Abstract
Facioscapulohumeral muscular dystrophy (FSHD) is an autosomal dominant disease involving shortening of D4Z4, an array of tandem 3.3-kb repeat units on chromosome 4. These arrays are in subtelomeric regions of 4q and 10q and have 1–100 units. FSHD is associated with an array of 1–10 units at 4q35. Unambiguous clinical diagnosis of FSHD depends on determining the array length at 4q35, usually with the array-adjacent p13E-11 probe after pulsed-field or linear gel electrophoresis. Complicating factors for molecular diagnosis of FSHD are the phenotypically neutral 10q D4Z4 arrays, cross-hybridizing sequences elsewhere in the genome, deletions including the genomic p13E-11 sequence and part of D4Z4, translocations between 4q and 10q D4Z4 arrays, and the extremely high G + C content of D4Z4 arrays (73%). In this study, we optimized conditions for molecular diagnosis of FSHD with a 1-kb D4Z4 subfragment probe after hybridization with p13E-11. We demonstrate that these hybridization conditions allow the identification of FSHD alleles with deletions of the genomic p13E-11 sequence and aid in determination of the nonpathogenic D4Z4 arrays at 10q. Furthermore, we show that the D4Z4-like sequences present elsewhere in the genome are not tandemly arranged, like those at 4q35 and 10q26.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Facioscapulohumeral muscular dystrophy (FSHD) is a unique dominant disorder involving shortening of D4Z4, a subtelomeric array of tandem 3.3-kb repeat units (Wijmenga et al. 1992). This copy-number polymorphic repeat (1–100 units per array) is present at both 4q35 and 10q26, but only short (1–10 unit) arrays at 4q35 are linked to FSHD (Upadhyaya et al. 1997; Wijmenga et al. 1993). The mechanism for this unusual relationship, which seems to be a cis effect (Ehrlich 2004), is still unknown (Gabellini et al. 2002; Jiang et al. 2003; Winokur 2003). The 4q-specific nature of this disease is impressive given the 99% homology between the 4q and 10q D4Z4 repeat units (GenBank AF117653 and NT_017795) and the >95% homology between the 42-kb region proximal to D4Z4 at 4q35 and 10q26 and the homology distal to the array (van Geel et al. 2002). At 4q35, there are two polymorphic forms of the D4Z4-distal sequence, 4qA and 4qB (Fig. 1), which are found at similar frequencies (van Geel et al. 2002). Only D4Z4 contractions on 4qA are associated with the disease but short 4q35 alleles only very infrequently have a distal, nonpathogenic 4qB sequence (Lemmers et al. 2002, 2004).
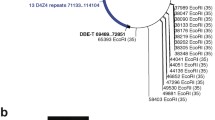
A schematic of the DNA sequences and diagnostically important restriction sites in the distal portion of 4q35 and 10q26 (not to scale). The D4Z4 arrays are depicted as containing two 3.3-kb KpnI repeat units (white and black triangles for canonical 4q35 and 10q26 units, respectively), although FSHD patients can have 1–10 units at their FSHD-causing short 4q35 D4Z4 array and the other arrays can have up to 100 units. D4Z4 has partial KpnI repeat units (small triangle and trapezoid) at its borders. The two polymorphic forms (4qA and 4qB) of sequences between the end of D4Z4 and the TTAGGG arrays of the telomere of 4q are indicated. The 6.2-kb β-satellite DNA in 4qA and 10q contains tandem 68-bp repeats. Restriction cleavage sites that are important for molecular diagnosis (E, EcoRI; H, HindIII, B, BlnI; X, XapI) and the >5-kb restriction fragments are shown for the type of digest indicated on the left. There is infrequently one additional HindIII site proximal to D4Z4 that is polymorphic and close to the proximal BlnI site. The NotI site that is occasionally used to determine the chromosomal location of a translocated D4Z4 array is on 4q proximal to the depicted sequence; NotI digests subjected to PFGE are blot-hybridized to a proximal probe that gives 4q35 fragments between 200 and 500 kb (Lemmers et al. 2003)
Unambiguous clinical diagnosis of FSHD depends on DNA analysis (Lemmers et al. 2001; Lunt 2000). Moreover, there is some correlation between the size of short, FSHD-causing arrays at 4q35 (1–10 D4Z4 repeat units) and the severity of the disease (Lunt et al. 1995; Tawil et al. 1996). For molecular diagnosis of FSHD, the 4q35 D4Z4 arrays have to be distinguished from the almost identical ones at 10q26, which are also highly polymorphic in array size. In addition, D4Z4 must be detected under conditions in which there is no interference from many homologous sequences elsewhere in the genome, especially in the acrocentric chromosomes (Hewitt et al. 1994). The probe for analysis of the length of each D4Z4 array is almost always the nonrepeated p13E-11 sequence, which is located 0.1 kb proximal to D4Z4 (Hewitt et al. 1994; Wright et al. 1993) (Fig. 1). The sizes of D4Z4 arrays are determined by blot hybridization to a 0.8-kb p13E-11 probe after electrophoresis of genomic DNA digested with restriction endonucleases that excise the intact array linked to the genomic p13E-11. The enzymes for the excision are EcoRI for linear gel electrophoresis (LGE) or EcoRI plus HindIII for pulsed-field gel electrophoresis (PFGE; Fig. 1). The use of HindIII as well as EcoRI for PFGE digests minimizes the electrophoretic distortion of the D4Z4 bands by yielding more extensive cleavage of the bulk DNA.
One and, preferably, two digests are also made with restriction endonucleases that cleave only canonical 10q26 D4Z4 repeat units (BlnI in an EcoRI/BlnI double digest) or only canonical 4q35 units (XapI) to distinguish short, FSHD-causing D4Z4 arrays on 4q35 from phenotypically neutral, short D4Z4 arrays on 10q26 (Fig. 1) (Deidda et al. 1996; Lemmers et al. 2001). To resolve the above-mentioned 4qA and 4qB polymorphism distal to D4Z4 on 4q35, HindIII digests can be used for blot hybridization with 4qA- or 4qB-specific oligonucleotide probes (Fig. 1). All FSHD alleles of 1–10 repeat units were shown to have the 4qA type of distal sequence (Lemmers et al. 2002).
Another complication in molecular diagnosis of FSHD stems from the homology between the D4Z4 arrays at 4q35 and 10q26. Subtelomeric exchanges between chromosomes 4 and 10 are found in about 20% of individuals (Lemmers et al. 1998; Matsumura et al. 2002; van Deutekom et al. 1996). Consequently, FSHD-sized D4Z4 repeats can have some canonical 10q-type units, which contain one BlnI site each, replacing some of the canonical 4q-type units, which do not have a BlnI site. For research on how contraction of D4Z4 causes FSHD, it is necessary to not only size D4Z4 alleles, but also to analyze any subtelomeric exchanges between 4q and 10q arrays to identify which BlnI-resistant D4Z4 repeat units actually reside on 4q.
In addition, DNA-based diagnosis of FSHD is complicated by deletions of the p13E-11 probe region from the genome (Lemmers et al. 2003). These deletions may or may not extend into the array itself, which is only 0.1 kb distal (Fig. 1). Deletions of p13E-11 are estimated to occur in about 3% of FSHD patients and of the control population. They do not influence the severity of the disease but they can result in a false-negative molecular diagnosis for FSHD (Lemmers et al. 2003).
A recent report describes a polymerase chain reaction (PCR) method to detect D4Z4 arrays of less than six repeat units specifically from 4q35 with a special Taq polymerase (Goto et al. 2006). Primers are used from sequences just outside the array where there are 1-bp differences between 4q and 10q. PCR gives a specific product only from small D4Z4 arrays on 4q. No product is obtained from unaffected individuals. However, such a negative result without an internal control to eliminate technical problems could produce false negatives. The method only worked for FSHD patients with 1–5 repeat units but many Caucasian patients have FSHD alleles of 6–10 repeat units (Butz et al. 2003; Orrell et al. 1999; Upadhyaya et al. 1997; Lemmers et al. 2006), including occasional patients with severe and early-onset symptoms (Wu et al. 2004). In addition, some somatic mosaics do not have the FSHD phenotype (Lemmers et al. 2004), but because their short 4q D4Z4 arrays could also be amplified, their DNA samples would produce false positives. Lastly, deletions of the p13E-11 region cannot be identified using PCR for molecular diagnosis.
These complications are not shared by Southern blot hybridization with p13E-11 probes supplemented with a new D4Z4 subfragment probe, as described in this report. Previously, we used a 0.7-kb subfragment of the 3.3-kb unit of D4Z4 arrays (Fig. 2) as a probe to analyze D4Z4 methylation and showed that most of the signal obtained by blot hybridization of KpnI digests of human DNA was in a 3.3-kb band (Tsien et al. 2001). There is one KpnI site per tandem D4Z4 unit at 4q35 and 10q26. In contrast, at least several YACs containing D4Z4-like sequences from acrocentric chromosomes do not seem to have these sequences in strict tandem arrays (Lyle et al. 1995). This indirect evidence suggested, but did not prove, that the 0.7-kb D4Z4 subfragment probe was specific. In the present study, we have designed a 1-kb subfragment of D4Z4 as a hybridization probe and rigorously tested it and the 0.7-kb probe for their specificity. We describe the use of the 1-kb probe in the molecular diagnosis of FSHD.
A diagram drawn to scale of D4Z4 subrepeats and hybridization probes. DUX4 is the open reading frame with two almost identical homeodomains to which the 9B6A probe hybridizes. The LSau and HHSPM3 subregions are partially homologous to repeats: GenBank X59423 and X06587, respectively. The positions of the 1-kb D4Z4 probe are given relative to the beginning of the recognition site of KpnI in GenBank AF117653
Materials and methods
Patients and samples
Clinically confirmed patients were obtained via one of the Dutch neuromuscular centers and informed consent was given. For rodent-human somatic cell hybrid (SCH) DNAs, cell lines were used (Coriell Institute for Medical Research). The hybrids with a Chinese hamster genetic background were as follows (the identity of the human chromosome(s) and then the cell line are given: del(4)(p16.2) and 5, GM14193; del(4p16.1), 5, and 8, GM11448; der(X)t(X;4)(p21;q35), GM14221A; 10, GM10926; 9, GM10611; 11, GM10927; 13, GM10898; 22, GM10888; Y, GM06317. The hybrids with a mouse background were as follows: 4, GM11687A; 1, GM13139; 3, GM11713; 14, GM10479; 15, GM11715; 16, GM10567; 21 and 22, GM10322.
To lyse red blood cells from 10 ml of peripheral blood supplemented with EDTA, the sample was incubated on ice for 15 min with 40 ml of 155 mM NH4Cl, 10 mM KHCO3, 1 mM EDTA, pH 8.0. After centrifugation, the pellet was treated once more with 15 ml of the above solution, and pelleted again. The final pellet was resuspended in 1 ml of 75 mM NaCl, 25 mM EDTA, pH 8.0 (SE). For PFGE, the suspension was quickly mixed with 1 ml of melted 1.4% agarose (InCert agarose, FMC) in SE at 50°C, and added to a plastic mold to form ∼0.1-ml plugs, each containing ∼1–1.5 × 106 cells for PFGE. From 10 ml of blood, ∼20 plugs can be obtained. After leaving at 4°C for 30 min, the ejected plugs were incubated in 10 ml of SE containing 1% sarcosyl and 0.6 mg/ml pronase (Sigma) for 2 days at 37°C. Then the plugs were rinsed with TE-4 (10 mM Tris–HCl, pH 7.5, 0.1 mM EDTA), and stored in 0.5 M EDTA, pH 8, at 5°C. For diagnostic LGE, DNA was isolated by the method of Miller et al. (1988) or by melting DNA from a DNA-agarose plug.
Enzyme digestion, electrophoresis, and blotting
DNA-agarose plugs containing ∼8 μg of DNA were rinsed in a rotator for 15 min with water; twice for ∼1.5 h with TE-4; and 3 h in the reaction buffer for the enzymes to be used. Half of a plug was incubated for 6 h with 30 U of each enzyme (EcoRI plus HindIII, EcoRI plus BlnI, or XapI; enzymes from Roche except XapI from Fermentas) under the manufacturers’ conditions but with an additional 3.3 mM spermidine (Sigma) and 1 mM dithiothreitol. The digests and markers (λ midi-range concatemer; New England Biolabs) were subject to PFGE in a 0.85% agarose gel (MP agarose, Roche; 20 × 20 or 14 × 21 cm gels) in 44.5 mM Tris–borate buffer, 1 mM EDTA, pH 8.3 with 0.15 μg/ml ethidium bromide (gel and electrophoresis buffer). Electrophoresis was done in a homemade device at 8.5 V/cm and 21°C with 4 cycles (5.5 h each) and a switch time increasing linearly from 1 to 20 s or in a commercial apparatus (CHEF Mapper XA, Bio-Rad) at 6 V/cm and 22°C for 22 h, with the switch time increasing linearly from 1 to 20 s. DNA was nicked by UV irradiation in a commercial crosslinking apparatus, denatured in 0.4 M NaOH, 0.6 M NaCl for 30 min, and transferred in that solution to a membrane (Hybond XL, Amersham or Zeta-GT, BioRad), neutralized, and cross-linked to the membrane by UV irradiation in the same apparatus. For LGE of D4Z4, samples were loaded onto a thin strip of 1% agarose on top of a 0.5% agarose gel and electrophoresed using 40 mM Tris–base, 1.1% acetic acid, 1 mM EDTA, pH 7.6, with 0.15 μg/ml ethidium bromide as the gel and running buffer. For LGE of KpnI digests, electrophoresis was in a 1% agarose gel at 120 V for 4 h.
Probes and hybridization
The 1-kb D4Z4 probe was generated by PCR using 100 ng of a plasmid containing the 3.3-kb D4Z4 unit in a Bluescript KS vector (Winokur et al. 1994). The primers (5′-3′) were GGAAGGCGGGCAGAGATG and CCGGACCTCTCCAGGGAT. The PCR conditions were 94°C for 15 min followed by 35 cycles of 94°C for 45 s, 62°C for 45 s, and 72°C for 1 min, with a final extension at 72°C for 5 min. The 1-kb PCR product was purified by gel extraction (QIAGEN). From the same plasmid, the 3.3-kb D4Z4 probe was excised with KpnI. The 0.7-kb D4Z4 subfragment probe was excised from its cloning vector (pBC KS +/−) using EagI and EcoRV (Kondo et al. 2000).The p13E-11 sequence (Wright et al. 1993) was excised as a 810-bp fragment with EcoRI and SacI from a plasmid or obtained by PCR. For 9B6A, the DUX4 homeodomain region was amplified by PCR from a plasmid (Hewitt et al. 1994) using vector primers (M13-20 Forward and M13 Reverse, Invitrogen) to give a product containing 320 bp of the DUX4 homeodomain region. About 25 ng of each DNA was labeled with 50 μCi of [α-32P]dCTP (3,000 Ci/mmol) by random priming (Megaprime, Amersham or NEBlot kit, New England Biolabs followed by Spin Column 30, Bio-Rad). For the λ probe, 1.5 ng of template was labeled with 5 μCi of [α-32P]dCTP. About one fifth of the product was used per blot and could be cohybridized with the p13E-11 probe.
For the 1-kb D4Z4 probe, the blots were prehybridized with rotation in a hybridization tube for 1 h at 60°C with 20 ml of 0.125 M sodium phosphate, pH 7.2, 0.25 M NaCl, 7% sodium dodecyl sulfate (SDS), 100 μg/ml of denatured salmon sperm DNA, and 50% freshly added formamide. The hybridization tubes were rated to withstand at least 180°C (e.g., Stovall hybridization bottle, Sigma) and were tightly capped. After addition of probe (∼2 × 107 cpm), the bottle was similarly rotated overnight. The blots were washed with rotation at 65°C for 10 min each with 50 or 100 ml (depending on the size of the bottle) of prewarmed 2 × SSC, 0.1% SDS; 0.5 × SSC, 0.1% SDS; and 0.1 × SSC, 0.1% SDS. For the p13E-11 probe, the hybridization solution was 0.125 M sodium phosphate, pH 7.2, 0.25 M NaCl, 1 mM EDTA, 7% SDS, 10% PEG 6000, 100 μg/ml of denatured salmon sperm DNA; prehybridization, hybridization, and washing were at 65°C but were done in a 100-ml volume in a covered tray with gentle shaking; and three 15-min washes were with ∼0.3 l of 2 × SSC, 1% SDS. Alternatively, a commercial hybridization solution (PerfectHyb, Sigma) could be used at 65°C in a hybridization tube followed by 10-min washes at 65°C with 2× SSC, 0.1% SDS, 1× SSC; 0.1% SDS, and 0.5× SSC, 0.1% SDS, preferably in covered trays. Phosphorimager exposure was for 2 days. For rehybridization, the blot was first stripped by pouring boiling 0.1× SSC, 0.1% SDS on top and cooling to room temperature or by soaking it in 0.4 M NaOH for 30 min at room temperature followed by rinsing with 2× SSC. Stripped membranes were kept moist until reuse. For diagnostic purposes, after probing with p13E-11 and phosphorimaging, the blot can be used without stripping for rehybridization with the 1-kb probe.
For the 9B6A probe, we used the hybridization conditions for p13E-11 followed by washing at 65°C twice with 0.3× SSC, 0.1% SDS for 15 min and then once with 0.1× SSC, 0.1% SDS for 30 min. For the 0.7-kb D4Z4 pro, a commercial solution (ExpressHyb, Clontech) was used at 72°C followed by two washes each at 72°C with 2× SSC, 0.05% SDS and 0.1× SSC, 0.1% SDS. The 3.3-kb D4Z4 probe was hybridized with the homemade solution and washed as for the 1-kb probe except at 63°C. For SCH DNAs, 3 μg/ml of denatured rat liver DNA was included in the hybridization solution. The blots containing KpnI digests were exposed to X-ray film for 2 days.
Results
Rationale for designing a new D4Z4 hybridization probe
We designed a new optimized D4Z4 probe and tested its specificity directly. One of the D4Z4 subregions (Fig. 2) that is repeated elsewhere in the genome independently of the rest of the D4Z4 sequence is the 0.3-kb LSau-homology sequence (Hewitt et al. 1994) with ∼90% homology to this part of the repeat unit. It is present on the short arms of the acrocentric chromosomes and the pericentromeric regions of chromosomes 1, 3, and 9, as well as on Yq (Meneveri et al. 1993). The 0.7-kb probe overlapped that sequence (Fig. 2). Therefore, by PCR, we generated a 1-kb D4Z4 subsequence that excludes the LSau region of D4Z4 repeat units and almost all of the HHSPM3 region (Sp-0.3–8), another low-copy repeat region (Hewitt et al. 1994; Zhang et al. 1987). This probe was tested for its specificity by PFGE and LGE as described below.
Initial tests of the specificity of the 1-kb D4Z4 probe
D4Z4 arrays are unusual because of their extremely high G + C content, 73% (69% for the 1-kb subfragment), compared with the overall human genomic G + C content of 42%. D4Z4’s high G + C content makes it more difficult to get specific conditions for hybridization. We tested the specificity of the 1-kb D4Z4 probe by blot hybridization of KpnI digests of human DNA and DNA from human-rodent SCHs containing one or a few human chromosomes. With a hybridization buffer containing 50% formamide and annealing conditions that gave a temperature for hybridization that was 15°C below the T m, only human DNA and SCH DNA containing human chromosome 4 or 10 gave a signal with the 1-kb D4Z4 probe (Fig. 3a, 1-kb probe). Essentially, the entire signal was in the expected band composed of 3.3-kb D4Z4 repeat units. Under nonstringent conditions (changing the temperature to 49°C), a high-molecular-weight smear was observed from human DNA and DNA from SCHs containing the acrocentric human chromosomes (Fig. 3b). The 0.7-kb D4Z4 probe (Fig. 2) that we had used in an earlier study (Tsien et al. 2001) gave only a slight increase in >3.3-kb signal from total human DNA under our standard conditions (see “Materials and methods”; data not shown) using nonstringent conditions for annealing with a commercial hybridization solution but very stringent conditions for the final wash (T m minus wash temperature of only 12°C). This validates its previous use as a hybridization probe in detection of DNA methylation within the D4Z4 array by Southern blotting DNA digested with CpG methylation-sensitive enzymes (Tsien et al. 2001). Surprisingly, even the entire 3.3-kb D4Z4 repeat unit could be used as a probe with a similar high degree of specificity for D4Z4 in chromosomes 4 and 10 as seen for the 0.7-kb probe, when the hybridization was under stringent conditions (Fig. 3c).
Determination of the specificity of D4Z4 probes with somatic cell hybrid DNAs. DNA from human lung or juvenile thymus, mouse, Chinese hamster ovary (CHO) cells, or SCHs was digested with KpnI and subject to LGE and repeated hybridization with the indicated D4Z4 probe and stripping. The SCH DNAs had only the indicated one human chromosome (horizontal labels over lanes) or several (vertical labels) and were in a murine or CHO genetic background. The asterisked SCH had a single der(X)t(X;4)(p21;q35); therefore, the only part of chromosome 4 present was the 4q35 region. After high-stringency hybridization with the 1-kb D4Z4 probe (a), the faint low-molecular-weight band seen at low stringency in human DNA (b) was not observed, even with longer exposures
The 1-kb probe for analysis of p13E-11 deletions
An important advantage of using a D4Z4 probe for molecular diagnosis of FSHD is that it allows the visualization of a D4Z4 allele distal to a p13E-11 deletion in FSHD patients as well as in unaffected individuals. Upon PFGE and blot hybridization with this probe, all four D4Z4 arrays, including the one with a deletion of p13E-11, can be seen in a sample from a patient with an FSHD-associated allele (Fig. 4b). As expected, the p13E-11 probe failed to reveal the FSHD-causing allele with the proximal deletion of the genomic p13E-11 sequence (Fig. 4b). Although PFGE is much more informative than LGE for FSHD molecular diagnosis, LGE is often employed and without the useful XapI digest. Figure 5 shows examples (FB3 and FB4) of how a D4Z4 probe, but not a p13E-11 probe, reveals a short D4Z4 array that lacks a p13E-11 sequence distal to it upon Southern blotting after LGE.
Comparison of different probes for PFGE diagnosis of FSHD. DNA samples from control blood (CB1 and CB2) or FSHD blood (FB1 and FB2, or FB p13EΔ for p13E-11 deletion patients) were digested with EcoRI/HindIII, EcoRI/BlnI, or XapI (ApoI) and subjected to PFGE. They were then blot-hybridized with the indicated probe (Fig. 2) either under the standard hybridization conditions for p13E-11 or 9B6A or the more stringent conditions for the 1-kb D4Z4 probe (see “Materials and methods”). The Y chromosome-derived fragments that cross-hybridize with the p13E-11 probe are indicated. These are seen in all males and not in females. The sizes of markers (in kilobase) for the first gel are given on the left and the sizes of D4Z4-containing bands within the blots. The small d indicates that there was a deletion of p13E-11 immediately proximal to that array, which is only visualized with the D4Z4 probe. The brackets denote the nonspecific signal from the 9B6A probe. Blots were hybridized first to the p13E-11 probe, stripped, and then hybridized to a D4Z4 probe
Identification of FSHD alleles in patients with p13E-11 deletions by LGE. The same terminology is used as in Fig. 4. Note that for the full molecular diagnostic test by LGE, XapI digests should used in addition to EcoRI single digests and EcoRI/BlnI double digests. These samples had previously been characterized by PFGE as in Fig. 4
Molecular diagnosis of FSHD with D4Z4 vs p13E-11 probes
Using the recommended digests for molecular diagnosis, EcoRI/HindIII, EcoRI/BlnI, and XapI, we did PFGE followed by blot hybridization to the p13E-11 probe and obtained the typical D4Z4 array patterns for unaffected individuals and FSHD patients (Fig. 4a). The blot was stripped and hybridized with the new 1-kb D4Z4 probe. One of the advantages of using the 1-kb D4Z4 probe is the increased sensitivity of detection of D4Z4 arrays in XapI digests (Fig. 4; CB2, FB1, and FB2), which allow visualization of only the 10q-type (XapI-resistant) arrays, thus facilitating identification of 4q arrays in the EcoRI/HindIII digests (Lemmers et al. 2001). When blots were probed with p13E-11, there was usually a decrease in the signal in XapI bands relative to the analogous EcoRI/HindIII bands. This decrease is due to the XapI sites in the genomic p13E-11 (Fig. 1). Avoiding decreases in signal intensity can be especially important when there are only small amounts of DNA in peripheral blood-derived DNA-agarose plugs.
Previously, when a D4Z4 probe was desired for PFGE (Lemmers et al. 1998), 9B6A (Fig. 2), a probe from the homeodomain region of D4Z4 (Hewitt et al. 1994), was used. This D4Z4 subfragment probe gave much nonspecific signal that interfered with detecting low-molecular-weight D4Z4 arrays under previously employed conditions (Lemmers et al. 1998) because of the absence of formamide to lower the T m during hybridization. (Fig. 3a). With the 9B6A probe from a DUX4 subregion of D4Z4, which is much more highly repeated in the human genome than is D4Z4, the stringent hybridization and washing conditions used for the 1-kb probe nonetheless gave interfering background. In contrast, even under low-stringency conditions (differential of the T m and the temperature for the annealing step was 35 and 29°C for the washing steps), the p13E-11 probe gave only specific bands, except for a Y chromosome-derived, noninterfering band of <10 kb (Fig. 4).
A flowchart of recommended procedures for molecular diagnosis incorporating the use of the 1-kb D4Z4 probe is shown in Fig. 6. No one simple set of procedures will cover all cases because of the inherent complexity of FSHD molecular diagnosis due to the 4q and 10q homology between D4Z4 arrays and adjacent sequences, translocations between 4q and 10q D4Z4 arrays, mitotic D4Z4 contractions, deletions encompassing p13E-11, and the wide range of sizes of D4Z4 arrays combined with the need for high resolution of bands in the 30- to 45-kb range. Although PFGE with DNA-agarose plugs is recommended, LGE is often used throughout the analysis or just to confirm the size of short D4Z4 arrays with more accuracy. Analysis should start with the p13E-11 probe and EcoRI/HindIII (PFGE) or EcoRI (LGE) digests, EcoRI/BlnI digests, and XapI digests. For a patient suspected of having FSHD, a short array that displays a reduction of 3 kb (Figs. 1 and 5) in EcoRI/BlnI vs EcoRI digests upon LGE confirms FSHD with >99% confidence. If a reduction of more than 3 kb is seen or the EcoRI/HindIII or EcoRI band is no longer visible in EcoRI/BlnI digests, XapI digests provide discrimination between a short D4Z4 array on 10q (no FSHD) and a short hybrid array containing 4q- and 10q-type repeat units on chromosome 4 (FSHD) (Lemmers et al. 2001). A short hybridizing XapI fragment corresponding to the short EcoRI/HindIII or EcoRI fragment indicates no FSHD, and while its absence indicates FSHD. Upon PFGE with high-molecular-weight DNA in agarose plugs, usually all four alleles can be visualized with the p13E-11 probe. This facilitates detection of hybrid arrays and mosaicism and can signal the need for further testing in the case of D4Z4 translocations between chromosomes 4 and 10 that interfere with the analysis (Lemmers et al. 1998). For all LGE analyses or when less than four alleles are seen upon PFGE, rehybridization of the membrane should been done with the 1-kb D4Z4 probe to look for FSHD alleles with a deletion of the p13E-11 probe region. When the neurologist is not completely sure of FSHD and the genetic analysis reveals a short D4Z4 array on 4q, then additional 4qA/4qB testing is needed (Fig. 6, asterisk). Rehybridization to the 1-kb D4Z4 probe also clarifies the hybrid composition of arrays, although this is not necessary for molecular diagnosis.
Flowchart for the molecular diagnosis of FSHD for confirmation (clinical FSHD) as well as exclusion (clinically no FSHD) purposes. Recommended enzyme digestions and hybridization with three sets of hybridization probes (three shaded circles), starting with the p13E-11 probe, are shown for molecular diagnosis of FSHD. The chromosomal location (4 or 10) of the D4Z4 array is indicated and whether the size of the array is within the range for FSHD linkage (short). Most frequent outcomes for either PFGE or LGE analysis for FSHD are shown with thick arrows. The dotted arrows indicate recommended rehybridization for confirmation of FSHD with the 1-kb D4Z4 probe for all LGE tests and for PFGE tests that reveal less than four arrays
If individuals that are not suspected of having FSHD (unaffected spouses or healthy family members of patients) are included in the genetic test and they do not display any FSHD-like allele, no analysis is needed beyond that described above. However, if these individuals unexpectedly display an allele that mimics an FSHD allele, extra tests have to be performed to explain these apparently false-positive results. If a short homogeneous 4q-type or hybrid array is detected, then an additional Southern blot with HindIII digested DNA is needed. Subsequent hybridization with 4qA-and 4qB-specific probes (Fig. 1) will discriminate pathogenic short 4qA alleles from rare nonpathogenic, short 4qB alleles (Lemmers et al. 2002).
Discussion
The p13E-11 sequence, which is adjacent to D4Z4 at 4q35 and 10q26, has been almost the only blot hybridization probe used for determining whether the 4q35 arrays had contracted to a size that is linked to FSHD. We identified a subfragment of the D4Z4 sequence and stringent hybridization conditions that allow detection of D4Z4 at 4q35 and 10q26 without interference from cross-hybridizing sequences elsewhere in the genome. Use of the 1-kb D4Z4 probe avoids the p13E-11 probe-associated loss in sensitivity for detection of 10q26 D4Z4 arrays (Fig. 1) in one of the recommended digests for molecular diagnosis of FSHD, the XapI digest (Lemmers et al. 2001) (Fig. 1). However, the 1-kb D4Z4 probe is not a substitute for the p13E-11 probe but rather should be used for rehybridization of certain samples (Fig. 6).
An important advantage of using a D4Z4 probe for molecular diagnosis of FSHD in conjunction with the p13E-11 probe is that it permits the identification of about 3% of FSHD patients who have a deletion of the p13E-11 genomic sequence next to a short 4q D4Z4 array (Lemmers et al. 2003). Upon hybridization to a p13E-11 probe, a probable deletion encompassing p13E-11 can be noticed by PFGE, but not LGE, which does not resolve all the D4Z4 arrays. It is therefore advisable to rehybridize routinely with a D4Z4 probe during LGE diagnosis of FSHD or to do PFGE, rather than LGE, in which case the need for additional probing with D4Z4 is usually apparent (Fig. 6). Even for PFGE, unusual combinations of D4Z4 alleles that could include an FSHD allele may only be detected upon rehybridization with the D4Z4 probe. Other examples of the superiority of PFGE for molecular diagnosis are cases with somatic mosaicism. Somatic mosaicism, which can affect the severity of FSHD, is present in almost half of the de novo FSHD patients (Lemmers et al. 2004) and requires PFGE, in which all the D4Z4 arrays are visualized, for assessment.
In addition, for research purposes, PFGE is needed to determine the composition of hybrid arrays containing both canonical 4q-type (BlnI-resistant, XapI-sensitive) and 10q-type (BlnI-sensitive, XapI-resistant) D4Z4 repeat units. This is not necessary for diagnosis because it is the length of the array on 4q35, and not whether it contains 4-type D4Z4 repeat units, which determines the clinical status. To characterize the composition of hybrid D4Z4 arrays, it is necessary to detect in EcoRI/BlnI and XapI digests not only the D4Z4 arrays linked to p13E-11 with the p13E-11 probe, but also to visualize all subfragments from hybrid arrays in a subsequent hybridization with a D4Z4 probe. About half of the translocation products between D4Z4 repeat units on 4q35 and 10q26 contain hybrid arrays.
Other advantages of a specific probe for the D4Z4 arrays at 4q35 and 10q26 for research on FSHD accrue from the lack of PCR primer pairs from within the array that amplify only 4q35 and 10q26 D4Z4 sequences without amplifying partially homologous sequences and subsequences elsewhere, especially on the acrocentric chromosomes (Lyle et al. 1995). The 1-kb subfragment of D4Z4 and even the only slightly less specific 0.7- and 3.3-kb D4Z4 hybridization probes permit a variety of experiments examining the nature of D4Z4 arrays. Using the 1-kb D4Z4 probe, the expected 3.3-kb KpnI fragments were observed only from human-rodent SCHs that contained chromosome 4 or 10 or a translocated subtelomeric portion of human 4q. Under nonstringent conditions, KpnI digests of DNA from the SCHs that had human acrocentric chromosomes gave a high-molecular-weight smear rather than a 3.3-kb repeat-unit band in 1% agarose LGE (Fig. 3b). Analogous blots of DNA digested just outside the array by EcoRI and HindIII gave a <3-kb smear from the human acrocentric chromosomes under low-stringency conditions, while the hybridizing signal from chromosomes 4 and 10 had the expected high molecular weight (data not shown). This indicates that the D4Z4-like sequences in the acrocentric chromosomes are not tandemly arranged, although they seem to be clustered from Southern blot analysis of several acrocentric chromosome-derived YACs (Lyle et al. 1995). In summary, the use of the 1-kb D4Z4 probe can aid in molecular diagnosis of FSHD (Fig. 6), advance our knowledge of D4Z4 chromatin structure, and reveal the composition of hybrid D4Z4 arrays for research on FSHD.
References
Butz M, Koch MC, Muller-Felber W, Lemmers RJ, van der Maarel SM, Schreiber H (2003) Facioscapulohumeral muscular dystrophy. Phenotype–genotype correlation in patients with borderline D4Z4 repeat numbers. J Neurol 250:932–937
Deidda G, Cacurri S, Piazzo N, Felicetti L (1996) Direct detection of 4q35 rearrangements implicated in facioscapulohumeral muscular dystrophy (FSHD). J Med Genet 33:361–365
Ehrlich M (2004) Exploring hypotheses about the molecular etiology of FSHD: loss of heterochromatin spreading and other long-range interaction models. In: Cooper DN, Upadhyaya M (eds) FSHD facioscapulohumeral muscular dystrophy: molecular cell biology and clinical medicine. BIOS Scientific Publishers, New York, NY, pp 253–276
Gabellini D, Green MR, Tupler R (2002) Inappropriate gene activation in FSHD: a repressor complex binds a chromosomal repeat deleted in dystrophic muscle. Cell 110:339–348
Goto K, Nishino I, Hayashi YK (2006) Rapid and accurate diagnosis of facioscapulohumeral muscular dystrophy. Neuromuscul Disord 16:256–261
Hewitt JE, Lyle R, Clark LN, Valleley EM, Wright TJ, Wijmenga C, van Deutekom JC, Francis F, Sharpe PT, Hofker M, Frants RF, Williamson R (1994) Analysis of the tandem repeat locus D4Z4 associated with facioscapulohumeral muscular dystrophy. Hum Mol Genet 3:1287–1295
Jiang G, Yang F, Van Overveld PG, Vedanarayanan V, Van Der Maarel S, Ehrlich M (2003) Testing the position–effect variegation hypothesis for facioscapulohumeral muscular dystrophy by analysis of histone modification and gene expression in subtelomeric 4q. Hum Mol Genet 12:2909–2921
Kondo T, Bobek MP, Kuick R, Lamb B, Zhu X, Narayan A, Bourc’his D, Viegas-Pequignot E, Ehrlich M, Hanash SM (2000) Whole-genome methylation scan in ICF syndrome: hypomethylation of non-satellite DNA repeats D4Z4 and NBL2. Hum Mol Genet 9:597–604
Lemmers RJ, van der Maarel SM, van Deutekom JC, van der Wielen MJ, Deidda G, Dauwerse HG, Hewitt J, Hofker M, Bakker E, Padberg GW, Frants RR (1998) Inter- and intrachromosomal sub-telomeric rearrangements on 4q35: implications for facioscapulohumeral muscular dystrophy (FSHD) aetiology and diagnosis. Hum Mol Genet 7:1207–1214
Lemmers RJL, de Kievit P, van Geel M, van der Wielen MJ, Bakker E, Padberg GW, Frants RR, van der Maarel SM (2001) Complete allele information in the diagnosis of facioscapulohumeral muscular dystrophy by triple DNA analysis. Ann Neurol 50:816–819
Lemmers RJ, de Kievit P, Sandkuijl L, Padberg GW, van Ommen GJ, Frants RR, van der Maarel SM (2002) Facioscapulohumeral muscular dystrophy is uniquely associated with one of the two variants of the 4q subtelomere. Nat Genet 32:235–236
Lemmers RJ, Osborn M, Haaf T, Rogers M, Frants RR, Padberg GW, Cooper DN, van der Maarel S, Upadhyaya M (2003) D4F104S1 deletion in facioscapulohumeral muscular dystrophy: phenotype, size, and detection. Neurology 61:178–183
Lemmers RJ, van der Wielen MJ, Bakker E, Padberg GW, Frants RR, van der Maarel SM (2004) Somatic mosaicism in FSHD often goes undetected. Ann Neurol 55:845–850
Lemmers RJ, van der Wielen MJ, Bakker E, Frants RR, van der Maarel SM (2006) Rapid and accurate diagnosis of facioscapulohumeral muscular dystrophy. Neuromuscul Disord 16:615–617
Lunt P (2000) Facioscapulohumeral muscular dystrophy: diagnostic and molecular aspects. In: Deymeer F (ed) Neuromuscular diseases: from basic mechanisms to clinical management, vol 18. Monogr Clin Neurosci (Karger, Basel) pp 44–60
Lunt PW, Jardine PE, Koch MC, Maynard J, Osborn M, Williams M, Harper PS, Upadhyaya M (1995) Correlation between fragment size at D4F104S1 and age at onset or at wheelchair use, with a possible generational effect, accounts for much phenotypic variation in 4q35-facioscapulohumeral muscular dystrophy (FSHD). Hum Mol Genet 4:951–958
Lyle R, Wright TJ, Clark LN, Hewitt JE (1995) The FSHD-associated repeat, D4Z4, is a member of a dispersed family of homeobox-containing repeats, subsets of which are clustered on the short arms of the acrocentric chromosomes. Genomics 28:389–397
Matsumura T, Goto K, Yamanaka G, Lee J, Zhang C, Hayashi YK, Arahata K (2002) Chromosome 4q;10q translocations; Comparison with different ethnic populations and FSHD patients. BMC Neurol 2:7
Meneveri R, Agresti A, Marozzi A, Saccone S, Rocchi M, Archidiacono N, Corneo G, Della Valle G, Ginelli E (1993) Molecular organization and chromosomal location of human GC-rich heterochromatic blocks. Gene 123:227–234
Miller SA, Dykes DD, Polesky HF (1988) A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res 16:1215
Orrell RW, Tawil R, Forrester J, Kissel JT, Mendell JR, Figlewicz DA (1999) Definitive molecular diagnosis of facioscapulohumeral dystrophy. Neurology 52:1822–1826
Tawil R, Forrester J, Griggs RC, Mendell J, Kissel J, McDermott M, King W, Weiffenbach B, Figlewicz D (1996) Evidence for anticipation and association of deletion size with severity in facioscapulohumeral muscular dystrophy. The FSH-DY Group. Ann Neurol 39:744–748
Tsien F, Sun B, Hopkins NE, Vedanarayanan V, Figlewicz D, Winokur S, Ehrlich M (2001) Hypermethylation of the FSHD syndrome-linked subtelomeric repeat in normal and FSHD cells but not in ICF syndrome cells. Mol Genet Metab 74:322–331
Upadhyaya M, Maynard J, Rogers MT, Lunt PW, Jardine P, Ravine D, Harper PS (1997) Improved molecular diagnosis of facioscapulohumeral muscular dystrophy (FSHD): validation of the differential double digestion for FSHD. J Med Genet 34:476–479
van Deutekom JC, Bakker E, Lemmers RJ, van der Wielen MJ, Bik E, Hofker MH, Padberg GW, Frants RR (1996) Evidence for subtelomeric exchange of 3.3 kb tandemly repeated units between chromosomes 4q35 and 10q26: implications for genetic counselling and etiology of FSHD1. Hum Mol Genet 5:1997–2003
van Geel M, Dickson MC, Beck AF, Bolland DJ, Frants RR, van der Maarel SM, de Jong PJ, Hewitt JE (2002) Genomic analysis of human chromosome 10q and 4q telomeres suggests a common origin. Genomics 79:210–217
Wijmenga C, Hewitt JE, Sandkuijl LA, Clark LN, Wright TJ, Dauwerse HG, Gruter AM, Hofker MH, Moerer P, Williamson R et al (1992) Chromosome 4q DNA rearrangements associated with facioscapulohumeral muscular dystrophy. Nat Genet 2:26–30
Wijmenga C, Frants RR, Hewitt JE, van Deutekom JC, van Geel M, Wright TJ, Padberg GW, Hofker MH, van Ommen GJ (1993) Molecular genetics of facioscapulohumeral muscular dystrophy. Neuromuscul Disord 3:487–491
Winokur ST, Bengtsson U, Feddersen J, Mathews KD, Weiffenbach B, Bailey H, Markovich RP, Murray JC, Wasmuth JJ, Altherr MR, Schutte BC (1994) The DNA rearrangement associated with facioscapulohumeral muscular dystrophy involves a heterochromatin-associated repetitive element: implications for a role of chromatin structure in the pathogenesis of the disease. Chromosome Res 2:225–234
Winokur ST, Chen YW, Masny PS, Martin JH, Ehmsen JT, Tapscott SJ, van der Maarel SM, Hayashi Y, Flanigan KM (2003) Expression profiling of FSHD muscle supports a defect in specific stages of myogenic differentiation. Hum Mol Genet 12:2895–2907
Wright TJ, Wijmenga C, Clark LN, Frants RR, Williamson R, Hewitt JE (1993) Fine mapping of the FSHD gene region orientates the rearranged fragment detected by the probe p13E-11. Hum Mol Genet 2:1673–1678
Wu ZY, Wang ZQ, Murong SX, Wang N (2004) FSHD in Chinese population: characteristics of translocation and genotype–phenotype correlation. Neurology 63:581–583
Zhang XY, Loflin PT, Gehrke CW, Andrews PA, Ehrlich M (1987) Hypermethylation of human DNA sequences in embryonal carcinoma cells and somatic tissues but not in sperm. Nucleic Acids Res. 15:9429–9449
Acknowledgements
We thank Yang Fan and Chunbo Shao for help with the manuscript and Drs. Robert Bloch, Meredith Bond, and Mimi Blitzer for generously hosting us at the University of Maryland Medical School in the aftermath of the Hurricane Katrina flood in New Orleans. This research was supported in part by the National Institutes of Health, USA (NIH NS048859 to M. Ehrlich); the FSH Society (to R.J.F.L. Lemmers); and Eurogendis, Fundación Kutxa and Fundación Ilundain (to P. Camano).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by E.A. Nigg
Rights and permissions
About this article
Cite this article
Ehrlich, M., Jackson, K., Tsumagari, K. et al. Hybridization analysis of D4Z4 repeat arrays linked to FSHD. Chromosoma 116, 107–116 (2007). https://doi.org/10.1007/s00412-006-0080-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-006-0080-6