Error bars are standard deviation. b Percentage of samples containing bacterial groups as divided by use of hand sanitiser. The dashed line indicates where 50% of samples contain a bacterial group
Potential association between factors was examined using the chi-squared test of association (Supplementary Table S2). It was found that due to the skew in the distribution of participants in variables within the factors e.g. occupation, there were many false associations. Of the factors identified by ANOSIM, associations were observed between ethnicity and diet type, ethnicity and hand sanitiser use and hand sanitiser and gender.
Discussion and concluding comments
The use of microbial profiles for acquiring information within the field of forensic science is still in its infancy, although its potential as a powerful tool in forensic investigations has been recognised [27,28,29,30,31]. A study showing the transfer of microbiome from the genital area after a sexual encounter has raised the idea that the microbiome could assist in investigation of sexual assault cases [32]. Additionally, similar to forensic examination of traces found on the soles of shoes, a recent study examined the microbiome from the soles of shoes and surrounding areas to determine the previous locations of the shoe based on its microbiome [16].
This study aimed to expand upon the use of the microbiome in forensic science, not only as a tool for linking between a suspect and a surface, but its potential for donor profiling. In terms of donor profiling, we asked whether the presence of specific bacteria that could be used as presumptive biomarker for donor characteristics, such as gender, biogeographical ancestry, preferable diet and use of hand sanitizers.
This study has identified two potential bacterial biomarkers, Lactococcus and Alloiococcus, which can be used to provide investigative leads on donor characteristics. The absence of Lactococcus from a bacterial trace could be indicative of a donor who recently consumed a primarily Chinese diet. As Lactococcus is commonly used in the dairy industry, it is speculated that the absence of Lactococcus in these samples may be due to a reduction in consumption of dairy products, which is typical of a Chinese diet, compared to a varied or mainly Western diet. An association was observed between diet type and ethnicity (Supplementary Table S2), but considering that Lactococcus was only identified as a biomarker for diet type, it is unlikely that ethnicity influenced the abundance of Lactococcus observed.
The presence of Alloiococcus, a bacteria genus present in the normal microbiota of the adult nasopharynx and ear canal [33], was found to be a potential indicator for three donor characteristics: Caucasian ethnicity, male gender and not using hand sanitisers. Interestingly, a characteristic which is often different between ethnic groups is the production of wet or dry earwax [34], wherein the wet earwax phenotype is more prevalent in European populations, while the dry earwax phenotype is more common amongst East Asian populations. Based on our results, we hypothesise that the presence of Alloiococcus in the majority of bacterial traces from donors of Caucasian and mixed ethnicities and its absence in the majority of bacterial traces from Asian descent is due to the wet/dry earwax phenotype. This observation may be due to potentially easier succession of the bacteria in the liquid/wet rather than dry medium. Notably, the earwax wet/dry phenotype is caused by a polymorphism in a single ABCC11 gene, which has been linked to increased/decreased sebum production and can be used for indirect Asian ancestry prediction [34, 35]. Moreover, males are known to produce more sebum than females [36]. As a result, higher sebum levels may also contribute to the increased presence of the Alloiococcus in the media, allowing it to be a potential indicator for a male donor.
Considering the association between gender and ethnicity, and the identification of Alloiococcus as a potential biomarker for both, it is likely that the two factors influence the presence of Alloiococcus. Indeed, the presence of Alloiococcus in bacterial traces can be ranked on the likelihood of its presence. Asian females are the least likely to have Alloiococcus, present in only 28% of the Asian female samples, followed by Asian males and Caucasian females, both 50%, followed by mixed ethnicity females (75%), Caucasian males (88%) and mixed ethnicity males (100%). It should however be noted that there was only one mixed ethnicity (male) subject in the current study and a follow-up study with a larger sample size is required to validate these results.
Interestingly, subject 45 (Asian female) was found to have Alloiococcus only on their dominant hand. It was noted that before collection, the dominant hand of subject 45 was in contact with the non-dominant hand of subject 42 (Caucasian male). As subject 42 was found to have Alloiococcus on both hands, it is possible that subject 42 transferred the Alloiococcus to subject 45 via hand contact. While this result may be represented as a potential downfall of using single bacteria as indicators of donor characteristics, i.e. the presence of Alloiococcus in subject 45’s trace would give a false indication of a Caucasian male; it does highlight how readily bacteria can be transferred from one person to another. This subsequently demonstrates the potential of microbial forensics in physical assault cases and reiterates the requirement for a “whole microbiome” approach, rather than interpretation based on a single bacterium.
In regard to Alloiococcus as a discriminator for hand sanitiser use, it is possible that the data is skewed due to the sampling size. Of the 15 subjects who used hand sanitiser, 10 were female of which 6 were of Asian descent. Comparatively of the 30 subjects who did not use hand sanitiser, 21 were male of which 14 were of Caucasian descent. Thus it is possible that the reduced presence of Alloiococcus in bacterial traces from donors who had used hand sanitisers was more a reflection of the donor gender and ethnicity, as was observed with the association between hand sanitiser, gender and ethnicity (Supplementary Table S2). However, it is also possible that given the low abundance of Alloiococcus (0.0024 relative abundance), it is more susceptible to removal by hand sanitisers.
In this study, participants were not asked to wash their hands prior to sampling, so it is likely that a number of bacteria examined were from the transient rather the core hand microbiome. While examination of transient microbiome was used in this study to determine if the previous locations/objects touched by participants could be determined by the microbiome, this study was unable to show such a link, except for the single example described above. Additionally, links between the sampling location and participants were unable to be ascertained. However, considering that for the majority of hands examined (63/90) the last object touch was a personal object such as a phone or computer, it is unsurprising that these associations could not be made. Additionally, few of the subjects who did touch public objects prior to sampling e.g. door knobs and handrails, touched objects in proximity to let alone the same object as other subjects. As such, the number of subjects whose last object touched was in proximity to another subject was too small for any similarity to between microbiomes to be observed.
This preliminary study has a few limitations. As seen in Figs. 1, 2, 3 and 4, there is a large standard deviation in samples. This lack of a clear distinction between groups within factors is also reflected in the ANOSIM values which are close to zero. Additionally, the robustness of the predictive models for most features tested (apart from the gender prediction) is low, as indicated by the prediction accuracy rates. We also note that the outcome of the LOOCV analysis could be misleading due to the small sample size and the uneven breakdown of groups within each examined factor (Supplementary Table S3). As such, the findings of this study need to be confirmed with a larger cohort.
Due to preliminary nature of this study, a number of questions could not be answered. Firstly, detection of fingermarks often relies on physical or chemical reactions. Whether these detection methods would interfere with obtaining a bacterial trace is vital in the validation of the technique. Furthermore, subjects were asked to touch playing cards with their whole hand. Whether the volume of bacteria from a single fingermark would be sufficient to provide a usable bacterial profile also needs to be considered. Secondly, this study used playing cards as the substrate, but would similar results be seen if using a different substrate such as coins or fabrics? Thirdly, as this study was conducted at a university, there was a bias in the subjects towards early post–graduate research students. Fourthly, we did not examine whether any of the subjects were co-habitants or if they owned pets, which is known to influence skin microbiota [9]. Finally, as this study examined a single time point, it is unknown whether any of the identified bacteria would remain or be in similar abundance in subsequent sampling. Expanding the sample size, diversity of the subjects and temporal scope would yield a greater wealth of information on the potential links between microbial signatures and donor characteristics of forensic interest.
This study looked at traces from the skin microbiome and whether they could be used for forensic investigative purposes rather than suspect identification. It showed that donor traits useful in criminal investigations such as gender and ethnicity could potentially be determined from a bacterial trace. It has identified two potential biomarkers for predicting donor characteristics. Specifically, the absence of Lactococcus was found to be an indicator of a primarily Chinese diet, while the absence Alloiococcus was found useful in estimating female gender, Asian ethnicity and the use of hand sanitisers. However, it should be noted that these two bacteria alone cannot be used as indicators of gender, ethnicity, diet or use of hand sanitisers. Rather like with predictive forensic DNA analysis (e.g. biogeographic ancestry and pigmentation estimation), a conjunction of the absence/abundance of key bacteria would be used as an indicator of factors of forensic relevance, providing valuable investigative leads on the person of interest. Through the creation and expansion of a database of microbial profiles, it is likely that other biomarkers will emerge. Additionally, much like the existing databases of fingerprints, trademarks, DNA and paint profiles, to name a few, a microbial database could be useful in solving or linking cases.
The use of the microbial traces to augment forensic investigations still requires significant development and testing. However, this study and others have shown that the microbiome could be a powerful tool for future investigators in not only the identification, but also profiling of suspects and generating investigative leads.
References
Stoilovic M, Lennard C (2012) Fingermark detection and enhancement, 6th edn. National Centre for Forensic Studies, Canberra
Champod C (2016) Fingerprints and other ridge skin impressions. CRC Press, Taylor & Francis Group, Boca Raton
van Dam A, van Beek FT, Aalders MC, van Leeuwen TG, Lambrechts SA (2016) Techniques that acquire donor profiling information from fingermarks—a review. Sci Justice 56:143–154
Human Microbiome Project Consortium (2012) Structure, function and diversity of the healthy human microbiome. Nature 486:207–214
Costello EK, Lauber CL, Hamady M, Fierer N, Gordon JI, Knight R (2009) Bacterial community variation in human body habitats across space and time. Science 326:1694–1697
Gao Z, Tseng CH, Pei Z, Blaser MJ (2007) Molecular analysis of human forearm superficial skin bacterial biota. Proc Natl Acad Sci U S A 104:2927–2932
Fierer N, Hamady M, Lauber CL, Knight R (2008) The influence of sex, handedness, and washing on the diversity of hand surface bacteria. Proc Natl Acad Sci U S A 105:17994–17999
Grice EA, Kong HH, Renaud G, Young AC, Program NCS, Bouffard GG, Blakesley RW, Wolfsberg TG, Turner ML, Segre JA (2008) A diversity profile of the human skin microbiota. Genome Res 18:1043–1050
Song SJ, Lauber CL, Costello EK, Lozupone CA, Humphrey G, Berg-Lyons D, Caporaso JG, Knights D, Clemente JC, Nakielny S, Gordon JI, Fierer N, Knight R (2013) Cohabiting family members share microbiota with one another and with their dogs. eLife 2:e00458
Hospodsky D, Pickering AJ, Julian TR, Miller D, Gorthala S, Boehm AB, Peccia J (2014) Hand bacterial communities vary across two different human populations. Microbiology 160:1144–1152
Leung MH, Wilkins D, Lee PK (2015) Insights into the pan-microbiome: skin microbial communities of Chinese individuals differ from other racial groups. Sci Rep 5:11845
Lax S, Smith DP, Hampton-Marcell J, Owens SM, Handley KM, Scott NM, Gibbons SM, Larsen P, Shogan BD, Weiss S, Metcalf JL, Ursell LK, Vazquez-Baeza Y, Van Treuren W, Hasan NA, Gibson MK, Colwell R, Dantas G, Knight R, Gilbert JA (2014) Longitudinal analysis of microbial interaction between humans and the indoor environment. Science 345:1048–1052
Fierer N, Lauber CL, Zhou N, McDonald D, Costello EK, Knight R (2010) Forensic identification using skin bacterial communities. Proc Natl Acad Sci 107:6477–6481
Yao X, Liu W, Han J, Pei G, Tong Y, Luo Y (2016) Analysis of microbiome DNA on frequently touched items and from palm prints. J Forensic Sci Med 2:74–77
Meadow JF, Altrichter AE, Green JL (2014) PeerJ 2:e477. https://doi.org/10.7717/peerj.7447
Lax S, Hampton-Marcell J, Gibbons SM, Colares GB, Smith D, Eisen JA, Gilbert JA (2015) Forensic analysis of the microbiome of phones and shoes. Microbiome 3:21
Lee S, Woo S, Choi G, Hong Y, Eom Y (2015) Microbial forensic analysis of bacterial fingerprint by sequence comparison of 16S rRNA gene. J Forensic Res 6:297
Schmedes SE, Woerner AE, Budowle B (2017) Forensic human identification using skin microbiomes. Appl Environ Microbiol 83:e01672–e01617
Lee S, Woo S, Lee S, Eom Y (2016) Forensic analysis using microbial community between skin bacteria and fabrics. Toxicol Environ Heal Sci 8:263–270
Turner S, Pryer KM, Miao VP, Palmer JD (1999) Investigating deep phylogenetic relationships among cyanobacteria and Plastids by small subunit rRNA sequence analysis. J Eukaryot Microbiol 46:327–338
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Gonzalez Pena A, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Lozupone CA, Hamady M, Kelley ST, Knight R (2007) Quantitative and qualitative diversity measures lead to different insights into factors that structure microbial communities. Appl Environ Microbiol 73:1576–1585
Segata N, Izard J, Waldron L, Geveres D, Miropolsky L, Garrett WS, Huttenhower C (2011) Metagenomic biomarker discovery and explanation. Genome Biol 12:R60
Multinu F, Harrington SC, Chen J, Jeraldo PR, Johnson S, Chia N, Walther-Antonio MR (2018) Systematic bias introduced by genomic DNA template dilution in 16S rRNA gene-targeted microbiota profiling in human stool homogenates. mSphere 3:e00560–e00517
Cosseau C, Romano-Bertrand S, Duplan H, Lucas O, Ingrassia I, Pigasse C, Roques C, Jumas-Bilak E (2016) Proteobacteria from the human skin microbiota: species-level diversity and hypotheses. One Health 2:33–41
Bobulsky GS, Al-Nassir WN, Riggs MM, Sethi AK, Donskey CJ (2008) Clostridium difficile skin contamination in patients with C. difficile-associated disease. Clin Infect Dis 46:447–450
Budowle B, Murch R, Chakraborty R (2005) Microbial forensics: the next forensic challenge. Int J Legal Med 119:317–330
Clarke TH, Gomez A, Singh H, Nelson KE, Brinkac L (2017) Integrating the microbiome as a resource in the forensics toolkit. Forensic Sci Int Genet 30:141–147
Gunn A, Pitt SJ (2012) Microbes as forensic indicators. Trop Biomed 29:311–330
Hampton-Marcell J, Lopez JV, Gilbert JA (2017) The human microbiome: an emerging tool in forensics. Microb Biotechnol 10:228–230
Leake SL (2013) Is human DNA enough?—potential for bacterial DNA. Front Genet 4:1–3
Tidico SR, Murray DC, Addison J, Kirkbride KP, Bunce M (2014) Metagenomic analyses of bacteria on human hairs: a qualitative assessment for applications in forensic science. Investig Genet 5:16
Tano K, von Essen R, Eriksson P-O, Sjostedt A (2008) Alloiococcus otitidis-otitis media pathogen or normal bacterial flora? Acta Pathol Microbiol Immunol Scand 116:785–790
Yoshiura K, Kinoshita A, Ishida T, Ninokata A, Ishikawa T, Kaname T, Bannai M, Tokunga K, Sonoda S, Komaki R, Ihara M, Saenko VA, Alipov GK, Sekine I, Komatsu K, Takahashi H, Nakashima M, Sosonkina N, Mapendano CK, Ghadami M, Nomura M, Liang DS, Miwa N, Kim DK, Garidkhuu A, Natsume N, Ohta T, Tomita H, Kaneko A, Kikuchi M, Russomando G, Hirayama K, Ishibashi M, Takahashi A, Saito N, Murray JC, Saito S, Nakamura Y, Niikawa N (2006) A SNP in the ABCC11 gene is the determinant of human earwax type. Nat Genet 38:324–330
Sumikawa Y, Takahashi H, Sumikawa M, Kusatake K, Kaneko S, Yamashita T, Morita E (2013) J Clin Exp Dermatol Res 4:100193
Luebberding S, Krueger N, Kerscher M (2013) Skin physiology in men and women:in vivoevaluation of 300 people including TEWL, SC hydration, sebum content and skin surface pH. Int J Cosmet Sci 35:477–483
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
All subjects were made aware of the nature of the study and gave written consent before any information gathering or sampling occurred. This study has been approved by the University of Technology Sydney Human Research Ethics committee (UTS HREC approval #2015000296).
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
The following files are available online: Table S1 – Bacterial groups identified as potential biomarkers using the LefSe pipeline; Table S2 – Association between Factors; Table S3 – Participant Intrinsic and Extrinsic Factors.
ESM 1
(DOCX 47 kb)
Rights and permissions
About this article
Cite this article
Phan, K., Barash, M., Spindler, X. et al. Retrieving forensic information about the donor through bacterial profiling. Int J Legal Med 134, 21–29 (2020). https://doi.org/10.1007/s00414-019-02069-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00414-019-02069-2