Abstract
In Drosophila melanogaster, the chromatin of interphase polytene chromosomes appears as alternating decondensed interbands and dense black or thin gray bands. Recently, we uncovered four principle chromatin states (4НММ model) in the fruit fly, and these were matched to the structures observed in polytene chromosomes. Ruby/malachite chromatin states form black bands containing developmental genes, whereas aquamarine chromatin corresponds to interbands enriched with 5′ regions of ubiquitously expressed genes. Lazurite chromatin supposedly forms faint gray bands and encompasses the bodies of housekeeping genes. In this report, we test this idea using the X chromosome as the model and MSL1 as a protein marker of the lazurite chromatin. Our bioinformatic analysis indicates that in the X chromosome, it is only the lazurite chromatin that is simultaneously enriched for the proteins and histone marks associated with exons, transcription elongation, and dosage compensation. As a result of FISH and EM mapping of a dosage compensation complex subunit, MSL1, we for the first time provide direct evidence that lazurite chromatin forms faint gray bands. Our analysis proves that overall most of housekeeping genes typically span from the interbands (5′ region of the gene) to the gray band (gene body). More rarely, active lazurite chromatin and inactive malachite/ruby chromatin may be found within a common band, where both the housekeeping and the developmental genes reside together.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Drosophila salivary gland polytene chromosomes represent one of the most convenient experimental models of interphase chromosomes (reviewed in Zhimulev et al. 2004). Besides their obvious giant size, these chromosomes display a very fine banding pattern that is composed of black and gray bands with light loosely compacted interbands found in between (Bridges 1935, 1938). These structures are probably best visualized at the level of electron microscopy (EM) and can be distinguished by a number of features. For instance, in contrast to black bands, the gray bands replicate early, display loosely condensed chromatin and high density of genes, most of which are classified as housekeeping (Zhimulev 1999; Zhimulev et al. 1982, 2004, 2014; Kozlova et al. 1994; Belyakin et al. 2005; Belyaeva et al. 2012; Kolesnikova et al. 2018).
Comprehensive studies of protein composition and genetic organization of the Drosophila genome have paved the way to our better understanding of the interplay between different chromatin states and the structural and functional organization of different regions of the Drosophila interphase chromatin. Presently, four major chromatin classifications have been put forward, with individual chromatin types each represented by a unique combination of proteins, and displaying functional specificity, as well as characteristic genomic position. (i) Using DamID of 53 chromatin proteins, five major chromatin states were established to exist in the Drosophila genome (Filion et al. 2010). In this model, transcriptionally active chromatin is represented by the RED and YELLOW types. Notably, YELLOW chromatin is enriched with housekeeping genes and histone mark H3K36me3, a histone modification associated with transcription elongation and localized in gene bodies, predominantly in exons (Kolasinska-Zwierz et al. 2009; Schwartz et al. 2009; Wagner and Carpenter 2012). (ii) Whole-genome analysis of the distribution of 18 histone modifications in the chromatin of two Drosophila male cell lines has uncovered 9 chromatin states (Kharchenko et al. 2011). Active chromatin here is represented by the states 1 and 2, and in the context of the X chromosome, by the state 5 that marks the regions associated with dosage compensation machinery (see below). (iii) One more chromatin classification model is based on the differential sensitivity of chromatin to DNAse I and broadly classifies the chromatin into open, neutral, and closed (Milon et al. 2014). (iv) It has long remained unclear how the molecular and genetic organization of the genome relates to the morphological structures found in polytene chromosomes. This was largely due to the inability to accurately locate the band/interband borders at the molecular map. One approach to address this issue was developed in the late 1980s and was geared towards identification of DNA sequences that flanked the insertions of P-element-based transposons that have landed into interbands (Semeshin et al. 1989). Combined with the genome-wide chromatin profiling data, this has allowed identification of a set of non-histone proteins specific for the interbands (Vatolina et al. 2011; Demakov et al. 2011). Using this protein ensemble as a marker, we developed a bioninformatic pipeline—four chromatin state models (4HMM) that classify the chromatin into four basic chromatin states referred to as aquamarine, lazurite, malachite, and ruby (Zhimulev et al. 2014; Boldyreva et al. 2017). It was for the first time that chromatin states having distinct degrees of condensation and transcription competence have been matched to the banding pattern of polytene chromosomes. This in turn has allowed us to propose the idea that positions of such chromatin types along the physical map closely correspond to the chromomeres of interphase chromosomes (Zhimulev et al. 2014). The key finding was that aquamarine fragments enriched with H3K4me and chromo domain protein interacting with Z4 (CHRIZ) (in the four cell lines analyzed) and containing transcription start sites (TSSs) of ubiquitously active genes correspond to the interbands of polytene chromosomes (Zhimulev et al. 2014; Boldyreva et al. 2017). Further, ruby/malachite chromatin was shown to form black bands harboring developmentally regulated genes and corresponding to the silent chromatin domains identified across different cell types (Zhimulev et al. 1982; Belyakin et al. 2005; Belyaeva et al. 2012; Khoroshko et al. 2016; Kolesnikova et al. 2018).
Lazurite chromatin likely corresponds to the thin gray bands and encompasses the coding parts of housekeeping genes, whose 5′-ends are found in the adjacent interbands (Zhimulev et al. 2014; Demakova et al. 2016; Zykova et al. 2018). Yet, this has not been explicitly demonstrated. Existing data do not provide an unequivocal evidence, as they either do not relate the cytology data with molecular map (Semeshin et al. 2002, 2003; Kotlikova et al. 2006) or rely on light microscopy (LM) (Zeilke et al. 2015; Demakova et al. 2016), which is insufficient to accurately localize relatively short lazurite fragments. A combined approach is needed that would take into account both the molecular and genetic makeup of the chromosome region, and its mapping at electron microscope (EM)-level resolution.
For this reason, in the present work, we performed mapping of lazurite fragments using not only LM-FISH but also EM-based immunodetection of the lazurite chromatin marker. On a genome-wide scale, lazurite chromatin contains gene bodies and 3′-untranslated terminal regions (3′-UTR); it is also enriched with histone modification H3K36me3 (Boldyreva et al. 2017). We hypothesized dosage compensation complex (DCC) proteins may serve as possible markers of lazurite chromatin, given that DCC targets are largely housekeeping genes, and that DCC predominantly associates with coding sequences marked with H3K36me3 (Alekseyenko et al. 2006; Philip and Stenberg 2013). DCC functions to provide a twofold enhancement of transcription in a single chromosome X of the Drosophila males and is composed of five protein subunits (MSL1, MSL2, MSL3, MOF, MLE) and two ncRNAs (roX1 and roX2) (rev. in Lucchesi and Kuroda 2015). It binds to numerous sites along the male X chromosome leading to H4K16 acetylation, resulting in the relaxation of the chromatin folding and transcription upregulation (Shogren-Knaak and Peterson 2006; Robinson et al. 2008). Current model (Kuroda et al. 2016) postulates that the strongest DCC peaks correspond to so-called chromatin entry sites (CESes) (Kelley et al. 1999; Alekseyenko et al. 2008) or “high-affinity sites” (HASs) (Straub and Becker 2008; Straub et al. 2013), wherein the complex robustly binds the X chromosome in a sequence-dependent manner. Finally, DCC spreads to the nearby H3K36me3-marked bodies of active genes (Larschan et al. 2007; Gorchakov et al. 2009) and boosts transcriptional elongation (rev. in Kuroda et al. 2016).
In the present study using the 4HMM model, we show that much as was observed for the male cell line S2, only lazurite chromatin (gene bodies) of the X chromosome is simultaneously enriched with DCC, particularly MSL1, and with histone marks associated with dosage compensation (DC), gene bodies, and transcription elongation. With this in mind, we decided to restrict our analysis to the X chromosome and to use MSL1 as the marker of lazurite chromatin. Since DCC binding pattern is largely invariant both in cell lines (Alekseyenko et al. 2006) and in polytene chromosomes (Sass et al. 2003; Kotlikova et al. 2006), we expect MSL1 to serve as a good marker of lazurite chromatin for EM immunodetection in the polytene X chromosome, as well.
We show here, that lazurite chromatin forms thin gray bands. From the gene regulation standpoint, this indicates that housekeeping genes are hosted by two morphological structures, i.e., interbands (5′ gene region) and gray bands (gene body). Significantly less frequently, lazurite chromatin forms the edges of dark black bands. In such cases, lazurite chromatin harbors housekeeping genes that have their 3′-ends oriented towards the inactive malachite/ruby part of the band.
Materials and methods
Fly strains and genetic crosses
Flies were raised on standard cornmeal-yeast-agar-molasses medium. Descriptions of all mutants and rearrangements can be found in FlyBase (www.flybase.org). Transgenic w; msl3 [w1; H83M2-61]/TM6, Tb (NOPU) and w; msl2 cn [w+M2SXB1-2]/TM6, Tb stocks with different levels of ectopic MSL2 production were described in Kelley et al. (1995), Semeshin et al. (2002), and Demakova et al. (2003). The Suppressor of Underreplication (SuURES) mutation was originally detected in the In(1)scV2 stock (Belyaeva et al. 1998). w, ru h SuURES stock was constructed in our laboratory; w, ru h SuURES female larvae were used for fluorescence in situ hybridization (FISH) mapping experiments. These mutants lack underreplication in polytene chromosomes, yet display normal banding pattern, which translates into chromosome squashes of exquisite morphology. w, ru h SuURES female larvae were grown at 25 °C in uncrowded vials on standard fly medium.
Light microscopy methods
Fluorescence in situ hybridization
Salivary glands were dissected in Ephrussi–Beadle solution, fixed in 3:1 ethanol/acetic acid mixture for about 1 h at – 20 °C, squashed in 45% acetic acid, snap frozen in liquid nitrogen and stored in 70% ethanol at – 20 °C. FISH on polytene chromosomes was performed as described (Ashburner et al. 2005). Random-primed labeling of 22 DNA probes (selected according to the four chromatin states) with TAMRA or fluorescein (Biosan, Russia) was done using Klenow enzyme. All the probes (probes from bands and interbands predicted by the four chromatin state model) used in this study are described in Table S1. FISH preparations were analyzed under the Olympus BX50 fluorescence microscope.
Immunostaining of polytene chromosomes
Salivary glands were dissected from third instar larvae reared at 18 or 25 °C. The procedure for chromosome squashing and immunostaining was done as described previously (Kolesnikova et al. 2013) with minor modifications. Dilutions of the primary antibodies/antisera were as follows: mouse monoclonal anti-CHRIZ (DSHB#6H11-4ES; 1:200), rabbit polyclonal anti-MSL1 (provided by Prof M. Kuroda, 1:150). Chromosome squashes were incubated with secondary anti-rabbit and anti-mouse IgG-specific conjugates (Thermo scientific, Alexa Fluor 488 or Alexa Fluor 568; 1:800). Chromosomes were examined using epifluorescence optics (Olympus BX50 microscope) and photographed with CCD Olympus DP50. In each experiment, 50 ± 100 nuclei with well-spread chromosomes were examined on several slides. Representative cases of cytological localization patterns are shown on the figures in the manuscript in the “Results” section.
Combined immunostaining and FISH on polytene сhromosomes
First, the procedure for chromosome squashing and immunostaining was done as described above. The slides were examined using epifluorescence optics and photographed; coordinates of the nuclei were recorded. Then, the same slide was processed for FISH, as described above. Two DNA probes were selected according to the four chromatin states model (4HMM) and the localization sites for CHRIZ and MSL1 proteins on polytene chromosomes. Labeling of DNA probes with Tamra (Biosan, Russia) was done using Klenow enzyme. The probes are described in the Table S1. FISH preparations were analyzed using fluorescence microscopy and photographed. The results of immunostaining and FISH for the same chromosomes were merged.
Electron microscopy methods
Electron microscopy slides
Salivary gland polytene chromosome squashes were prepared for electron microscopy analysis and examined as described earlier (Semeshin et al. 1979). The sections (120–150 nm) were cut using an LKB-IV ultratome (LKB Bromma, Sweden) and examined under a JEM-100C (JEOL, Japan) electron microscope at 80 kV.
Electron microscopy immunostaining
For these experiments, the transgenic w; msl3 [w1; H83M2-61]/TM6, Tb females were used, since their two X chromosomes in most of the regions demonstrate the DCC pattern resembling that of the wild-type males. Primary affinity-purified rabbit anti-MSL1 antibodies were used at a dilution of 1:50. Immunostaining with gold-labeled antibodies was as in Semeshin et al. (2002). Then, chromosomes were treated with Silver Enhancement reagent (Boehringer Mannheim, Indianapolis) for 20 min to increase the size of gold particles. Thereafter, slides were thoroughly washed in PBT and double-distilled water, dehydrated in a graded ethanol series (20, 35, 50, and 70%) for 5 min in each, and left overnight in a 1.5% solution of uranyl acetate in 70% ethanol for staining. Further procedures of dehydration and embedding in epoxy resin have been described elsewhere (Semeshin et al. 1998). Ultra-thin sections were examined using JEM-100C electron microscope at 80 kV. To localize chromosome regions, we referred to the revised cytological maps of polytene chromosomes by Bridges (reproduced in Lindsley and Zimm 1992), Saura et al. (1993), and our own EM mapping data.
Bioinformatics methods
Data sources and common bioinformatics methods
FlyBase database BDGP R5/dm3 was used to retrieve chromosomal positions of the genes, transcripts, and structural parts analyzed in the study (www.flybase.org). The modENCODE ChIP-chip datasets (Roy et al. 2010) were extracted via the InterMine (Smith et al. 2012). R programming 3.4.4 version and Bioconductor 3.8 version libraries were used to analyze and visualize the data (http://www.R-project.org/ R Development Core Team 2015; Gentleman et al. 2004; Lawrence et al. 2009, 2013): GenomicRanges, IRanges, dplyr, gplot2.
Comparative analysis of MSL1, JIL1, and H3K36me3 enrichment levels in the four chromatin states
In order to measure the coverage of various chromatin types by the chromatin marks MSL1, JIL1, and H3K36me3 (according to modENCODE datasets: male S2 cell line -modENCODE_3293, modENCODE_3038 modENCODE_303 and female Kc cell line modENCODE_3037, modENCODE_302 was taken for the 5st model (Filion et al. 2010)), we calculated the ratio of overlap between an individual region and the mark, to the average coverage over the X chromosome. The percentage of coverage was calculated as the ratio of the total length of the desired protein enrichment regions within the each chromatin type to the total length of each chromatin type over the X chromosome. The significance of differences between the chromatin types was estimated using Mann–Whitney U test. To correct p values in multiple comparisons, the Holm–Bonferroni method was used. Random permutation test for 9999 iterations was also performed as described before (Boldyreva et al. 2017) with the same data. Differences were considered significant at p value < 0.001.
Overlap between the four and nine chromatin states (4st vs 9st models)
The contribution of each of the desired chromatin states (Kharchenko et al. 2011) to the four chromatin states was defined as a sum of the lengths of chromatin types overlap (9st vs 4st and vice versa) normalized to the total DNA length of particular chromatin state excluding the length of the internal data gaps.
Mapping MSL1, H3K36me3, and JIL1 to the structural parts of genes of the male X chromosome in 4НММ chromatin types
We used in analysis three described above S2 cell line datasets for MSL1, H3K36me3, and JIL1 from ModENCODE database. For each dataset, the percentage of coverage was measured as the ratio of the total length of transcripts overlapping with the protein enrichment regions of interest, to the total length of protein enrichment regions over the X chromosome. For each dataset we compiled two subsets of genomics regions that encompassed (i) 5′- regions of transcripts (-500 bp upstream TSS and entire 5′UTR) and (ii) coding sequences (CSs) defined as regions of transcripts between the translation start and stop. Also, each dataset was overlaid with the 4-chromatin types (Zhimulev et al. 2014), i.e. for 5′-regions and CSs 4 × 2 = 8 additional subsets for each protein were compiled. Thus, in total, 10 subsets were formed. For each of these three datasets (MSL1, H3K36me3, and JIL1), on the one hand, as well as for each of the 10 subsets above on the other hand, we applied the full-genome permutation test as we described before (Boldyreva et al. 2017).Whenever significant enrichment or depletion of the overlap between the two track analyzed, p value is calculated, which corresponds to the probability of a random overlap above/below the value observed in reality.
Results
MSL1, H3K36me3, and JIL1 proteins are abundant in the lazurite chromatin
We asked whether MSL1 is enriched in the lazurite chromatin of the X chromosome, compared with other chromatin types. Also, we cross-compared the MSL1-enrichment regions (areas of confidence) with data of other models of chromatin clasterization in S2 cells (Kharchenko et al. 2011; Milon et al. 2014). Similar analyses were done for JIL1 and H3K36me3 that are shown to be abundant in the actively transcribed gene bodies (Kharchenko et al. 2011).
Previously, we localized MSL1 among the open chromatin markers in male polytene X chromosome (Demakova et al. 2003). In genomic scale, MSL1 was found enriched in the open aquamarine and lazurite chromatin (Zhimulev et al. 2014; Boldyreva et al. 2017). We focused here at the X chromosome-wide enrichment profiles of MSL1, JIL1, and H3K36me3 (significantly enriched regions as defined by modENCODE (Roy et al. 2010)) comparison with different chromatin types produced by 4HMM (Zhimulev et al. 2014), 5st (Filion et al. 2010), 9st S2 (Kharchenko et al. 2011), GCSDI 2, and GCSDI 3 models (Milon et al. 2014) (Fig. 1). To assess the significance of differences, Holm–Bonferroni method was used for running Wilcoxon test, and p values below 0.001 were considered statistically significant.
Overlap between the MSL1-, JIL1-, and H3K36me3-enriched regions (modENCODE (Roy et al. 2010)) with various chromatin states produced by 4HMM (Zhimulev et al. 2014), 5st (Filion et al. 2010), 9st (S2 cells) (Kharchenko et al. 2011), GCSDI 2, and GCSDI 3 (Milon et al. 2014) chromatin partitioning models, shown as the fraction of the cumulative length of each chromatin type. Y axis: degree of overlap for MSL1: 0, no overlap; 1, complete overlap. n* is the number of fragments of the given chromatin type that overlap with the protein-enriched region by at least one basepair. Boxplots: lines in the middle of the boxes indicate the medians of each distribution. Top of the boxes represent the 75th percentile. Bottom of the boxes represent the 25th percentile. Whiskers indicate the maximum or minimum, observation within 1.5 times of the box height from the top, or the bottom of the box, respectively at p < 0.001, based on the Wilcoxon U test. Circles represent outliers
This test indicated that both lazurite and aquamarine fragments of the X chromosome are significantly enriched with MSL1, compared with the two other chromatin types (malachite and ruby) (p < 2.2E-16). We observed a virtually complete overlap between lazurite chromatin and MSL1-bound regions in the X chromosome (median overlap value is 0.96), which is 1.8 times above the median overlap of aquamarine chromatin with MSL1-bound chromatin, this difference being highly significant (p value < 2.2E-16) (Fig. 1). Compared with the two other genome partitioning models (9st S2 (Kharchenko et al. 2011), GCSDI 2, and GCSDI 3 (Milon et al. 2014)), lazurite chromatin in our 4HMM model shows the strongest overlap with MSL1 in the X chromosome (p value < 0.001) (Fig. 1).
Localization of JIL1 in the X chromosome likewise shows extensive overlap with lazurite fragments. Median overlap between significantly enriched JIL1 regions with lazurite chromatin is 0.91, which is significantly above the median overlap with aquamarine chromatin (0.49) (p value < 2.2E-16). For the malachite and ruby chromatin of the X chromosome, the median fraction of overlap with JIL1-bound regions is zero. The observed extensive overlap between lazurite chromatin and JIL1 localization in the X chromosome is in contrast to the overlap of JIL1 with other chromatin types produced by other chromatin-classifying models (p value < 0.001) (Fig. 1), except for GREEN chromatin of the 5st model (Filion et al. 2010) (p value = 1) (this chromatin state having very few overlaps n* = 77) and state 1 chromatin of the 9st S2 model (Kharchenko et al. 2011) (p value = 1).
In the X chromosome, histone methylation mark H3K36me3 displays significant (p value < 0.001), albeit compared with MSL1 and JIL1, less extensive, overlap with lazurite fragments. The median overlap between H3K36me3 and lazurite fragments on the X chromosome is 0.47; this value being significantly higher than the degree of overlap with aquamarine, malachite, and ruby fragments of the X chromosome (p value < 2.2E-16) (Fig. 1). Similarly, this overlap within the territory of the X chromosome is the strongest, compared with other chromatin states produced by alternative chromatin classification models (p value < 0.001).
The 9st model of Kharchenko et al. (2011) namely chromatin state 5 is maximally presented in active genes in the male X (Fig. 1). For this reason, we performed an intersection of lazurite chromatin state (4HMM) with these 9 chromatins (Kharchenko et al. 2011) and reciprocally state 5 of 9st model (Kharchenko et al. 2011) against four states HMM (Zhimulev et al. 2014). The data obtained indicate that the degree of overlap between lazurite chromatin and the state 5-chromatin type is indeed very high (82%) (Fig. 2a) and is above the rest of the chromatin states by Kharchenko et al. (2011). In a reciprocal intersection, the state 5, although partially overlapping with the malachite chromatin (32%), still displays predominant association with the lazurite chromatin (55%). State 5 of 9st model areas overlap with aquamarine and ruby chromatin states by a mere of 6–7% (Fig. 2b).
Overlap of lazurite chromatin (4HMM model) with nine chromatin states (9st model, S2 cells) (a) and state 5 chromatin (9 state model, S2 cells) with four chromatin states (4HMM model) (b) in the chromosome X (as % of the DNA length). a The fraction of lazurite chromatin state overlapped by each of the nine chromatin types (Kharchenko et al. 2011). b The fraction of chromatin state 5 overlapped by each of the four chromatin states (Zhimulev et al. 2014)
MSL1, H3K36, and JIL1 are located in the structural parts of genes of the male X chromosome in the 4НММ chromatin types
In order to understand how MSL1, H3K36me3, and JIL1 enrichment regions are arranged relatively to the structural elements of genes on the male X chromosome (adjusted by the four chromatin states defined by the 4HMM), we calculated the fraction of transcripts covered by these protein binding sites (Fig. 3). For this purpose, 29,721 transcripts of 13,753 D. melanogaster protein-coding genes annotated in FlyBase r.5.57 were used. Of these, 5318 transcripts map to the X chromosome (FlyBase r.5.57 ftp://ftp.flybase.net/releases/FB2014_03/dmel_r5.57/gff/dmel-all-no-analysis-. r5.57.gff.gz, all annotation marked as “mRNA”).
We focused our analysis on two gene parts, namely the 5′ region spanning 500-bp upstream of the TSS and the entire 5′-UTR (-500TSS; AUG) (indicated as the TSS in Fig. 3) and the region from start codon to stop codon (labeled as the GENE BODY in Fig. 3).
Of the genomic fragments mapping within TSS, only MSL1- and JIL1-bound regions within aquamarine chromatin are enriched (p value < 1E-24, p value < 2E-20) (Fig. 3a, c). In contrast, GENE BODY shows enrichment with MSL1, JIL1, and H3K36me3 associated with the lazurite chromatin (p value < 4E-55, p value < 9E-52, and p value < 2E-27) (Fig. 3a–c), as well as with MSL1, JIL1, and H3K36me3 regions in general, regardless of their chromatin identity (p value < 3E-37, p value < 6E-33, and p value < 2E-21). GENE BODY regions also display moderate enrichment for malachite-overlapping MSL1 binding sites (p value < 4E-7) (Fig. 3a). TSS enrichment with H3K36me3 embedded within aquamarine chromatin (Fig. 3b) is not statistically different from the value observed for random fragments, as assessed by the whole-genome permutation test (p value < 0.04).
Thus, MSL1, JIL1, and H3K36me3 predominantly associate with the coding part of the genes (GENE BODY) embedded in lazurite chromatin; for the TSS part, MSL1 and JIL1 localize to the aquamarine chromatin.
Cytology mapping and protein profiling of the X chromosome region encompassing alternating lazurite and aquamarine chromatin
Previously, we proposed that active lazurite chromatin corresponds to the thin gray bands of polytene chromosomes (Zhimulev et al. 2014). We set out to test this idea for the large fragment of the X chromosome that is exclusively represented by alternating aquamarine/lazurite chromatin and appears to correspond to a series of thin gray bands on the polytene chromosome map. Further, we aimed at describing the composition of the associated proteins, histone marks, and genomic elements (according to the published data). The region selected for our analysis (chrX: 16,163,665-16,352,204) corresponds to the cytological location 14B-D, as inferred from the FlyBase and 4HMM-data. According to the 4HMM model, this region is flanked on both sides with the groups of black bands 14В1-2 and 14D1-2 that are largely composed of the malachite/ruby chromatin enriched with SUUR protein (Fig. 4b–d; Fig. S1s, t). Using FISH with the probes located at the proximal side of 14B1-2 (kat80 gene) and the distal part of 14D1-2 (para gene), we mapped the borders of the bands flanking the aquamarine-lazurite tract. Cytology-based borders of the aquamarine-lazurite tract indeed showed an excellent match to the 4HMM-predicted borders (Fig. 4b–f; Fig. S1b). Within this region, 16 individual lazurite chromatin islands embedded in the aquamarine chromatin are found (Fig. 4b). This is exactly the number of thin gray bands observed under the EM by Saura and colleagues (Saura et al. 1993), who specified that of 12 bands in this region of the X chromosome (Fig. 4g, h) depicted on the Bridges map (Bridges 1938) four doublets are in fact represented by eight solitary bands.
Cytological mapping of the 14B1-2–14D1-2 region of the X chromosome exclusively composed of the alternating aquamarine and lazurite fragments. a Genomic coordinates. b Map of the four chromatin states. c Positions of the DNA probes used for FISH analysis (genes kat80 and para). d Borders of the dense black bands 14B1-2 and 14D1-2, as per 4HMM. e, f FISH mapping of the extensive stretch of aquamarine-lazurite -chromatin in the polytene chromosome X. e Phase contrast. f Merged image: FISH, phase contrast. g EM map of the region (Saura et al. 1993). h Revised Bridges’ map (1938). Across all figures, bar denotes 2 μm
Next, we compiled a map summarizing the distribution of proteins, histone modifications, and genetic elements in between the marker genes kat80 and para, based on the data extracted from the literature (see Fig. S1). First of all, the region begins with the band 14B1-2, whereas FlyBase indicates a different band, 14B7 (Fig. 4e). Next, FlyBase does not provide the information on interband positions in general (Fig. S1a) that play nonetheless a very important role in the organization and functioning of the genome. As a consequence, the borders of the bands and interbands in our model are distinct from the ones provided by the FlyBase (Fig. S1a). The same was found in other regions (Vatolina et al. 2011; present paper). In the region of interest, i.e., between the genes kat80 and para (border positions delimited by vertical red and green lines in Fig. S1) lazurite chromatin fragments are shaded blue. All the aquamarine fragments (spacers between the blue stripes) share the characteristic molecular and genetic features of interbands (Zhimulev et al. 2014), such as state 1 (Kharchenko et al. 2011) chromatin (Fig. S1c), promoter regions (Fig. S1f), enhancer elements, DNAse I hypersensitive sites (DHS), Origin Recognition Complex (ORC)-associated proteins, paused RNA polymerase II, non-specific lethals (NSL) 1-3 (Fig. S1e-j); histone marks associated with active promoters (Fig. S1k), as well as insulator proteins, boundary element-associated factor (BEAF), and CHRIZ (Fig. S1n) that are specifically found in interbands (Zhao et al. 1995; Gortchakov et al. 2005). In contrast, DCC proteins (MSL1, MSL2, and MOF), JIL1, as well as histone modifications associated with elongation (most notably, H3K36me3, and H3K79me), map to the lazurite chromatin (Fig. S1o-r). In line with our hypothesis, most of the genes in this region have their coding parts and 3′-ends in the lazurite chromatin (Fig. S1d).
Distribution of CHRIZ and MSL1 in the region of the polytene X chromosome encompassing a series of thin gray bands and interbands
We proceeded to comparing the mutual distribution of proteins characteristic for aquamarine and lazurite chromatin states versus banding pattern in yet another region of the X chromosome encompassing a series of thin bands and interbands. We expected that immunostaining for these proteins in the region 2C-2E would appear as the alternation of signals and follow the banding pattern.
First, we performed FISH mapping of two aquamarine chromatin fragments using CR45478/east fragment and the 5′-end of the pn gene as the probes (Fig. 5a, b). Both probes were predictably located in the interbands: proximal to 2C1-2 and distal to the band 2E1-2 (Fig. 5c, d). Secondly, simultaneous detection of aquamarine-specific DNA probes and CHRIZ shows complete co-localization (yellow signal) (Fig. 5e). To detect lazurite chromatin using MSL1 as a marker, chromosomes from the M2SXB1-2 transgenic females were used (see “Materials and Methods”). These chromosomes display a profile of MSL1 binding retaining the strongest MSL1 binding sites (Demakova et al. 2003). In both instances, MSL1 is clearly detectable proximal to the marked interbands, i.e., in the thin gray bands 2С4-5 and 2Е1-2 (Fig. 5f, denoted as black and white triangles in Fig. 5), encompassing east and pn gene bodies, respectively (Fig. 5a, b). Immuno-FISH data underscore the differences between the two cases: in the vicinity of the CG45478 gene, the signals for the aquamarine chromatin DNA probe/CHRIZ (interband) and MSL1 (gray bands) are non-overlapping, whereas in the pn gene, they overlap (yellow signal), as the DNA probe harbors part of the lazurite fragment (Fig. 5g). At the scale of the entire male X chromosome, where many more MSL1 sites are present, signal overlap is detected significantly more frequently (data not shown). Therefore, to accurately map short lazurite chromatin fragments relatively to the fine banding pattern, EM analysis is required.
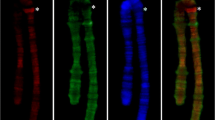
Distribution of CHRIZ (aquamarine chromatin) and MSL1 (lazurite chromatin) immunostaining signals in the region 2C-E, composed of the alternating interbands and thin gray bands in the chromosome of the M2SXB1-2 female.a Genomic coordinates, position of the CG45478 gene (the rest of the genes are not shown), FISH probes, and 4HMM-derived chromatin states.b Genomic coordinates, position of the prune gene (the rest of the genes are not shown), FISH DNA probes, and 4HMM-derived chromatin states. c, d FISH mapping of the aquamarine fragments (genes CR45478 and pn):c Phase contrast. d Overlay of FISH and phase contrast images. e Immuno-FISH of the DNA probes from aquamarine chromatin (genes CR45478 and pn) and the interband protein CHRIZ. Labeling sites appearing yellow indicate co-localization. f Double immunolocalization of MSL1 and CHRIZ proteins. Red/green bars denote lack of co-localization in this chromosome region. g Immuno-FISH of the proteins of interest and aquamarine fragments (CR45478 and pn genes). Black and white triangles mark thin gray bands 2С4-5 and 2Е1-2
EM mapping of lazurite fragments marked by MSL1 along the polytene X chromosome
Previously, we reported on the results of EM analysis of MSL1 immunodetection in polytene chromosome, where gold particles were localized over the decompacted gray bands and at the edges of large black bands (Semeshin et al. 2002, 2003). This analysis involved just two regions of the X chromosomes and could not relate the cytological mapping data to the molecular map.
Here, for high-resolution cytological mapping of lazurite chromatin, we focused on the fragments marked by strong MSL1 binding (modENCODE data for S2 cells) and designed the DNA probes for FISH as close to the MSL1 peaks as possible (Figs. 6, 7, 8, 9, 10, 11, and 12). Taking into account that male polytene X chromosome has diffuse morphology unsuitable for EM mapping, we used transgenic females of the w/w; msl3 [w1; H83M2-61]/TM6, Tb (NOPU) genotype (Kelley et al. 1997). In this stock, ectopic expression of the male-specific MSL2 protein (Kelley et al. 1997) leads to the assembly of DCC in females and results in the X chromosome binding pattern very similar to that of the wild-type males, yet with less diffuse X chromosome morphology (Kotlikova et al. 2006). Moreover, under the EM-immunostaining conditions used, only strong MSL1 binding sites are detected, which greatly simplifies mapping of the lazurite fragments of interest on the band/interband map.
Cytology mapping of lazurite fragments in the region 1B of polytene X chromosome. a Genomic coordinates (bp). b Gene map. c 4HMM-based chromatin map (Zhimulev et al. 2014). d Enrichment profiles of CHRIZ and DCC subunits in S2 cells (modENCODE project), and in salivary glands of wandering larvae (wl) (Conrad et al. 2012). e Relative expression levels in female SGs of wandering larvae (Conrad et al. 2012). Red asterisk above MSL1 peak denotes position of a CES. f Relative expression values in male SG of wandering larvae (Conrad et al. 2012). g Position of the FISH probe based on the Exp6 sequence. h Proximal border of the band 1B1-4, as defined by the 4HMM. i, n FISH mapping of the lazurite fragment (gene Exp6). i Overlay of FISH and phase contrast images. n Phase contrast. j-m MSL1 and CHRIZ mapping against the banding pattern of the 1B region. k, l Double immunostaining for MSL1 and CHRIZ proteins. Red/green bars denote lack of co-localization in this chromosome region. j, m Phase contrast. o–r EM immunodetection of antibodies against MSL1 in the 1B region. Black triangles denote a stretch of grains of gold on the edge of the band 1В1-4. s EM map of the region
Cytology mapping of the lazurite fragments in the region 5D-E. a Genomic coordinates (bp). b MSL1 binding in the chromatin of S2 cells. Red asterisks above the MSL1 profile denote positions of CES. c 4HMM-derived chromatin map (Zhimulev et al. 2014). d Positions of DNA probes used for FISH (from genes CG12236, raptor, and CG14446). e Band borders (4HMM). f–m FISH mapping of lazurite fragments marked by high-MSL1 peaks. f, g, h Overlay of FISH and phase contrast images. i, j, k Phase contrast. l EM immunodetection of antibodies against MSL1. m EM map of the region
Cytology mapping of the lazurite fragments in the region 7E-8A. a Genomic coordinates (bp). b MSL1 binding in the chromatin of S2 cells. Red asterisks above the MSL1 profile denote positions of CES. c 4HMM-derived chromatin map (Zhimulev et al. 2014). d Positions of DNA probes used for FISH (from genes CG11190, Trf-2, Trf4-1, and caf1-180). e Band borders (4HMM). f–m FISH mapping of lazurite fragments marked by high-MSL1 peaks. f-h Overlay of FISH and phase contrast images. j-m Phase contrast. n EM immunodetection of antibodies against MSL1. o EM map of the region
Cytology mapping of the lazurite fragments in the polytene region 9A-B. a Genomic coordinates (bp). b MSL1 binding in the chromatin of S2 cells. Red asterisks above the MSL1 profile denote positions of CES. c 4HMM-derived chromatin map (Zhimulev et al. 2014). d Positions of DNA probes used for FISH (from genes CG1354, ZAP3, and CG34408). e Band borders (4HMM). f–m FISH mapping of lazurite fragments marked by high-MSL1 peaks. f, j Overlay of FISH and phase contrast images. h, i Phase contrast. j–m Mapping MSL1 and CHRIZ binding relatively to the fine banding pattern of the region. j, m Phase contrast. k, l Double immunolocalization of MSL1 and CHRIZ proteins. Red/green bars denote lack of co-localization in this chromosome region; red/yellow/green bars correspond to the partial co-localization. n, o EM immunodetection of antibodies against MSL1. Red bar points to thin interband 9A1/9A2. p EM map of the region
Cytology mapping of the lazurite fragments in the polytene region 9F11-10B1-2. a Genomic coordinates (bp). b MSL1 binding in the chromatin of S2 cells. Red asterisks above the MSL1 profile denote positions of CES. c 4HMM-derived chromatin map (Zhimulev et al. 2014). d Band borders (4HMM). Black triangles mark the positions of DNA probes used for FISH analysis in earlier reports (Vatolina et al. 2011; Zhimulev et al. 2014). e–h Distribution of MSL1 and CHRIZ proteins relatively to the fine banding pattern in the region. e, h Phase contrast. f, g Double immunolocalization of MSL1 and CHRIZ proteins. Red/green bars denote lack of co-localization in this chromosome region. i, j EM immunodetection of antibodies against MSL1. k EM map of the region
Cytology mapping of the lazurite fragments in the polytene region 11A. a Genomic coordinates (bp). b MSL1 binding in the chromatin of S2 cells. Red asterisks above the MSL1 profile denote positions of CES. c 4HMM-derived chromatin map (Zhimulev et al. 2014). d Positions of DNA probes used for FISH (from genes CG1492, Tango4, and Rab40). e Band borders (4HMM). f–i FISH mapping of lazurite fragments marked by high-MSL1 peaks. f, g Overlay of FISH and phase contrast images. h, i Phase contrast. j EM immunodetection of antibodies against MSL1
Cytology mapping of the lazurite fragments in the polytene region 19A-C. a Genomic coordinates (bp). b MSL1 binding in the chromatin of S2 cells. Red asterisks above the MSL1 profile denote positions of CES. c 4HMM-derived chromatin map (Zhimulev et al. 2014). d Positions of the DNA probes used for FISH (form genes DR2, sw, and HERC2). e Band borders (4HMM). f–k FISH mapping of the lazurite fragments marked by high-MSL1 peaks. f–h Overlay of FISH and phase contrast images. i–k Phase contrast. l, m Distribution of MSL1 and CHRIZ proteins relatively to the fine banding pattern in the region. l Double immunolocalization of MSL1 and CHRIZ proteins. Red/green bars denote lack of co-localization in this chromosome region. m Phase contrast. n EM immunodetection of antibodies against MSL1. o EM map of the region
Region 1B1-4
Region 1В1-4 harbors two black ruby/malachite bands. From the proximal side, it is neighbored by the coding sequence of Exp6 gene. This gene is localized in the lazurite chromatin enriched with DCC proteins and H3K36me3 (Fig. 6a–h) and corresponds to the proximal part of the band 1B1-4 (Fig. 6i, j). Results of double immunostaining with antibodies against CHRIZ and MSL1 are shown for the wild-type male (Fig. 6j, k) and M2SXB1-2-transgenic female (Fig. 6l, m). Figure 6 j–m display incomplete overlap between the two channels, with green signal (CHRIZ, 5′-part of the Exp6 gene) overlapping with the interband, and red signal (MSL1, gene body) associating with the band (Fig. 6j, k). EM shows that MSL1 signal is detected exactly at the proximal part of the band 1B1-4, this pattern being highly reproducible across multiple X chromosomes analyzed (Fig. 6n–p). Thus, a cytologically stand-alone band is in fact “hybrid” in nature. It encompasses both inactive and active chromatin, and in genetic terms harbors both developmental genes characteristic of the black bands, and housekeeping genes (e.g., Exp6).
Region 5С-F
This region hosts three MSL1 peaks in the lazurite chromatin (Fig. 7a–c). Our FISH data indicate that these peaks correspond to the gray band 5С7-8, region between 5D3-4 and 5D5-6 (Fig. 7d–k), and the gray band 5E1-2 (which is particularly well seen on the stretched chromosome shown in Fig. 7 h, k). In the region between 5D3-4 and 5D5-6 bands, lazurite chromatin is immediately adjacent to the malachite/ruby chromatin. Strong MSL1 signals decorate the gray bands 5С7-8, 5Е1-2, 5F1-2, and 5F5-6; weak MSL1 signal is detectable over the black “hybrid” band 5D3-4 (Fig. 7l, m).
Region 7E-8A
Four MSL1 peaks associated with lazurite chromatin are present within this region (Fig. 8b, c). FISH analysis indicates that these sequences are found in the gray bands 7Е3, 7Е6, 7F7, and 7F10 (Fig. 8c–m). EM data confirm that these lazurite fragments are found specifically in the gray bands rather than in the neighboring interbands (Fig. 8n, o).
Region 9AВ
This region harbors three MSL1 peaks (Fig. 9a, b) mapping within lazurite fragments. Here, all FISH probes were derived from the coding sequences of genes (lazurite chromatin). At the level of light microscopy, the most distal MSL1 peak (gene CG1354) maps between two black dense bands 9A1 and 9A2 (Fig. 9f, h) that are separated by a very narrow interband (Fig. 9h, p; see also Bridges 1938). EM immunodetection places this MSL1 peak (and the corresponding lazurite fragment) to the very edge of the band 9A2 (Fig. 9n–p, black triangle), as one can see a thin strip of decompacted interband material distal to the label (Fig. 8n, shown red).
The second MSL1 peak marks the border of the lazurite fragment occupied by the coding part of the gene ZAP3 (Fig. 9b–e). FISH maps it to the black dense band 9A4 (Fig. 9g, i). Double immunostaining also shows MSL1 (lazurite chromatin, red signal) to be localized in the black band 9A4 (Fig. 9j–m), distal to the CHRIZ binding site found in the interband (aquamarine chromatin, green signal). Finally, EM immunodetection demonstrates the band border localization of MSL1-labeling (Fig. 9n, o, black triangle).
The third MSL1 peak (CG34408 gene) is found in the decompacted band 9В5-6 (Fig. 9b–e, g, i). CHRIZ (aquamarine chromatin) and MSL1 (lazurite chromatin) signals show no overlap in this region, with red MSL1 signal positioned more proximal, in the band material (Fig. 9j–m). EM analysis indicates that lazurite fragment belongs to the distal part of the decompacted lazurite/malachite band (Fig. 9n, o).
Region 9F11-10B1-2
No FISH analysis was performed for this region, because this site is particularly well studied at the level of light and electron microscopy, and so positions of all bands here are well established (Kozlova et al. 1994; Vatolina et al. 2011; Zhimulev et al. 2014). This region harbors four strong MSL1 binding sites (Fig. 10b). At the physical map, these sites correspond to lazurite chromatin fragments. Accurate matching of the EM signal to bands is not always feasible (Fig. 10i), as three closely spaced gray bands 9F11, 9F12, and 9F13 are found proximal to the 10A1/A2 band (Bridges 1938; Zhimulev et al. 2014). Nevertheless, sections of well-spread chromosomes (Fig. 10j) have gold particles exactly at thin gray bands 9F11 and 9F12, with 9F13 remaining unlabeled (Fig. 10). Thus, the EM label follows the pattern of lazurite fragments marked by the strongest MSL1 peaks (Fig. 10b–d).
The second localization site of gold-labeled antibodies is found in the region of super fine gray bands 10А6-7 and 10А8-10 (Fig. 10i–k). This region is particularly challenging for mapping purposes due to its highly decompacted nature, yet the strongest DCC site maps exactly to the band 10А8-10 (Zhimulev et al. 2014). Finally, the last lazurite fragment hosting a strong MSL1 site is found next to the extensive stretch of ruby/malachite chromatin (Fig. 10b, c). Using co-immunostaining (red label, MSL1 found at the distal part of 10В1-2; green label, CHRIZ found more distal, in the interband), one can observe transitions from red to green in the adjacent regions (Fig. 10f, g). At the EM level, MSL1 label is detectable at the distal edge of the black band 10В1-2 (Fig. 10i, j). Thus, this example corresponds to yet another “hybrid” band situation.
Region 11A
This region has three strong MSL1 peaks localized within the lazurite chromatin. One is supposedly found in the thin gray band, and two other peaks map to the edges of bands composed of the ruby chromatin and flanked by malachite fragments (Fig. 11b–e). FISH mapping of these lazurite fragments (Fig. 11d) places the coding part of Tango gene to the decompacted gray band 11А5, coding part of CG1492—to the proximal end of the band 11A1-2-(Fig. 11f, h), and coding part of Rab40—to the proximal edge of the black dense band 11А6-9 (Fig. 11g, i). EM analysis clearly indicates that the label is found in the band material, rather than in the adjacent interbands (Fig. 11j).
Region 19A-C
In the region 19AC, three strong MSL1 binding sites are present (Fig. 12a, b). The most distal site maps to the aquamarine fragment, negative for both CHRIZ and TSS (Fig. 12b, c). In agreement with the 4HMM-model, this aquamarine fragment (found within the intron of DR2 gene) does not correspond to the interband and maps to the middle of the thick dense band 19A1-4 (Fig. 12f, i). Two other peaks map to the lazurite chromatin (genes sw and HERC2) and are found in the thin gray bands 19B1-2 and 19C4, respectively (Fig. 12g, h, j, k).
Next, we performed immunodetection of CHRIZ and MSL1 on the X chromosome of a M2SXB1-2 male, to assess the degree of overlap between the interband-specific protein CHRIZ, and supposedly, a gray band-specific protein MSL1. We could detect a narrow strip of a weak signal in the central portion of a black dense band 19A1-4, which remains overall label-free (Fig. 12l, m). In the region 19B1-2, the green MSL1 signal is supposed to be bracketed from both sides by the red CHRIZ signals, which corresponds to the thin gray band 19B1-2 and the two flanking interbands (Fig. 12l, m). However, in the region of a decompacted band 19B1-2, the red and green signals are not clearly set apart, and finally strong yellow signal is seen under the band 19С4 (Fig. 12l, m). Yet, at the EM level, the three MSL1 peaks clearly correspond to the bands 19А1-4, 19В1-2, and 19С4 (Fig. 11n–o).
Discussion
The goals of the present study were to understand how the gene activity (i.e., transcription elongation) relates to the chromatin state (produced by the 4HMM), and whether these two parameters are associated with a specific type of bands and interbands in the polytene chromosomes. To do so, we compared four genome-wide chromatin partitioning models that have been published over the past 9 years, in terms of the identification and localization of proteins and histone marks associated with DCC-mediated transcription elongation in the context of male X chromosome as well as with gene body decompactization. According to the model developed by Kharchenko et al. (2011), localization of the DCC subunits perfectly matches the chromatin state 5 (male X chromosome genes and transcription elongation) (Kharchenko et al. 2011). The 4НММ model (Zhimulev et al. 2014) posits that these features are largely associated with lazurite chromatin, where the gene bodies are located.
According to the 4HMM, two chromatin types, lazurite and aquamarine, belong to chromatin open for transcription and are decondensed in the sense of accessibility for DNAse I (Milon et al. 2014). Genetically, these chromatin states are represented by housekeeping genes, with aquamarine chromatin hosting promoter regions of such genes, and lazurite chromatin encompassing the coding parts (Fig. 3, Fig. S1, Figs. 5, 6, 7, 8, 9, 10, 11, and 12, Table 1).
Nonetheless, these two chromatin types are distinct in many ways (Table 1). Genome-wide bioinformatic analysis of chromatin profiling data indicate that aquamarine chromatin is invariably positioned upstream of the lazurite chromatin. Also, aquamarine chromatin is rich in DHS and ORCs in two cell types (Table 1). Lazurite and aquamarine fragments show distinctive localization of histone modifications mostly associated with transcription elongation and exonic features, which includes Н3К36me3, H2b-ubq, H4K20me1, H3K79me1, H4K16ac, H3K27me,1 and H3K79me2 (Table 1). In addition to the peculiar histone modification patterns, we also note the presence of several exon- and elongation-associated proteins, such as MSL1, MLE, JIL1, MRG15, and RNA pol II. Both groups of markers are largely associated with the lazurite chromatin (Table 1). The present study also addresses the question which parts of the X-linked genes contain Н3К36me3, MSL1, and JIL1. Following X chromosome wide analysis of how these proteins are distributed relatively to the gene 5′-end and the coding sequence, it became clear that MSL1, JIL1, and H3K36me3 are typically found in the coding sequences, i.e., in the exonic portion of lazurite chromatin (Fig. 3). Yet, their localization is not restricted to the lazurite chromatin, and they are rarely detectable in the aquamarine chromatin, as well.
The EM analysis of MSL1 immunolocalization shows that the signals are present exclusively in the male X chromosome and that two-label distribution patterns are identifiable. The first one involves the situation wherein the label covers decompacted gray bands. The second one has the label at the edges of black bands. By relating the exact positions of the chromatin states and the two types of bands on the molecular map, one can explain how these two labeling patterns may form. In when the label is found in the decondenced bands, we can see complete coincidence in localizations of MSL1, lazurite chromatin, and decondensed bands (Fig. 13a).
Black “hybrid” band formation upon fusion of the lazurite chromatin with the black band edge. а Typical arrangement of chromatin fragments, wherein black and gray bands are separated by the aquamarine chromatin (i.e., interbands). 5′-end of the gene 1 is found next to the black band material. Grains of gold on EM images are denoted as white circles and map to the gray bands (lazurite chromatin). b When the 3′-end of the gene 2 is juxtaposed to the black band, a “hybrid” black band is formed encompassing the typical black band and the coding sequence of a gene from the gray band
Our retrospective analysis of the EM of immunogold staining data presented by Semeshin et al. (2002, 2003) indicates that the label was found at thin “puffing” gray bands 16B3, 16B10-11, 16D3-4, 16E4-5, 17A9-10, and 17F1-2 (Fig. 1 in Semeshin et al. 2002 and Semeshin et al. 2003), although back then, the label positions could not be mapped relatively to the lazurite chromatin, other proteins, or histone modifications. It can therefore be concluded that whenever the MSL1 label is found in the gray bands, this likely corresponds to the situation of a housekeeping gene whose promoter is found in the interband and whose coding part maps to the thin gray band (gene 1 on the Fig. 13a).
This is unlike the situation with the label found of the edges of the bands. Both the present study and the reports by Semeshin et al. (2002, 2003) indicate that this type of labeling is exclusively associated with large black bands. In the papers referenced above, these are 15D1-2, 16D1-2, 16F7-8, and 17F7-8, and numerous cases are shown in the Figs. 5, 6, 7, 8, 9, 10, 11, and 12. It is known that big black bands differ from gray decondensed bands in sense of chromatin decompaction (Kozlova et al. 1994; Vatolina et al. 2011), replication timing, and even in underreplication (Zhimulev et al. 1982). Moreover, both sides of the black band composed of the ruby chromatin are bordered by the short stretch of malachite chromatin which shows an intermediate timing of replication completion, as well as stronger chromatin decondensation (Khoroshko et al. 2016, 2018).
Formally, two possibilities for the mutual arrangement of the aquamarine/lazurite chromatin and black bands (ruby/malachite chromatin) may exist: aquamarine/lazurite chromatin may contact by the 5′-end or 3′-end of the gene to the malachite chromatin of the black band (Fig. 13. In the former scenario, black band (malachite chromatin) is neighbored by the interband (5′-end of the gene in the aquamarine chromatin) (Fig. 13a); in the latter scenario, the 3′-end of the gene in the lazurite chromatin (gene 2 in Fig. 13b) joins with material of the black, forming the whole band (Fig. 13b). This black dense band is “hybrid” in nature, as it contains a densely packed material peculiar for the black bands and developmentally regulated genes, as well as the coding part of a housekeeping gene and the material of a loose gray band (Fig. 13b) (see also Khoroshko et al. 2018). Here we would like to make it clear that both genetic content, morphology of the band materials, and chromatin state in parts of the hybrid bands are the same as in respective gray and black bands (lazurite chromatin/housekeeping genes and malachite chromatin/genes of development) (see Fig. 13b). Because the sizes of gray bands are only several kilobases and black bands—between several dozens and several hundred kilobases—existence in hybrid band of something other except of the black band material is not visible, and the band looks like usual black band. Accepting the existence of “hybrid” bands, we can explain finding the strong labeling of the black band border when MSL1 antibodies were used: it labels the gene body joined to the border of the black band.
It is therefore interesting to know whether these “hybrid” bands are relatively widespread. Earlier, attempts to enumerate them were done. Demakova et al. (2016) analyzed the distribution of chromatin states in 29 finely mapped bands and observed that in 60% of cases, the bands were found adjacent to the aquamarine chromatin. In other words, a classical pattern of alternating bands and interbands was observed, with the rest of the cases apparently corresponding to the “hybrid” bands. Khoroshko et al. (2018) have focused on the analysis of edges of 62 late-replicating black bands (124 edges). Of these, 116 edges were amenable to analysis, 44 (30%) of which could be classified as “hybrid.” Thus, the percentage of “hybrid” bands could be rather high. Our bioinformatic analysis indicates that there are 810 (403 forward + 407 reverse DNA strand) ruby/malachite/lazurite chromatin combinations (borders of the “hybrid” bands) in the genome. However, this expected number of hybrid joinings (ruby-malachite-lazurite chromatins) can reflect only maximal number of hybrid bands because this kind of joinings exists in the middle part of the black bands as well (Khoroshko et al. 2016). New studies with usage of new approaches could solve this question.
So, using the antibodies against proteins of the gene body (MSL1), we obtained experimental data proving that the gene bodies are located in the gray bands (lazurite chromatin) and discovered a new type of the band combining both genes of housekeeping and genes of development and the number of these bands could be pretty high.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Alekseyenko AA, Larschan E, Lai WR, Park PJ, Kuroda MI (2006) High-resolution ChIP–chip analysis reveals that the Drosophila MSL complex selectively identifies active genes on the male X chromosome. Genes Dev 20:848–857
Alekseyenko AA, Peng S, Larschan E, Gorchakov AA, Lee OK et al (2008) A sequence motif within chromatin entry sites directs MSL establishment on the Drosophila X chromosome. Cell 134:599–609
Ashburner M, Golic KG, Hawley RS (2005) Drosophila: a laboratory handbook, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Belyaeva ES, Zhimulev IF, Volkova EI, Alekseyenko AA, Moshkin YM, Koryakov DE (1998) Su(UR)ES a gene suppressing DNA underreplication in intercalary and pericentric heterochromatin of Drosophila melanogaster polytene chromosomes. Proc Natl Acad Sci U S A 95:7532–7537
Belyaeva ES, Goncharov FP, Demakova OV, Kolesnikova TD, Boldyreva LV, Semeshin VF et al (2012) Late replication domains in polytene and non-polytene cells of Drosophila melanogaster. PLoS One 7:e30035
Belyakin SN, Christophides GK, Alekseyenko AA, Kriventseva EV, Belyaeva ES, Nanayev RA, Makunin IV, Kafatos FC, Zhimulev IF (2005) Genomic analysis of Drosophila chromosome underreplication reveals a link between replication control and transcriptional territories. Proc Natl Acad Sci U S A 102:8269–8274
Boldyreva LV, Goncharov FP, Demakova OV, Zykova TY, Levitsky VG, Kolesnikov NN, Pindyurin AV, Semeshin VF, Zhimulev IF (2017) Protein and genetic composition of four chromatin types in Drosophila melanogaster. Cell Lines Curr Genom 18:214–226
Braunschweig U, Hogan GJ, Pagie L, van Steensel B (2009) Histone H1 binding is inhibited by histone variant H3.3. EMBO J 28:3635–3645
Bridges CB (1935) Salivary chromosome maps with a key to the banding of the chromosomes of Drosophila melanogaster. J Hered 26:60–64
Bridges CB (1938) A revised map of the salivary gland X-chromosome of Drosophila melanogaster. J Hered 29:11–13
Chlamydas S, Holz H, Samata M, Chelmicki T, Georgiev P, Pelechano V, Dündar F, Dasmeh P, Mittler G, Cadete FT, Ramírez F, Conrad T, Wei W, Raja S, Manke T, Luscombe NM, Steinmetz LM, Akhtar A (2016) Functional interplay between MSL1 and CDK7 controls RNA polymerase II Ser5 phosphorylation. Nat Struct Mol Biol 23(6):580–589. https://doi.org/10.1038/nsmb.3233
Conrad T, Cavalli FM, Holz H, Hallacli E, Kind J, Ilik I, Vaquerizas JM, Luscombe NM, Akhtar A (2012) The MOF chromobarrel domain controls genome-wide H4K16 acetylation and spreading of the MSL complex. Dev Cell 22:610–624
Cugusi S, Kallappagoudar S, Ling H, Lucchesi JC (2015) The Drosophila helicase maleless (MLE) is implicated in functions distinct from its role in dosage compensation. Mol Cell Proteomics 14(6):1478–1488
Demakov SA, Vatolina TY, Babenko VN, Semeshin VF, Belyaeva ES, Zhimulev IF (2011) Protein composition of interband regions in polytene and cell line chromosomes of Drosophila melanogaster. BMC Genomics 12:566
Demakova OV, Kotlikova IV, Gordadze PR, Alekseyenko AA, Kuroda MI, Zhimulev IF (2003) The MSL complex levels are critical for its correct targeting to the chromosomes in Drosophila melanogaster. Chromosoma 112:103–115
Demakova OV, Boldyreva LV, Demakov SA, Goncharov FP, Antonenko OV, Zhimulev IF (2016) Characteristic of the chromatin type corresponding to thin “grey” bands in polythene chromosomes of Drosophila melanogaster. Tsitologiya 58:248–252
Eaton ML, Prinz JA, MacAlpine HK, Tretyakov G, Kharchenko PV, MacAlpine D (2011) Chromatin signatures of the Drosophila replication program. Genome Res 21(2):164–174
Figueiredo MLA, Kim M, Philip P, Allgardsson A, Stenberg P, Larsson J (2014) Non-coding roX RNAs prevent the binding of the MSL-complex to heterochromatic regions. PLoS Genet 10(12):e1004865. https://doi.org/10.1371/journal.pgen.1004865
Filion GJ, van Bemmel JG, Braunschweig U, Talhout W, Kind J, Ward LD, Brugman W, de Castro IJ, Kerkhoven RM, Bussemaker HJ, van Steensel B (2010) Systematic protein location mapping reveals five principal chromatin types in Drosophila cells. Cell 143:212–224
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, Ellis B, Gautier L, Ge Y, Gentry J et al (2004) Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 5:R80
Gorchakov AA, Alekseyenko AA, Kharchenko P, Park PJ, Kuroda MI (2009) Long-range spreading of dosage compensation in Drosophila captures transcribed autosomal genes inserted on X. Genes Dev 23:2266–2271
Gortchakov AA, Eggert H, Gan M, Mattow J, Zhimulev IF, Saumweber H (2005) Chriz, a chromodomain protein specific for the interbands of Drosophila melanogaster polytene chromosomes. Chromosoma 114(1):54–66
Hoskins RA, Landolin JM, Brown JB, Sandler JE, Takahashi H, Lassmann T, Yu C, Booth BW, Zhang D, Wan KH, Yang L, Boley N, Andrews J, Kaufman TC, Graveley BR, Bickel PJ, Carninci P, Carlson JW, Celniker SE (2011) Genome-wide analysis of promoter architecture in Drosophila melanogaster. Genome Res 21(2):182–192
Ivaldi MS, Karam CS, Corces VG (2007) Phosphorylation of histone H3 at Ser10 facilitates RNA polymerase II release from promoter-proximal pausing in Drosophila. Genes Dev 21(21):2818–2831
Joshi AA, Struhl K (2005) Eaf3 chromodomain interaction with methylated H3-K36 links histone deacetylation to Pol II elongation. Mol Cell 20:971–978
Kapoor-Vazirani P, Vertino PM (2014) A dual role for the histone methyltransferase PR-SET7/SETD8 and histone H4 lysine 20 monomethylation in the local regulation of RNA polymerase II pausing. J Biol Chem 289(11):7425–7437
Kelley RL, Solovyeva I, Lyman LM, Richman R, Solovyev V, Kuroda MI (1995) Expression of msl-2 causes assembly of dosage compensation regulators on the X chromosomes and female lethality in Drosophila. Cell 81:867–877
Kelley RL, Wang J, Bell L, Kuroda MI (1997) Sex lethal controls dosage compensation in Drosophila by a non-splicing mechanism. Nature 387(6629):195–199
Kelley RL, Meller VH, Gordadze PR, Roman G, Davis RL, Kuroda MI (1999) Epigenetic spreading of the Drosophila dosage compensation complex from roX RNA genes into flanking chromatin. Cell 98:513–522
Kharchenko PV, Alekseyenko AA, Schwartz YB, Minoda A, Riddle NC, Ernst J, Sabo PJ, Larschan E, Gorchakov AA, Gu T, Linder-Basso D, Plachetka A, Shanower G, Tolstorukov MY, Luquette LJ, Xi R, Jung YL, Park RW, Bishop EP, Canfield TK, Sandstrom R, Thurman RE, MacAlpine D, Stamatoyannopoulos JA, Kellis M, Elgin SC, Kuroda MI, Pirrotta V, Karpen GH, Park PJ (2011) Comprehensive analysis of the chromatin landscape in Drosophila melanogaster. Nature 471:480–485
Khoroshko VA, Levitsky VG, Zykova TY, Antonenko OV, Belyaeva ES, Zhimulev IF (2016) Chromatin heterogeneity and distribution of regulatory elements in the late-replicating intercalary heterochromatin domains of Drosophila melanogaster chromosomes. PLoS One 11:e0157147. https://doi.org/10.1371/journal.pone.0157147
Khoroshko VA, Zykova TY, Popova OO, Zhimulev IF (2018) Border structure of intercalary heterochromatin bands of Drosophila melanogaster polytene chromosomes. Dokl Biochem Biophys 479(1):114–117
Kolasinska-Zwierz P, Down T, Latorre I, Liu T, Liu XS, Ahringer J (2009) Differential chromatin marking of introns and expressed exons by H3K36me3. Nat Genet 41:376–381
Kolesnikova TD, Posukh OV, Andreyeva EN, Bebyakina DS, Ivankin AV, Zhimulev IF (2013) Drosophila SUUR protein associates with PCNA and binds chromatin in a cell cycle-dependent manner. Chromosoma 122(1-2):55–66
Kolesnikova TD, Goncharov FP, Zhimulev IF (2018) Similarity in replication timing between polytene and diploid cells is associated with the organization of the Drosophila genome. PLoS One 13:e0195207
Kotlikova IV, Demakova OV, Semeshin VF, Shloma VV, Boldyreva LV, Kuroda MI, Zhimulev IF (2006) The Drosophila dosage compensation complex binds to polytene chromosomes independently of developmental changes in transcription. Genetics 172:963–974
Kozlova TY, Semeshin VF, Tretyakova IV, Kokoza EB, Pirrotta V, Grafodatskaya VE et al (1994) Molecular and cytogenetical characterization of the 10A1-2 band and adjoining region in the Drosophila melanogaster polytene X chromosome. Genetics 136:1063–1073
Kuroda MI, Hilfiker A, Lucchesi JC (2016) Dosage compensation in Drosophila—a model for the coordinate regulation of transcription. Genetics 204(2):435–450
Lam KC, Muhlpfordt F, Vaquerizas JM, Raja SJ, Holz H et al (2012) The NSL complex regulates housekeeping genes in Drosophila. PLoS Genet 8(6):e1002736
Larschan E, Alekseyenko AA, Gorchakov AA, Peng S, Li B et al (2007) MSL complex is attracted to genes marked by H3K36 trimethylation using a sequence-independent mechanism. Mol Cell 28:121–133
Lawrence M, Gentleman R, Carey V (2009) rtracklayer: an R package for interfacing with genome browsers. Bioinformatics 25:1841–1842
Lawrence M, Huber W, Pagès H, Aboyoun P, Carlson M, Gentleman R, Morgan MT, Carey VJ (2013) Software for computing and annotating genomic ranges. PLoS Comput Biol 9:e1003118
Lindsley DL, Zimm GG (1992) The genome of Drosophila melanogaster. Academic Press, San Diego
Lucchesi JC, Kuroda MI (2015) Dosage compensation in Drosophila. Cold Spring Harb Perspect Biol 7(5):a019398. https://doi.org/10.1101/cshperspect.a019398
Milon B, Sun Y, Chang W, Creasy T, Mahurkar A, Shetty A et al (2014) Map of open and closed chromatin domains in Drosophila genome. BMC Genomics 15:988
Nechaev S, Fargo DC, dos Santos G, Liu L, Gao Y, Adelman K (2010) Global analysis of short RNAs reveals widespread promoter-proximal stalling and arrest of Pol II in Drosophila. Science 327(5963):335–338. https://doi.org/10.1126/science.1181421
Philip P, Stenberg P (2013) Male X-linked genes in Drosophila melanogaster are compensated independently of the male-specific lethal complex. Epigenetics Chromatin 6:35
Posukh OV, Maksimov DA, Laktionov PP, Koryakov DE, Belyakin SN (2017) Functional dissection of Drosophila melanogaster SUUR protein influence on H3K27me3 profile. Epigenetics Chromatin 10:56. https://doi.org/10.1186/s13072-017-0163-z
R Development Core Team (2015) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Raja SJ, Charapitsa I, Conrad T, Vaquerizas JM, Gebhardt P, Holz H, Kadlec J, Fraterman S, Luscombe NM, Akhtar A (2010) The nonspecific lethal complex is a transcriptional regulator in Drosophila. Mol Cell 38(6):827–841
Robinson PJ, An W, Routh A, Martino F, Chapman L, Roeder RG, Rhodes D (2008) 30 nm chromatin fibre decompaction requires both H4-K16 acetylation and linker histone eviction. J Mol Biol 381:816–825
Roy S, Ernst J, Kharchenko PV, Kheradpour P, Negre N et al (2010) Identification of functional elements and regulatory circuits by Drosophila modENCODE. Science 330:1787–1797
Sass GL, Pannuti A, Lucchesi JC (2003) Male-specific lethal complex of Drosophila targets activated regions of the X chromosome for chromatin remodeling. Proc Natl Acad Sci U S A 100:8287–8291
Saura AO, Tapio IH, Sorsa V (1993) Electron microscopic analysis of the banding pattern in the salivary gland chromosomes of Drosophila melanogaster. Divisions 11 through 20 of X. Herediras 119:123–141
Schubeler D, MacAlpine DM, Scalzo D, Wirbelauer C, Kooperberg C et al (2004) The histone modification pattern of active genes revealed through genome-wide chromatin analysis of a higher eukaryote. Genes Dev 18:1263–1271
Schwartz S, Meshorer E, Ast G (2009) Chromatin organization marks exon-intron structure. Nat Struct Mol Biol 16(9):990–995
Semeshin VF, Zhimulev IF, Belyaeva ES (1979) Electron microscope autoradiographic study on transcriptional activity of Drosophila melanogaster polytene chromosomes. Chromosoma 73:163–177
Semeshin VF, Demakov SA, Alonso PM, Belyaeva ES, Bonner JJ, Zhimulev IF (1989) Electron microscopical analysis of Drosophila polytene chromosomes. Chromosoma 97(5):396–412
Semeshin VF, Artero R, Perez Alonso M, Shloma VV (1998) Electron microscopic in situ hybridization of digoxigenin-dUTP-labelled DNA probes with Drosophila melanogaster polytene chromosomes. Chromosom Res 6:405–410
Semeshin VF, Andreyeva EN, Shloma VV, Saumweber H, Zhimulev IF (2002) Immunogold electron microscope localization of proteins in Drosophila polytene chromosomes: applications and limitations of the method. Chromosom Res 10:429–433
Semeshin VF, Shloma VV, Andreyeva EN, Saumweber H, Zhimulev IF (2003) Use of immunogold labelling technique for immunoelectron microscope localization of proteins in Drosophila polytene chromosomes. Tsitologiya 45(3):235–243
Sher N, Bell GW, Li S, Nordman J, Eng T, Eaton ML, Macalpine DM, Orr-Weaver TL (2012) Developmental control of gene copy number by repression of replication initiation and fork progression. Genome Res 22(1):64–75
Shogren-Knaak M, Peterson CL (2006) Switching on chromatin: mechanistic role of histone H4-K16 acetylation. Cell Cycle 5(13):1361–1365
Smith RN, Aleksic J, Butano D, Carr A, Contrino S, Hu F, Lyne M, Lyne R, Kalderimis A, Rutherford K, Stepan R, Sullivan J, Wakeling M, Watkins X, Micklem G (2012) InterMine: a flexible data warehouse system for the integration and analysis of heterogeneous biological data. Bioinformatics 28:3163–3165
Steiner LA, Schulz VP, Maksimova Y, Wong C, Gallagher PG (2011) Patterns of histone H3 lysine 27 monomethylation and erythroid cell type-specific gene expression. J Biol Chem 286(45):39457–39465. https://doi.org/10.1074/jbc.M111.243006
Straub T, Becker PB (2008) DNA sequence and the organization of chromosomal domains. Curr Opin Genet Dev 18:175–180
Straub T, Zabel A, Gilfillan GD, Feller C, Becker PB (2013) Different chromatin interfaces of the Drosophila dosage compensation complex revealed by high-shear ChIP-seq. Genome Res 23(3):473–485
Vatolina TY, Boldyreva LV, Demakova OV, Demakov SA, Kokoza EB, Semeshin VF et al (2011) Identical functional organization of nonpolytene and polytene chromosomes in Drosophila melanogaster. PLoS One 6:e25960
Wagner EJ, Carpenter PB (2012) Understanding the language of Lys36 methylation at histone H3. Nat Rev Mol Cell Biol 13:115–126
Zabidi MA, Arnold CD, Schernhuber K, Pagani M, Rath M, Frank O, Stark A (2015) Enhancer-core-promoter specificity separates developmental and housekeeping gene regulation. Nature 518(7540):556–559
Zeilke T, Glotov A, Saumweber H (2015) High-resolution in situ hybridization analysis on the chromosomal interval 61C7-61C8 of Drosophila melanogaster reveals interbands as open chromatin domains. Chromosoma 125:423–435. https://doi.org/10.1007/s00412-015-0554-5
Zhao K, Hart CM, Laemmli UK (1995) Visualization of chromosomal domains with boundary element-associated factor BEAF-32. Cell 81(6):879–889
Zhimulev IF (1999) Genetic organization of polytene chromosomes. Adv Genet 39:1–589
Zhimulev IF, Semeshin VF, Kulichkov VA, Belyaeva ES (1982) Intercalary heterochromatin in Drosophila. I Localization and general characteristics. Chromosoma 87:197–228
Zhimulev IF, Belyaeva ES, Semeshin VF, Koryakov DE, Demakov SA, Demakova OV, Pokholkova GV, Andreyeva EN (2004) Polytene chromosomes: 70 years of genetic research. Int Rev Cytol 241:203–275
Zhimulev IF, Zykova TY, Goncharov FP, Khoroshko VA, Demakova OV, Semeshin VF et al (2014) Genetic organization of interphase chromosome bands and interbands in Drosophila melanogaster. PLoS One 9:e101631
Zykova TY, Levitsky VG, Belyaeva ES, Zhimulev IF (2018) Polytene chromosomes—a portrait of functional organization of the Drosophila genome. Curr Genom 19(3):179–191
Acknowledgements
We express our deep gratitude to M. Kuroda and R. Kelley for antibodies against MSL1 and transgenic stocks of Drosophila.
Funding
The study was supported by the grant from the Russian Science Foundation (grant number 19-14-00051, bioinformatics analysis), the Fundamental Scientific Research program (0310-2019-0003, immunofluorescence analysis), and the RFBR (grant number 17-00-00284, EM mapping of antibodies and data processing). Data analysis contributed by V.G. Levitsky was supported by the RFBR grant 18-29-13040.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Fig 1.
Localization of various protein and genetic features in the region 14B1-2 - 14D1-2 of X chromosome. a Localization of bands within the region 14B1-2 - 14D1-2 of polytene X chromosome according to FlyBase version R 5.57. b Localization of the fragments of marker genes kat80 and para is shown as vertical red and green highlight. с Chromatin states as defined by the chromatin classification models (top-bottom: Zhimulev et al. 2014; Kharchenko et al. 2011 in S2 and BG3 cell lines; Milon et al. 2014; Filion et al. 2010). d Positions of the annotated genes (RefSeq Genes). e Housekeeping and developmental enhancers (Zabidi et al. 2015). f Peak and Broad promoters (Hoskins et al. 2011). g DNAse I Hypersensitive Sites in S2, Kc and BG3 cells (Kharchenko et al. 2011). h Profiles for the ORC2 subunits in the chromosomes from S2, Kc, and BG3 cell lines (Eaton et al. 2011), as well as larval salivary glands (Sher et al. 2012). i Short RNAs derived from stalled RNA polymerase II in Drosophila cells (Nechaev et al. 2010). j Enrichment profiles of NSL complex components: NSL1 binding profile from salivary glands (Raja et al. 2010), NSL3 in S2 cells (Lam et al. 2012). k Transcription initiation-associated histone marks (culture cells, modENCODE data). l Localization of histone H1 dips in Kc cells (Braunschweig et al. 2009). m RNA polymerase II (modENCODE data). n Interband-specific proteins CHRIZ and BEAF (modENCODE data). o MSL1 protein, S2 (modENCODE data). p DCC subunits in wild-type and mutants (Figueiredo et al. 2014). q Gene body-, exon- and transcription elongation-associated histone marks (modENCODE data). r JIL1 protein enrichment across different cell types (modENCODE data). s SUUR distribution in salivary glands obtained by DamID method (Posukh et al. 2017). t SUUR distribution in Kc cells obtained by DamID method (Filion et al. 2010). Blue highlights correspond to grey bands, spaces between the bars correspond to interbands. (PNG 1705 kb)
Table S1
(DOCX 22 kb)
Rights and permissions
About this article
Cite this article
Demakova, O.V., Demakov, S.A., Boldyreva, L.V. et al. Faint gray bands in Drosophila melanogaster polytene chromosomes are formed by coding sequences of housekeeping genes. Chromosoma 129, 25–44 (2020). https://doi.org/10.1007/s00412-019-00728-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-019-00728-2