Abstract
In mammals, X-inactivation process is achieved by the cis-spreading of long noncoding Xist RNA over one of the female X chromosomes. The Xist binding accumulates histones H3 methylation and H4 hypoacetylation required for X inactivation that leads to proper dosage compensation of the X-linked genes. Co-transcription of Tsix, an antisense copy of Xist, blocks the Xist coating on the Xi. In mice ES cells, an RNase III enzyme Dicer1 disrupts Xist binding and methylated H3K27me3 accumulation on the Xi. Later, multiple reports opposed these findings raising a question regarding the possible role of Dicer1 in murine X silencing. Here, we show that reduction of DICER1 in human female cells increases XIST transcripts without compromising the binding of the XIST and histone tail modifications on the Xi. Moreover, DICER1-depleted cells show differential upregulation of many human X-linked genes by binding different amounts of acetylated histone predominantly on their active promoter sites. Therefore, X-linked gene silencing, which is thought to be coupled with the accumulation of XIST and heterochromatin markers on Xi can be disrupted in DICER1 depleted human cells. These results suggest that DICER1 has no apparent effect on the recruitment of heterochromatic markers on the Xi but is required for inactivation of differentially regulated genes for the maintenance of proper dosage compensation in differentiated cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The equalization of X-linked gene expression between two sexes despite their differences in copy number is mediated by the transcriptional silencing of one of the X chromosomes in female cells (Gribnau and Grootegoed 2012). X inactivation is a multistep process, where choice of the active X chromosome (Xa), establishment of X inactivation in early development, and its maintenance in subsequent cell divisions occur sequentially (Chan et al. 2011). The inactive state of X chromosome is associated with painting of noncoding Xist RNA synthesized from the same chromosome (Hall and Lawrence 2003). The cis-spreading of Xist RNA on the X establishes facultative heterochromatin formation of the inactive X chromosome (Xi), which is orchestrated by accumulation of several modified histones, including H3K27me3, H3K9me2, H4K20me, as well as accumulation of histone variant macroH2A (Payer and Lee 2008). Conversely, Xi is depleted for positive histone marks such as H3K4me2 and H3 and H4 acetylated histones inferring that Xi heterochromatin is not a homogenous entity. In human somatic cells, at least two distinct classes of heterochromatin marks were found, either the presence of XIST RNA, macroH2A, H3K27me3, H4K20me, or the occurrence of constitutive heterochromatin signatures, including late replication timing, H3K9me3, H4K20me, etc. (Payer and Lee 2008). These different heterochromatin domains appear to remain spatially distinct during the metaphase and the interphase (Chadwick and Willard 2004). In an instance, they might act together for bringing mitotically stable higher-order chromatin packaging through several rounds of cell divisions. However, the mechanism of recruitment of epigenetic marks on Xi and stable maintenance of those marks in subsequent cell divisions by XIST RNA still remains unknown.
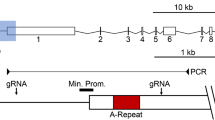
The regulation of Xist expression in random X inactivation is complex in mouse. Tsix, the antisense copy of murine Xist, produces a long (>30 kb) noncoding RNA (ncRNA) reading through same template of Xist coding sequence in spatial and temporal manner, which covers the entire Xist locus (Stavropoulos et al. 2001; Zhao et al. 2008). In mouse, Tsix blocks Xist sense RNA coating on the Xa chromosome despite two major unanswered questions, as follows: how Tsix stably silences Xist RNA on the Xa and how Xist induces X inactivation on the Xi. Recent studies demonstrated that Tsix transcripts particularly read through Xist promoter is essential for Xist silencing, which leads to the modification of chromatin organization (Ohhata et al. 2011). This routine job of TSIX in human dosage compensation still remained inconclusive due to the truncation of the TSIX transcripts during primate evolution (Migeon et al. 2002). Basically, human TSIX differs in its expression pattern and complementarity from its murine counterpart (Migeon et al. 2001). Human TSIX transcripts truncate abruptly before the four exons and the promoter of the XIST locus (Migeon et al. 2001; Sado et al. 2002; Tian et al. 2010; Deuve and Avner 2011) (Fig. 1a). These findings reasonably established that TSIX may not counteract XIST binding on the human Xa through conventional mechanism described in mice. Such species variations in XIST/TSIX structures provide different operational mechanism and accumulation of distinct components of Xi for silencing pathway in humans (Migeon 2003). The repeat A sequence in exon 1 of the Xist locus plays a crucial role in silencing functions of the mouse (Zhao et al. 2008; Royce-Tolland et al. 2010) and for the localization of the XIST RNA in humans (Chow et al. 2007). In mouse RNAi pathway, it was shown to regulate the Xist localization on the X chromosome probably through Xist/Tsix-derived small RNA (Ogawa et al. 2008). The balance of sense and antisense RNA is important in determining the probability that a given Xist allele will be expressed (Nesterova et al. 2008). The formation of these Xi RNAs needed Tsix transcription and in Tsix-deficient cells, their levels dropped significantly (Lee and Lu 1999; Ogawa and Lee 2003). Dicer1 knockout female ES cells eliminated Xi RNA production and disrupted Xist binding and H3K27 methylation on the Xi (Kanellopoulou et al. 2005). In rare case, 5 % depletion of Dicer1 from the normal endogenous level reduced the XiRNA level that leads to a 5–10-fold increase in Xist RNA in the undifferentiated ES cells (Kota 2009). However, further investigations oppose such findings by demonstrating no role of Dicer1 in either Xist RNA coating or H3K27 methylation in mouse cells, which raise questions against their specific role in murine X silencing (Kanellopoulou et al. 2009; Wutz et al. 2002). Moreover, both sense and antisense transcripts modulate Xist promoter DNA methylation in undifferentiated ES cells but do not have any role on Xist binding (Nesterova et al. 2008) In addition, a procedural change in dosage compensation in humans raised further possibility whether RNAi process has any operational role on human X silencing because repeat-associated small interefering RNA (siRNA) plays an instructive role in transcriptional silencing and heterochromatin formation in Drosophila (Yin and Lin 2007). In undifferentiated embryonic mouse cells, the absence of the Tsix or other chromatin-modulating factors often leads to a nominal increase in the Xist expression (Chang and Brown 2010). The deletion in Tsix or DNA methylation up regulates Xist RNA inappropriately during the onset of cell differentiation (Lee and Lu 1999). It suggests an existence of alternate possibly redundant mechanism to separate the regulation of Xist expression from the binding of the Xist and heterochromatin markers on Xi.
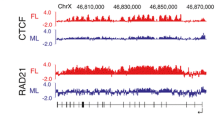
Effect of DICER1 deficiency in cells. A Schematic diagram showing exon 1 region of Xist locus. XIST (red) and TSIX (green) transcripts (in no scale) differ between human and mouse. In mouse Tsix overlaps Xist exon 1 and promoter regions while in humans TSIX transcript truncates before exons 1–4 and promoter regions (p: promoter region, a, b, c, d: conserved repeat elements present within the Exon-1). B The amount of DICER1 transcript was estimated relative to GAPDH mRNA by semiquantitative RT-PCR. The mean values of the triplicate ratios (DICER1/GAPDH) were presented as a bar diagram. *P < 0.05, values significantly different from the no-siRNA and control siRNA transfected control. C Western blot hybridization was performed in no siRNA, control siRNA, and DICER1 siRNA-treated cells, and protein levels were estimated in comparison to the expression of β-ACTIN. D DICER1-dependent microRNA; miR-93 levels were analyzed in no siRNA, control siRNA, and DICER1-transfected HEK-293T cells, and Northern blot analysis was performed. The mean values, as presented by the bar diagram, is determined by triplicate measurements. *P < 0.05, the average values that differ from the values of no siRNA-transfected cells at the 95 % confidence level. E The control siRNA and DICER1 siRNA-transfected human HEK-293T (1Xa, 2Xi) cells were hybridized with fluorescence XIST RNA probe (Wutz and Jaenisch 2000). DNA was also counterstained with DAPI. Scale (white bar), 10 μM
In mammalian RNAi, siRNA is produced by the cleavage of long dsRNA. Dicer1 cleaves long RNA duplex into small noncoding RNA, which are later bound to Argonaute-associated RNAi silencing complex (Bernstein et al. 2001; Okamura et al. 2004). RNAi has also been implicated in the chromatin modifications such as DNA and histone methylations particularly histone H3 lysine methylation in plants (Stroud et al. 2012) and animals (Sinkkonen et al. 2008; Benett et al. 2008). The preferential localization of lysine 9 methylated histone H3 and associated proteins, HP1 that are required for assembly and maintenance of centromeric heterochromatin, were disrupted by the defective RNAi components in Schizosaccharomyces pombe, Drosophila, and in vertebrate cells (Peng and Karpen 2007; Kagansky et al. 2009). Loss of Dicer1 in chicken–human hybrid cells accumulated mitotic cells due to malfunctioning of pericentric heterochromatin in which HP1 was strongly delocalized (Ohfuchi et al. 2006). Therefore, RNAi machinery might be involved in female X silencing. It offers a novel model for determining the role of RNAi in human X silencing in a manner typical to its role in centromeric heterochromatin.
To address an operational role of RNAi on the Xist RNA and heterochromatin proteins binding on the Xi and in human X-linked gene silencing, we have examined two above aspects in the DICER1-depleted human cells. The ability of XIST RNA to bind with heterochromatin factors on the Xi was also monitored. Here, we also uncoupled silencing of certain X-linked genes from the XIST binding on the Xi chromosome during the maintenance of human dosage compensation by the DICER1 enzyme. Our analyses also demonstrated how DICER1 upregulates X chromosomal genes expression without influencing XIST coating and heterochromatin maintenance of the human Xi chromosome.
Materials and methods
SiRNA-mediated depletion
For downregulation of human DICER1 genes by transfecting siRNA, three sets of siRNAs (as previously used, Murchison et al. 2005) were designed and synthesized (Dharmacon, USA). Three siRNAs were efficiently transfected in human culture cells HEK-293T (3A; Xa + 2Xi > 90 % of cells), HeLa (female Xa), and Jurkat (male Xa) using HiPerFect (Qiagen, Germany) reagent according to manufacturer’s protocol (Qiagen, Germany). Transfection of the construct that reduces DICER1 transcripts maximally was selected for further study. The sequences of DICER1 siRNA for most effective DICER1 depletion and control siRNA were noted below.
-
DICER1 siRNA (Murchison et al. 2005) GCUUGAAGCAGCUCUGGAdTdT
-
Control siRNA (without DICER1) AGGUAGUGUAAUCGCCUUGdTdT
Cell culture and transfection
The human cell lines HeLa and HEK-293T cells were grown in Dulbecco’s modified Eagle’s media (DMEM) with 10 % FBS and maintained at 37 °C with 5 % CO2. Jurkat cells were grown in RPMI media supplemented with 10 % FBS under same conditions. Transfections in HeLa, Jurkat, and HEK-293T cells were carried out with HiPerFect Transfection Reagent (Qiagen, Germany). All culture conditions, including the serum batch and seeding density, were kept constant for the entire experiment.
Human cells were plated at 60 % confluency and transfected twice on alternate days, and RNA was isolated using TRIZOL reagent (Invitrogen) after 5 days. Cells were trypsinized and replated in chamber slides after second transfection.
Fluorescent in situ hydridization and immunofluorescence
Human cells were fixed with 3.7 % paraformaldehyde in PHEM buffer (60 mM PIPES, 21 mM HEPES, 10 mM EGTA, 2 mM MgCl2, 685 mM NaCl) for 10 min and permeabilized with 1 % Triton X-100. Three types of cells were immediately used for staining or stored in 70 % alcohol at 4 °C until further use. XIST RNA FISH was carried out using Cy3-labeled random-primed probes as described (Wutz and Jaenisch 2000) that were synthesized using the DECAprime™ II Kit (Ambion) at 37 °C overnight. Cells were washed at 39 °C using 2× sodium citrate/sodium chloride (SSC) and deionized formamide and later with 2× SSC. Cells were counterstained with 4′,6-diamidino-2-phenylindole (DAPI; Vectashield) and visualized under ×20 and ×100 objectives in confocal microscope (FV1000, Olympus). The specificity of XIST RNA probes (Wutz and Jaenisch 2000) was confirmed by destruction of XIST signals, when samples were pretreated with RNase.
For immunofluorescence, fixed cells were immunostained using 1:100 or 1:200 dilution of the following primary antibodies: rabbit or mouse H3K27me3 (Upstate, Abcam), rabbit H3K9 acetyl (Upstate) mouse H3K4 di- and trimethyl (Abcam), rabbit H3K9me3 (Upstate), rabbit BRCA1 (Upstate), glyceraldehyde 3-phosphate dehydrogenase (GAPDH; Santa Cruz). The 1:100 or 1:200 dilutions of secondary anti-rabbit or anti-mouse antibodies conjugated to FITC, Cy3, or Cy5 (Jackson Pvt Ltd) were used and finally counterstained with DAPI or propidium iodide (PI). The cells were observed under confocal or fluorescent microscopes.
Western blots
The control and DICER1 siRNA-transfected and nontransfected HEK-293T and HeLa cells were washed with ice-cold phosphate-buffered saline (PBS), lysed with 1× Laemmli buffer. The protein concentration was estimated by Lowry’s method. The 50 μg protein/per lane was fractionated in 12 % SDS-PAGE gel. Separated proteins were transferred to a PVDF membrane. The membranes were probed with anti-rabbit H3K9me2, H3K27me3, H3K4me3, and H3K9Ac (1:1000; Upstates), anti-rabbit histone H3 antibodies (1:1000), anti-mouse histone 3 (1:1000 dilution), and/or β-ACTIN antibody (1:500 dilution; Progen, USA). The blots were treated with goat anti-rabbit or goat anti-mouse alkaline phosphatase-conjugated secondary antibody (Roche, Germany) in 1:2000 dilutions and developed for color detection using NBT and BCIP.
Total RNA from cells and real-time PCR
A population of siRNA-transfected and nontransfected HEK-293T, HeLa and Jurkat cells was trypsinized and resuspended in DMEM. Cells were collected and washed with PBS and used for RNA isolation. Total cellular RNA was isolated by TRIZOL reagent (Invitrogen) and treated with RNase-free DNase I (Ambion) for 1 h at 37 °C. Two micrograms of DNase-free RNA were added as template for complementary DNA (cDNA) synthesis using Moloney Murine Leukemia Virus (MMLV) reverse transcriptase (Promega) and random hexamers. Further amplification of cDNA was performed by PCR using gene-specific primers. The following primers were used for amplification of each gene.
-
XIST:
-
Forward primer: 5′-AGCTCCTCGGACAGCTGTAA-3′,
-
Reverse primer: 5′-CTCCAGATAGCTGGCAACC-3′;
-
GAPDH:
-
Forward primer: 5′-TGAAGGTCGGAGTCAACGGATTTG-3′,
-
Reverse primer: 5′-TGATGGCATGGACTGTGGTCATGA-3′;
-
Col A45:
-
Forward primer: 5′-ACAGCTTTTGGCTGGCAACT-3′,
-
Reverse primer: 5′-CGGCTAATTCGTGTCCTCAAG-3′;
-
MAGE H1:
-
Forward primer: 5′-CCTTTGTAGCTGAGAACGGC-3′,
-
Reverse primer: 5′-CCTCTTCAGGAAGCAACAGC-3′;
-
Hemophilia AF8:
-
Forward primer: 5′-GTGGCAGACTTATCGAGGAAA-3′,
-
Reverse primer: 5′-TCGAGCAATAATTGGAGGGT-3′;
-
SMARCA 1:
-
Forward primer: 5′-ACGAAATGCTCCCCAGTTTA-3′,
-
Reverse primer: 5′-TCTCTCTCTTTCCTCAATTTCCA-3′;
-
KIAA1166:
-
Forward primer: 5′-CCCTTGGCTGGTGTATTTGT-3′,
-
Reverse primer: 5′-CTCCATCTGCAGGGTCTTGT-3′;
-
MOSPD2:
-
Forward Primer: 5′-GATCATGGCAGAGAATCACG-3′,
-
Reverse primer: 5′-GATCATGGCAGAGAATCACG-3′.
Quantitative real-time PCR was carried out using the Roche Lightcycler® 480 II system using Sybr Fast Universal Ready Mix (KAPA Biosystems) according to manufacturer’s protocol. The software in Lightcycler 480 II was used to analyze and quantify the PCR products.
Northern transfer and hybridization
Total cellular RNA from no siRNA, control siRNA, and DICER1 siRNA-transfected cells were prepared, electrophoresed, and hybridized with antisense miRNA 93-synthesized probe. Blots were prepared in triplicate for each probe. As a loading control, the blots were reprobed with β1-tubulin RNA. Quantification was performed using a Fuji 2000 Bas Phosphorimager.
Chromatin immunoprecipitation
RNA-ChIP was performed as reported earlier (Cavalli et al. 1999; Sun et al. 2006). For a detailed method, please see Electronic supplementary material.
Results
Production of DICER1-depleted human cells
Since RNase III enzyme DICER1 has a central role in RNA interference pathway, we were interested to see whether it has any influence on human dosage compensation and maintenance of X inactivation. For this, we had generated three cell types, HEK-293T (a female embryo lineage, 3A; Xa + 2Xi), Jurkat (male Xa), and HeLa (female Xa) deficient for DICER1 gene. Initially, DICER1-depleted human cell lines were generated by incubating cells in DICER1 gene-specific siRNA-containing media as described earlier (Murchison et al. 2005). To ensure whether the siRNA dramatically reduced the DICER1 transcripts, quantitative real-time PCR analysis was conducted. Data showed a multifold (6-fold) reduction of DICER1 messenger RNA (mRNA) within 5 days by incorporating siRNA at a regular interval (alternate days) relative to the same mRNA from untreated (no siRNA) and control siRNA-transfected HEK-293T (3A; Xa + 2Xi) cells (Fig. 1b). The reduction of DICER1 expression was also quantitated in the protein (Fig. 1c) levels (Murchison et al. 2005) using Western blot hybridization. In both cases, treatment of DICER1 inhibitory siRNA reduced transcripts and DICER1 proteins dramatically as verified by quantitative RT-PCR and Western blot analysis respectively. To verify whether this reduction of DICER1 protein is sufficient to negate its function, an ideal DICER1-dependent miRNA, miR-93 levels were analyzed in DICER1 siRNA-treated cells by northern blot analysis. Considerable decrease of miR-93 in DICER1-deficient HEK-293T cells (Fig. 1d) indicated a successful functional knockdown of DICER1. Cells at this stage were immediately fixed and used for further study. To test a potential link between RNAi, X chromosomal identity of noncoding RNA and X chromosomal dosage compensation in humans, three human cell lines, HEK-293T(3A; Xa + 2Xi), HeLa (with no Xi-stained chromosome), Jurkat (Only male Xa) were verified by fluorescence in situ hybridization (FISH) using XIST RNA probe (Supplementary Fig. 1; Wutz and Jaenisch 2000). The current dogma is that noncoding XIST RNA is transcribed from inactive center of the X (XIC) and is only coated on silent X chromosome in females; however, its binding on active male X was not detected (Gribnau and Grootegoed 2012; Chan et al. 2011). Generally, two inactive X chromosomes coated with XIST RNA were only present in HEK-293T cells (3A; Xa + 2Xi) while XIST RNA fails to hybridize with existing X chromosome in Jurkat and HeLa cells (Supplementary Fig. 1).
Binding of XIST RNA
To test whether DICER1 affects XIST painting on Xi, DICER1-depleted cells were hybridized with fluorescent XIST RNA probe. In mouse ES cells, loss of Dicer1 failed to paint Xist RNA on Xi in the interphase nuclei (Ogawa et al. 2008). The region of the X heterochromatin in transfected nuclei showed an intense DAPI staining as a signature for strong XIST RNA localization (Supplementary Fig. 2). Because high chromatin compaction in heterochromatin always shows intense DAPI staining, a similar level of condensation was also noticed in DICER1-depleted HEK-293T cells (Fig. 1e). Thus, DICER1 has no apparent effect on the Xi condensation in female human cells. For better understanding of the XIST RNA localization on the entire inactive X, an in situ hybridization analysis was conducted using fluorescent XIST RNA probe in human cultured cells. The silent Xs of HEK-293T cells (3A; Xa + 2Xi) were coated with two XIST RNA (Fig. 2a) but HeLa (female Xa) and Jurkat (male Xa) cells do not carry any XIST- coated X chromosome (Supplementary Fig. 1). Once the cells are transfected with DICER1 siRNA, XIST still binds on the inactive Xs (Fig. 2a). Enumeration of XIST foci in no siRNA, control siRNA, and DICER1 siRNA-transfected HEK-293T (3A; Xa + 2Xi) cells showed nearly the same numbers. There is no significant difference in XIST foci number in three transfected cell types (Fig. 2b). Therefore, depletion of DICER1 does not compromise intense XIST RNA coating on the Xi chromosomes.
Localization of XIST RNA by degradation of human DICER1 mRNA on multiple inactive Xs in the human HEK-293T cells. a Fluorescence in situ hybridization of XIST RNA on control siRNA or DICER1 siRNA-transfected HEK-293T cells. Scale (white bar), 10 μM. b Bar diagram showing enumeration of XIST foci in no siRNA, control siRNA, or DICER1 siRNA-transfected cells. The mean values ± S.D. from the transfected cells (n = 460 cells) were presented by the bar diagram (P < 0.01). The types of siRNA are noted in the diagram
Recruitment of chromatin modifications
Heterochromatinization of human Xi chromosome is characterized by accumulation of more repressive histone codes associated with XIST RNA binding. XIST RNA coating is the first event in a cascade of changes on the Xi chromosome. Several early histone modifications including enrichment of macro-H2A deposition (Li 2002), histone H3K27me3 (Chan et al. 2011), and H3K9me2 (Rougeulle et al. 2004) impart profound effects on Xi heterochromatin formation coupled with X-linked gene silencing. Using immunofluorescence technique, an enrichment of modified histone isoforms including H3K27me3 on the Xi chromosome was noticed as a Xi marker (Fig. 3a) in control and DICER1 siRNA-transfected HEK-293T cells (3A; Xa + 2Xi). The statistical analysis on H3K27me3-bound Xi revealed that nearly equal number of Xi-bound H3K27me3 foci existed in variant cells (Fig. 3b).
Localization of X silencing histone markers H3K27me3 in DICER1 knockdown cells. a Immunostaining was performed on control siRNA and DICER1 siRNA-treated HEK-293T cells using antibodies against H3K27me3 (middle panel, green) or DAPI (left panel, blue). Merged image is shown in the right. Scale (white bar), 10 μM. Inset, the histogram of H3K27me3+ve foci in DICER1-transfected cells, compared with cells having control siRNA and cell with no siRNA. The mean values are derived from at least three measurements of the foci. The average value with S.D. are presented by a histogram. The total (N = 375 cells) were measured in each case
Apart from early chromatin modifications, H3K9 methylation and XIST RNA overlaps on the inactive X chromosomes suggesting that enrichment of histone H3 lysine 9 methylation might also be involved in the maintenance of stable transcriptional silencing. In most cases, major heterochromatin marks in mammalian nuclei, H3K9me2, and DNA methylation ensure complete nucleolar organization, which is essential for RNAi machinery (Peng and Karpen 2007; Kondo et al. 2008; Kagansky et al. 2009). The level of H3K9me2 and other major histone marks were estimated by Western blots from HEK-293T cells treated with no siRNA, control siRNA, and/or DICER1 siRNA (Fig. 4a). We also measured the amount of histone H3 by the Western blot hybridization as a loading control. To avoid saturation of the gel blots, we also rehybrized the same blots with β-ACTIN probe as a gel control. The level of methylated histone H3 proteins (H3K9me2 and H3K27me3) in DICER1 siRNA-transfected cells was changed at a marginal level relative to no siRNA and control siRNA transfected cells. However, the total amount of extractable histone H3 and β-ACTIN proteins in both cases appear to be similar, which suggests that H3 and β-ACTIN modifications were unaffected in the DICER1, control, and no siRNA-incorporated cells. Thus, DICER1 is not important for targeting XIST RNA and histone methylation although H3K9me2 level remained unaltered on the Xi chromosomes, which readily accounts for higher order chromosome packaging needed for Xi maintenance.
Analysis of localization of heterochromatic markers with no siRNA, with control siRNA and DICER1 knockdown cells. a Western blot analysis of H3K9me2, H3K27me3, H3K4me3, and H3K9Ac in no siRNA, control siRNA, and DICER1 siRNA-treated HEK-293T cells. Histone 3 and β-ACTIN (nonsaturated) were also used as positive control. b Bar diagram showing the ratio of expression of H3K9me2, H3K4me3, H3K27me3, and H3K9Ac with respect to the expression of β-ACTIN and in no siRNA, control siRNA, and DICER1 siRNA-transfected cells. *P < 0.01, the average values (triplicate mean values ± S.D.) that differ from the no siRNA-transfected cells at the 95 % confidence level. c FISH analysis in no siRNA, control siRNA and DICER1-transfected cells stained with propidium iodide (PI, left panel, red) or α-H3me2K9 (middle panel, green). Right, merged pictures. Scale (white bar), 10 μM. Inset, comparison of H3K9me2 in no siRNA, control siRNA, and DICER1 siRNA-transfected HEK-293T cells
When mean ratios of H3K27me3/H3K9me2 were calculated in comparison to β-ACTIN control in no, control, and DICER1 siRNA-treated cells, almost a similar pattern was observed for those heterochromatin markers (Fig. 4b). On the other hand, both the euchromatin-activating factors, such as H3K4me3 and H3K9Ac showed nearly 1.5–2-fold enhancement in DICER1-depleted cells (Fig. 4b), which suggested that expression and localization of histone H3K9 and H3K27 methylations were not affected only by the depletion of DICER1 human mRNA. The histone H3K9me2 was compared between cells containing intact and reduced DICER1 mRNA (Fig. 4c). The antibodies were accumulated mainly at the centromeric regions of the chromosomes in HEK-293T (Fig. 4c) as isolated spots. The strong H3K9me2 accumulation in control siRNA-transfected HEK-293T cells does not reduce markedly with DICER1 siRNAs (Fig. 4c). The level of histone H3 lysine 9 methylation associated with RNAi machinery has no apparent role on stabilization of the XIST RNA coating and/or maintenance of the X inactivation. These findings do not reflect the loss of higher order of chromosomal condensation in inactive X chromosome. These results ruled out the possibility that accumulation of lysine 9-methylated histone H3 on the Xi and its negligible delocalization by the loss of DICER1 activity might be due to higher nuclear density of the silent X chromatin.
Conversely, the Xi is depleted for marks commonly linked to transcriptional activators, for example, H3K4me3 methylation (Fig. 5). These results revealed that Xi silencing is not a homogeneous entity. In human somatic cells, a consistent reduction of H3K4me2, H3K4me3 binding on the Xi was detected as colorless foci in the immunostained nuclei. Double staining with H3K4me3 and H3K27me3 (Fig. 5) antibodies showed that reduction of DICER1 did not increase the level of H3K4me3 accumulation on the Xi chromosome, which was detected by the excess H3K27me3 deposition in female cells.
Immunofluorescence of H3K4me3 in control and DICER1 knockdown human cells. Control siRNA and DICER1 siRNA-treated cells were stained with DAPI (left, blue), H3K27me3 (2nd from left, red), and H3K4me3 (3rd from left, green) antibodies. Right, H3K27me3 and H3K4me3 colocalized image. Here, H3K27me3 was used as inactive X chromosome marker. The localization of two methylations (H3K27me3 and H3K4me3) in the cells does not overlap. Scale (white bar), 10 μM
Colocalization of BRCA1 and Xist
A XIST-interacting partner, BRCA1 functions in DNA damage pathways as a ubiquitin ligase (Stucki et al. 2005; Wu et al. 2010). BRCA1 mutations predispose carriers of hereditary breast or ovarian cancer. Many breast cancers related to BRCA1 deletions no longer show a Barr body, suggesting a loss or reactivation of the inactive X chromosome (Stone et al. 2003; Pageau et al. 2007). It is established that BRCA1 protein is required for XIST RNA localization on the Xi (Ganesan et al. 2002). Colocalization of BRCA1 and XIST RNA on the Xi indicated that BRCA1 is a transfactor agent involved in XIST localization. Later, a similar study failed to reproduce the overlapping binding of the XIST RNA and BRCA1 in the same nuclei (Pageau et al. 2007). In another study, X-linked gene expression and XCI were not disrupted in BRCA1-deficient mice (Sirchia et al. 2005). These findings are clearly hard to reconcile with each other.
To reinvestigate the overlapping binding BRCA1 and XIST RNA on Xi, tumor MCF7 cells as control and nonneoplastic HEK-293T (3A; Xa + 2Xi) cells were immunostained with Xi marker H3K27me3 and BRCA1 antibodies (Fig. 6). The BRCA1 antibody is associated with different subnuclear regions in more than 95 % of the nuclei in control MCF 7 cells and HEK-293T cells. But these bindings fail to overlap with H3K27me3-enriched region or Xi chromosome, which are marked by the enrichment of H3K27me3 antibody in both tumor and normal cells (Fig. 6). The absence of overlapping BRCA1 and H3K27me3 binding on the Xi suggest that BRCA1 has no definitive role in the X-inactivation process. DICER1-depleted cells increased BRCA1 binding in comparison to the control siRNA or no siRNA-transfected cells; the amount of DICER1 depletion which enhanced the BRCA1 binding throughout the nuclei suggested (Supplementary Fig. 3) that DICER1 intervenes BRCA1 binding of the human nuclei.
Analysis of binding BRCA1 and XIST RNA on Xi. Tumor MCF7 cells (prominent for BRCA1 marker used as control) and nonneoplastic HEK-293T (3A; Xa + 2Xi) cells were immunostained with Xi marker H3K27me3 and BRCA1 antibodies. The BRCA1 antibody is associated with different subnuclear regions in more than 95 % of the nuclei. Scale (white bar), 10 μM
X-linked gene expression
X silencing compensates the dosage of X-linked gene expression between male and females in mammals. But some genes escape silencing and are thus expressed for both alleles in females. In mouse, nearly 3–4 % mouse genes are expressed from the inactive X chromosome (Yang et al. 2010), whereas about 15 % of X-linked genes in human escape inactivation to some degree (Carrel and Willard 2005). Such genes potentially contribute to the sexual dimorphic traits to phenotypic variation.
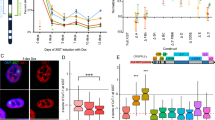
To test whether the X-linked gene repression is maintained in the cells with reduced expression of DICER1, we performed real-time PCR analysis of select transcripts that are located in many loci along the X chromosome. Consistent with the role of Dicer1 in regulating the mouse Xist levels, an increase in human XIST levels in DICER1 knockdown cells was detected relative to no siRNA and control siRNA-transfected cells in both HEK-293T (3A; Xa + 2Xi) and Jurkat (Only male Xa) (Fig. 7a). Since the increase is nearly 2–2.5-fold in case of HEK-293T cells, it suggested that XIST loci in inactive X chromosome in HEK-293T cells was expressed. This involves upregulation of XIST transcripts in DICER1-depleted HEK-293T cells. DICER1 is involved in siRNA and miRNA biogeneses in humans. The reduction of DICER1 in these two cell lines delimits the influence of siRNA and endogenous microRNA in both cases on the X-linked transcripts. We used these cells for further experiments.
Upregulation of X-linked gene expression: a quantitative estimation XIST mRNA (real-time PCR) from no siRNA, control siRNA, and DICER1 siRNA-transfected HEK-293T (3A; Xa + 2Xi) and Jurkat (only male Xa) (top) cells were presented. The relative ratios (n = 3) of gene/GAPDH of triplicate values were plotted in a bar diagram. The standard deviations of each mean values were present at the top of each bar. The name of each gene was sited at the bottom of the histogram. *P < 0.05, values were significantly different from the respective control. b The amount of DICER1 transcript was assayed relative to GAPDH mRNA by semiquantitative RT-PCR in the Jurkat cells. The mean values of the triplicate ratios (DICER1/GAPDH) were presented as a bar diagram. *P < 0.05, values are significantly different from the no siRNA-transfected control. c Expression of six X-associated transcripts, COL4A5, MAGEH1, hemophilia A F8, SMARCA1, KIAA1166, and MOSPD2 analyzed by RT-PCR in Jurkat (top) and HEK-293T(bottom) cell lines. The quantitative values of six different transcripts of control siRNA and DICER1 siRNA-transfected cells relative to no siRNA cells with S.D. noted in the key, is represented by the bar diagram. Each ratio is determined by triplicate measurements. *P < 0.05, 95 % confidence interval
To determine whether repression of RNAi factor has no impact on the inactive X chromosome restoring its activity, we have measured six different X-linked transcripts, viz., collagen alpha-5(IV) chain (COL4A5), melanoma-associated antigen H1 (MAGEH1), hemophilia A factor VIII (hemophilia A F8), SWI/SNF-related, matrix-associated, actin-dependent regulator of chromatin, subfamily a, member 1 (SMARCA1), hepatocellular carcinoma-associated antigen 127 (KIAA1166), motile sperm domain-containing 2 (MOSPD2), from the DICER1-transfected, no siRNA, and control siRNA-transfected cells (Fig. 7a, b). Selection of six X chromosomal genes is important, and they are not involved in sexual dimorphism or phenotipic variation in humans. Six X-associated transcripts, COL4A5, MAGEH1, hemophilia A F8, SMARCA1, KIAA1166, and MOSPD2 were increased significantly in DICER1-transfected HEK-293T cells relative to the amount in no siRNA and control siRNA-transfected cells (Fig. 7b) However, the amount was almost similar in three cases of no siRNA, control siRNA, and DICER1 siRNA-transfected Jurkat cells. It indicated that 2Xi chromosome containing female cells (HEK-293T, 3A; Xa + 2Xi) increases the transcription compared with Xa-containing Jurkat cells. The elevated X transcript in general revealed that the genes were transcriptionally active regardless of their influence of siRNA and miRNA in DICER1-reduced cells (Fig. 7a, b).
A consistent but different degree of upregulation of X-linked mRNA was observed in DICER1-depleted HEK-293T cells relative to DICER1-reduced Jurkat cells. A comparison with control siRNA-transfected cells demonstrating that DICER1 is involved in a post-establishment Xi process for maintaining the balance of certain X-linked gene dosage in human somatic cells revealed that DICER1 is a transcriptional target for individual gene expression; therefore, DICER1 uncoupled functionally the X-linked gene expression from the chromatin modifications including XIST binding on the silent X chromosome during maintenance of human dosage compensation. These outcomes are consistent with earlier findings that DICER1 depletion in fact had a global effect on gene expression, including upregulation of nearly >2000 genes in HEK-293T cells (Tang et al. 2007).
Recruitment of chromatin modifiers on X-linked gene promoters
To correlate possible chromatin changes occurring gene activation, we determined by a series of ChIP assays coupled with real-time PCR the histone methylation state at the regulatory regions of the four X-linked genes, COL4A5, MAGEH1, KIAA1166, and MOSPD2 (Fig. 8a, b). We have analyzed major histone methylation, histone H3K27me3 and H3K9me2, whose excess accumulation is required for chromatin changes of Xi chromosome. The enrichment of histone methylation H3K27me3 and H3K9me2 is commonly associated with gene repression. Results showed that H3K9me2 expression was nominally decreased in two genes (COL4A5 and MOSPD2) in the DICER1-deficient cells relative to the no siRNA and nonspecific siRNA-transfected controls in a sex-specific manner, while no significant changes were observed of their accumulation at the promoter region of MAGEH1 and KIAA1166 genes (Fig. 8a, Supplementary Fig. 4). Furthermore, we analyzed the levels of trimethylated H3K27, a histone modifier that has shown an enriched accumulation on the Xi. However, their enriched accumulation in Xi does not lead to their higher accumulation at the promoters of all four tested genes in DICER1-depleted cells (Fig. 8a, b). Therefore, DICER1 has no contribution on the localized chromatin modification related to the upregulated genes. The upregulation of X-linked gene activities and chromatin modulations via histone methylation are two coexisting but separate processes, which might be differentiated by the DICER1 activity in humans. However, when we tested whether the euchromatin-activating factors H3K9Ac and H3K4me3 are associated with the expression of the abovementioned foue X-linked genes, we found a great enhancement of expression of all the four genes in both H3K9Ac and H3K4me3 (Fig. 8b).
Variable enrichment of four histone codes in four X-linked gene promoters. a Accumulation of histone H3K27me3 and H3K9me2 proteins and b histone H3K9Ac and H3K4me3 proteins in COL4A5, MAGEH1, and KIAA1166 MOSPD2 gene promoters in no siRNA, control siRNA, and DICER1 siRNA-transfected HEK-293T cells were shown. The quantitative data with respect to the X-linked gene as compared to the accumulation of GAPDH were represented by the bar diagrams. Each ratio (mean values ± S.D.) is determined by triplicate measurements. *P < 0.01, average values that differ from the no siRNA-transfected cells at the 95 % confidence level
Discussion
In different ways, human dosage compensation is distinct from its mouse counterpart. These differences in origin and function of X-inactivation process might underlie the subtle variations of the dosage compensation mechanism between primate and nonprimate mammals. In humans, truncation of TSIX during evolution and its negligible role in Xi imprinting supports this view (Migeon et al. 2001). Contrary to the earlier observations in mice ES cells (Shekar et al. 2011), our results indicate DICER1 has no role in the maintenance of XIST or H3K27 marks on the Xi. It is likely that general epigenetic marks including XIST RNA and other histone modifications once established during embryonic development allows DICER1-independent maintenance of Xi chromosomes. On the other hand, the existence of temporally regulated Xist and Tsix duplexes and probable processing of these duplexes into small XiRNA and that the absence of functional Dicer1 in mouse ES cells leads to loss of Xist and H3K27 trimethylation on Xi (Shekar et al. 2011). These changes in Dicer1-depleted mouse ES cells may be caused by amenability of ES cell chromatin to short-term changes and recruitment of epigenetic activators of the Xi genes. Surprisingly, in agreement with our present report that DICER1 has no significant role in binding of heterochromatin marks on human Xi, several groups demonstrated that mouse Xi is faithfully propagated in Dicer1-deficient mature T cells (Liston et al. 2008), in mouse embryos (Kanellopoulou et al. 2005), and in mouse male and female ES cells (Murchison et al. 2005). Even Dicer1 hypomorphic mice also showed no differences in genomic imprinting in tested loci (Fukasawa et al. 2006). These contrasting results relating to the function of mammalian Dicer1 in nuclear-related transcriptional processes are surprising considering the evolutionary conservation in function of these enzymes in post-transcriptional gene regulation. Knowledge regarding the specific cofactors that aid or mediate the functions of Dicer1 in distinct cellular compartments will definitely help in dissecting the roles of small RNA in epigenetic processes.
Together, our results show that XIST binding and hypermethylation of histone H3K27 contributed in Xi establishment. However, DICER1 knockdown resulted in elevation of X-linked gene expression. Some genes (XIST, MOSPD2, COl4A5, and MAGEH1) are more sensitive than others, suggesting a selective and preferential effect of DICER1 in human dosage compensation (Supplementary Fig. 3). Gene regulation profiles in mouse and humans in the Xi chromosome is quite distinct. Mice have significantly fewer escaper genes compared to humans. Furthermore, escape genes do not cluster in mouse, unlike the large escape domains in humans, suggesting that expression is regulated at the level of individual genes (Yang et al. 2010), whereas the proportion of genes in human escaping inactivation differs dramatically between different regions of the X chromosome, reflecting the evolutionary history of the sex chromosome (Carrel and Willard 2005). Even a whole genome expression analysis in DICER1-deficient cells also showed upregulation of more than 2000 genes (Tang et al. 2007). Therefore, it is now an open question whether DICER1 has any role in sex chromosome evolution or not.
Dicer1 has been shown to aid in establishment of proper DNA methylation by regulating repressors of DNA methyl transferases (DNMTs) through miRNA-290 cluster-mediated inhibition in mouse ES cells and through unknown mechanism in human cancer cell lines. The functional role of Dicer1-dependent RNAi process is a global increase of X-linked gene expression that may be uncoupled from the binding of Xist and heterochromatin markers on the Xi chromosome. Therefore, the study only showed the maintenance of silencing and related chromatin modifications on the Xi chromosome in consecutive cell divisions. It is functionally different from its individual control on specific gene expression. The influence of DICER1-associated complex on gene expression might be contributed by the direct or indirect miRNA-mediated functions of Dicer1 rather than through the modification of chromatin state. Similarly, it is also possible that upregulation of gene expression is also an effect of disruption of microRNA-mediated gene regulation. Studies with human female ES cells will be useful in determining their specific functions in DICER1 in Xi initiation and establishment and also provide a new platform for investigating the role of nuclear RNAi in heritability and maintenance of higher-order Xi chromatin structure in primates.
Xist locus controls the process of X inactivation and recruitment of heterochromatin markers on the silent X chromosome during initiation phase of X inactivation (Wutz and Jaenisch 2000). Its role is thought to be redundant for maintenance of X inactivation. Our data shows that XIST locus in humans is indispensible for successive maintenance of X-mediated gene expression and genome stability. Moreover, it would be interesting to see the expression and the binding role of XIST in humans at the initiation of X-inactivation process, where XIST RNA along with its partner TSIX is known to mediate the counting and choice of Xi. Consistent with our presumption, whole genomic microarray analysis of XIST/TSIX small RNA <200 nt revealed extensive presence of small RNA including −35 and −50 nt small RNA at several gene loci (Kapranov et al. 2007), some of which are conserved between mouse and humans, hinting at plausible important regulatory roles played by these classes of RNA in genome maintenance and organization.
References
Benett R, Gonzalo S, Jaco I et al (2008) A mammalian microRNA cluster controls DNA methylation and telomere recombination via Rbl2-dependent regulation of DNA methyltransferases. Nat Struct Mol Biol 15:268–279
Bernstein E, Caudy AA, Hammond SM, Hannon GJ (2001) Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409:363–366
Carrel L, Willard H (2005) X-inactivation profile reveals extensive variability in X-linked gene expression in female. Nature 434:400–404
Cavalli G, Orlando V, Paro R (1999) Mapping DNA target sites of chromatin-associated proteins by formaldehyde cross-linking in Drosophila embryos. In: Bickmore WA (ed) Chromosome structural analysis: a practical approach. Oxford University Press, UK, pp 20–37
Chadwick BP, Willard HF (2004) Multiple spatially distinct types of facultative heterochromatin on the human inactive X chromosome. Proc Natl Acad Sci U S A 101:17450–17455
Chan K, Zhang H, Malureanu L, van Deursen J, Zhang Z (2011) Diverse factors are involved in maintaining X chromosome inactivation. Proc Natl Acad Sci U S A 108:16699–17704
Chang SC, Brown CJ (2010) Identification of regulatory elements flanking human XIST reveals species differences. BMC Mol Biol 11:20
Chow JC, Hall LL, Baldry SE, Thorogood NP, Lawrence JB, Brown CJ (2007) Inducible XIST-dependent X-chromosome inactivation in human somatic cells is reversible. Proc Natl Acad Sci U S A 104:10104–10109
Deuve JL, Avner P (2011) The coupling of X-chromosome inactivation to pluripotency. Annu Rev Cell Dev Biol 27:611–629
Fukasawa M, Morita S, Kimura M, Horii T, Ochiya T, Hatada I (2006) Genomic imprinting in Dicer1-hypomorphic mice. Cytogenet Genome Res 113:138–143
Ganesan S, Silver DP, Greenberg RA, Avni D, Drapkin R et al (2002) BRCA1 supports XIST RNA concentration on the inactive X chromosome. Cell 111:393–405
Gribnau J, Grootegoed JA (2012) Origin and evolution of X chromosome inactivation. Curr Opin Cell Biol 24:397–404
Hall LL, Lawrence JB (2003) The cell biology of a novel chromosomal RNA: chromosome painting by XIST/Xist RNA initiates a remodelling cascade. Semin Cell Dev Biol 14:369–378
Kagansky A, Folco HD, Almeida R, Pidoux AL, Boukaba A et al (2009) Synthetic heterochromatin bypasses RNAi and centromeric repeats to establish functional centromeres. Science 324:1716–1719
Kanellopoulou C, Muljo SA, Kung AL, Ganesan S, Drapkin R et al (2005) Dicer-deficient mouse embryonic stem cells are defective in differentiation and centromeric silencing. Genes Dev 19:489–501
Kanellopoulou C, Muljo SA, Dimitrov SD, Chen X, Colin C et al (2009) X chromosome inactivation in the absence of Dicer. Proc Natl Acad Sci U S A 106:1122–1127
Kapranov P, Willingham AT, Gingeras TR (2007) Genome-wide transcription and the implications for genomic organization. Nat Rev Genet 8:413–423
Kondo Y, Shen L, Ahmed S, Boumber Y, Sekido Y et al (2008) Downregulation of histone H3 lysine 9 methyltransferase G9a induces centrosome disruption and chromosome instability in cancer cells. PLoS One 3:e2037
Kota SK (2009) RNAi in X-inactivation: contrasting findings on role of interference. BioEssays 31:1280–1283
Lee JT, Lu N (1999) Targeted mutagenesis of Tsix leads to nonrandom X inactivation. Cell 99:47–57
Li E (2002) Chromatin modification and epigenetic reprogramming in mammalian development. Nat Rev Genet 3:662–673
Liston A, Lu LF, O'Carroll D, Tarakhovsky A, Rudensky AY (2008) Dicer-dependent microRNA pathway safeguards regulatory T cell function. J Exp Med 205:1993–2004
Migeon BR (2003) Is Tsix repression of Xist specific to mouse? Nat Genet 33:337
Migeon BR, Chowdhury AK, Dunston JA, McIntosh I (2001) Identification of TSIX, encoding an RNA antisense to human XIST, reveals differences from its murine counterpart: implications for X inactivation. Am J Hum Genet 69:951–960
Migeon BR, Lee CH, Chowdhury AK, Carpenter H (2002) Species differences in TSIX/Tsix reveal the roles of these genes in X-chromosome inactivation. Am J Hum Genet 71:286–293
Murchison EP, Partridge JF, Tam OH, Cheloufi S, Hannon GJ (2005) Characterization of Dicer-deficient murine embryonic stem cells. Proc Natl Acad Sci U S A 102:12135–12140
Nesterova TB, Popova BC, Cobb BS, Norton S, Senner CE et al (2008) Dicer regulates Xist promoter methylation in ES cells indirectly through transcriptional control of Dnmt3a. Epigenetics Chromatin 1:2
Ogawa Y, Lee JT (2003) Xite, X-inactivation intergenic transcription elements that regulate the probability of choice. Mol Cell 11:731–743
Ogawa Y, Sun BK, Lee JT (2008) Intersection of the RNA interference and X-inactivation pathways. Science 320:1336–1341
Ohfuchi E, Kato M, Sasaki M, Sugimoto K, Oma Y, Harata M (2006) Vertebrate Arp6, a novel nuclear actin-related protein, interacts with heterochromatin protein 1. Eur J Cell Biol 85:411–421
Ohhata T, Senner CE, Hemberger M, Wutz A (2011) Lineage-specific function of the noncoding Tsix RNA for Xist repression and Xi reactivation in mice. Genes Dev 25:1702–1715
Okamura K, Ishizuka A, Siomi H, Siomi MC (2004) Distinct roles for Argonaute proteins in small RNA-directed RNA cleavage pathways. Genes Dev 18:1655–1666
Pageau GJ, Hall LL, Lawrence JB (2007) BRCA1 does not paint the inactive X to localize XIST RNA but may contribute to broad changes in cancer that impact XIST and Xi heterochromatin. J Cell Biochem 100:835–850
Payer B, Lee JT (2008) X chromosome dosage compensation: how mammals keep the balance. Annu Rev Genet 42:733–772
Peng JC, Karpen GH (2007) H3K9 methylation and RNA interference regulate nucleolar organization and repeated DNA stability. Nat Cell Biol 9:25–35
Rougeulle C, Chaumeil J, Sarma K, Allis CD, Reinberg D et al (2004) Differential histone H3 Lys-9 and Lys-27 methylation profiles on the X chromosome. Mol Cell Biol 24:5475–5484
Royce-Tolland ME, Andersen AA, Koyfman HR, Talbot DJ, Wutz A et al (2010) The A-repeat links ASF/SF2-dependent Xist RNA processing with random choice during X inactivation. Nat Struct Mol Biol 17:948–954
Sado T, Li E, Sasaki H (2002) Effect of TSIX disruption on XIST expression in male ES cells. Cytogenet Genome Res 99:115–118
Shekar PC, Naim A, Sarathi DP, Kumar S (2011) Argonaute-2-null embryonic stem cells are retarded in self-renewal and differentiation. J Biosci 36:649–657
Sinkkonen L, Hugenschmidt T, Berninger P et al (2008) MicroRNAs control de novo DNA methylation through regulation of transcriptional repressors in mouse embryonic stem cells. Nat Struct Mol Biol 15:259–267
Sirchia SM, Ramoscelli L, Grati FR, Barbera F, Coradini D et al (2005) Loss of the inactive X chromosome and replication of the active X in BRCA1-defective and wild-type breast cancer cells. Cancer Res 65:2139–2146
Stavropoulos N, Lu N, Lee JT (2001) A functional role for Tsix transcription in blocking Xist RNA accumulation but not in X-chromosome choice. Proc Natl Acad Sci U S A 98:10232–10237
Stone C, McCabe N, Ashworth A (2003) X-chromosome inactivation: X marks the spot for BRCA1. Curr Biol 13:63–64
Stroud H, Otero S, Desvoyes B, Ramírez-Parra E, Jacobsen SE, Gutierrez C (2012) Genome-wide analysis of histone H3.1 and H3.3 variants in Arabidopsis thaliana. Proc Natl Acad Sci U S A 109:5370–5375
Stucki M, Clapperton JA, Mohammad D, Yaffe MB, Smerdon SJ, Jackson SP (2005) MDC1 directly binds phosphorylated histone H2AX to regulate cellular responses to DNA double-strand breaks. Cell 123:1213–1226
Sun BK, Deaton AM, Lee JT (2006) A transient heterochromatic state in Xist preempts X inactivation choice without RNA stabilization. Mol Cell 21:617–628
Tang KF, Wang Y, Wang P, Chen M, Chen Y et al (2007) Upregulation of PHLDA2 in Dicer knockdown HEK293 cells. Biochim Biophys Acta 1770:820–825
Tian D, Sun S, Lee JT (2010) The long noncoding RNA, Jpx, is a molecular switch for X chromosome inactivation. Cell 143:390–403
Wu J, Lu LY, Yu X (2010) The role of BRCA1 in DNA damage response. Protein Cell 1:117–123
Wutz A, Jaenisch R (2000) A shift from reversible to irreversible X inactivation is triggered during ES cell differentiatin. Mol Cell 5:695–705
Wutz A, Rasmussen TP, Jaenisch R (2002) Chromosomal silencing and localization are mediated by different domains of Xist RNA. Nat Genet 30:167–174
Yang F, Babak T, Shendure J, Disteche CM (2010) Global survey of escape from X inactivation by RNA-sequencing in mouse. Genome Res 20:614–622
Yin H, Lin H (2007) An epigenetic activation role of Piwi and a Piwi-associated piRNA in Drosophila melanogaster. Nature 450:304–308
Zhao J, Sun BK, Erwin JA, Song JJ, Lee JT (2008) Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science 322:750–756
Acknowledgments
We thank T. Jenuwein for histone antibodies, J. Lee for Xist DNA clone, and N. Rangaraj for confocal facilities. The work was funded by HFSP Young Investigator grant and CSIR network project to U.B.
Authors’ contributions
SKK conducted majority of cytological experiments except Fig. 1e and initial write up. LKR contributed in the concept of repeat association of Xi. DRC and VP repeated the experiments and RT-PCR data. UB designed and analyzed the experimental data, compiled the final correction, and final write up. UB and LS commented on the final write up. All authors discussed the results and approved the final version of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 1060 kb)
Rights and permissions
About this article
Cite this article
Kota, S.K., Roy Chowdhury, D., Rao, L.K. et al. Uncoupling of X-linked gene silencing from XIST binding by DICER1 and chromatin modulation on human inactive X chromosome. Chromosoma 124, 249–262 (2015). https://doi.org/10.1007/s00412-014-0495-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-014-0495-4