Abstract
Sex chromosomes have evolved many times from morphologically identical autosome pairs, most often presenting several recombination suppression events, followed by accumulation of repetitive DNA sequences. In Orthoptera, most species have an X0♂ sex chromosome system. However, in the subfamily Melanoplinae, derived variants of neo-sex chromosomes (neo-XY♂ or neo-X1X2Y♂) emerged several times. Here, we examined the differentiation of neo-sex chromosomes in a Melanoplinae species with a neo-XY♂/XX♀ system, Ronderosia bergi, using several approaches: (i) classical cytogenetic analysis, (ii) mapping via fluorescent in situ hybridization of some selected repetitive DNA sequences and microdissected sex chromosomes, and (iii) immunolocalization of distinct histone modifications. The microdissected sex chromosomes were also used as sources for Polymerase chain reaction (PCR) amplification of RNA-coding multigene families, to study variants related to the sex chromosomes. Our data suggest that the R. bergi neo-Y has become differentiated after its formation by a Robertsonian translocation and inversions, and has accumulated repetitive DNA sequences. Interestingly, the ex autosomes incorporated into the neo-sex chromosomes retain some autosomal post-translational histone modifications, at least in metaphase I, suggesting that the establishment of functional modifications in neo-sex chromosomes is slower than their sequence differentiation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sex chromosomes have evolved independently several times in a wide range of plant and animal species, originating from a pair of morphologically identical chromosomes (Ohno 1967; Bull 1983). It is widely assumed that the gradual lack of recombination between the newly formed sex chromosomes, X and Y or W and Z, leads to their morphological and genetic divergence (Rice 1996; Filatov et al. 2000). The ultimate fate of this process is genetic degeneration, leading to the inactivation/loss of many genes on the Y or W chromosomes and the accumulation of repetitive DNA sequences, which is strong after the abolition of recombination. The genetic erosion of Y or W chromosomes may even lead to the complete disappearance of these chromosomes (Charlesworth et al. 1994; Rice 1996; Steinemann and Steinemann 1997, 2005; Navajas-Pérez et al. 2005, 2009; Bachtrog 2006; Kejnovsky et al. 2009; Pokorná et al. 2011a).
There are some well-documented examples of the accumulation of repetitive DNA sequences and gene degeneration as important steps for the generation of morphological and genetic differences between sex chromosomes. For example, the relatively ancient human Y chromosome carries only 27 distinct protein-coding genes, of which only 16 have homologs on the X chromosome. Approximately half of the human Y chromosome is entirely heterochromatic, while the euchromatic portion is a mosaic of X degenerated, X transposed, and ampliconic sequences (Skaletsky et al. 2003). Similarly, the Drosophila melanogaster Y chromosome is almost completely heterochromatic and distinct from the X chromosome, carrying only approximately 15 genes (Carvalho 2002). In the plant Silene latifolia, the XY sex chromosomes evolved recently from autosomes approximately 5–10 million years ago (mya) (Nicolas et al. 2005; Bergero et al. 2007) but also demonstrate occurrence of repetitive DNA sequences (Hobza et al. 2007; Kubat et al. 2008; Kejnovsky et al. 2009; Matsunaga 2009).
Among insects, it is assumed that the ancestral X0♂/XX♀ sex chromosome system in certain groups, such as Orthoptera, Blattodea, Mantodea, and Phasmatodea, evolved from an XY♂/XX♀ system (White 1973; Hewitt 1979; Castillo et al. 2010a; Charlesworth and Mank 2010; Kaiser and Bachtrog 2010). In contrast, the Z0♀/ZZ♂ sex chromosome system, which is thought to be ancestral for Trichoptera and Lepidoptera, evolved into the WZ♀/ZZ♂ system during the evolution of Lepidoptera (Traut et al. 2007). On the other hand, in some species, a loss of the W chromosome resulted in a derived Z0♀/ZZ♂ system (Yoshido et al. 2013). In grasshoppers, the X0♂/XX♀ system has been observed in most studied species, with males being the heterogametic sex (X0), although in some groups, mainly in Melanoplinae species, derived systems with neo-XY♂ or neo-X1X2Y♂ evolved several times by repeated autosome-sex chromosome Robertsonian translocations (Rb-translocations) (White 1973; Hewitt 1979; Bidau and Martí 2001; Castillo et al. 2010a,b; Palacios-Gimenez et al. 2013).
In Melanoplinae and in Orthoptera as a whole, as revised by Castillo et al. (2010a), although relative ages for neo-XY sex chromosomes were not determined until now, it is largely accepted that neo-XY sex chromosomes with recent evolutionarily history present (i) synapsis at Pachytenes between neo-Y and XR from neo-X that corresponds to the autosome involved in the Rb-translocation, (ii) a lack of heterochromatinization and (iii) interstitial chiasmata formation, e.g., Boliviacris noroestensis, Mariacris viridipes, and Neuquenina fictor. On the other hand, advanced “old” systems most often involve (i) contact/chiasmata restricted to distal regions, (ii) heterochromatinization and to a lesser extent (iii) structural rearrangements, including inversions. Advanced systems with terminal contact and heterochromatinization were observed for example, in Zygoclistron species and Dichroplus vittigerum. Moreover, among the 75 species with neo-XY, at least in ten structural rearrangements involving distinct regions of neo-X or neo-Y took place, e.g., Ronderosia ommexechoides and four Aleuas species with short pericentric inversion in neo-Y, Spathalium audouini with pericentric inversion changing the morphology of neo-Y to metacentric and three Dichroplus species, Dichroplus silveiraguidoi, Dichroplus vittatus, and Dichroplus maculipennis with pericentric inversion in neo-X. Among these extreme conditions, neo-XY sex system demonstrating a mixture of the above cited characteristics have also been reported (Mesa and de Mesa 1967; Díaz and Sáez 1968; White 1973; Cardoso and Dutra 1979; Castillo et al. 2010a; 2014; Bidau et al. 2011).
To improve our understanding of the poorly explored differentiation of the neo-sex chromosomes in grasshoppers (see Palacios-Gimenez et al. 2013), we selected for this study as a model species Ronderosia bergi (Acrididae, Melanoplinae), which has a karyotype of 2n = 22, neo-XY♂ (Castillo et al. 2010b). We examined the neo-sex chromosomes of this species using the following approaches: (i) classical cytogenetic analysis; (ii) physical mapping of DNA sequences by fluorescence in situ hybridization (FISH) with various probes, such as 5 multigene families, 16 microsatellites, a telomeric probe, C 0 t DNA fractions, and probes that were prepared from microdissected X and Y chromosomes; and (iii) immunolocalization of different histone methylations, acetylations, and phosphorylation in metaphase I. Moreover, the microdissected chromosomes were used as sources for the specific amplification of four RNA-coding multigene families with the aim of addressing variations of these sequences in the sex chromosomes to infer their evolution.
Materials and methods
Animals, chromosome obtaining, and DNA extraction
Males and females of R. bergi adults were collected at the campus of the Univ Estadual Paulista—UNESP and Parque Estadual Edmundo Navarro de Andrade (Rio Claro, São Paulo State, Brazil) from December 2012 to March 2014 with the authorization of ICMBio SISBIO (process number 16009-1). The male testes were dissected and fixed in a 3:1 ethanol:acetic acid solution. For immunolabeling, fixation was performed using 2 % formaldehyde (see below). Additionally, some females were kept in captivity until female oviposition occurred to obtain embryos, which were cytologically prepared as described in Webb et al. (1978) with slight modifications. The residues of the specimens were stored in 100 % ethanol for DNA extraction.
To describe the general aspects of chromosomes, conventional staining with 5 % Giemsa for each individual was used. In meiosis, White’s (1973) terminology was considered, the arms of the neo-X chromosome will be referred to as XL, the arm from the original X chromosome, and XR, the arm that shares homology with the neo-Y. The C-banding procedure was conducted according to Sumner (1972). The genomic DNA was extracted using the phenol-chloroform procedure (Sambrook and Russel 2001).
Probes for multigene families, telomeric repeats, and C 0 t DNA fractions
The DNA probes for four multigene families were obtained by polymerase chain reaction (PCR) from the genomes of Abracris flavolineata (5S ribosomal DNA (rDNA) and H3 histone gene) and Rhammatocerus brasiliensis (U1 and U2 snDNA) using the primers designated by Cabral-de-Mello et al. (2010, 2012), Colgan et al. (1998) and Bueno et al. (2013). The identified DNA sequences are deposited in GenBank under the accession numbers KC936996 (5S rDNA), KC896792 (H3 histone gene), KC896793 (U1 snDNA) and KC896794 (U2 snDNA). For the 18S rDNA, the probe was obtained from a cloned fragment previously isolated from Dichotomius semisquamosus (Cabral-de-Mello et al. 2010, GenBank accession number GQ443313). The telomeric probe was obtained by non-template PCR using the self-complementary primers (TTAGG)5 and (CCTAA)5 following the conditions that were described by Ijdo et al. (1991).
Repetitive DNA-enriched samples were obtained based on the renaturation kinetics of C 0 t DNA (DNA fraction that is enriched for highly and moderately repetitive DNA sequences) according to the protocol of Zwick et al. (1997) with modifications from Cabral-de-Mello et al. (2010). It was denaturated 150 ng/μL of fragmented genomic DNA at 95 °C. The reassociation times were as follows: for C 0 t-1, 10 min and 48 s, and for C 0 t-100, 18 h.
Microdissection of sex chromosomes and amplification of RNA multigene families
Microdissections of neo-X and neo-Y chromosomes from male meiotic cells were performed. Before microdissection, a cell suspension of four testicular follicles in 100 μl of 50 % acetic acid was spread onto a coverslip using a warm plate at approximately 50 °C. The neo-sex chromosomes were easily recognized due to their C-shaped configuration at metaphase I (Castillo et al. 2010b), and their early segregation avoided cross contamination.
The microdissection was conducted using an Eppendorf 5171 micromanipulator coupled with a Nikon Axiphot inverted microscope. Ten neo-Y chromosomes and eight neo-X chromosomes were microdissected separately and then amplified using the GenomePlex Single Cell Whole Genomic Amplification kit WGA4 (Sigma-Aldrich, St Louis, MO, USA) followed by reamplification using the GenomePlex WGA3 kit (Sigma-Aldrich). We used the DNAs that were amplified by the GenomePlex WGA3 kit to generate probes of the neo-X (μX-DNA) and neo-Y (μY-DNA) chromosomes.
To verify the presence of multigene families for rDNAs (18S and 5S) and snDNAs (U1 and U2) in the neo-sex chromosomes, PCRs were performed using the microdissected chromosomes separately as templates. Moreover, for a comparative analysis, the repeats successfully amplified from microdissected sex chromosomes (5S rDNA, U1 and U2 snDNAs) were also amplified from female genomic DNA (♀gDNA). The primers that were used are described in Cabral-de-Mello et al. (2010) for 18S and 5S rDNAs, Cabral-de-Mello et al. (2012) for U1 snDNA, and Bueno et al. (2013) for U2 snDNA. PCR was performed using 10× PCR Rxn Buffer, 0.2 mM MgCl2, 0.16 mM dNTPs, 2 mM each primer, 1 U of Taq Platinum DNA Polymerase (Invitrogen, San Diego, CA, USA) and 50–100 ng/μl template DNA. The PCR conditions included an initial denaturation at 94 °C for 5 min and 30 cycles at 94 °C (30 s), 55 °C (30 s), and 72 °C (80 s), plus a final extension at 72 °C for 5 min. The PCR products were visualized on a 1 % agarose gel, and the bands were isolated and purified using the Zymoclean™ Gel DNA Recovery Kit (Zymo Research Corp., The Epigenetics Company, USA) according to manufacturer’s recommendations.
The purified PCR products were cloned using a cloning kit pGEM-T (Promega, Madison, WI, USA) and DH5α Escherichia coli competent cells. A total of 95 recombinant colonies were chosen for DNA sequencing using Macrogen Inc. (Korea), including 33 clones of U1 snDNA, 34 clones of U2 snDNA and 27 clones of 5S rDNA.
Sequence analysis
The quality of the sequences was determined using the Geneious 4.8.5 software (Drummond et al. 2009). The consensus sequences were subjected to BLAST (Altschul et al. 1990) searches on the NCBI website (http://www.ncbi.nlm.nih.gov/blast) and, as expected, were recognized as 5S ribosomal RNA (rRNA), U1, and U2 small nuclear RNA (snRNA) genes. For all of the sequences, we checked the similarity with human repeats to avoid contaminants. The sequences were deposited into the NCBI database under the following accession numbers: KP213271-KP213276 for 5S rDNA, KP213277-KP213281 for U1 snDNA and KP213282-KP213286 for U2 snDNA.
For DNA sequence analyses, we computed the basic sequence statistics with the program DnaSP v.5.10.01 (Librado and Rozas 2009). Phylogenetic and molecular evolutionary analyses were conducted using MEGA v.5 (Tamura et al. 2011). We discarded the primer region at both ends for every sequence and used Drosophila virilis (Diptera) as an outgroup.
Fluorescence in situ hybridization
The plasmid containing the 18S rRNA gene, the PCR products from the H3 histone gene, the C 0 t DNA fractions, and the μX-DNA and μY-DNA probes were labeled using biotin-14-dATP (Invitrogen, San Diego, CA, USA). Additionally, we used 16 synthetic oligonucleotide probes that were directly labeled with biotin-14-dATP during their synthesis at the 5′ end: (A)30, (C)30, (CA)15, (CG)15, (TA)15, (AG)10, (CAA)10, (CAC)10, (TAA)10, (GAA)10, (CGG)10, (GAC)10, (CAT)10, (GAG)10, (GACA)4, and (GATA)8 (Sigma-Aldrich). The 5S rDNA, U snDNAs (U1 and U2) and telomeric probes were PCR-labeled with digoxigenin-11-dUTP (Roche, Mannheim, Germany).
Single- or two-color FISH was performed according to Pinkel et al. (1986) with modifications (Cabral-de-Mello et al. 2010) using meiotic and mitotic chromosome preparations. The post hybridization washes were performed as follows: two times in 2× SSC at 42 °C for 5 min, two times in 0.1× SSC at 42 °C for 5 min, one time in 2× SCC at 42 °C for 5 min and finally in 2× SSC at room temperature for 10 min. The probes that were labeled with digoxigenin-11-dUTP were detected using anti-digoxigenin rhodamine (Roche), while the probes that were labeled with biotin-14-dATP were detected using Streptavidin Alexa Fluor 488-conjugated (Invitrogen).
The preparations were counterstained using 4′,6-diamidine-2′-phenylindole (DAPI) and mounted in VECTASHIELD (Vector, Burlingame, CA, USA). The chromosomes and hybridization signals were observed using an Olympus microscope BX61 that was equipped with a fluorescent lamp and appropriate filters. Fluorescent images were recorded using a DP71 cooled digital camera in grayscale. The images were pseudo-colored in blue (chromosomes) and red or green (signals), merged and optimized for brightness and contrast using Adobe Photoshop CS2.
Immunolabeling
The testes from two adult males were removed and fixed in 2 % formaldehyde in phosphate-buffered saline (PBS), which was freshly prepared from paraformaldehyde, containing 0.05 % Tween for 15 min. Subsequently, the testes were immersed in a small droplet of the fixative on a glass slide and gently flattened under a coverslip, which was then removed after immersing the preparation in liquid nitrogen. The slides were then immediately transferred to cold PBS. Immunolabeling followed the technique described by Cabrero et al. (2007). The primary antibodies (rabbit polyclonal IgG, Upstate Biotechnology, USA) anti-H3K4me2, anti-H3K9me2, anti-H3K4me3, anti-H3K27me3, anti-H4K5ac, anti-H3K27ac, anti-H3K9ac, and anti-H3S10ph were diluted 1:600 in 1 % bovine serum albumin (BSA) in PBS. The slides were incubated with the antibodies overnight at 4 °C and, after washing, were detected with FITC-conjugated anti-rabbit IgG (Sigma-Aldrich) that was diluted 1:60 in PBS, 1 % BSA for 60 min. After final washing in PBT (1× PBS, 0.01 % Tween 20), the preparations were counterstained with DAPI and mounted in VECTASHIELD. Finally, the chromosome images were recorded using the same equipment as mentioned above for FISH.
Results and discussion
Differentiation between the sex chromosomes of R. bergi
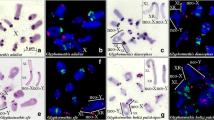
R. bergi presented a karyotype composed of 2n = 22♂/22♀, with ten pairs of acrocentric autosomes and a neo-XY♂/XX♀ sex chromosome pair that was formed by a metacentric neo-X and an acrocentric neo-Y easily recognizable because its short arm is longer than those observed in autosomes (Fig. 1a). This karyotype is identical to that previously published (Díaz and Sáez 1968; Cardoso and Dutra 1979; Castillo et al. 2010b). In Pachytene and Diplotene the sex chromosomes were terminally associated, and pairing involved the short arm of the neo-Y and XR arm of the neo-X (Fig. 1b). This is consistent with the occurrence of chiasmata at the chromosomal ends as previously described (Díaz and Sáez 1968; Cardoso and Dutra 1979; Castillo et al. 2010b).
Classical cytogenetic analysis (a–c) and FISH mapping of repetitive DNA sequences (d–h) on the mitotic and meiotic metaphasic chromosomes of Ronderosia bergi. Each technique and probe type used is indicated directly on the images. The neo-XY sex chromosomes and some autosomes that carry positive FISH signals are indicated. b The C-shaped oriented neo-XY sex bivalent in metaphase I. The chromosome arms of the neo-sex chromosomes that are involved in the Rb-translocation are indicated. XL arm derived from the original X chromosome fused to an autosome; XR autosomal arm of the neo-X that shares ancient homology with the neo-Y chromosome. Note in c that the neo-Y is entirely heterochromatic and the inset in d shows the dispersion of the C 0 t–1 DNA fraction throughout the long arm of the neo-Y chromosome. In e, f note the absence of interstitial telomeric signals on the metacentric neo-X chromosome. The green and red arrowheads in b, f point to the terminal contacts between neo-X chromosome and the neo-Y chromosome and the centromere of the neo-X chromosome, respectively. g, h observe the multigene families mapped in sex chromosomes. Bar 5 μm
The neo-sex chromosomes apparently caused by an X-A Rb-translocation are observed in all species of the genus Ronderosia (Castillo et al. 2010b). However, the Rb-translocation was followed by a pericentric inversion in the neo-Y in only two species, R. bergi and in R. ommexechoides (Carbonell and Mesa 2006), as inferred from the C-shaped sex bivalent at metaphase I (Castillo et al. 2010a,b). These rearrangements confine contact and chiasmata between the chromosomes to the distal region, reducing crossing over between the neo-XY pair, a common placement to old neo-sex chromosomes among grasshoppers, contrasting with recent neo-XY in which chiasmata could occurs interstitially and in multiple points (Díaz and Sáez 1968; Cardoso and Dutra 1979; Castillo et al. 2010a,b, 2014). There is no published estimation of the time of origin of the Ronderosia species to estimate the time of these rearrangements. However, the genus is exclusive to South America (Cigliano 1997) and members of the Melanoploid lineage have been diverging at least 46 mya in this region (Chintauan-Marquier et al. 2011), suggesting that neo-sex chromosomes originated more recently than this date.
Telomeric DNA was seen only on at the two ends of each chromosome. The long arm of the neo-Y chromosome had intense hybridization signals (Fig. 1e, f), confirming that the neo-X and neo-Y association involves the short arm of the neo-Y (Fig. 1f). This data suggest that telomeric repeats were either lost in the Rb-translocation event or eliminated later during sex chromosome differentiation, as in other Melanoplinae species (Palacios-Gimenez et al. 2013), though the possibility that interstitial telomeric sites exist, but were not detected by FISH, cannot be completely ruled out.
The heterochromatin was concentrated in the pericentromeric regions, and the neo-X chromosome also had terminals blocks, while heterochromatin was observed throughout the neo-Y (Fig. 1c). FISH using the C 0 t-DNA fractions C 0 t-1 and C 0 t-100 yielded strong signals in most centromeric heterochromatic regions, and dispersed faint signals along chromosomal arms were also observed; they were more intense especially in heterochromatin of the neo-Y long arm. These signals were slightly more intense for the C 0 t-100 probe (Fig. 1d, Supplementary material 1a–f). Our FISH mapping of multigene families yielded sex chromosome signals only for H3 histone (in the centromeric region of the neo-X, Fig. 1g) and U2 snDNA (in the interstitial region of the neo-Y, Fig. 1h). H3 histone signals were also seen on autosomal pairs 1–5, 9, and U2 snDNA signals on pair 1. The 18S and 5S rDNAs and U1 snDNA were located exclusively on autosomes (Supplementary material 1g-i). The accumulation of highly/moderately repetitive DNAs and heterochromatinization are common features of many sex chromosome systems with suppressed recombination (Steinemann and Steinemann 1997; Charlesworth et al. 1994, 2005), and are reported in several well studied insect species, such as in the Y of Drosophila melanogaster (Carvalho 2002) and Drosophila miranda (Steinemann and Steinemann 2005; Kaiser and Bachtrog 2010, Zhou et al. 2013) and in the W chromosomes of many lepidopteran species (e.g., Vítková et al. 2007). The accumulation of repetitive sequences can increasing the heteromorphism of sex chromosomes (Charlesworth et al. 2005). The expansion of DNA repeats can also produce novel gene functions, restrict recombination, and lead to chromosomal rearrangements and gene erosion, leading to further morphological and size differences between sex chromosome pairs (Kaiser and Bachtrog 2010).
In R. bergi, C-banding and FISH with C 0 t DNA and some multigene families, reveal similar changes as in other derived sex-systems, i.e., accumulation of repetitive DNAs in the neo-Y chromosome and in the centromere of neo-X and this could account for the subsequent change to the heterochromatic structure. The neo-sex chromosome systems of three other Melanoplinae, Eurotettix minor, Dichromatos lilloanus, and Dichromatos schrottkyi, do not show this change to the heterochromatic condition, with C-positive blocks and C 0 t-1 fractions restricted to the centromeric region (Palacios-Gimenez et al. 2013). This difference could be related to more recent origin of these sex chromosomes or slow rate of repetitive DNAs accumulation and diversification in the three cited species. These conjectures will be better understood only when data concerning the origin of distinct genus and species is available.
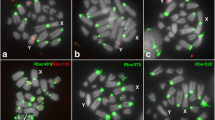
The probes for several microsatellite arrays, (CA)15, (AG)10, (TA)15, (CAC)10, (CAA)10, (CAT)10, (GAC)10, and (GAG)10, showed clustered location revealing a band like hybridization signal at the centromere of neo-X chromosome and some autosomes, besides scattered signals for (CA)15, (CAC)10, (CAA)10, (GAC)10, and (GAG)10 (Fig. 2). The other microsatellite probes, (A)30, (C)30, (CG)15, (CGG)10, (GAA)10, (TAA)10, (GACA)4, and (GATA)8 had no sex chromosome-specific hybridization signals, presenting mainly scattered signals and some clustered sites restrict to autosomes (Supplementary material 2). The observation that some microsatellites have accumulated in the pericentromeric region of neo-X, but not the neo-Y, while the chromosomal arms present similar distribution patterns for these sequences, suggests some differentiation between these chromosomes affecting the pericentromeric region. This resembles microsatellite expansions also observed in the relatively young Y chromosomes of the plants Rumex acetosa (Kejnovský et al. 2013), 12–13 mya (Navajas-Pérez et al. 2005) and S. latifolia (Kubat et al. 2008), 5–10 mya (Bergero et al. 2007). However, this was not detected in the ancient Y chromosomes of the liverwort Marchantia polymorpha or humans (Kejnovský et al. 2013), and, in animals, such as lizards, the distribution of microsatellites on sex chromosomes is independent of the level of heteromorphism (Pokorná et al. 2011b; Matsubara et al. 2013).
FISH for microsatellite probes in male mitotic metaphases of Ronderosia bergi. Each probe that was used is indicated directly on the images. Note the specific signals for microsatellite arrays in a, c, and e and specific and dispersed signals in b, d, and f–h. Also note the specific centromeric signals of microsatellites on the neo-X chromosome but their absence in the centromeric region of the neo-Y chromosome. Bar 5 μm
The differentiation between the R. bergi neo-sex chromosomes is corroborated by the use of probes from the microdissected sex chromosomes. The μY-DNA probe painted exclusively the entire neo-Y chromosome (Fig. 3a,c), demonstrating that this element is enriched in from repetitive DNAs almost no shared with the ex-autosomal homolog (XR arm) of the neo-X involved in the Rb-translocation. The μX-DNA probe showed hybridization signals in the centromeric regions of autosome pairs 3, 4, and 5 and in the neo-X chromosome (Fig. 3b, d), indicating that we have failed to develop a neo-X-painting probe, most likely due to its euchromatic nature. Despite the absence of neo-X-painting, the hybridization pattern of the neo-X probe confirms differentiation of the neo-sex chromosomes. The labeling of some autosomes by our probe derived from the isolated neo-X indicates that the neo-X chromosome carries similar centromeric sequences as autosomes. Accumulation of repetitive DNAs, heterochromatinization in neo-Y and differentiation from neo-X revealed by some probes (Fig. 4) highlight an ancient history of the R. bergi neo-XY sex chromosomes, which could be favored by the large pericentric inversion of neo-Y that involved more than 90 % of its length, including the heterochromatin, except the actual short arm and termini of the long arm (Díaz and Sáez 1968; Cardoso and Dutra 1979; Castillo et al. 2010a,b).
Chromosome painting of the neo-Y and neo-X chromosomes with the μY-DNA and μX-DNA probes, respectively, which were obtained by microdissection in Ronderosia bergi. Each probe that was used is indicated directly on the images. a shows meiotic metaphase I, b shows meiotic metaphase II, and c and d show the neo-XY sex chromosome bivalents that were selected from meiotic metaphase I. Bar 5 μm
FISH signals and histone modifications in the Ronderosia bergi neo-XY sex chromosomes that were analyzed in this study (upper panel). The probes, histone modifications and their relative position on the neo-XY bivalent are indicated using lines and colors. The lower panel shows the selected sex chromosomes, demonstrating the positions of some of the markers that were mapped by FISH
rRNA and snRNA evolution in the sex chromosomes of R. bergi and their relationship with autosomes
Of the four RNA-coding multigene families mapped by FISH, the 5S rDNA and U snDNAs were successfully amplified by PCR from the DNA of the microdissected chromosomes. Alignments of the sequences corresponding to the different haplotypes for U1 and U2 snDNA and 5S rDNA are shown in Supplementary material 3. Concordant with the FISH results, U2 snDNA was amplified by PCR from the μY-DNA. In addition, U1 snDNA and 5S rDNA, which were not detected in the neo-sex chromosomes by FISH, were amplified from μY-DNA and μY-DNA/μX-DNA, respectively. The recovery of U1 snDNA and 5S rDNA from microdissected chromosomes, despite their being undetectable by FISH, suggests that they are present in these elements in a few copies, or that their arrays are scattered. Thus, these sequences could also be involved in the differentiation of the R. bergi neo-XY pair.
We obtained 102 nucleotide partial U1 snDNA sequences from 17 clones from ♀ genomic DNA, and 16 from μY-DNA. Five haplotypes were observed, three exclusive to the female sequences, and two seen only in the μY-DNA. The whole set of U1 snDNA sequences yielded a tree with two clearly separated clades, ♀gDNA and μY-DNA (Fig. 5a). The nucleotide diversity per site (Pi) for ♀gDNA was almost three times higher than in μY-DNA (Table 1). The absence of the U1 snRNA gene from the neo-X chromosome suggests that this gene may have been transposed onto the neo-Y after its origin, and subsequently diverged from autosomal copies, explaining the presence of haplotypes exclusive to the neo-Y.
Neighbor-joining trees showing the relationships between the sequences that were obtained from the genomic DNA of a female (♀gDNA) and from the microdissected neo-X (μX-DNA) and neo-Y (μY-DNA) chromosomes. a Relationship of the U1 snDNA sequences that were derived from ♀gDNA and μY-DNA, b relationship of the U2 snDNA sequences that were derived from ♀gDNA and μY-DNA, and c relationship of the 5S rDNA sequences that were derived from ♀gDNA, μY-DNA and μX-DNA. The haplotypes from ♀gDNA, μY-DNA and μX-DNA are shown in blue, red, and green, respectively; the numbers in parentheses next to each sample type indicate the number of similar sequences for each situation. The trees are drawn to scale, with the branch lengths corresponding to the number of substitutions. The corresponding sequences of Drosophila virilis (U1 snDNA accession number XR049161, U2 snDNA accession number XR049229, and 5S rDNA accession number XR049459) were used as outgroups
For the 141 nucleotides U2 snDNA partial sequences, we found two haplotypes among the 18 ♀gDNA clones analyzed, and three in the 16 μY-DNA clones. Of the total of 34 sequences, 29 were haplotype 1, which was seen in both ♀gDNA and μY-DNA, and the tree from the U2 snDNA sequences shows only one clade (Fig. 5b). Taken together, the presence of U2 copies in the neo-Y and their absence from the neo-X chromosome, based on FISH, our results imply that the neo-Y shares haplotype 1 with autosomal U2 snDNA loci, suggesting transposition of this gene too after the neo-sex chromosomes’ origin, in this case followed by amplification to form a Y cluster detectable by FISH. This is also consistent with the possibility of concerted evolution proposed for this gene (Nei and Rooney 2005). Nucleotide diversity per site was four times higher in the sequences from the μY-DNA than from the ♀gDNA (Table 2), suggesting that some sequences specific to the neo-Y have also evolved faster, possibly as a consequence of the relaxed purifying selection in this chromosome. An alternative interpretation for the results concerning snDNA repeats is that they could have been present in the autosome involved in the Rb-translocation, and then eliminated from the neo-X during sex chromosome differentiation. There is some support for this situation from the 5S rDNA sequences (see below).
For 5S rDNA a more intense sequence diversification was noticed that could be attributed to the multiplicity of sites. Three haplotypes were found in our 13 clones from ♀gDNA, 3 in our 4 μX-DNA clones and 5 in our 10 μY-DNA clones, but there were only 6 haplotypes in total, indicating that haplotypes are shared between autosomes, and neo-X and neo-Y. Tree analysis using the whole set of 5S rDNA sequences revealed only one main clade (Fig. 5c). The nucleotide diversity was more than three times lower in the ♀gDNA than in μY- or μX-DNA (Table 3). Shared haplotypes between the sex chromosomes and autosomes indicate that the 5S rDNA sequences present in both the neo-sex chromosomes could be sequences inherited from the former autosome involved in the X-A Rb-translocation, with some of the sequences having remained after the rearrangement, and some having diverged.
Post-translational histone modifications in R. bergi sex chromosomes
Histone post-translational modifications in R. bergi chromosomes were analyzed to better understand the structure and differentiation between the sex chromosomes and compare with the autosomes. Although we analyzed histone modifications in meiosis I, we focused mainly on metaphase I, in which the sex chromosomes are easily recognized, allowing their precise analysis and a comparison with the autosomes. In other meiotic phases and mitosis, histone modifications could be different from those reported here.
Immunostaining for H3K4me2, H3K4me3, and H4K5ac revealed differences between the autosomes and sex chromosomes at metaphase I. These histone post-translational modifications were detected on the autosomes, the neo-Y and the XR (neo-X) arm, but not the XL (ancestral X) arm of the neo-X (Fig. 4, 6a–c). In contrast, for H3K9me2 and H3S10ph, we detected labeling in both the autosomes and the sex chromosomes (Fig. 4, 6d, e), and for H3K27me2, H3K27ac and H3K9ac there were no signals in either the autosomes or the sex chromosomes (Fig. 6f–h).
Chromatin post-translational modifications on the neo-XY bivalent of Ronderosia bergi selected from metaphase I. Each type of histone modification is indicated directly on the image. XL ancestral arm of the neo-X chromosome; XR originally autosomal part of the neo-X chromosome; Y neo-Y chromosome. Bar 5 μm
The histone post-translational modifications analyzed here are ones expected to be mainly associated with transcriptionally inactive euchromatin or heterochromatin (methylation), with transcription and DNA replication, recombination, and repair (acetylation), or related to chromatin condensation (phosphorylation) (see for example Cobb et al. 1999a,b; Turner et al. 1992, 2000; Manzanero et al. 2000, 2002; Houben et al. 2003, 2013; Kouzarides 2007; Fuchs and Schubert 2012; Page et al. 2012; Oliver et al. 2013). Three types of histone modifications, i.e., H3K4me2, H3K4me3, and H4K5ac, observed in the autosomes are maintained unchanged after the X-A Rb-translocation (appearing similar in the neo-Y and XR arm of neo-X), suggesting little change in post-transcriptional control. The retention of some post-translational histone modifications in regions corresponding to the pre-Rb-translocation autosome suggests that histone modifications change slower than sequence diversification of neo-sex chromosomes.
However, some sequences in the sex chromosomes could have changed post-translational histone modifications that were not detected, due to the intense chromosomal condensation. The absence of signals for H3K4me2 and H3K4me3, and for H4K5 acetylation, from the XL arm of the sex chromosomes may be due this part of the neo-X chromosome is facultatively heterochromatic. Hypoacetylation of H3K9 was also observed in the grasshoppers Eyprepocnemis plorans and Heteracris adspersa and, according to Cabrero et al. (2007), might be a general feature in grasshoppers. The pattern for H3S10ph for the XL arm, which is facultatively heterochromatic and corresponds to the ancestral X, is similar to that described for E. plorans (Manzanero et al. 2000), but differs from two other grasshoppers with X0 systems (Sotero-Caio et al. 2011), suggesting dynamism in this histone modification depending of the species.
Abbreviations
- 2n :
-
Diploid number
- ♀gDNA:
-
Female genomic DNA
- BSA:
-
Bovine serum albumin
- C 0 t-1 DNA:
-
C 0 is the initial concentration of single-stranded DNA in moles per liter and t is the reannealing time in seconds
- DAPI:
-
4′,6-Diamidine-2′-phenylindole
- FISH:
-
Fluorescence in situ hybridization
- mya:
-
Million years ago
- PBS:
-
Phosphate-buffered saline
- PCR:
-
Polymerase chain reaction
- Rb-translocation:
-
Robertsonian translocation
- rDNA:
-
Ribosomal DNA
- rRNA:
-
Ribosomal RNA
- snRNA:
-
Small nuclear RNA
- μX-DNA:
-
X chromosome DNA obtained by microdissection
- μY-DNA:
-
Y chromosome DNA obtained by microdissection
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Bachtrog D (2006) A dynamic view of sex chromosome evolution. Curr Opin Genetic Dev 16:578–585
Bergero R, Forrest A, Kamau E, Charlesworth D (2007) Evolutionary strata on the X chromosomes of the dioecious plant Silene latifolia: evidence from new sex-linked genes. Genetics 175:1945–1954
Bidau CJ, Martí DA (2001) Meiosis and the Neo-XY of Dichroplus vittatus (Melanoplinae, Acrididae): a comparison between sexes. Genetica 110:185–194
Bidau CJ, Martí DA, Castillo ER (2011) Inexorable spread: inexorable death? The fate of neo-XY chromosomes of grasshoppers. J Genet 90:397–400
Bueno D, Palacios-Gimenez OM, Cabral-de-Mello DC (2013) Chromosomal mapping of repetitive DNAs in Abracris flavolineata reveal possible ancestry for the B chromosome and surprisingly H3 histone spreading. PLoS One 8:e66532
Bull JJ (1983) Evolution of sex determining mechanisms. Benjamin Cummings, Menlo Park
Cabral-de-Mello DC, Moura RC, Martins C (2010) Chromosomal mapping of repetitive DNAs in the beetle Dichotomius geminatus provides the first evidence for an association of 5S rRNA and histone H3 genes in insects, and repetitive DNA similarity between the B chromosome and A complement. Heredity 104:393–400
Cabral-de-Mello DC, Valente GT, Nakajima RT, Martins C (2012) Genomic organization and comparative chromosome mapping of the U1 snRNA gene in cichlid fish, with an emphasis in Oreochromis niloticus. Chromosome Res 20:279–292
Cabrero J, Teruel M, Carmona FD, Camacho JPM (2007) Histone H2AX phosphorylation is associated with most meiotic events in grasshopper. Cytogenet Genome Res 116:311–315
Carbonell CS, Mesa A (2006) Ronderosia ommexechoides: a new species of Brazilian Dichroplini (Orthoptera: Acrididae, Melanoplinae). Neotrop Entomol 35:632–637
Cardoso H, Dutra A (1979) The Neo-X Neo-Y sex pair in Acrididae, its structure and association. Chromosoma 70:323–336
Carvalho AB (2002) Origin and evolution of the Drosophila Y chromosome. Curr Opin Genet Dev 12:664–668
Castillo ER, Martí DA, Bidau CJ (2010a) Sex and neo-sex chromosomes in Orthoptera: a review. J Orthopt Res 19:213–231
Castillo ERD, Bidau CJ, Martí DA (2010b) Neo-sex chromosome diversity in neotropical melanopline grasshoppers (Melanoplinae, Acrididae). Genetica 138:775–786
Castillo ERD, Tafarel A, Martí DA (2014) The early evolutionary history of neo-sex chromosomes in Neotropical grasshoppers, Boliviacris noroestensis (Orthoptera: Acrididae: Melanoplinae). Eur J Entomol 111:321–327
Charlesworth D, Mank JE (2010) The birds and the bees and the flowers and the trees: lessons from genetic mapping of sex determination in plants and animals. Genetics 186:9–31
Charlesworth B, Sniegowski P, Stephan W (1994) The evolutionary dynamics of repetitive DNA in eukaryotes. Nature 371:215–220
Charlesworth D, Charlesworth B, Marais G (2005) Steps in the evolution of heteromorphic sex chromosomes. Heredity 95:118–128
Chintauan-Marquier IC, Jordan S, Berthier P, Amédégnato C, Pompanon F (2011) Evolutionary history and taxonomy of a short-horned grasshopper subfamily: The Melanoplinae (Orthoptera: Acrididae). Mol Phyl Evol 58:22–32
Cigliano MM (1997) Ronderosia, a new genus of South American Melanoplinae (Orthoptera: Acrididae). J Orthoptera Res 6:1–19
Cobb J, Cargile B, Handel MA (1999a) Acquisition of competence to condense metaphase I chromosomes during spermatogenesis. Dev Biol 205:49–64
Cobb J, Miyaike M, Kikuchi A, Handel MA (1999b) Meiotic events at the centromeric heterochromatin: histone H3 phosphorylation, topoisomerase IIα localization and chromosome condensation. Chromosoma 108:412–425
Colgan DJ, McLauchlan A, Wilson GDF, Livingston SP, Edgecombe GD, Macaranas J, Cassis G, Gray MR (1998) Histone H3 and U2 snRNA DNA sequences and arthropod molecular evolution. Austral J Zool 46:419–437
Díaz MO, Sáez FA (1968) DNA synthesis in the neo-X neo-Y sex determination system of Dichroplus bergi (Orthoptera: Acrididae). Chromosoma 24:10–16
Drummond AJ, Ashton B, Cheung M, Heled J, Kearse M, Moir R et al. (2009) Geneious v4.8.5, Available from. http://www.geneious.com
Filatov DA, Moneger F, Negrutiu I, Charlesworth D (2000) Low variability in a Y linked plant gene and its implications for Y-chromosome evolution. Nature 404:388–390
Fuchs J, Schubert I (2012) Chromosomal distribution and functional interpretation of epigenetic histone marks in plants. In: Bass HW, Birchler JA (ed.). Plant cytogenetics, plant genetics and genomics: crops and models 4. Springer Science+Business Media, LLC, pp 231–253
Hewitt GM (1979) Grasshoppers and crickets. Animal Cytogenetics. vol 3: Insecta 1. Orthoptera. Gebrüder Borntraeger, Berlin
Hobza R, Kejnovsky E, Vyskot B, Widmer A (2007) The role of chromosome rearrangements in the evolution of Silene latifolia sex chromosomes. Mol Genet Genomics 278:633–638
Houben A, Demidov D, Gernand D, Meister A, Leach CR, Schubert I (2003) Methylation of histone H3 in euchromatin of plant chromosomes depends on basic nuclear DNA content. Plant J 33:967–973
Houben A, Demidov D, Karimi-Ashtiyani R (2013) Epigenetic control of cell division. Springer, Berlin
Ijdo JW, Wells RA, Baldini A, Reeders ST (1991) Improved telomere detection using a telomere repeat probe (TTAGGG)n generated by PCR. Nucleic Acids Res 19:4780
Kaiser VB, Bachtrog D (2010) Evolutions of sex chromosome in insects. Annu Rev Genet 44:91–112
Kejnovsky E, Hobza R, Cermak T, Kubat Z, Vyskot B (2009) The role of repetitive DNA in structure and evolution of sex chromosomes in plants. Heredity 102:533–541
Kejnovský E, Michalovova M, Steflova P, Kejnovska I, Manzano S, Hobza R, Kubat Z, Kovarik J, Jamilena M, Vyskot B (2013) Expansion of microsatellites on evolutionary young Y chromosome. PLoS One 8:e45519
Kouzarides T (2007) Chromatin modifications and their function. Cell 128:693–705
Kubat Z, Hobza R, Vyskot B, Kejnovsky E (2008) Microsatellite accumulation on the Y chromosome in Silene latifolia. Genome 51:350–356
Librado P, Rozas J (2009) DnaSP v5: A software a comprehensive analysis of DNA polymorphism data. Biogeosciences 25:1451–1452
Manzanero S, Arana P, Puertas MJ, Houben A (2000) The chromosomal distribution of phosphorylated histone H3 differs between plants and animals at meiosis. Chromosoma 109:308–317
Manzanero S, Rutten T, Kotseruba V, Houben A (2002) Alterations in the distribution of histone H3 phosphorylation in mitotic plant chromosomes in response to cold treatment and the protein phosphatase inhibitor cantharidin. Chromosome Res 10:467–476
Matsubara K, Knopp T, Sarre SD, Georges A, Ezaz T (2013) Karyotypic analysis and FISH mapping of microsatellite motifs reveal highly differentiated XX/XY sex chromosomes in the pink-tailed worm-lizard (Aprasia parapulchella, Pygopodidae, Squamata). Mol Cytogenet 6:60
Matsunaga S (2009) Junk DNA promotes sex chromosome evolution. Heredity 102:525–526
Mesa A, de Mesa RS (1967) Complex sex-determining mechanisms in thre species of South American grasshoppers (Orthoptera, Acridoidea). Chromosoma 21:163–180
Navajas-Pérez R, de la Herrán R, Jamilena M, Lozano R, Ruiz Rejón C, Ruiz Rejón M, Garrido-Ramos MA (2005) Reduced rates of sequence evolution of Y-linked satellite DNA in Rumex (Polygonaceae). J Mol Evol 60:391–399
Navajas-Pérez R, Quesada del Bosque ME, Garrido-Ramos MA (2009) Effect of location, organization, and repeat-copy number in satellite-DNA evolution. Mol Genet Genomics 282:395–406
Nei M, Rooney AP (2005) Concerted and birth-and-death evolution of multigene families. Annu Rev Genet 39:121–152
Nicolas M, Marais G, Hykelova V, Janousek B, Laporte V, Byskot V, Mouchiroud D, Negrutiu I, Charlesworth D, Monéger F (2005) A gradual process of recombination restriction in the evolutionary history of the sex chromosomes in dioecious plants. PLoS Biol 3:e4
Ohno S (1967) Sex chromosomes and sex linked genes. Springer, Berlin
Oliver C, Pradillo M, Corredor E, Cuñado N (2013) The dynamics of histone H3 modifications is species-specific in plant meiosis. Planta 238:23–33
Page J, de la Fuente R, Manterola M, Parra MT, Viera A, Berríos S, Fernández-Donoso R, Rufas JS (2012) Inactivation or non-reactivation: what accounts better for the silence of sex chromosomes during mammalian male meiosis? Chromosoma 121:307–326
Palacios-Gimenez OM, Castillo ER, Martí DA, Cabral-de-Mello DC (2013) Tracking the evolution of sex chromosome systems in Melanoplinae grasshoppers through chromosomal mapping of repetitive DNA sequences. BMC Evol Biol 13:167
Pinkel D, Straume T, Gray JW (1986) Cytogenetic analysis using quantitative, high sensitivity, fluorescence hybridization. Proc Natl Acad Sci U S A 83:2934–2938
Pokorná M, Giovannotti M, Kratochvíl L, Kasai F, Trifonov VA, O’Brien PCM, Caputo C, Olmo E, Ferguson-Smith MA, Rens W (2011a) Strong conservation of the bird Z chromosome in reptilian genomes is revealed by comparative painting despite 275 million years divergence. Chromosoma 120:455–468
Pokorná M, Kratochvíl L, Kejnovský E (2011b) Microsatellite distribution on sex chromosomes at different stages of heteromorphism and heterochromatinization in two lizard species (Squamata: Eublepharidae: Coleonyx elegans and Lacertidae: Eremias velox). BMC Genet 12:90
Rice WR (1996) Evolution of the Y sex chromosome in animals. Bioscience 46:331–343
Sambrook J, Russel DW (2001) Molecular cloning. A laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, New York
Skaletsky H, Kuroda-Kawaguchi T, Minx PJ et al (2003) The male-specific region of the human Y chromosome is a mosaic of discrete sequence classes. Nature 423:825–837
Sotero-Caio CG, de Souza MJ, Cabral-de-Mello DC, Brasileiro-Vidal AC, Guerra M (2011) Phosphorylation of histone H3S10 in animal chromosomes: is there a uniform pattern? Cytogenet Genome Res 135:111–117
Steinemann M, Steinemann S (1997) The enigma of Y chromosome degeneration: TRAM, a novel retrotransposon is preferentially located on the neo-Y chromosome of Drosophila miranda. Genetics 145:261–266
Steinemann S, Steinemann M (2005) Retroelements: tools for sex chromosome evolution. Cytogenet Genome Res 110:134–143
Sumner AT (1972) A simple technique for demonstrating centromeric heterochromatin. Exp Cell Res 75:304–306
Tamura K, Peterson D, Stecher G, Nei M, Kumar S (2011) Molecular evolutionary genetics using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Traut W, Sahara K, Marec F (2007) Sex chromosomes and sex determination in Lepidoptera. Sex Dev 1:332–346
Turner BM, Birley AJ, Lavender J (1992) Histone H4 isoforms acetylated at specific lysine residues define individual chromosomes and chromatin domains in Drosophila polytene nuclei. Cell 69:375–384
Turner JMA, Shantha K, Mahadevaiah RB, Offenberg HH, Heyting C, Burgoyne PS (2000) Analysis of male meiotic “sex-body” proteins during XY female meiosis provides new insights into their functions. Chromosoma 109:426–432
Vítková M, Fukova I, Kubíčková S, Marec F (2007) Molecular divergence of the W chromosomes in pyralid moths (Lepidoptera). Chromosome Res 15:917–930
Webb GC, White MJD, Contreras N, Cheney J (1978) Cytogenetics of the parthogenetic grasshopper Warramaba (formely Moraba) virgo and its bisexual relatives. IV. Chromosome banding studies. Chromosoma 67:309–339
White MJD (1973) Animal cytology and evolution. Cambridge University Press, Cambridge
Yoshido A, Šíchová J, Kubíčková S, Marec F, Sahara K (2013) Rapid turnover of the W chromosome in geographical populations of wild silkmoths, Samia cynthia ssp. Chromosome Res 21:149–164
Zhou Q, Ellison CE, Kaiser VB, Alekseyenko AA, Gorchakov AA, Bachtrog D (2013) The epigenome of evolving Drosophila neo-sex chromosomes: dosage compensation and heterochromatin formation. PLoS Biol 11:e1001711
Zwick MS, Hanson RE, McKnight TD, Nurul-Islam-Faridi M, Stelly DM (1997) A rapid procedure for the isolation of C 0 t–1 DNA from plants. Genome 40:138–142
Acknowledgments
The authors are grateful to Frantisek Marec for critical reading of the manuscript and to the anonymous reviewers for their substantial contributions, and to “Parque Estadual Edmundo Navarro de Andrade” administration for sample collecting authorization. OMPG acknowledges scholarship obtained from Fundação de Amparo a Pesquisa do Estado de São Paulo-FAPESP (process number 2012/01421-7). This study was partly supported by FAPESP (process number 2014/11763-8), Coordenadoria de Aperfeiçoamento de Pessoal de Nível Superior-CAPES, Conselho Nacional de Desenvolvimento Científico e Tecnológico-CNPq, and the Programa Primeiros Projetos-PROPE/UNESP from Brazil. DAM was supported by Consejo Nacional de Investigaciones Científicas y Técnicas-CONICET from Argentina. The authors are grateful to Antonio Sergio Pascon for technical assistance in obtaining embryos.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary material 1
FISH analysis for the C 0 t-1 and C 0 t-100 DNA fractions and three multigene families in the male mitotic metaphase complements of Ronderosia bergi. Each probe that was used and the neo-XY sex chromosome are indicated directly on the images. Note the absence of signals for multigene families in the sex chromosomes and the propagation of highly and moderately repetitive DNA sequences throughout the long arm of neo-Y chromosome; however, no differences in the distribution of the hybridization signals were observed with the distinct probes. Bar = 5 μm. (GIF 466 kb)
Supplementary material 2
FISH of the microsatellite probes in the male mitotic metaphase complements of Ronderosia bergi. Each probe that was used is indicated directly on the images. Note the specific and dispersed signals of the microsatellite arrays. The sex chromosomes are indicated. Bar = 5 μm. (GIF 509 kb)
Supplementary material 3
Alignment of the multigene family sequences that were isolated from the autosomes and sex chromosomes of Ronderosia bergi and Drosophila virilis. (GIF 169 kb)
Rights and permissions
About this article
Cite this article
Palacios-Gimenez, O.M., Marti, D.A. & Cabral-de-Mello, D.C. Neo-sex chromosomes of Ronderosia bergi: insight into the evolution of sex chromosomes in grasshoppers. Chromosoma 124, 353–365 (2015). https://doi.org/10.1007/s00412-015-0505-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-015-0505-1