Abstract
To structurally dissect mitotic chromosomes, we aim to position along the folded chromatin fiber proteins involved in long-range order, such as topoisomerase IIα (topoIIα) and condensin. Immuno-electron microscopy (EM) of thin-sectioned chromosomes is the method of choice toward this goal. A much-improved immunoprocedure that avoids problems associated with aldehyde fixation, such as chemical translinking and networking of chromatin fibers, is reported here. We show that ultraviolet irradiation of isolated nuclei or chromosomes facilitates high-level specific immunostaining, as established by fluorescence microscopy with a variety of antibodies and especially by immuno-EM. Ultrastructural localizations of topoIIα and condensin I component hBarren (hBar; hCAP-H) in mitotic chromosomes were studied by immuno-EM. We show that the micrographs of thin-sectioned chromosomes map topoIIα and hBar to the center of the chromosomal body where the chromatin fibers generally converge. This localization is defined by many clustered gold particles with only rare individual particles in the peripheral halo. The data obtained are consistent with the view that condensin and perhaps topoIIα tether chromatin to loops according to a scaffolding-type model.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Topoisomerase IIα (topoIIα) and condensin are abundant components of mitotic chromosomes and are required for their assembly from interphase nuclei. The involvement of topoIIα in chromosome condensation is based on genetic evidence (Uemura et al. 1987) and assembly studies in Xenopus egg extracts (Adachi et al. 1991; Hirano and Mitchison 1993). The condensin complex was first described in the Xenopus system (Hirano et al. 1997) and has since been characterized in different organisms (McHugh and Heck 2003; Hirano 2000). Condensin is composed of five different subunits that, in the human system, are called SMC4 (hCAP-C) and SMC2 (hCAP-E; a heterodimer) and three additional subunits called hCAP-D2 (hEg7), hCAP-G, and hCAP-H (hBarren or hBar) (McHugh and Heck 2003; Hirano 2000). The SMC proteins of condensin were originally identified genetically in budding yeast as genes involved in the “structural maintenance of chromosomes” (Strunnikov et al. 1993, 1995). They formed the SMC family of DNA-dependent adenosine triphosphatases that were well conserved from bacteria to mammals (reviewed by Cobbe and Heck 2000; Hirano 2002). Recently, it was shown that there were two types of condensin (condensin I and II) in vertebrates (Ono et al. 2003, 2004), which appeared to have distinct functions (Hirota et al. 2004).
TopoIIα and condensin harbor adenosine-triphosphate (ATP)-dependent catalytic activities that can alter DNA topology in different ways. TopoIIα, a homomeric dimer, removes positive or negative DNA supercoils and can catenate or decatenate DNA strands (Wang 1996). Conversely, condensin introduces positive supercoils (Kimura et al. 1999; Bazett-Jones et al. 2002). How do these enzymatic activities apply their shape-determining role to carve out from the homogenous mass of interphase mitotic chromosomes? Do these activities also exert a structural shape-maintaining function in assembled chromosomes? The conspicuous subchromosomal location of both topoIIα and condensin, which is discussed next, supports this notion.
Immunolocalization studies, initially of swollen and lately of near-native chromosomes, establish that topoIIα and condensin are localized to an axially restricted longitudinal chain that extends through the entire chromosomal body (Earnshaw and Heck 1985; Gasser et al. 1986; Boy de la Tour and Laemmli 1988; Taagepera et al. 1993; Maeshima and Laemmli 2003; Ono et al. 2003; Kireeva et al. 2004). Interestingly, double staining for topoIIα and condensin I components generates a striking “barber pole” pattern, which appears to arise from two axial “chains” composed of longitudinally out-of-phase regions (beads) alternatively enriched for topoIIα or condensin (Maeshima and Laemmli 2003). The structural significance of the barber pole is not understood.
Immunofluorescence (IF) studies are consistent with the scaffolding loop model of chromosomes, where a longitudinal (continuous) network that somehow crossties chromatin into loops is proposed (Laemmli et al. 1978). In line, topoIIα and condensin constitute the main components of the chromosomal-shaped scaffolding, whose structural stability is ATP-dependent (Maeshima and Laemmli 2003). This ATP dependence inspires confidence that the biochemical scaffolding is reflective of genuine interactions that are manifested by IF as an axial element. Electron microscopy (EM) studies of histone-depleted, swollen, and native-like compact chromosomes are also consistent with this scaffolding model (Paulson and Laemmli 1977; Marsden and Laemmli 1979; Earnshaw and Laemmli 1983). Particularly telling are micrographs of chromosomal cross sections that show a star-like radial orientation of the chromatin fibers (Marsden and Laemmli 1979; Adolph 1981).
Clearly, although the abovementioned studies support some kind of scaffolding model, the structural details thereof remain an enigma. Toward a better comprehension of chromosome structure, at a higher resolution, we need to localize scaffolding proteins in chromosomes by immuno-EM. Major questions have to be answered: Which scaffolding proteins define the bases of the chromatin loops? How are neighboring loops structurally linked? What is the structural relationship of topoIIα and condensin?
Immuno-EM of chromatin antigens is experimentally challenging since it is often afflicted with high background (spurious gold particles) and poor definition (few gold particles) of the antigens. One major difficulty is finding conditions of aldehyde fixation that preserve both the antigenicity and the structural integrity of chromosomes. Toward this goal, we search for an experimental procedure omitting chemical fixation. Surprisingly, we demonstrate here that ultraviolet (UV) irradiation of isolated chromosomes and nuclei allows high-level specific immunolabeling (i.e., the location of chromosomal antigens is defined by many gold particles and low background, while maintaining chromosome structure well). This UV method was explored by IF and was then applied to localize the condensin protein hBar (hCAP-H, a component of condensin I) and topoIIα in compact mitotic chromosomes using immuno-EM. The obtained micrographs demonstrate that these proteins localize to a central region of chromosomal cross section to where chromatin loops converge. This observation supports the loop model, but it also illustrates the remaining formidable challenge of understanding chromosome structure at a higher resolution.
Materials and methods
IF staining of isolated chromosomes and HeLa cells fixed in paraformaldehyde or glutaraldehyde
HeLa S3 cells were maintained in a suspension in RPMI 1640 medium (Gibco-BRL) containing 5% newborn bovine serum (Biochrom, Switzerland), 100 U/ml penicillin, and 100 μg/ml streptomycin (Biochrom) at 37°C under a 5% CO2 atmosphere (Figs. 1, 2, and 3). For chromosome isolation, one liter of suspension culture of exponentially growing cells at a density of 3×105 cells/ml was blocked with 0.06 μg/ml colcemide for 12 h and then used for chromosome isolation. Chromosomes were isolated in polyamine–ethylenediaminetetraacetic acid (EDTA) buffers, as described previously (Lewis and Laemmli 1982; Gasser et al. 1986), except that the detergent used for cell lysis was 0.05% Empigen (Calbiochem) instead of digitonin. The final purification step in a Percoll gradient was omitted for the experiments reported in this paper. Isolated chromosomes were stored at −20°C in a buffer containing 3.75 mM Tris–HCl (pH 7.5), 20 mM KCl, 0.5 mM K-EDTA, 0.05 mM spermine, 0.125 mM spermidine, 1% Trasylol, 1% thioglycol, 0.1 mM phenylmethylsulphonylfluoride (PMSF), 0.05% Empigen, and 60% glycerol. Typically, yields ranged from 15 to 20 absorbance units at 260 nm (A260) of chromosomes from one liter of culture of arrested HeLa S3 cells.
Immunofluorescence staining of a condensin component hBar in isolated chromosomes under various fixation conditions. Isolated HeLa chromosomes were spun onto coverslips, fixed as indicated, immunostained for the condensin component hBar (red), and counterstained with DAPI (blue). a and b Fixation with 0.8% PAF. c Fixation with 0.001% GLU. d No fixation. Note that unfixed and GLU-fixed chromosomes do not immunostain, while PAF-fixed chromosomes highlight the beaded axial scaffolding (red) within the DAPI (blue) body in each chromatid
Immunofluorescence staining of several chromosomal proteins in isolated chromosomes after UV irradiation. Isolated chromosomes were spun onto coverslips, irradiated for 5 min with the UV Stratalinker 2400 (Stratagene), and then conventionally immunostained for several proteins indicated at the top of each column. a, e, and i αTopoIIα. b, f, and k αCENP-F. c, g, and l αhBar. d, h, and m αTRF-1. Alexa-conjugated IgG (Molecular Probes) was used as a secondary AB. Fixation: Top row, UV; middle row, PAF; bottom row, no fixation. Note that UV-irradiated chromosomes immunostain similarly well as PAF-fixed ones. In contrast, unfixed chromosomes stain poorly for all these proteins
Comparison of staining properties of conventional fluorescent IgG and FluoroNanogold ABs. a and b Mitotic cells were fixed in 2% PAF and immunostained with hBar AB (αhBar). As secondary ABs, either Alexa594 IgG or FluoroNanogold AB was used. Note that the staining signal of FluoroNanogold, although adequate, lacks the crisp appearance observed with the IgG control supposedly due to the higher general background. c–g Immunostaining of hBar with FluoroNanogold after UV irradiation. Spread chromosomes were immunostained for hBar following UV irradiation for different lengths of times (indicated). Note that 5–10 min of UV irradiation produced strong scaffolding signals within the DAPI-stained chromosomal bodies. This staining is similar to that obtained with conventional fluorescent IgGs
IF staining was carried out as described previously (Boy de la Tour and Laemmli 1988) with the following adaptations. A suspension of 0.1 A260 of chromosomes was diluted about ten times into 100 μl of XBE2 buffer [10 mM 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES)–KOH (pH 7.7), 1 mM MgCl2, 100 mM KCl, 5 mM ethyleneglycotetraacetic acid (EGTA), 0.5 mM ATPγS, and 0.1 mM PMSF]. After 15 min of incubation on ice, chromosomes were fixed for 15 min with freshly prepared 0.8% paraformaldehyde (PAF; Fluka) or an EM-grade 0.001% glutaraldehyde (GLU; Fluka) in XBE2 buffer. Samples were spun down onto polylysine-coated round coverslips (diameter, 12 mm) through 250 μl of a 30% sucrose cushion in XBE2 at 2,000 rpm for 5 min with a tabletop centrifuge (Sorvall GLC-1, HL4 rotor). Centrifugation was performed in 500-μl plastic buckets that held the coverslips and fitted the swing-out rotor. After centrifugation, the liquid over the coverslips was carefully removed, and 250 μl of XBE2 containing 0.5 mg/ml NaBH4 was overlaid for 5 min. The coverslips were carefully washed with 250 μl of XBE2 buffer and then removed from the buckets. All subsequent steps were carried out by floating the coverslips on drops of liquid (usually around 250 μl) on a piece of Parafilm. To reduce the background fluorescence signal, the coverslips were incubated with 3% normal goat serum (NGS; Nordic Immunology) in XBE2 buffer for 30 min at room temperature. The coverslips were then incubated with the first antibody (AB) in XBE2 buffer containing 1% NGS at room temperature for 1 h. The AB dilutions used were as follows: 3,000-fold dilution for anti-hBar rabbit serum (Maeshima and Laemmli 2003), 1,000-fold dilution for anti-human TopoIIα monoclonal AB (MBL, Japan), 2,000-fold dilution for anti-human Emerin rabbit serum (from K. Wilson), 1,000-fold dilution for anti-human CENP-F (Abcom), and 5,000-fold dilution for anti-human tumor-necrosis-factor-receptor-associated factor homolog (TRF-1) rabbit serum (from T. de Lange). After washing with XBE2 buffer (five times for 3 min), the coverslips were incubated with 1,000-fold-diluted secondary AB [goat anti-mouse Alexa488 or goat anti-rabbit Alexa594 (Molecular Probes)] in XBE2 buffer containing 1% NGS at room temperature for 1 h. After extensive washing with XBE2 buffer (six times for 3 min), the coverslips were mounted in PPDI [10 mM HEPES (pH 7.7), 1 mM MgCl2, 100 mM KCl, 5 mM EGTA, 78% glycerol, 1 mg/ml paraphenylene diamine (Sigma), and 0.5 μg/ml DAPI (Roche)] and sealed with a rapid epoxy glue (Araldit, Switzerland). Images were taken using a DeltaVision microscope (Applied Precision) and deconvoluted. Where no fixation was performed (Figs. 1d and 2i–m), the chromosome suspension diluted in XBE2 buffer was directly spun onto the polylysine-coated coverslips. The coverslips were processed as described above for immunostaining. hBar staining of mitotic HeLa S3 cells using PAF (Fig. 3a and b) was performed as described (Maeshima and Laemmli 2003). In Fig. 3b, FluoroNanogold (Nanoprobes) was used as a secondary AB (20-fold dilution).
UV procedure for light microscopy
A suspension of 0.1 A260 of isolated chromosomes was diluted about 25 times into 250 μl of XBE2 buffer (Figs. 2 and 3). After 15 min of incubation on ice, samples were spun down onto the polylysine-coated round coverslips as described above. After centrifugation, the plastic buckets were put on an ice-cold metal plate without removing the liquid, placed in a Stratalinker 2400 UV crosslinker (Stratagene), and subjected to UV irradiation for 5–10 min. This UV dose was equivalent to 4,000–8,000 J/m2. The coverslips were then carefully washed in the plastic buckets with XBE2 buffer, removed from the buckets, and immunostained as described above. In some cases (Fig. 3c–g), anti-rabbit FluoroNanogold (fluorescein isothiocyanate) was used as the secondary AB at a 20-fold dilution.
Isolation of chromosome clusters and nuclei for EM
We used a chromosome isolation procedure adapted from Marsden and Laemmli (1979) for EM experiments. Two hundred milliliters of the mitotic HeLa S3 cells, which were blocked as described above, was chilled in an ice bath for 30 min and collected by centrifugation. After centrifugation, the cells were gently resuspended in 50 ml of isotonic buffer containing 10 mM Tris–HCl (pH 7.5), 50 mM NaCl, 5 mM MgCl2, 0.1 mM PMSF, 0.1% Trasylol, and 10 nM Microcystin LR and then incubated for 10 min on ice. After another centrifugation, the cell pellet was suspended in 12 ml of buffer containing 10 mM Tris–HCl (pH 7.5), 50 mM NaCl, 5 mM MgCl2, 0.5 mM CaCl2, 0.5 M sucrose, 0.1 mM PMSF, 0.1% Trasylol, and 10 nM Microcystin LR. After a 2-min incubation on ice, a solution of 10% NP40 and 5% sodium deoxycholate (DOC) was diluted 100-fold into the cell suspension, and the cells were immediately disrupted by 15 vigorous strokes of a type A pestle in a Dounce homogenizer. The lysates were immediately layered onto 30 ml of sucrose cushion containing 10 mM Tris–HCl (pH 7.5), 50 mM NaCl, 5 mM MgCl2, 0.5 mM CaCl2, 40% sucrose, 0.1 mM PMSF, 0.1% Trasylol, 10 nM Microcystin LR, 0.1% NP40, and 0.05% DOC. Centrifugation was carried out for 30 min at 2,500×g at 4°C. Chromosomal pellets were resuspended in 0.2 ml of the same buffer. Typical yields of chromosomes were about 10 A260. Nuclei were isolated according to the same procedure, except that asynchronous cells were used without mitotic blocking.
Bovine serum albumin/polylysine-coated coverslips
For better attachment of chromosomes or nuclei to coverslips, bovine-serum-albumin (BSA)/polylysine-coated coverslips were used for EM samples. Ethanol-washed coverslips (diameter, 12 mm) were incubated with a solution of 10% BSA (Sigma) for 5 min at room temperature. After briefly washing with double-distilled H2O (ddH2O), the coverslips were baked at 65°C for 30 min. The BSA-coated coverslips were then incubated in a solution of 1 mg/ml poly-l-lysine (Sigma; average M w=150 kDa) for 5 min at room temperature and air-dried completely after briefly washing with ddH2O.
UV procedure for EM
A total of 0.5 A260 of isolated chromosome clusters or nuclei was diluted about 25 times into 250 μl of C10Mg1 buffer [10 mM cacodylate (pH 7), 1 mM MgCl2, 0.5 mM ATPγS, 0.1 mM PMSF, and 10 nM Microcystin LR] and incubated on ice for 20 min (Figs. 4, 5, and 6). The suspensions were spun at 0°C onto the BSA/polylysine-coated coverslips placed in the plastic buckets and UV-irradiated as described above for 5–10 min. Following a brief wash with C10Mg1 buffer, the chromosomes or nuclei were used for immunogold labeling as described below. For whole cell staining (Fig. 4a), HeLa S3 cells were grown on coverslips coated with polylysine. The coverslips were placed into the plastic buckets to treat the cells with 250 μl of an ice-cold extraction buffer containing 0.1% Triton X-100, 0.5 mM ATPγS, 0.1 mM PMSF, and 10 nM Microcystin LR in C10Mg1. After briefly washing the coverslips in the buckets with ice-cold C10Mg1, the plastic buckets were filled with 250 μl of C10Mg1 and UV-irradiated as described above. The coverslips were then processed for immunostaining using FluoroNanogold.
IF and immuno-EM studies of Emerin localization in UV-irradiated nuclei. a Cells or isolated nuclei (not shown) were UV-irradiated and immunostained for Emerin (α-Emerin) and FluoroNanogold ABs. The stained cells were observed with fluorescence microscope. b For EM observation, immunostained nuclei were fixed with GLU, postfixed with OSO4, and, finally, gold-enhanced to increase electron scattering. Following these steps, the samples were dehydrated and embedded in Epon resin. Note that the nuclear periphery is densely labeled with numerous gold particles, while the nucleoplasm is nearly free of background signals
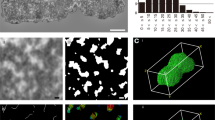
Immuno-EM of chromosomal scaffolding proteins in UV-irradiated chromosomes. Micrographs of thin-sectioned chromosomes immunostained for hBar (αhBar) or topoIIα (αtopoIIα) as indicated and thin-sectioned according to the UV-complete protocol. a–c and e αhBar. d αTopoIIα. All images, except e (longitudinal), are cross-sectional images of the chromosomes. Note that the immunogold particles for both proteins map toward the center of the chromosomal body and that the chromatin fibers are predominantly radial. f A chromosomal cross section prepared by the PAF/GLU combination protocol. In this case, chromosomes were first prefixed with PAF (instead of UV) and followed by the UV-complete procedure. Note that this protocol leads to networking and a poor preservation of the structures. Section thickness is around 90 nm
Omission of the GLU and OsO4 steps yields chromosomes with excellent global morphology and chromatin structure. a–c UV-only procedure: Chromosomes were UV-irradiated and processed for dehydration by ethanol without immunostaining and fixation steps by GLU and OS. d No-UV procedure: Chromosomes were processed as described above, but, in addition, by omitting the UV irradiation step. Note that these chromosomes diluted in C10Mg1 were directly subjected to ethanol dehydration with fine steps essentially as described by Marsden and Laemmli (1979) prior to embedding as described for the UV-complete protocol. Note that both procedures produced well-preserved, rather homogeneous 30-nm fibers that are predominantly star-like, oriented around these chromosomal cross sections. Section thickness is around 90 nm
Immunogold labeling of isolated chromosome clusters and nuclei
The following steps were carried out by floating the coverslips on drops of solution placed on a piece of Parafilm. The UV-irradiated chromosomes or nuclei were incubated with 3% BSA and 2% NGS in C10Mg1 for 15 min at room temperature. The coverslips were incubated with the first AB [3,000-fold-diluted anti-hBar rabbit serum (Maeshima and Laemmli 2003), 1,000-fold-diluted human topoIIα mouse monoclonal AB (MBL), or 2,000-fold-diluted anti-Emerin rabbit serum (from K. Wilson)] in C10Mg1 buffer containing 0.2% acetylated BSA (Aurion) for 1 h. After washing with C10Mg1 containing 0.2% acetylated BSA (five times for 3 min), the coverslips were incubated at room temperature for 1 h with 20-fold-diluted anti-rabbit or anti-mouse FluoroNanogold (Nanoprobes) in C10Mg1 containing 0.2% acetylated BSA. For light microscopy, after extensive washing with C10Mg1 containing 0.2% acetylated BSA (six times for 3 min), the coverslips were mounted on PPDI and observed with a DeltaVision microscope. For EM (UV-complete protocol; Figs. 4b and 5a–e), the chromosomes on the coverslips were postfixed in plastic buckets filled with 250 μl of 0.4% GLU in C10Mg1 for 12 h at 0°C. After extensively washing the coverslips in buckets with C10Mg1 and subsequently with ddH2O, gold enhancement was performed for 2 min according to the manufacturer's instructions (Nanoprobes). Gold-enhanced chromosomes on the coverslips were fixed with 1% osmium tetraoxide (OsO4) in C10Mg1 for 1 h on ice. Fine-step dehydration with ethanol was carried out exactly as described (Marsden and Laemmli 1979). The Epon mixture used in this study contained 25 g of Glycid ether 100 (Serva), 13 g of DDSA (Serva), 12 g of MNA (Serva), and 0.6 ml of DMP-30 (Serva). Infiltration was carried out according to the following schedule: 1:1 ethanol/Epon for 1 h; 3:1 ethanol/Epon for 1 h; and pure Epon for 1 h. Polymerization was allowed to proceed at 60°C for 3 days. After removing the coverslips, sectioning was performed with a diamond knife (DiATOME, Switzerland) using a Leica EM FCS apparatus (Leica, Austria). Around 90-nm thin sections were collected and transferred to 200-mesh hexagonal grids using Perfect Loop (DiATOME). The sections were stained with 3% uranyl acetate (Fluka) followed by 0.3% lead citrate (Fluka). Electron micrographs were taken with a Philips EM-310 electron microscope operated at 60 keV. For immunogold labeling following PAF fixation (Fig. 5f), the diluted chromosomes in C10Mg1 (volume, 250 μl) were spun onto the BSA/polylysine-coated coverslips in plastic buckets as described above. The chromosomes on the coverslips were fixed with freshly prepared 0.8% PAF in C10Mg1 for 20 min on ice and then immunolabeled as described above. For UV-only and no-UV protocols (Fig. 6), immunostaining was omitted. The chromosomes on coverslips with or without UV irradiation in C10Mg1 were directly subjected to a series of fine ethanol dehydration steps exactly as described (Marsden and Laemmli 1979) and then embedded in Epon as described above.
Results
Immuno-EM of mitotic chromosomes
One long-term goal of our laboratory is to deduce the organizational principles that fold the chromatin fiber in chromosomes. Toward this end, we wish to localize the scaffolding proteins in chromosomes using immuno-EM. Chromosomes are too large for whole-mount EM studies, hence necessitating thin-sectioning techniques. Ideally, this work would be carried out with cryosectioned, vitrified native (flash-frozen) chromosomes. Technical progress toward such an approach was recently reported (Al-Amoudi et al. 2004). But prior to attempting this difficult technique, it was necessary to develop a specific immunolabeling protocol that was also compatible with cryo-EM. Such a procedure has to be efficient as to “immunopaint” (with many gold particles) organizational principles into the complex, fibrous network of chromatin. We report here on an improved protocol for immuno-EM and evaluate the results using classical resin-embedded chromosomes.
Two possibilities for the immunolabeling of embedded biological specimens exist: either ABs are applied prior to embedding, or staining is attempted on thin sections. Since initial experiments established that thin sections of chromosomes immunostain poorly, supposedly due to loss of antigenicity and reduced accessibility, we explore preembedded staining.
Problems with PAF and GLU fixation: antigenicity, penetration, and networking
Fixation with PAF is the preferred method for IF studies since, empirically, it appears to preserve well the integrity and antigenicity of biological specimens. Conversely, GLU is the preferred fixative for EM studies. This treatment, however, often abolishes antigenicity—an observation made with all specific ABs studied here.
Figure 1 (a and b) shows that PAF-fixed chromosomes immunostained for a condensin component hBar display a beaded axial scaffolding (red) within the blue (DAPI) body in each chromatid (Maeshima and Laemmli 2003). In contrast, fixation with GLU at concentrations as low as 0.001% results in loss of staining (c). Unexpectedly, unfixed chromosomes also do not immunostain for hBar (d) and other antigens such as topoIIα (see below; Fig. 2). Hence, PAF fixation is an important step that not only fixes the structure, but also “renders” chromosomes antigenic. We do not understand the rationale of this observation, but speculate that this aldehyde facilitates penetration of ABs, as one can note a partial unfolding of chromosomes and the chromatin fiber following treatment (see below).
IF following UV irradiation
We aimed to develop a procedure that omits aldehyde fixation but allows immunostaining under conditions where the global structures of chromosomes and the chromatin fiber are preserved. Different chemical crosslinkers, including bifunctional reagents such as diimidoesters and N-hydroxysuccimide (NHS) esters, were explored with unsatisfactory results. Unexpectedly, we found that UV irradiation of chromosomes and nuclei promoted specific immunostaining using conventional IF (shown next) or the FluoroNanogold ABs that were needed for immuno-EM (below).
UV irradiation with 295-nm germicidal lamps of chromatin introduces predominantly DNA lesions, such as thymidine dimers, and, to a much lower level, some DNA protein crosslinks. In contrast, protein–protein crosslinks should be very rare. Indeed, we observe that the protein pattern of chromosomes on sodium dodecyl sulfate gels remains unaltered following irradiation by the optimal UV dose (not shown).
The UV protocol is simple; native chromosomes or nuclei are spun onto coverslips, irradiated for different lengths of time with the UV Stratalinker 2400 (Stratagene), and then conventionally immunostained. Figure 2 shows that spread chromosomes irradiated for 5 min stain equally well (top row) as PAF-fixed ones (middle row) for a number of antigens indicated. These are the scaffolding proteins topoIIα and hBar and the centromeric/telomeric proteins CENP-F and TRF-1, respectively. In contrast, unfixed chromosomes (bottom row) stain poorly for all these proteins. Some topoIIα staining, though, is noted at the telomeres and centromeres, suggesting that this antigen may be more accessible at these regions.
FluoroNanogold secondary ABs
Immuno-EM demands secondary ABs tagged with an electron-dense group to allow detection by electron scattering. Colloidal gold is classically used; this bulky tag, however, often results in low-level staining and significant background, presumably arising from the negative charge and poor penetration of the bulky colloidal gold (Humbel et al. 1998). Recent studies illustrated improved properties, with better penetration, of the commercial ABs called FluoroNanogold (Nanoprobes). These are Fab fragments containing both a fluorescent group and a small cluster of gold atoms. The FluoroNanogold used here contains 64 gold atoms (diameter, 1.4 nm) and requires a gold enhancement step for the detection by EM (Nanoprobes). Importantly, the dual tags of FluoroNanogold allow correlative fluorescence and EM, thus permitting examination of immunolabeling results by IF prior to a more cumbersome EM analysis (Robinson et al. 2001).
We compared the staining properties of conventional fluorescent immunoglobulin G (IgG) and FluoroNanogold ABs using PAF-fixed mitotic cells stained for hBar. Figure 3a shows impressive staining results obtained with a conventional fluorescent IgG secondary AB. This micrograph has to be compared to that obtained with FluoroNanogold (b). This comparison establishes that the staining signal of FluoroNanogold, although adequate, lacks the crisp appearance observed with the IgG control, supposedly due to a much higher general background. We show next that UV fixation of chromosomes significantly improves immunostaining with FluoroNanogold.
Spread chromosomes were immunostained for hBar following UV irradiation for different lengths of times (indicated). Figure 3c–g shows that chromosomes irradiated for a few minutes stain very well for hBar using FluoroNanogold for detection (d–g). In contrast, nonirradiated chromosomes (c; 0 min) display a strong background signal at the chromosomal periphery that arises from a nonspecific interaction of FluoroNanogold with the sticky chromosomal periphery. Specific staining is noted over a large time range of UV irradiation (3–20 min) by the beaded appearance of the signal, but is judged optimal around 5 min. This corresponds to a UV dose of about 4,000 J/m2. Examination of these micrographs obtained by UV irradiation indicates that, under these conditions, FluoroNanogold ABs stain similarly well as conventional fluorescent IgGs.
Immuno-EM and UV irradiation
Immunosignals observed by immuno-EM are composed of individual gold particles. Such localization studies appear convincing when antigen positions are defined by multiple clustered gold particles. Consequently, a high-level definition of antigens with many gold particles and a low background is a major experimental objective. We demonstrate next that UV irradiation is an important experimental step used to achieve this goal for chromatin antigens.
Emerin
We explored the UV FluoroNanogold procedure using an antigen that should yield an easily interpretable pattern. Emerin is an integral protein of the inner nuclear membrane; loss of its function causes Emery–Dreifuss muscular dystrophy (Bengtsson and Wilson 2004). In confirmation, IF studies of UV-irradiated cells or isolated nuclei (not shown) localize Emerin to the nuclear periphery with high specificity (Fig. 4a). Isolated nuclei were further processed for embedding and thin sectioning by the following protocol, termed UV-complete procedure: Isolated nuclei were UV-irradiated, immunostained using FluoroNanogold as the secondary AB, fixed with GLU, postfixed with OsO4, and, finally, gold-enhanced to increase the electron scattering power of the 1.4-nm gold cluster. Following these steps, the samples were dehydrated and embedded in Epon resin. The right panel (b) of Fig. 4 reveals that the nuclear periphery is densely labeled with numerous gold particles, while the nucleoplasm is nearly free of background signals. This observation demonstrates that the UV-complete protocol can yield immuno-EM data of excellent specificity and low background.
Chromosomal scaffolding proteins
Previous structural studies by EM of thin-sectioned chromosomes showed a radial, star-like orientation for the chromatin fiber in a cross section (Marsden and Laemmli 1979). The data suggest that chromosomes consist of radial chromatin loops that centrally are somehow tethered by scaffolding proteins. These tethers are structurally invisible and chromatin-like, but are inferred from the convergence of the fibers toward a longitudinal axis both in swollen and compact chromosomes. IF studies of topoIIα, hBar, and other condensin components are consistent with this radial loop model, and the immuno-EM data presented next provide additional support at a higher resolution.
Figure 5 shows a number of micrographs of thin-sectioned chromosomes immunostained for hBar or topoIIα (indicated) using the UV-complete protocol (a–e). Examination of chromosomal cross sections (a–d) establishes that the immunogold particles for both proteins (indicated) map toward the center of the chromosomal body. This localization is defined by a number of clustered gold particles with only rare individual particles in the peripheral halo (background). This clustering inspires confidence that the results are genuine.
The gold particles in cross sections appear frequently arranged in a C-shape (a and d); this may reflect coiling of the scaffolding. In more oblique sections (b), gold particles appear more linearly arranged or follow a wider arc. Examination of these panels confirms our previous observation that chromatin is star-like, oriented around the long chromosomal axis. Fold-back chromatin loops are sometimes observed at the periphery and can be traced toward the center, where individual fibers can no longer be followed with certainty. These scaffolding proteins are expected to define the bases of the chromatin loops, but the compact nature of chromosomes obscures a direct demonstration of this notion.
Fig. 5f shows a chromosomal cross section prepared by the PAF/GLU combination protocol. In this case, chromosomes were first prefixed with PAF (instead of UV) and were processed thereafter by the UV-complete procedure. This micrograph and further extensive studies established that PAF fixation generated chromosomes that appeared disorganized. This is inferred from a reduced radial orientation of the fibers and a broader distribution of the immunosignals, which may be due to swelling of chromosomes. PAF also poorly preserves the 30-nm chromatin fibers, as evidenced by its more heterogeneous (thin and thick) width. We further suggest that PAF fixation and, to a lower extent, GLU fixation (below) lead to chromatin “networking,” where chromatin can only be traced for short distances. This is proposed to arise through a chemical interlinking of chromatin fibers by PAF (see below); such artifactual crossties might obscure the radial look of chromatin. Based on these observations, we consider it reasonable to assume that PAF is an unsuitable fixative chromatin structure.
Radial loop model revisited: fixation artifact?
The radial loop model proposes that scaffolding proteins mediate interchromatin crossties. Could this model be an experimental artifact where chromatin crossties are induced by aldehyde fixation? It is certainly conceivable that chemical fixation could translink neighboring chromatin fibers into artifactual loops. Indeed, our studies with PAF-fixed chromosomes suggest that this chemical led to a networking of chromatin. Although networking was less evident with GLU fixation, we addressed this problem by omitting fixation steps by GLU and OsO4 in the UV-complete protocol (i.e., chromosomes were irradiated with UV and, while omitting GLU and OsO4 fixation, directly dehydrated in fine steps with ethanol and embedded in Epon resin; UV-only method). As a control, nonirradiated chromosomes were processed identically (no-UV). Note that these chromosomes were not immunostained (see below).
Unexpectedly, we observe that omission of the GLU and OsO4 steps yields chromosomes with an excellent global morphology and chromatin structure both with the UV-only and the no-UV protocols (Fig. 6). Examination of the micrographs of Fig. 6 shows chromatin as well-preserved, rather homogeneous 30-nm fibers that are predominantly radially oriented. Interestingly, the chromosome structure appears globally less complex than those prepared with the UV-complete protocol shown above. This is manifested by a better traceability of the chromatin fiber over longer distances toward the center. Sometimes entire radially oriented loops can be traced from the tips to their presumptive bases near the chromosomal center (arrow heads, a and b). In contrast, entire loops are never observed in sectioned chromosomes prepared with the UV-complete protocol. As has been said, the micrographs obtained with the UV-only and no-UV procedures are similar. This is not surprising since UV irradiation is not a protein–protein fixative, although it serves as an immunostaining method that is omitted in the discussion here.
The experiments argued against the concern that the radial look of chromosomal cross sections arose from aldehyde fixation. One drawback of the UV-only and no-UV protocols, though, was that we were unable to find conditions to combine with FluoroNanogold staining. In a series of experiments, we learned that the buffers for gold enhancement were incompatible with maintaining the morphology of unfixed chromosomes.
Discussion
To dissect mitotic chromosomes structurally, it is necessary to understand the folding principles of the chromatin fiber in three dimensions and to position along its length proteins involved in long-range chromosome structure, such as topoIIα and condensin. EM is the instrument of choice for such a study, where the large dimensions of mitotic chromosomes demand thin sectioning combined with immuno-EM. We report here on a much improved immuno-EM procedure that avoids aldehyde fixation. This procedure was developed because GLU fixation generally caused loss of antigenicity, while PAF fixation allowed immunostaining but poorly preserved the global structure of chromosomes and chromatin. In addition, both fixation procedures, most notably PAF fixation, led to chromatin networking, where chromatin fibers were chemically translinked by aldehyde-mediated crossties.
UV irradiation
We showed that UV irradiation of isolated nuclei and chromosomes facilitates immunostaining, as established by fluorescence microscopy, with a variety of ABs directed against chromosomal proteins (Fig. 2). We do not understand why UV irradiation works. Conceivably, penetration of ABs into untreated chromosomes is impeded, perhaps by side-by-side aggregated and torsionally stressed chromatin strands that are relaxed by UV-induced DNA breaks. Whatever the rationale might be, what is of importance is that UV irradiation results in immunostaining of similar specificity to that obtained with conventional PAF fixation.
The main advantage of UV irradiation, however, is that it can be extended to immuno-EM using classical thin-sectioning procedures and supposedly to cryo-EM (below). This is experimentally exemplified by the immuno-EM studies of Emerin in nuclei (Fig. 4), as well as topoIIα and hBar in chromosomes (Fig. 5). All these micrographs show impressive substructural immunosignals with little general background (i.e., the Emerin signals are strongly concentrated at the nuclear periphery, while only rare gold particles are observed in the nuclear lumen). Similarly, the location of topoIIα and hBar is defined with many gold particles to be a central subchromosomal region with little peripheral noise. This contrasts with observations where the UV step is replaced by PAF prefixation. As has been said, PAF led to chromatin networking, dispersed immunosignals, and poor preservation of the chromatin fiber and the global chromosome structure (Fig. 5f).
Chromosome structure
Although this report has a methodological focus, the results obtained extend aspects of chromosome structure. The immuno-EM data reinforce the notion that chromosome structure involves interactions, presumably crossties of chromatin loops, mediated by a central longitudinal organizer called scaffolding. This is inferred from the predominantly radial orientation of the chromatin fiber in cross-sectional views and the restricted axial location of topoIIα and hBar. The compactness of the chromosomal center, however, obscures the important details. The main questions remain unanswered. Do these proteins define the bases of the chromatin loops? What is the structural relationship of topoIIα and hBar? Do they form a protein-linked skeleton? Toward this aim, it will be necessary to apply double immunostaining techniques and to avoid potential artifacts of the classical EM.
UV-only and no-UV protocols
As mentioned above, PAF fixation and, to a much lower extent, GLU fixation led to networking (chemical translinking). Since GLU (also OsO4) fixation was used in the UV-complete protocol, we also embedded chromosomes by omitting both these fixation steps. Interestingly, the micrographs obtained were even more consistent with a radial-type model, since it was often possible to trace entire loops (Fig. 6). Unexpectedly, chromosomes and chromatin appeared to be well preserved by this simplified procedure even if UV irradiation was omitted. Supposedly, fixation was achieved in these protocols by the fine dehydration steps with ethanol.
Artifacts
While we consider UV irradiation and immunostaining, a near-native procedure carried out in physiological buffer, the same does not apply to plastic embedding protocols. Despite this reservation, IF, immuno-EM, and live imaging (Tavormina et al. 2002; Hirota et al. 2004) experiments congruently establish that complexes involved in long-range chromosomes structure (topoIIα and condensin) map to a longitudinal organizer that we have called scaffolding.
Further progress in chromosome structure will demand the cryo-EM of vitrified native chromosomal sections. We believe that it should be possible to adapt the UV procedure to cryo-EM, where immunostained chromosomes are then processed for cryosectioning. Toward this goal, it will be necessary to replace the FluoroNanogold with another tag, such as colloidal gold, since the former demands gold enhancement under nonphysiological conditions. Clearly, the next experimental hurdle is to master the cryosectioning of vitrified chromosomes and to combine this with immunostaining.
Abbreviations
- TopoIIα:
-
Topoisomerase IIα
- AB:
-
antibody
- IF:
-
immunofluorescence
- GLU:
-
glutaraldehyde
- PAF:
-
paraformaldehyde
- hBar:
-
hBarren
- EM:
-
electron microscopy
- UV:
-
ultraviolet
- OsO4 :
-
osmium tetraoxide
References
Adachi Y, Luke M, Laemmli UK (1991) Chromosome assembly in vitro: topoisomerase II is required for condensation. Cell 64:137–148
Adolph KW (1981) A serial sectioning study of the structure of human mitotic chromosomes. Eur J Cell Biol 24:146–153
Al-Amoudi A, Chang JJ, Leforestier A, McDowall A, Salamin LM, Norlen LP, Richter K, Blanc NS, Studer D, Dubochet J (2004) Cryo-electron microscopy of vitreous sections. EMBO J 23:3583–3588
Bazett-Jones DP, Kimura K, Hirano T (2002) Efficient supercoiling of DNA by a single condensin complex as revealed by electron spectroscopic imaging. Mol Cell 9:1183–1190
Bengtsson L, Wilson KL (2004) Multiple and surprising new functions for emerin, a nuclear membrane protein. Curr Opin Cell Biol 16:73–79
Boy de la Tour E, Laemmli UK (1988) The metaphase scaffold is helically folded: sister chromatids have predominantly opposite helical handedness. Cell 55:937–944
Cobbe N, Heck MM (2000) Review: SMCs in the world of chromosome biology—from prokaryotes to higher eukaryotes. J Struct Biol 129:123–143
Earnshaw WC, Heck MM (1985) Localization of topoisomerase II in mitotic chromosomes. J Cell Biol 100:1716–1725
Earnshaw WC, Laemmli UK (1983) Architecture of metaphase chromosomes and chromosome scaffolds. J Cell Biol 96:84–93
Gasser SM, Laroche T, Falquet J, Boy de la Tour E, Laemmli UK (1986) Metaphase chromosome structure. Involvement of topoisomerase II. J Mol Biol 188:613–629
Hirano T (2000) Chromosome cohesion, condensation, and separation. Annu Rev Biochem 69:115–144
Hirano T (2002) The ABCs of SMC proteins: two-armed ATPases for chromosome condensation, cohesion, and repair. Genes Dev 16:399–414
Hirano T, Mitchison TJ (1993) Topoisomerase II does not play a scaffolding role in the organization of mitotic chromosomes assembled in Xenopus egg extracts. J Cell Biol 120:601–612
Hirano T, Kobayashi R, Hirano M (1997) Condensins, chromosome condensation protein complexes containing XCAP-C, XCAP-E and a Xenopus homolog of the Drosophila Barren protein. Cell 89:511–521
Hirota T, Gerlich D, Koch B, Ellenberg J, Peters JM (2004) Distinct functions of condensin I and II in mitotic chromosome assembly. J Cell Sci 117:6435–6445
Humbel BM, de Jong MD, Muller WH, Verkleij AJ (1998) Pre-embedding immunolabeling for electron microscopy: an evaluation of permeabilization methods and markers. Microsc Res Tech 42:43–58
Kimura K, Rybenkov VV, Crisona NJ, Hirano T, Cozzarelli NR (1999) 13S condensin actively reconfigures DNA by introducing global positive writhe: implications for chromosome condensation. Cell 98:239–248
Kireeva N, Lakonishok M, Kireev I, Hirano T, Belmont AS (2004) Visualization of early chromosome condensation: a hierarchical folding, axial glue model of chromosome structure. J Cell Biol 166:775–785
Laemmli UK, Cheng SM, Adolph KW, Paulson JR, Brown JA, Baumbach WR (1978) Metaphase chromosome structure: the role of nonhistone proteins. Cold Spring Harb Symp Quant Biol 42:351–360
Lewis CD, Laemmli UK (1982) Higher order metaphase chromosome structure: evidence for metalloprotein interactions. Cell 29:171–181
Maeshima K, Laemmli UK (2003) A two-step scaffolding model for mitotic chromosome assembly. Dev Cell 4:467–480
Marsden MP, Laemmli UK (1979) Metaphase chromosome structure: evidence for a radial loop model. Cell 17:849–858
McHugh B, Heck MM (2003) Regulation of chromosome condensation and segregation. Curr Opin Genet Dev 13:185–190
Ono T, Losada A, Hirano M, Myers MP, Neuwald AF, Hirano T (2003) Differential contributions of condensin I and condensin II to mitotic chromosome architecture in vertebrate cells. Cell 115:109–121
Ono T, Fang Y, Spector DL, Hirano T (2004) Spatial and temporal regulation of Condensins I and II in mitotic chromosome assembly in human cells. Mol Biol Cell 15:3296–3308
Paulson JR, Laemmli UK (1977) The structure of histone-depleted metaphase chromosomes. Cell 12:817–828
Robinson JM, Takizawa T, Pombo A, Cook PR (2001) Correlative fluorescence and electron microscopy on ultrathin cryosections: bridging the resolution gap. J Histochem Cytochem 49:803–808
Strunnikov AV, Larionov VL, Koshland D (1993) SMC1: an essential yeast gene encoding a putative head–rod–tail protein is required for nuclear division and defines a new ubiquitous protein family. J Cell Biol 123:1635–1648
Strunnikov AV, Hogan E, Koshland D (1995) SMC2, a Saccharomyces cerevisiae gene essential for chromosome segregation and condensation, defines a subgroup within the SMC family. Genes Dev 9:587–599
Taagepera S, Rao PN, Drake FH, Gorbsky GJ (1993) DNA topoisomerase II alpha is the major chromosome protein recognized by the mitotic phosphoprotein antibody MPM-2. Proc Natl Acad Sci U S A 90:8407–8411
Tavormina PA, Come MG, Hudson JR, Mo YY, Beck WT, Gorbsky GJ (2002) Rapid exchange of mammalian topoisomerase II alpha at kinetochores and chromosome arms in mitosis. J Cell Biol 158:23–29
Uemura T, Ohkura H, Adachi Y, Morino K, Shiozaki K, Yanagida M (1987) DNA topoisomerase II is required for condensation and separation of mitotic chromosomes in S. pombe. Cell 50:917–925
Wang JC (1996) DNA topoisomerases. Annu Rev Biochem 65:635–692
Acknowledgements
We are very grateful to T. Durussel for her competent technical assistance, and Drs. E. Käs and C. Bauer for comments on the manuscript. We thank Drs. K. Wilson and T. de Lange for the anti-Emerin and TRF-1 sera, respectively. The Louis-Jeantet Medical Foundation, the Swiss National Fund, and the Canton of Geneva supported this work.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Communicated by E. A. Nigg
Rights and permissions
About this article
Cite this article
Maeshima, K., Eltsov, M. & Laemmli, U.K. Chromosome structure: improved immunolabeling for electron microscopy. Chromosoma 114, 365–375 (2005). https://doi.org/10.1007/s00412-005-0023-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-005-0023-7