Abstract
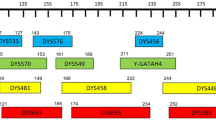
The PowerPlex® Y 23 System (Promega) is a short tandem repeat (STR) multiplex that allows co-amplification of 23 gonosomal Y-STRs, combining 17 loci commonly included in commercially available kits (DYS389I, DYS448, DYS389II, DYS19, DYS391, DYS438, DYS437, DYS635, DYS390, DYS439, DYS392, DYS393, DYS458, DYS385a/b, DYS456, and Y-GATA-H4) and six new loci (DYS481, DYS549, DYS533, DYS643, DYS576, and DYS570) with the last two being rapidly mutating Y-STRs (RM Y-STRs). In order to assess the possible gain in forensic efficiency provided by the six additional markers, a population sample of 410 unrelated healthy males originating from Northeast Italy (Veneto, Trentino Alto Adige, Lombardia, and Friuli Venezia Giulia regions) was typed. The data (335 of the 410 samples) are available in the Y chromosome haplotype reference database under accession number YA003327. Overall, 410 unique haplotypes were found corresponding to a global haplotype diversity (HD) of 0.999994 with a discriminatory capacity (DC) of 100 %. Allelic microvariants, null alleles, and duplications were detected. Pairwise genetic distances (R ST) calculated among neighboring European reference populations revealed no significant differences. Furthermore, for studying Y-STR mutation rates, 90 father-son pairs, in which the fathers were already included in the full dataset, were tested. On a total of 2,070 meioses considered, eight single-step mutational events were observed, two of which within the same father-son pair and the average mutation rate was 3.38 × 10−3 per locus per generation (95 % confidence interval, 1.36 × 10−3–6.95 × 10−3).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Analysis of short tandem repeats from the non-recombining region of the Y-chromosome is a powerful tool in forensic genetics to establish the paternal lineage, to discriminate the male component of DNA mixed stains (especially in rape cases) and in population genetics for evolutionary studies [1, 2]. Recently, a new multiplex kit, the PowerPlex® Y 23 System (PPY23, Promega, Madison, WI), that allows co-amplification of 23 Y-STRs has been developed and validated [3]. This kit includes a panel of 17 Y-STRs routinely used in forensic genetic analysis comprising 9 Y-STR markers of the European minimal haplotype (minHt), two loci recommended by the SWGDAM, six of the extended haplotype (ExtHt), and six additional new markers (DYS481, DYS549, DYS533, DYS643, DYS576, and DYS570) two of which have termed rapidly mutating Y-STRs [4]. In order to evaluate the potential suitability in forensic genetics of these six additional new markers, here, we reported allelic frequencies and basic forensic parameters from a population sample of 410 healthy males, carefully selected based on unrelated pedigrees, different surnames, and genealogical data of residence in the sampling area of Northeast Italy (Veneto, Trentino Alto Adige, Lombardia and Friuli Venezia Giulia) with at least three generations, after obtaining written informed consent. Moreover, 90 father-son pairs in which the fathers were already included in the full dataset were also studied to evaluate mutation rates useful for the interpretation of data in kinship analysis [5]. Each biological relationship was confirmed by autosomal STRs typing using various panels of DNA markers (AmpFl STR Identifiler™ kit, PowerPlex® ESX and ESI System, and PowerPlex® Fusion System) with a paternity probability >99.99 %.
Genomic DNA was extracted from buccal swabs with the QIAamp DNA Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instruction. The quantity of recovered DNA was determined using Quantifiler® Duo DNA Quantification Kit (Applied Biosystems) on the 7500 Real-Time PCR System with HID Analysis software v1.1. The samples were amplified using PowerPlex® Y 23 System (Promega), following the manufacturer’s instructions. PCR products were detected by capillary electrophoresis in an ABI Prism 3130 Genetic Analyzer (Applied Biosystems). Data analysis and genotyping were automatically assigned by GeneMapper® ID-X Software v.1.2 (Applied Biosystems).
Allele and haplotype frequencies were calculated by counting method. Gene Diversity (GD) was calculated using the formula GD = n(1 − ∑p i 2)/(N − 1) where N is the sample size and p i is the allelic frequency. Haplotype Diversity (HD) was computed with the same equation using haplotype frequencies instead of allele frequencies. Statistical calculations of standard diversity indices were performed with the Arlequin v. 3.5 software [6]. Discriminatory capacity (DC) was determined by dividing the number of different haplotypes by the total number of samples in a given population. The haplotype match probability (HMP) was calculated as HMP = 1 − HD [7].
In order to examine the genetic relationships between the Y-chromosomal landscape of Northeast Italy and neighboring European reference populations (Switzerland [YP000395, n = 138], Germany [YP000148, n = 444], Austria [YP000217, n = 259], Hungary [YP000061, n = 202], Bosnia and Herzegovina [YP000824, n = 200], Croatia [YP000916, n = 1489], Slovenia [YP000217, n = 257]) pairwise genetic distances (R ST) and associated probability values (p values, 10,000 permutations) were determined using the analysis of molecular variance (AMOVA) tool of the YHRD website (www.yhrd.org) [8]. The selection of reference populations was done to evaluate the presence of any shared male heritage since, at the beginning of the nineteenth century, these were all part of the Austro-Hungarian Empire held by the Hapsburg Monarchy. Their empire extended from actual Ukraine and Belarus to Northeast Italy, and comprised the population sample here studied. The R ST calculations were performed excluding the DYS385a/b marker, samples showing duplications, null, and microvariants alleles and tested for significance using 10,000 permutations. Moreover, to reduce the probability of falsely rejecting the null hypothesis (type I error), an a priori significance value (α) was determined by applying the Bonferroni’s correction (α = 0.0018) [9]. The genetic distances were subsequently used to generate multi-dimensional scaling (MDS) plot.
Mutation rates and their exact confidence intervals (CIs) from binomial probability distribution were calculated using the StatCalc v.3.0 software (http://www.ucs.louisiana.edu/~kxk4695/StatCalc.htm). Furthermore, to determine if the fathers’ age at the time of son’s birth could significantly influence the occurrence of mutational events, a Mann–Whitney U test was performed using a web-based tool available at http://scistatcalc.blogspot.it/2013/10/Mann–Whitney-u-test-calculator-work-in.html. Our Y-STR data (335 of the 410 samples) were submitted to the YHRD after passing the quality control requirements for population genetic data and an accession number for the sample was received (YA003327) [10].
No typing differences were detected in the samples previously analyzed with the Yfiler kit [11] and the PPY23 used in this study; however, novel intermediate alleles, copy number variations, and null alleles were revealed. Thirteen rare microvariant alleles not included in the bin set of the allelic ladder were observed at loci DYS458 (16.2, 17.2, 18.2, 19.2, and 22.2) and DYS385a/b (17.2). Within the six additional PPY23-specific loci, one intermediate allele (22.2) at locus DYS481 was here found for the first time. All the microvariants were confirmed by repeating the amplification and genotype process. Copy number variations such as duplications were detected at DYS19 (n = 3), DYS635 (n = 1) loci, and in two of the six PPY23-specific markers specifically at DYS481 (n = 1) and DYS549 (n = 1). Based on our observations, the average duplication rate across the 23 Y-STRs was 6.36 × 10−4. However, in our opinion, such rate is likely to be underestimated since duplications yielding homoallelic combinations (no length differences between the two alleles) may remain undetected in the electropherogram in comparison to heteroallelic combinations. Null alleles at DYS448 and DYS458 were observed as singletons. Also, at one of the six PPY23-specific markers (DYS549), an additional null allele was found. The sequencing performed with external primers spanning the entire region of the locus DYS549 showed a point mutation presumably located in the reverse primer binding site [12].
With regard to forensic parameters, allelic frequencies and gene diversity values for PPY23 System are shown in Table 1 of Electronic Supplementary Material (ESM). It was possible to establish that the haplotype diversity calculated for PPY23 was of 0.999994 and the corresponding HMP was of 0.000006, values significantly higher compared to the 17 Y-STRs of the Yfiler kit [11]. Evidently, such an increase in HD and HMP values could be reliably ascribed to the presence, within the PPY23 System, of the additional new markers. This was remarked by the evidence that the six markers alone provided a HD value of 0.998982, slightly higher than that obtained by nine Y-STRs of the minimal haplotype (HD = 0.998316). On the other hand, in our sample, 410 unique haplotypes (UH) were detected; therefore, the six loci in PPY23 accounted for an increased DC value over other Y-STR panels that raised from 79.02 % minHt loci to 98.70 % for Yfiler kit, up to 100 % for the PPY23 System. It should however be pointed out that, within our carefully selected male population sample in a few cases, the haplotypes differed at only one marker, specifically Y-GATA H4 or DYS439 as shown in ESM Table 2.
Examination of pairwise genetic distances (R ST) calculated, among all anciently comprised Austro-Hungarian Empire populations (as shown in ESM Table 3 and ESM Fig. 1), revealed that the Northeast Italy sample tightly clustered with Austria and Germany, while appeared more distant to Croatian and Bosnian, with Switzerland, Slovenia, and Hungary locating within an intermediate distance between the two groups. These findings resulted in line with previous studies, based on both Y-STR and Y-SNPs haplogroup distribution patterns [13, 14] that showed the relative genetic similarity among Northeast Italy, North Croatian, and Slovenian Littoral region populations, while appearing more distant to Central and Southern Croatian as well as Bosnian.
The mutation rate study performed on a total of 2,070 meiotic events showed that the overall average mutation rate estimated for the PP23Y System was 3.38 × 10−3 (95 % CI 1.36 × 10−3–6.95 × 10−3). Eight single-step mutational events were found with the 23 Y-STR loci in PP23Y of which five revealed in this study and three previously reported at DYS389I/II, DYS456, and DYS458 loci [11] (ESM Table 4a-b).
If the interest of the forensic community toward RM Y-STRs hinge on the potential that these markers offer in distinguishing between male relatives of the same family tree, it should also be emphasized their inadequacy in the establishment of paternity. Therefore, with the aim of a combined use of Y-STR and RM Y-STRs, as in the PP23Y System, in which the mutation rate is also increased by the RM Y-STR component, in our opinion, it would seem appropriate to revise exclusion criteria for paternity testing. Previously, Ballantyne et al. [4] indicated that, with regard to the sole RM Y-STR set, the limit of three mutations is not sufficient for paternity exclusion (criterion evaluated for autosomal STR [15]). Accordingly, we do not believe that the analysis of Y-STR markers with different intrinsic characteristics (especially with different mutation rates) should be associated within the same biostatistic calculation for paternity testing purposes.
In addition, it was then examined whether the father's age, at the time of son’s birth, may influence the occurrence of a mutational event. The age of the seven fathers carrying a mutation was comprised between 20–70 years (average 35.57 ± 16.02 SD) while for the other 82 fathers without mutations ranged between 17–70 years (average 34.19 ± 10.92 SD). A nonparametric test (Mann–Whitney U test) showed no significant differences between the age of subjects in the two groups (p = 0.7568).
From our results, it could be evinced that the PowerPlex® Y 23 System proved to be extremely discriminatory and these additional loci had a positive significant impact on the global forensic parameters, increasing PPY23 HD and DC values in comparison to other multiplex Y-STR sets, which is reflected beneficially in forensic casework through a reduction of the chance of unrelated male subjects sharing the same haplotype. However, considering their mutation rate, we would suggest the adoption of a careful approach in kinship testing, in order to avoid misinterpretation of results.
This paper follows the guidelines for publication of population data requested by the journal [16].
References
Roewer L (2009) Y chromosome STR typing in crime casework. Forensic Sci Med Pathol 5:77–84
Jobling MA, Tyler-Smith C (2003) The human Y chromosome: an evolutionary marker comes of age. Nat Rev Genet 4:598–612
Thompson JM, Ewing MM, Frank WE, Pogemiller JJ, Nolde CA, Koehler DJ, Shaffer AM, Rabbach DR, Fulmer PM, Sprecher CJ, Storts DR (2013) Developmental validation of the PowerPlex® Y23 System: a single multiplex Y-STR analysis system for casework and database samples. Forensic Sci Int Genet 7:240–250
Ballantyne KN, Keerl V, Wollstein A, Choi Y, Zuniga SB, Ralf A, Vermeulen M, de Knijff P, Kayser M (2012) A new future of forensic Y-chromosome analysis: rapidly mutating Y-STRs for differentiating male relatives and paternal lineages. Forensic Sci Int Genet 6:208–218
Kayser M, Roewer L, Hedman M, Henke L, Henke J, Brauer S, Krüger C, Krawczak M, Nagy M, Dobosz T, Szibor R, de Knijff P, Stoneking M, Sajantila A (2000) Characteristics and frequency of germline mutations at microsatellite loci from the human Y chromosome, as revealed by direct observation in father/son pairs. Am J Hum Genet 66:1580–1588
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Res 10:564–567
Bosch E, Lee AC, Calafell F, Arroyo E, Henneman P, de Knijff P, Jobling MA (2002) High resolution Y chromosome typing: 19 STRs amplified in three multiplex reactions. Forensic Sci Int 125:42–51
Willuweit S, Roewer L, International Forensic Y Chromosome User Group (2007) Y chromosome haplotype reference database (YHRD): update. Forensic Sci Int Genet 1:83–87
Bonferroni CE (1936) Teoria statistica delle classi e calcolo delle probabilità. Pubbl Ist Super Sci EconCommer Firenze 8:3–62
Purps J, Siegert S, Willuweit S, Nagy M, Alves C et al (2014) A global analysis of Y-chromosomal haplotype diversity for 23 STR loci. Forensic Sci Int Genet 28(12C):12–23
Turrina S, Atzei R, De Leo D (2006) Y-chromosomal STR haplotypes in a Northeast Italian population sample using 17plex loci PCR assay. Int J Legal Med 120:56–59
Turrina S, Caratti S, De Leo D (2013) Powerplex® Y 23 System: molecular characterization of a null allele at locus DYS549. Forensic Sci Int Genet Suppl Series 4:e117–e118
Mršić G, Gršković B, Vrdoljak A, Vrdoljak A, Popović M, Valpotić I, Anđelinović Š, Stenzl V, Ehler E, Urban L, Lacković G, Underhill P, Primorac D (2012) Croatian national reference Y-STR haplotype database. Mol Biol Rep 39:7727–7741
Zupan A, Vrabec K, Glavač D (2013) The paternal perspective of the Slovenian population and its relationship with other populations. Ann Hum Biol 40:515–526
Gjertson DW, Brenner CH, Baur MP, Carracedo A, Guidet F, Luque JA, Lessig R, Mayr WR, Pascali VL, Prinz M, Schneider PM, Morling N (2007) ISFG: recommendations on biostatistics in paternity testing. Forensic Sci Int Genet 1:223–231
Poetsch M, Bajanowski T, Pfeiffer H (2012) The publication of population genetic data in the International Journal of Legal Medicine: guidelines. Int J Legal Med 126:489–490
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Turrina, S., Caratti, S., Ferrian, M. et al. Haplotype data and mutation rates for the 23 Y-STR loci of PowerPlex® Y 23 System in a Northeast Italian population sample. Int J Legal Med 129, 725–728 (2015). https://doi.org/10.1007/s00414-014-1053-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00414-014-1053-6