Abstract
Mislocalization and abnormal deposition of TDP-43 into the cytoplasm (TDP-43 proteinopathy) is a hallmark in neurons of amyotrophic lateral sclerosis (ALS) and frontotemporal lobar degeneration (FTLD). However, the pathogenic mechanism of the diseases linked to TDP-43 is largely unknown. We hypothesized that the failure of mRNA transport to neuronal axons by TDP-43 may contribute to neurodegeneration in ALS and FTLD, and sought to examine the function of TDP-43 by identifying its target mRNA for axonal transport. We found that mRNAs related to translational function including ribosomal proteins (RPs) were decreased by shRNA-based TDP-43 knock-down in neurites of cortical neurons. TDP-43 binds to and transports the RP mRNAs through their 5′ untranslated region, which contains a common 5′ terminal oligopyrimidine tract motif and a downstream GC-rich region. We showed by employing in vitro and in vivo models that the RP mRNAs were translated and incorporated into native ribosomes locally in axons to maintain functionality of axonal ribosomes, which is required for local protein synthesis in response to stimulation and stress to axons. We also found that RP mRNAs were reduced in the pyramidal tract of sporadic ALS cases harboring TDP-43 pathology. Our results elucidated a novel function of TDP-43 to control transport of RP mRNAs and local translation by ribosomes to maintain morphological integrity of neuronal axons, and proved the influence of this function of TDP-43 on neurodegeneration in ALS and FTLD associated with TDP-43 proteinopathy.
Similar content being viewed by others

Avoid common mistakes on your manuscript.
Introduction
Amyotrophic lateral sclerosis (ALS) and frontotemporal lobar degeneration (FTLD) are neurodegenerative diseases that mainly affect motor neurons of the brain and spinal cord, and cortical neurons of the cerebrum to cause muscle weakness and dementia, respectively. A variety of causative genes have been identified in ALS and FTLD thus far, and most of the genes are involved in the pathogenesis of both diseases [29]. However, the molecular mechanism underlying these diseases is unclear and robust treatment strategy for them has not yet been developed.
However, one of the most important pathological hallmarks of the diseases is the disappearance of TAR DNA-binding protein (TDP)-43 from the nucleus and abnormal deposition into the cytoplasm of affected neurons (TDP-43 proteinopathy) [3, 34], which is observed in cases with many of the causative gene mutations [29]. Mutations of the TDP-43 gene TARDBP itself have been identified in some ALS and FTLD cases, which exhibit TDP-43 proteinopathy [23, 41, 54]. Furthermore, these pathological abnormalities of TDP-43 are also observed in sporadic cases of ALS and FTLD [29]. These suggest that the aberrant localization of TDP-43 may be a common pathogenic factor among ALS and FTLD of different etiologies to execute neurodegeneration.
TDP-43 is an RNA-binding protein that mainly exists in the nucleus, and is known to control gene transcription or alternative splicing of transcribed pre-mRNA [37]. TDP-43 is also supposed to regulate generation of some micro-RNAs [25]. Moreover, TDP-43 has been reported to shuttle between the nucleus and the cytoplasm, which implies that this protein is involved in the regulation of transport and translation of mature mRNA in the cytoplasm [37]. Thus far it is not confirmed whether a toxic gain or loss of function of TDP-43 is critical for the pathogenesis. Previous reports showed that both the overexpression of wild-type or disease-related mutant TDP-43 [18, 42, 50, 51, 53], and TDP-43 deficiency in neurons [19, 52] can cause neurodegeneration phenotype in animal models. Furthermore, TDP-43 deficiency has been reported to cause the same changes in the expression profile of a subset of mRNAs in motor neurons, as in those of ALS cases possessing TDP-43 proteinopathy [28]. The similar pathological phenotypes obtained from mislocalization and dysregulated expression levels of TDP-43 suggest that, although there is a possible neurotoxic effect of deposited TDP-43 itself, the functional abnormality of TDP-43 may be a definitive factor of neurodegeneration in ALS and FTLD.
Neurons are characterized by having long neurites consisting of axons and dendrites to communicate with each other via synapses in the neuronal network, which make them essential for transporting a variety of materials such as proteins and mitochondria to the neurites helping to maintain their structure and function. Recently, mRNAs were also reported to be direct cargo for transport within neurites in conjunction with some RNA-binding proteins to form neuronal RNA granules [30]. On the occasion of developmental axonal projection, regeneration, or synaptic remodeling, transported mRNAs undergo translation to proteins locally in the translational apparatus ribosomes that exist in neurites, by which neurons can respond to a stimulus or injury immediately to elongate axons or form synapses, and adapt themselves promptly to the surrounding environment [26]. Dysfunction of an RNA-binding protein supplying critical mRNAs to neurites can result in perturbation of neuronal homeostasis and result in neuronal dysfunction. For instance, the pathogenesis of spinal muscular atrophy, a motor neuron disease affecting juveniles, is linked to the disturbance of axonal mRNA transport by the RNA-binding protein SMN (survival motor neuron protein) in motor neurons [14].
A fraction of TDP-43 is known to be located in axons, suggesting that this protein may be involved in the axonal transport of mRNA by forming neuronal RNA granules similarly to SMN [2, 13]. Indeed, previous reports have shown that TDP-43 binds to some mRNAs and migrates within axons, and that some mRNAs in axons decrease due to reduced expression of TDP-43 in neurons [2, 10, 21]. However, its major target mRNAs transported to axons have not been determined thus far. To identify functionally relevant TDP-43 target mRNAs systematically, we hypothesized that, in ALS and FTLD, functional depletion of TDP-43 triggered by its integration into protein aggregates in the cytoplasm causes inhibition of the transport of critical mRNAs to axons, and results in structural and functional failure of neurons. We searched target mRNAs for axonal transport by TDP-43 comprehensively by using microarray analysis, and identified mRNAs encoding ribosomal proteins as the target. Subsequently, we confirmed that transported ribosomal protein mRNAs by TDP-43 contribute to the maintenance of ribosomal function in axons via their local translation and assembly. We also demonstrated that loss of TDP-43-dependent functional integrity of ribosomal proteins causes neuronal dysfunction and ultimately neurodegeneration. While our current analysis was focused on TDP-43 function and not on the toxic property of deposited TDP-43, it showed that TDP-43 plays an essential role in the regulation of translational machinery in axons, and suggests that abnormal translational function in axons may lead to neurodegeneration in ALS and FTLD.
Materials and methods
Animal experiments
All animals were maintained in accordance with the institutional guidelines of the National Center of Neurology and Psychiatry, Osaka University and Niigata University. The technical protocols for animal experiments in this study were approved by the Small Animal Welfare Committee in the National Center of Neurology and Psychiatry.
TARDBP Flox mice were generated using ES cells derived from 129 mice according to the previously reported method [1] with a targeting vector where loxP sequences with neo cassettes interposed exon 3 of TARDBP gene. Eno2-Cre Tg mice were obtained from the Jackson Laboratory (the strain name; Tg(Eno2-cre)39Jme/J, Cat. No. 005938). TARDBP Flox/Flox mice were crossed with Eno2-Cre Tg mice to obtain TARDBP Flox/+ , Eno2-Cre Tg mice, which were then crossed with TARDBP Flox/Flox mice to obtain TARDBP Flox/Flox, Eno2-Cre Tg mice, TARDBP Flox/Flox, Eno2-Cre(−) mice and TARDBP Flox/+ , Eno2-Cre Tg mice. These mice were used for analyses.
Culture of cortical neurons
Cerebral cortices were excised from C57BL/6 J mice of embryonic day 15 (E15), dissociated with papain (Worthington Biochemical Corp.), and then seeded into a chamber insert (6-well plate size, 1 μm pore, Greiner Bio-One), a microfluidic chamber (Xona microfluidics), a 3.5-cm glass-bottom dish (AGC Techno Glass) or a 24-well plate (Greiner Bio-One) at 3 × 106, 2.7 × 105, 3 × 105 or 5 × 105 cells, respectively. After cultured in media containing DMEM (Thermo Fisher Scientific), 10% FBS (HyClone), and 1 × penicillin/streptomycin (Thermo Fisher Scientific) for the first 24 h, neurons were cultivated in N1 medium (MACS Neuro Medium (Miltenyi Biotec) with 2% MACS NeuroBrew-21 (Miltenyi Biotec) + 1 mM sodium pyruvate (Thermo Fisher Scientific) + 27.5 μM 2-mercaptoethanol (Thermo Fisher Scientific) + 1 × penicillin/streptomycin) for 48 h, followed by N1 medium + 2 μM cytosine β-D-arabinofuranoside (Ara-C, Sigma-Aldrich). For generating samples for microarray analysis and quantitative PCR, the medium was changed to N1 medium without Ara-C 24 h before harvest.
Preparation of viral vectors
Constructs for lentiviral vector (pLKO.1) expressing control shRNA that has no endogenous target gene (Cat. No. SHC002) and two different shRNA against mouse TDP-43 gene (TARDBP) (Cat. No. TRCN0000175828 (TDP-43 shRNA#1) and TRCN0000174930 (TDP-43 shRNA#2)) were purchased from Sigma-Aldrich.
Lentivirus production was performed as previously described [5]. Briefly, 1.6 × 107 293FT cells (Thermo Fisher Scientific) were seeded in a 15-cm dish and cultured in a media containing DMEM with 10% FBS and 1 × penicillin/streptomycin. Δ8.9 (22.5 μg) and VSVG (7.5 μg) plasmids were co-transfected with a shRNA construct (30 μg) 6 h after plating the cells by the calcium phosphate method. The lentivirus-containing medium was harvested at 38 and 62 h after transfection. The media was ultracentrifuged at 70,000×g for 140 min, and the precipitate was resuspended in Neuro Medium and used as virus solution. For transduction, viral solution was applied to cortical neurons at 1 day in vitro (DIV) for 6 h.
Recombinant adeno-associated virus serotype 1 (rAAV1) to express EGFP or La (rAAV1-CAG-EGFP or rAAV1-CAG-FLAG-La) was prepared according to the methods previously reported [44]. For transduction, viral solution was applied to cortical neurons at 1 DIV for 6 h.
Microarray analysis
Total RNA sample was prepared using TRIZOL (Thermo Fisher Scientific) from neurite fractions of chamber inserts at 8 DIV. A mixture of RNA samples from three wells was used for each group. The extracted total RNA was analyzed by 3D-Gene Mouse Oligo Chip 24 K (Toray Industries) after confirming that there was no noticeable denaturing of RNA by Bioanalyzer (Agilent Technologies). Total RNA was labeled with Cy3 or Cy5 using Amino Allyl MessageAMP II aRNA Amplification Kit (Thermo Fisher Scientific). The Cy3- or Cy5-labeled aRNA pools were hybridized for 16 h. The hybridization signals were detected using 3D-Gene Scanner (Toray Industries) and analyzed with 3D-Gene Extraction software (Toray Industries). Detected signals for the respective genes were normalized by the global normalization method (Cy3/Cy5 ratio median = 1). Gene ontology analysis was performed using the manufacturer’s analytical algorithm.
Quantitative PCR
After the extraction of total RNA using TRIZOL from neurite fractions of chamber inserts at 6 DIV, reverse transcriptase reaction was carried out according to the protocol of ReverTra Ace (TOYOBO) using 100 ng of total RNA and 25 μM random primer. The reaction mixture was diluted fivefold (final 100 μL), and 2 μL was subjected to the quantitative PCR using a protocol of SYBR Green Master Mix (Roche Diagnostics) and the Applied Biosystems Prism model 7300 sequence detection instrument as previously described [49]. Quantification of each transcript was performed by the ΔΔCt method using β-actin as a reference gene. Primer sets used for amplification of each gene product are shown in Supplementary Table 1 (online resource).
mRNA quantification with nCounter analysis system
Total RNA was extracted from the cerebral white matter excised from six frozen-stored brain slices (100 μm thickness each) of mice (3 mice for each experimental condition) at 21 days of age. Excision was performed using an optical microscope for laser microdissection (LMD7, Leica Microsystems). RNA extraction was performed using TRIZOL and reverse transcriptase reaction was done as above. After 20 cycles of PCR from the cDNA with TaqMan Universal Master Mix II, no UNG (Applied Biosystems), hybridization with probes and counts of the signals were achieved using nCounter (NanoString Technologies) according to the protocol for single cell gene expression. The data were analyzed by nSolver Analysis Software 3.0 (NanoString Technologies). Sequences of primer and probe sets for the analysis are available on request.
In situ hybridization
In situ hybridization cytochemistry was performed essentially as previously described [4]. Antisense and sense RNA probes from the portion of the open reading frame in each gene were synthesized from DNA templates with T7 promoter using RiboMAX Large Scale RNA Production Systems T7 (Promega) and digoxigenin. Cortical neurons at 5 DIV plated on the microfluidic chamber were fixed with 4% paraformaldehyde (PFA) in PBS for 20 min, followed by inactivation of peroxidase with 1% hydrogen peroxide in methanol for 10 min. After acetylation with acetylation buffer (2.26 mL triethanolamine and 0.3 mL hydrogen chloride in 167 mL RNase-free water) for 10 min, permeabilization was performed with 0.3% Triton-X in PBS for 5 min. RNA probes were hybridized at 0.5 μg/mL in hybridization buffer (5 × SSC, 2% blocking reagents (Roche Diagnostics), 50% formamide, 0.1% N-lauroylsarcosine and 0.1% SDS) overnight at 65 °C. After washing, hybridized probes were detected by peroxidase-conjugated anti-digoxigenin antibody (Cat. No. 11207733910, Roche Diagnostics) and TSA Fluorescein Tyramide Reagent Pack (PerkinElmer) in parallel with the immunostaining of TDP-43 using anti-TDP-43 antibody (Cat. No. 10782-2-AP, Proteintech). After the images were photographed with a confocal fluorescence microscope (FLUOVIEW FV1000, Olympus), the intensities of each RP mRNA and TDP-43 signal, and the co-localizing ratios between them in axons were calculated using ImageJ (the National Institutes of Health) and its plugin (Coloc 2) defining TDP-43 signals as the ROI. The differences of Pearson’s R values between groups were statistically examined as previously described [12].
Visualization of RP mRNAs
Expression plasmids pcDNA-Rpl41 full length, pcDNA-Rpl41 lacking 5′UTR, pcDNA-Rplp1 full length, and pcDNA-Rplp1 lacking 5′UTR contained the bacteriophage MS2 coat protein binding RNA sequence (MS2bs) downstream of the open reading frame of each RP with or without its 5′UTR [7]. NLS-MS2-Venus plasmid expresses MS2 coat protein with the nuclear localization signal fused to Venus [6]. pcDNA-full length or 5′UTR(−)-Rpl41, Rplp1-MS2bs and NLS-MS2-Venus plasmids were co-transfected into cortical neurons at 6 DIV in a 24-well plate with lipofectamine 2000 (Thermo Fisher Scientific). For analysis of the transport of RP mRNA in axons, mCherry expression plasmid (pmCherry-C1) was also co-expressed for labeling, and the longest neurite was used to measure. To see the effect of TDP-43 knock-down, neurons transduced with TDP-43 shRNA#1 lentiviral vector at 1 DIV were used. In analyzing co-localization of RP mRNA with TDP-43 and/or 5′ terminal oligopyrimidine tracts (5′TOP) trans-factors (La, TIA-1 and AUF1), pmCherry-TDP-43-Myc, pFLAG-La, TIA-1 or AUF1 was also co-transfected. Analysis of RP mRNA transport and co-localization with TDP-43 was done with live cells, whereas co-localization with each trans-factor was examined by immunostaining of cells after fixation with 4% PFA in PBS(−) using a fluorescence microscope (Leica Microsystems). Transport distances from the nucleus and the number of NLS-MS2-Venus granules, or the co-localizing ratios between NLS-MS2-Venus and TDP-43 or trans-factor signals were measured in axons using ImageJ and its plugin Sholl Analysis, or Coloc 2, respectively, defining NLS-MS2-Venus signals as the ROI.
RNA immunoprecipitation
Neuro2a cells were seeded on a 10-cm dish, and the plasmids pEGFP-TDP-43-Myc and pcDNA-full length, 5′UTR(−) or 3′UTR(−)-Rpl41-MS2bs were co-transfected with lipofectamine 2000 (Thermo Fisher Scientific). Inositol triphosphate receptor (IP3R) 3′UTR-MS2bs was used as a negative control [6]. Forty-eight hours after transfection, 300 mJ of UV was irradiated to crosslink protein–RNA complexes and cells were harvested. The cells were lysed in 50 mM Tris–HCl, pH 7.4, 300 mM NaCl, 5 mM MgCl2, 0.5% NP40 and 1 mM DTT with protease and RNase inhibitors by sonication. Supernatants of the lysates were immunoprecipitated using anti-Myc antibody (9E10, Developmental Studies Hybridoma Bank) and Protein G Dynabeads (Thermo Fisher Scientific) in the presence of 10 mg/mL BSA and 100 μg/mL tRNA. After treatment of the beads with 9 U RQ1 DNase (Promega) and RNase inhibitor for 30 min at 37 °C, the precipitates were recovered by adding SDS (final conc. 1%) and incubating 15 min at 65 °C. After proteins in the precipitates were digested by 200 μg/mL proteinase K for 40 min at 52 °C, RNA was extracted by TRIZOL. Thereafter, cDNA was synthesized by a reverse transcriptase reaction as described above, and quantitative PCR was carried out using primers MS2bsfwd (CTGGTCGACTCTAGATCG) and BGH reverse (TAGAAGGCACAGTCGAGG). β-actin was used as a reference gene.
Immunoprecipitation
Neuro2a cells were seeded in a 6-well plate, and pEGFP-TDP-43-Myc and pFLAG-La, -TIA-1, -AUF1 or -GAPDH were co-transfected with lipofectamine 2000. Cells were harvested 48 h after transfection and sonicated in lysis buffer (50 mM Tris–HCl, pH 7.4, 150 mM NaCl, 1 mM MgCl2, 0.05% NP40 and 1% Triton-X with protease and RNase inhibitors). Supernatants of the lysates were immunoprecipitated with anti-FLAG antibody (M2, Sigma-Aldrich) and Protein G Dynabeads. The precipitated proteins were recovered by adding 1 × loading buffer with 100 mM DTT and incubating 5 min at 95 °C.
Analysis of protein synthesis in growth cones by photoconversion
Kikume expression plasmid (pRpl41 5′UTR( +) or (−)-hKikGRMNL) [16] was transfected into cortical neurons at 6 DIV with lipofectamine 2000. Twenty-four hours later, a field of view including a growth cone and neighboring three consecutive fields were irradiated with UV by a time-lapse confocal fluorescence microscope (FLUOVIEW FV1000, Olympus) at 60 × objective lens and 3 × zoom to convert green fluorescence from Kikume to red fluorescence. Green and red fluorescence in a growth cone was then recorded every minute for a total of 30 min. Green and red fluorescence intensities at every time point were analyzed using ImageJ, and the ratio of both fluorescences was calculated.
Analysis of novel protein synthesis using BDNF or puromycin
After the medium was removed from neurons in chamber inserts or microfluidics at 6 DIV, the conditioned medium containing 50 nM BDNF or 10 μg/mL puromycin and/or 100 μM cycloheximide was added, restricted to either the bottom side of the inserts or to the axonal or cell body side of the microfluidics, and incubated for 2 h. After incubation and washing once with PBS(−), the neurites were collected from the lower surface of the inserts or fixed with 4% PFA in the microfluidics, and were subjected to immunoblotting or immunocytochemistry with anti-Rpl26 (Cat. No. A300-686A, Bethyl Laboratories), anti-Rps6 (Cat. No. 2217S, Cell Signaling Technologies), anti-tau (Cat. No. MAB3420, Sigma-Aldrich), anti-βIII-tubulin (Cat. No. PRB-435P, BioLegend) or anti-puromycin (Cat. No. 3RH11, Kerafast) antibody. For immunocytochemistry, the intensities of the protein signals in axons on confocal fluorescent images by a confocal fluorescence microscope were measured using ImageJ.
Analysis of RP incorporation into ribosomes in axons
Each mRNA for EGFP or EGFP-Rpl10a were synthesized from the PCR-amplified cDNA by mMESSAGE mMACHINE T7 Ultra Kit (Thermo Fisher Scientific) according to the manufacturer’s protocol, and transfected confined to the axonal portion of cortical neurons at 7 DIV in the microfluidic chamber using lipofectamine messenger MAX (Thermo Fisher Scientific). For activation of translation, 30 mM KCl was added to both the cell body and axonal compartments. Twenty-four hours after transfection, the neurons were fixed with 4% PFA in PBS(−) and subject to the immunostaining with anti-GFP antibody (12B15, Developmental Studies Hybridoma Bank) and anti-Rpl26 antibody. After the images were photographed with a confocal fluorescence microscope, the co-localizing ratios of EGFP signals onto Rpl26 signals in axons were calculated using ImageJ plugin (Coloc 2) defining Rpl26 signals as the ROI.
Analysis of axonal outgrowth in cultured neurons
Control or mouse TDP-43-targeted shRNA, and pcDNA or pcDNA-RP-HA with or without 5′UTR or pmCherry-TDP-43-Myc or pFLAG-La, and pCAG-EGFP were co-transfected into cortical neurons at 3 DIV and incubated for 48 h. After the fixation with 4% PFA in PBS(−), immunostaining was performed with anti-GFP antibody (Cat. No. 598, MBL) and the images were photographed with a fluorescent microscope, and the length of the longest neurite was measured with ImageJ plugin (Simple Neurite Tracer). To confirm the translation of introduced RP in axons, pcDNA-5′UTR( +)-RP-HA was transfected into neurons as well, and immunostaining was done with anti-HA (Cat. No. MMS-101R, BioLegend) and βIII-tubulin antibodies.
In utero electroporation
The uterus of a pregnant ICR mouse at embryonic day 14 (E14) was exposed and 1.6 μg control or TDP-43-targeted shRNA, 0.4 μg pcDNA or pcDNA-5′UTR( +)-Rpl26-HA, 0.5 μg pCAG-EGFP in PBS(−) with 0.05% Fast Green FCF (Sigma-Aldrich) was injected into the left lateral ventricle of an embryo by a glass capillary (Narishige), and electroporation was carried out under the condition of 40 V, 50 ms pulse, 450 ms interval, 5 times by electroporator (CUY21, NepaGene). Pups were collected 5 days later (P1), and the cerebrum was fixed overnight with 4% PFA in PBS(−), replaced with PBS(−) containing 20% sucrose, embedded in OCT compound (Sakura Finetek), and frozen coronal sections were prepared with a thickness of 20 μm using a cryostat (HM550, Thermo Fisher Scientific). Immunostaining with anti-GFP antibody was performed, and the axonal length of cortical neurons was measured from the exit at the basement of the cerebral cortex to the tip of axons observed by fluorescence of EGFP with ImageJ plugin (Simple Neurite Tracer). The axonal length was normalized against the traverse diameter of the electroporated hemisphere on the brain section.
Analysis of ALS patient samples
All brain samples in this study were collected in the National Center of Neurology and Psychiatry (NCNP) Brain Bank and the Brain Bank for Aging Research (BBAR) in the Tokyo Metropolitan Geriatric Hospital and Institute of Gerontology (TMGHIG). This study was carried out according to the approval of the Ethics Committees of NCNP and TMGHIG. ALS cases used in this study (n = 7) were diagnosed based on the El Escorial and Airlie House revised criteria, and have been confirmed to have the pathological change in accordance with TDP-43 proteinopathy, i.e., the mislocalization and deposition of TDP-43 in the cytoplasm, in affected lesions. Control samples (n = 7) were selected from cases that had no neurological pathologies. There was no significant difference in age between these groups (ALS group 73.9 ± 8.7, Control group 74.1 ± 8.6). The background of each case is summarized in Supplementary Table 2 (online resource).
A portion of pyramidal tract was excised from the medulla oblongata of the samples by a blade, lysed in 1 mL of TRIZOL and total RNA was extracted according to the manufacturer’s protocol. Reverse transcriptase reaction and quantitative PCR were performed as described above. Primer sets used for amplification of each gene product are shown in Supplementary Table 1 (online resource). RNA integration number (RIN) of all samples was determined to be above 6.0 by Bioanalyzer.
Statistical analyses
Results are shown as mean ± SE. Differences between groups were examined for statistical significance using Prism 8 by the method indicated in figure legends. P value lower than 0.05 was determined to be significant.
Results
Identification of cytoplasmic ribosomal protein mRNAs as targets for axonal transport by TDP-43
To survey target mRNA candidates transported into neuronal axons by TDP-43, we first developed a method to obtain neurites separate from cell bodies of cultured cortical neurons. We defined the target mRNA candidates as the ones decreased in the neurites by RNAi-mediated down-regulation of TDP-43 expression in neurons. We seeded mouse cortical neurons on porous chamber inserts in culture dishes. Neurites having traversed the pores underneath to elongate processes were harvested from the bottom surface of the inserts. The tissue obtained from the bottom side did not include mRNA or protein derived from the nucleus (Fig. 1a), confirming that we can obtain pure neurites by this method. We also examined the presence of tau and MAP2A/2B, markers for axons and dendrites, respectively, in this fraction, and found that tau was easily detected, but the amount of MAP2A/2B was trivial (Fig. 1a), indicating that the obtained neurites exhibited characteristics of axons.
Identification of RP mRNAs as targets for axonal transport by TDP-43. a Confirmation of cell body–neurite separation by the porous chamber insert in the culture dish by reverse transcriptase (RT) PCR and immunoblot analysis. Lysates were prepared from the tissue collected from the top or bottom of the insert, and analyzed by RT-PCR and immunoblot to detect indicated molecules using 2 ng total RNA and 30 μg proteins of each sample, respectively. Lysates from the top of the inserts were cell bodies, and the bottom were neurites. Representative images of the detection are shown here, respectively. b The down-regulated expression of TDP-43 in the cell bodies of cortical neurons by two independent shRNAs used in this study was confirmed by immunoblot analysis. After collection of neurites for microarray analysis, the neuronal cell bodies in 3 culture dish wells for each condition were analyzed here as individual samples. c The number of mRNA transcripts dysregulated in neurites by TDP-43 down-regulation using the two independent shRNAs. d The expression levels of indicated genes in neurites (black bars) and cell bodies (white bars) of cortical neurons with shRNA-mediated TDP-43 down-regulation relative to the control shRNA-transduced neurons determined by quantitative RT-PCR (n = 4 in each group from 4 independent experiments). *P < 0.005, **P < 0.001 and ***P < 1.0 × 10–4 compared with the value of respective mRNA in the control shRNA-transduced neurons by multiple t-test. e Body and brain weights of neuron-specific TARDBP deficient mice at 21 days of age (n = 4–6 in each genotype). *P < 0.05 and **P < 0.001 compared with Flox/Flox, Eno2-Cre(–) and Flox/ + , Eno2-Cre Tg mice by one-way ANOVA test. Note that both the brain and body weights are significantly less in TARDBP Flox/Flox, Eno2-Cre Tg mice than Flox/Flox, Eno2-Cre(–) mice and Flox/ + , Eno2-Cre Tg mice. f The expression levels of indicated genes in the cerebral white matter of neuron-specific TARDBP deficient mice. The expression levels determined as hybridized signal counts of amplified transcripts using nCounter are shown relative to the levels in TARDBP Flox/Flox, Eno2-Cre(–) mice (n = 3 for each genotype). *P < 0.05 compared with TARDBP Flox/ + , Eno2-Cre(–) mice, and #P < 0.05 compared with TARDBP Flox/Flox, Eno2-Cre Tg mice by one-way ANOVA test. Note that the expression level of each RP mRNA is reduced in TARDBP Flox/Flox, Eno2-Cre Tg mice
To obtain candidate mRNAs for TDP-43-dependent transport in axons, we seeded cortical neurons on the chamber insert, and infected them with either of the two shRNA lentiviral vectors targeting two different sequences for TDP-43 in separate experiments. Neurites were harvested to compare the amounts of mRNA with those from the control to search for mRNAs reduced by TDP-43 down-regulation using microarray analysis. The suppressed expression of TDP-43 protein was confirmed by these targeted shRNAs (Fig. 1b). For the purpose of identifying target for TDP-43-dependent axonal transport, we focused our analysis on mRNAs down-regulated in axons by TDP-43 down-regulation. Among approximately 1000 mRNAs decreased in TDP-43 knock-down neurites by both shRNAs (Fig. 1c), we found that mRNA expression of cytoplasmic ribosomal proteins were significantly decreased by gene ontology analysis (Supplementary Table 3, online resource). Cytoplasmic ribosomes are constituted of ribosomal proteins and four ribosomal RNAs (28S, 18S, 5.8S and 5S in eukaryotes) [8]. We observed that expression of 46 among 80 total cytoplasmic ribosomal proteins as well as several eukaryotic elongation factors, both are translation-related genes, was reduced to less than half of the reference level by both TDP-43 shRNAs (Supplementary Table 4 and Supplementary Files 1 and 2, online resource). In contrast, only two of 80 mitochondrial ribosomal proteins, which have a different evolutional origin than cytoplasmic ones, were reduced by TDP-43 down-regulation, suggesting that the TDP-43 shRNA-dependent reduction is specific to the cytoplasmic type. Among approximately 2,400 mRNAs increased by TDP-43 knock-down using both shRNAs (Fig. 1c), glutathione peroxidase 4 was the most up-regulated one while no genetic group was indicated by gene ontology analysis (Supplementary File 3, online resource). In the following, we focus the analysis on the decreased mRNAs of cytoplasmic ribosomal proteins to see the direct link to ribosomal function and refer to the cytoplasmic ribosomal proteins simply as ribosomal proteins (RPs).
To validate the expression data obtained by microarray analysis, we examined the expression levels of the candidates in both cell body and neurite fractions by quantitative PCR. We confirmed that mRNAs of multiple RPs were commonly reduced in neurites, but not in cell bodies by TDP-43 down-regulation using shRNA#1, the more effective one of the two shRNAs (Fig. 1b, d). The amount of Bex2 mRNA, a control transcript that was not reduced in the microarray assay, was confirmed unchanged by TDP-43 knock-down in both cell body and neurite fractions (Fig. 1d). These results suggest that the reduction of RP mRNAs in neurites is due to decreased transport rather than transcriptional down-regulation.
In order to confirm whether the decrease in RP mRNA levels is observed in neuronal axons by TDP-43 down-regulation in vivo as well, we quantified the RP mRNAs of the cerebral white matter of mice deficient in the TDP-43 gene (TARDBP) in neurons. TARDBP was deleted specifically in neurons by generating TARDBP Flox mice expressing Cre recombinase under the enolase 2 (Eno2) promoter. As the amount of total RNA obtained by laser microdissection was limited, quantification was performed using the nCounter analysis system instead of quantitative PCR to detect target signals with fewer PCR cycles. TARDBP Flox/Flox, Eno2-Cre(–) mice and TARDBP Flox/+ , Eno2-Cre Tg mice exhibited no obvious phenotypic abnormalities, whereas TARDBP Flox/Flox, Eno2-Cre Tg mice had lower body and brain weights than the control (Fig. 1e), tremor and waddling gait immediately after birth (Movie 1, online resource), and a majority of the mice died within one month of age. Histological analysis of the cerebrum revealed that the number of cortical neurons was not altered (Supplementary Fig. 1a, b, online resource), implying that the phenotype was caused independent of neuronal death. The expression of TDP-43 mRNA was significantly reduced in TARDBP Flox/Flox, Eno2-Cre Tg mice than that in TARDBP Flox/Flox, Eno2-Cre(–) mice (Supplementary Fig. 1c, online resource), which indicates that the change of mRNA expression in neurites was well detected by this method. RP mRNAs were significantly decreased in TARDBP Flox/Flox, Eno2-Cre Tg mice, but not in TARDBP Flox/ + , Eno2-Cre Tg mice compared with TARDBP Flox/Flox, Eno2-Cre(–) mice (Fig. 1f). The expression levels of mRNAs that were not decreased by the microarray analysis (Gap43 and Gpx4) were maintained. These results from the animal model confirm that RP mRNAs in axons were reduced by down-regulation of TDP-43.
RP mRNAs are transported by TDP-43 in axons
The TDP-43-dependent axonal localization of RP mRNAs suggested that TDP-43 might be involved in axonal transport of RP mRNAs. To explore this possibility, we first examined the localization of TDP-43 protein and RP mRNAs in axons. Cortical neurons were seeded in microfluidic chambers, a device with grooves (150 μm of length) in the middle of the chamber, allowing only axons to extend to the opposite side (Supplementary Fig 3a, online resource), and in situ hybridization of RP mRNA along with immunostaining of TDP-43 were simultaneously performed on the axonal portion. We found that mRNAs of Rpl41, Rplp1, Rps7 and Rpl26, all of which were representative RP mRNAs decreased by RNAi-mediated TDP-43 down-regulation, were detected as a granular pattern along axons of the control neurons, which suggests that these mRNAs are located in axons as RNA granules (Fig. 2a and Supplementary Fig. 2, online resource). Importantly, we found that TDP-43 was co-localized with the mRNAs of Rpl41, Rplp1, Rps7 and Rpl26; and the co-localization ratios between them were significantly higher than that between TDP-43 and GAPDH mRNA (Supplementary Fig. 3b, online resource). The percentage of RP mRNAs co-localized with TDP-43 was around 80% in average, whereas that of GAPDH mRNA was 17.7% (Supplementary Fig. 3c, online resource). Moreover, both TDP-43 and RP mRNA signals were decreased significantly by RNAi-mediated TDP-43 down-regulation (Fig. 2a, Supplementary Fig. 2 and Supplementary Fig. 3d,e, online resource). These results suggest that TDP-43 is a component of the RNA granules and required for transporting RP mRNAs.
RP mRNAs are transported into axons as neuronal RNA granules by TDP-43. a Representative photomicrographs showing localization of Rpl41 mRNA (green) and TDP-43 protein (red) in axons of cortical neurons by in situ hybridization and immunostaining, respectively. Arrows indicate co-localized granule-like signals in control axons. Note that Rpl41 mRNA signal is decreased with TDP-43 down-regulation. Scale bar, 10 μm. b The levels of Rpl41 mRNA co-immunoprecipitated with TDP-43. TDP-43 and Rpl41 mRNA were expressed in Neuro2a cells and the amount of Rpl41 mRNA co-precipitated with TDP-43 was measured by quantitative PCR. The levels are shown as a value relative to the level of mRNA precipitated with control IgG (n = 3 for each group). *P < 0.0005 compared with the negative control, #P < 0.0001 compared with Rpl41 FL and Rpl41 Δ3′UTR by one-way ANOVA test. Note that TDP-43 binds to Rpl41 mRNA less efficiently without its 5′UTR. c–f Axonal transport of RP mRNAs in live cultured cortical neurons using MS2-MS2 binding RNA sequence system. RP mRNAs (with or without 5′UTR) bearing MS2 binding sequence are transfected with NLS-MS2-Venus. Representative photomicrographs of Venus fluorescent signal associated with Rpl41 mRNA are shown in (c) and (e); and the median transported distances of RP mRNA granules from the nucleus are shown in (d) and (f) (n = 30–36 for (d) and 35–43 for (f) in each group from 3 independent experiments). Scale bar, 20 μm in c and e. In e and f, the shRNA viral vector for TDP-43 down-regulation was transduced, followed by transfection with expression constructs for the indicated RP and NLS-MS2-Venus. *P < 0.0005 compared with the distance of the mock construct and #P < 0.05 compared with that of respective mRNA without 5′UTR by Welch’s ANOVA test in (d). #P < 0.05 and **P < 0.01 compared with the distance of respective mRNA in control shRNA-transduced neurons by unpaired t-test in (f). g Co-localization of Rpl41 mRNA with 5′UTR and TDP-43 in live neurons is demonstrated in cultured cortical neurons, by transfecting expression constructs for Rpl41 with 5′UTR bearing MS2 binding RNA sequence, NLS-MS2-Venus, and mCherry-TDP-43. Scale bar, 20 μm
To verify the direct involvement of TDP-43 in axonal transport of RP mRNAs, we examined whether TDP-43 associates with RP mRNA. For this purpose, both TDP-43 and RP mRNA were expressed in Neuro2a cells and the amounts of RP mRNAs co-precipitated with TDP-43 were measured by quantitative RT-PCR. Previous reports have indicated that nucleotide sequences called 5′ terminal oligopyrimidine tracts (5′TOP) are commonly found in the 5′ untranslated region (5′UTR) of RP and eukaryotic elongation factor mRNAs including those reduced by decreased expression of TDP-43 in the microarray analysis [17]. 5′TOP, together with a GC-rich region immediately downstream of it, is regarded important as a cis-factor for translational control of RP mRNAs [33]. Therefore, we also examined the contribution of 5′UTR in the mRNA in association with TDP-43 by RNA immunoprecipitation. We chose to examine Rpl41 and Rplp1 in subsequent experiments, since 5′TOP in the 5′UTR was well annotated in these mRNAs and the mRNAs were expected to show clear difference of binding and transport by TDP-43 by intervention in the 5′UTR. We found Rpl41 mRNA containing both 5′UTR and 3′UTR, but not the control mRNA in the TDP-43 immunoprecipitate (Fig. 2b and Supplementary Fig. 4, online resource). Furthermore, we found that Rpl41 mRNA lacking the 5′UTR was less observed in the TDP-43 immunoprecipitate, while that lacking the 3′UTR was indifferent. These results suggest that the 5′UTR of RP mRNAs plays a role in their binding with TDP-43.
To know whether the binding of TDP-43 with RP mRNAs via its 5′UTR is important for their transport within axons, we measured the extent of transport of RP mRNAs to axons using a method to visualize exogenous mRNA in living neurons. For this purpose, RP mRNA bearing the MS2 binding sequence at its 3′UTR was co-transfected with NLS-MS2-Venus plasmid in cultured cortical neurons [6]. We found that Rpl41 and Rplp1 containing 5′UTR were clearly well transported along axons—both in terms of the transport distance and the number of transported granules (Fig. 2c, d and Supplementary Fig. 5, online resource). The localization of the RP-MS2bs transcripts in this system was well correlated with that of RP mRNAs by in situ hybridization in axons (Supplementary Fig. 5a, online resource), indicating that the system can optimally detect the localization of target mRNAs. On the other hand, the same RP mRNAs deficient in 5′UTR exhibited significantly decreased transport distance and the number of granules, demonstrating the critical role of the 5′UTR for axonal transport of RP mRNAs (Fig. 2c, d and Supplementary Fig. 5b, online resource). Furthermore, we found that the axonal transport of RP mRNA containing 5′UTR was significantly reduced by RNAi-mediated TDP-43 down-regulation (Fig. 2e, f and Supplementary Fig. 5c, online resource). The transport of RP mRNAs deficient in 5′UTR was not inhibited by TDP-43 knock-down, demonstrating that their transport is independent of TDP-43 (Supplementary Fig. 5g, h, online resource). By co-expression of mCherry-TDP-43 at the same time in this visualization system, RP mRNA with a 5′UTR was confirmed to be more frequently co-localized with TDP-43 than inositol triphosphate receptor (IP3R) mRNA, which is known not to be a target of axonal transport by TDP-43 (Fig. 2g and Supplementary Fig. 5d, online resource). The RP mRNA and TDP-43 also showed identical movement in axons of live cells (Movie 2, online resource). These results indicate that the association of TDP-43 and RP mRNAs at their 5′UTR is required for their transport to axons.
Trans-factor La is associated with the RP mRNA-containing granules in axons
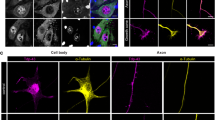
It is known that several RNA-binding proteins can bind to 5′TOP of RP mRNAs as trans-factors to regulate translation of the mRNA [11, 20, 24]. We therefore examined the possibility that some of these trans-factors may also be involved in the axonal transport of Rpl41 mRNA. For this purpose, we first examined the localization of three trans-factors, i.e., La, TIA-1 and AUF1 in neurons when each factor was co-expressed with Rpl41 mRNA. We found that while La, TIA-1 and AUF1 are all located mainly in the nucleus, La was also localized in axons, as previously reported [47] (Fig. 3a); and Rpl41 mRNA was mostly co-localized with La in axons (Fig. 3a, b and Supplementary Fig. 5e, f, online resource). In addition, we found that, when mCherry-TDP-43 was additionally co-expressed, mCherry-TDP-43 was co-localized with La and Rpl41 mRNA (Fig. 3b). To examine if these molecules are associated, we expressed TDP-43 and each trans-factor in Neuro2a cells to test if they could be co-immunoprecipitated from the cell lysate. TDP-43 was found in immunoprecipitate of La, as well as that of AUF1 (Fig. 3c). Endogenous RP mRNAs were also co-localized with endogenous La, as in the case of RP mRNAs and TDP-43 (Supplementary Fig. 6a, online resource). These results suggest that La is a good candidate molecule that mediates the axonal transport of RP mRNA via TDP-43. Although the detailed function of La on RP mRNAs has not been elucidated, La modestly increased RP translation in neurites (Supplementary Fig. 6b, c, online resource); but not the binding of RP mRNAs to TDP-43 nor transport of RP mRNAs to axons (Supplementary Fig. 6d, e, online resource), which may be a rationale of the protective effect of La described later.
La is associated with RP mRNA-containing granules in axons. a Representative photomicrographs of cultured cortical neurons transfected with constructs for Rpl41 with 5′UTR bearing MS2 binding sequence, NLS-MS2-Venus, and indicated FLAG-tagged trans-factor RNA-binding protein for 5′TOP. Note that the granular localization of FLAG-La overlaps well with Rpl41 mRNA. Scale bar, 20 μm. b Co-localization of Rpl41 mRNA with 5′UTR, TDP-43, and La is demonstrated in cultured cortical neurons using MS2-MS2 binding RNA sequence system, by transfecting expression constructs for Rpl41 with 5′UTR bearing MS2 binding RNA sequence, NLS-MS2-Venus, mCherry-TDP-43, and FLAG-La. Scale bar, 20 μm. c Association of TDP-43 with each trans-factor RNA-binding protein for 5′TOP is demonstrated by co-immunoprecipitation. EGFP-TDP-43-Myc was detected in immunoprecipitate of FLAG-La or FLAG-AUF1 using FLAG antibody (arrow), indicating that these components bind each other. IP immunoprecipitation, IB immunoblot
RPs in axons are translated locally to be assembled into ribosomes
Proteins are translated from mRNAs mostly in close proximity to the nucleus. In neurons, some mRNAs are known to be transported to axons and translated at the site to function as proteins. TDP-43 dependent transport of RP mRNAs to axons suggests that RP mRNAs are translated to proteins in axons. Translation of transported mRNA in axons of cultured neurons can be generally promoted by local stimulation with KCl or BDNF, which mimics neuronal activity [27, 55]. Therefore, to examine RP translation in axons, we cultured cortical neurons on a chamber insert or microfluidic chamber, and analyzed the change in RP translation after addition of BDNF to only the axonal side of the chamber. Here we chose to analyze Rpl26 and Rps6, because of the easy availability of well-functioning antibodies against them. We found by immunoblot analysis that BDNF stimulation increased the expression of RPs in neurites (Fig. 4a, b). The amounts of TDP-43, La, and RP mRNAs were not changed in neurites by BDNF stimulation (Fig. 4a, b and Supplementary Fig. 7a, b, d, online resource), which indicates that the increase of RPs was not because of enhanced transport of RP mRNAs to axons. We also found that BDNF enhanced the immunoreactivity of RPs in the axons plated in the microfluidic chamber (Fig. 4c, d). The increase in the protein content was counteracted by axonal administration of cycloheximide, a protein synthesis inhibitor (Fig. 4c, d). However, the administration of the reagent on the cell body side did not have this effect (Supplementary Fig. 7c, online resource). These results suggest that induction of RP expression by BDNF was due to the promotion of translation locally in axons.
RP mRNAs in axons are translated locally for assembly into ribosomes. a–d Expression of TDP-43, Rpl26 and Rps6 was assessed in response to BDNF administration locally to neurites. Representative immunoblots to detect endogenous TDP-43, Rpl26 and Rps6 in neurites of cortical neurons on chamber inserts are shown in a. The expression levels of each protein against that of GAPDH were quantified in neurites relative to the control levels in (b) (n = 3 in each group from 3 independent experiments). Expression of both RPs was significantly increased by local BDNF treatment, whereas that of TDP-43 was not altered. *P < 0.05 compared with the value in control neurites by unpaired t-test. Representative photomicrographs for immunocytochemical detection of Rpl26 in neurites of cortical neurons are shown in c. Note that Rpl26 signal is increased with BDNF administration, and that this increase is negated by cycloheximide (CHX). Scale bar, 10 μm. In d, immunocytochemical signal intensities of Rpl26 and Rps6 proteins were quantified in axons relative to the control levels (n = 27–31 in each group from 3 independent experiments). BDNF treatment in the axonal compartment significantly increased the amount of each RP. Cycloheximide (CHX), an inhibitor of protein synthesis by ribosomes, negated the effect of BDNF on both RPs. *P < 0.0005 and **P < 0.0001 compared with control and BDNF + CHX-treated axons by Welch’s ANOVA test. e, f Local translation of Kikume alone or Kikume with Rpl41 5′UTR of in growth cones was visualized and quantified after the photoconversion of green fluorescence of Kikume to red fluorescence, newly synthesized green fluorescence was captured by time-lapse imaging in parallel with red fluorescence every minute for 30 min. Representative photomicrographs of a growth cone at 0 and 30 min after the photoconversion are shown in (e), and quantified green fluorescence intensities against red fluorescence intensities at 3 indicated time points are shown in f (n = 11 in each group from 5 independent experiments). Scale bar, 10 μm. *P < 0.05 and **P < 0.005 compared with the signals of “Kikume without 5′UTR”-expressing growth cone at the same time point by Welch’s t-test. g, h Increased translation/assembly to ribosome of Rpl10a in response to KCl stimulation was visualized and quantified in cultured cortical neurons. Representative photomicrographs showing immunocytochemical localization of EGFP or EGFP-Rpl10a (green) and endogenous Rpl26 (red) in neurites are shown in (g). Scale bar, 10 μm. The co-localization tendency of EGFP and Rpl26 signals was estimated by Pearson’s R values in (g) (n = 6–12 in each group from 3 independent experiments). EGFP-Rpl10a with KCl treatment co-localized with Rpl26 significantly more than EGFP or EGFP-Rpl10a without KCl treatment. *P < 0.01 compared with EGFP and EGFP with KCl treatment, and #P < 0.001 compared with EGFP-Rpl10a without KCl treatment by one-way ANOVA test
As explained above, we demonstrated that the 5′TOP of RP mRNAs is required for their transport to axons. To examine if the 5′TOP of RP mRNAs plays a role in their translation in axons, we generated an expression construct, in which the 5′UTR of Rpl41 (including 5′TOP) is fused to the open reading frame of Kikume, a photoconvertible fluorescent protein. We expressed this construct in cultured neurons to analyze its translation in growth cones after photoconversion. We observed increased Kikume signal when it had the Rpl41 5′UTR; whereas, Kikume without the additional sequence was hardly detectable, suggesting that Rpl41 5′UTR promotes Kikume translation in growth cones. These results collectively suggest the importance of 5′TOP in the translational control of RP mRNA in axons (Fig. 4e, f).
Translated RPs from their mRNAs in axons need to be assembled into ribosomes locally to become functional components. To determine whether RP mRNAs in axons are translated and incorporated into functional ribosomes within axons, we employed Rpl10a fused to EGFP (EGFP-Rpl10a), which has previously been shown to be interchangeable with Rpl10a [55]. By transfecting EGFP-Rpl10a mRNA locally to axons in culture, we found that EGFP-Rpl10a and endogenous Rpl26 exhibited similar granular axonal distribution. Furthermore, co-localization of EGFP-Rpl10a with Rpl26 was significantly enhanced by KCl treatment. EGFP mRNA, on the other hand, caused a background level of EGFP co-localization with endogenous Rpl26 in axons, which was not altered even after the activation of translation by KCl (Fig. 4g, h). These results suggest that locally translated RPs can be assembled into axonal ribosomes to help maintain the integrity of the translation apparatus.
Reduction of TDP-43 disrupts translational activity of ribosomes in axons
We have shown that TDP-43 is involved in axonal transport of RP mRNAs, and that the transported RP mRNAs in axons are translated to be assembled into axonal ribosomes. To correlate TDP-43-dependent axonal transport of RP mRNAs and ribosomal function in axons, we investigated whether the decrease of RP mRNA transport by RNAi-mediated TDP-43 down-regulation affects ribosomal ability to synthesize proteins in axons. In accordance with the decrease of RP mRNA transport to axons by TDP-43 knock-down, the expression of RPs in neurites was reduced by RNAi-mediated down-regulation of TDP-43 (Supplementary Fig. 7e, f, online resource). To confirm that this observation is translation-dependent, a tRNA analogue puromycin was applied only to the neurite side of neurons seeded on the chamber insert to label newly synthesized proteins at the site; and subsequently, immunoblot analysis of the neuritic proteins was carried out to visualize puromycin-incorporated proteins representing proteins newly synthesized by ribosomes. We found that the puromycin-labeled proteins in neurites of control neurons appeared in a smear fashion; whereas in neurons with RNAi-mediated reduced TDP-43 expression, the puromycin-positive smear was significantly diminished, especially in the high molecular weight range (Fig. 5a, b). No band or smear was detected in unlabeled or cycloheximide-treated samples, which confirmed that this system detected only newly synthesized proteins. To visualize changes in puromycin incorporation in neurites from cultured neurons, we performed immunocytochemistry to detect the incorporated puromycin in neurons seeded in a microfluidic chamber. We found that the puromycin immunoreactivity was significantly lower in axons of TDP-43 down-regulated neurons than in control neurons, confirming the defective translational activity in axons by TDP-43 reduction (Fig. 5c, d). These results suggest that TDP-43 is involved in controlling the ribosomal function in axons.
Reduction of TDP-43 expression disrupts translational activity of ribosomes in axons. a–d Expression analysis of puromycin-incorporated proteins in neurites of cortical neurons with or without TDP-43 shRNA transduction. Representative images of immunoblots are shown in (a) and quantified intensities of puromycin immunoreactivity against tau immunoreactivity normalized to the control level are shown in (b) (n = 3 in each group from 3 independent experiments). *P < 0.05 and **P < 0.0001 compared with control shRNA-transduced neurons by one-way ANOVA test. Note that the band density of proteins, especially in high molecular weight ranges, weakened with shRNA-mediated down-regulation of TDP-43 expression. Puromycin-incorporated proteins were not detected in CHX-treated neurites. Representative photomicrographs of puromycin-incorporated proteins in neurites of cultured cortical neurons (stained in green; counterstained with βIII-tubulin in red) are shown in (c), and quantified levels of puromycin immunoreactivity against βIII-tubulin immunoreactivity normalized to the control level are shown in (d) (n = 27 in each group from 3 independent experiments). Scale bar, 20 μm in (c). *P < 0.05 compared with signals of control shRNA-expressing neurites by Welch’s t-test in (d). e The amounts of ribosomal RNAs (5′ETS and 18S) in the neurite and cell body fractions of neurons measured by quantitative PCR (n = 3 in each group from 3 independent experiments). The ratio is the value from TDP-43 shRNA-transduced neurons against that from control shRNA-transduced ones. The ratio of the amount of 5′ETS or 18S against that of β-actin in control shRNA-transduced neurons was determined as 1. 18S RNA, a mature ribosomal RNA, was decreased whereas 5′ETS, a premature form of ribosomal RNA, was increased confined to the neurite fraction by TDP-43 knock-down. *P < 0.05 compared with control shRNA-transduced neurons by unpaired t-test
RPs also have an essential function to convert premature long ribosomal RNAs into mature ones [15]. To examine if the TDP-43-dependent mechanism is also involved in the ribosomal RNA maturation, we analyzed the function of RPs in neurites expressing shRNA for TDP-43. We found that the ratio of 18S RNA, one of the mature ribosomal RNAs, was decreased; whereas 5′ETS RNA, a premature form of ribosomal RNA, was increased significantly in the neurite fraction compared with that in control neurons (Fig. 5e). There was no change in the amounts of these RNAs in the cell body fraction, suggesting that the ribosomal RNA maturation was only disturbed locally in neurites, namely axons. These results suggest that the TDP-43-dependent mechanism is involved in ribosomal RNA maturation.
RP has protective effects against TDP-43 dysfunction in axons
Dysregulation of TDP-43 expression was previously reported to cause defects in axonal outgrowth of neurons in vitro [45] and in vivo [22]. Our current results suggest that these effects may be mediated by failure of TDP-43 to transport RP mRNAs axonally. To prove this possibility, we performed RNAi-mediated TDP-43 down-regulation in neurons in culture, as well as, in vivo to examine its effect on axonal extension. Looking at cultured cortical neurons, we found that axonal extension was inhibited by TDP-43 reduction. We also found that this reduction was mitigated by overexpression of Rpl41, Rpl26 or Rps7 including 5′UTR (Fig. 6a, b, supplementary Fig. 8, online resource). Among them, Rpl26 was the most effective in rescuing the extension defect of axons; whereas Rplp1, another RP, did not improve the defect, suggesting that there is a redundancy and specificity for the effects of each RP. On the other hand, each RP lacking 5′UTR had no protective effect (Fig. 6b). The failure of axonal extension by TDP-43 knock-down was reversed by overexpression of human TDP-43, which is not targeted by the shRNA against mouse TDP-43, suggesting that the failure of axonal outgrowth was not due to off-target effects of the shRNA. Importantly, we were also able to demonstrate that La is effective in ameliorating decreased axonal extension by suppression of TDP-43 expression. These results suggest that TDP-43-dependent axonal transport/translation of RP mRNAs is relevant for axonal extension.
RP has a protective effect against TDP-43 dysfunction in axons. a, b Neurite outgrowth of cortical neurons transfected with TDP-43 shRNA together with indicated RP, hTDP-43 or La was assessed in culture. Representative photomicrographs of neurons expressing control or TDP-43 shRNA with or without Rpl26 including 5′UTR are shown in (a), and the lengths of the longest neurite in each neuron expressing indicated molecules were quantified relative to control shRNA + pcDNA neurons in (b) (n = 71–100 in each group from 3 independent experiments). Scale bar, 50 μm in (a). *P < 0.0001 compared with control shRNA + pcDNA-transfected neurons, #P < 0.005 and ##P < 0.0001 compared with TDP-43 shRNA + pcDNA-transfected neurons by one-way ANOVA test. c, d Axonal outgrowth of cortical neurons with or without TDP-43 shRNA and Rpl26 expression was assessed in vivo. TDP-43 shRNA and Rpl26 with 5′UTR expression constructs were transfected to cortical neurons of mouse embryos with in utero electroporation. Axonal extension from cortical neurons was traced by simultaneous transfection of an EGFP expression construct. Coronal sections at the corpus callosum level were immunostained with anti-GFP antibody. Representative photomicrographs of sections expressing indicated constructs are shown in (c), and normalized axonal length of neurons expressing indicated constructs quantified from the transfection site is shown in (d) (n = 3–11 in each group from 5 independent experiments). Scale bar, 500 μm in (c). *P < 0.01 compared with control shRNA + pcDNA-transfected brains, #P < 0.01 compared with TDP-43 shRNA + pcDNA-transfected brains by one-way ANOVA test in (d)
To examine the effects of TDP-43 down-regulation on axonal extension in vivo, we introduced shRNA for TDP-43 in cortical neurons of mouse embryos using in utero electroporation. We found that the length of axonal projection to the corpus callosum in EGFP-labeled cerebral cortical neurons was reduced by TDP-43 shRNA expression (Fig. 6c, d). We were also able to show that co-transfection of Rpl26 gene, which was the most effective against the defect of axonal extension by TDP-43 knock-down in vitro, was able to rescue the TDP-43 shRNA-induced defect (Fig. 6c, d). For neurons transfected with control shRNA, Rpl26 did not bring an apparent change in the axonal projection, suggesting that the protective effects of Rpl26 were specific to TDP-43 dysfunction. These results demonstrate that TDP-43-mediated RP mRNA transport/translation in axons promotes developmental axonal extension.
Finally, to investigate whether the impairment of RP mRNA transport/translation by TDP-43 down-regulation is relevant in the pathological mechanism of ALS/FTLD, we examined RP mRNA levels in the white matter of human samples. For this purpose, we determined the expression levels of RP mRNAs in the pyramidal tract of the medulla oblongata, where axons of motor neurons are located, in human samples with sporadic ALS in which the pathological change of TDP-43 had been confirmed. Among four RP mRNAs examined, the ALS group exhibited a significant decrease in Rplp1 and Rps6 mRNAs compared with the control group (Fig. 7). Rpl41 and Rpl26 mRNAs tended to decline in the ALS group, but they were not significantly different (P = 0.093 and 0.064, respectively). These results suggest that TDP-43 mislocalization associated with ALS/FTLD may also cause the impairment of axonal transport of RP mRNAs.
RP mRNAs are decreased in the pyramidal tract of ALS cases. The expression levels of indicated RP mRNAs in the pyramidal tract of medulla oblongata from control and sporadic ALS cases (n = 7 in each group). The expression level of each mRNA against that of β-actin mRNA was normalized relative to control brains. The plots and lines in boxes indicate individual values and medians of each RP mRNA, respectively, and the ends of whiskers and boxes mean maximum/minimum and upper/lower quartile of the values after exclusion of outliers in each group, respectively. All mRNAs examined had a trend to decrease in ALS cases, a part of which reduced significantly compared with those in control samples. *P < 0.005 and **P < 0.05 compared with control samples by unpaired t-test
Discussion
Many reports have indicated the collection of target genes for expression or splicing by TDP-43. However, the detailed mechanism for neurodegeneration by specific genes or transcripts among the large number of TDP-43 targets has not yet been determined [28, 35, 43]. There have also been some reports investigating the transport function of mRNA into neurites by TDP-43. TDP-43 transports mRNAs of neurofilament-L, PSD-95 and CaMKIIα into axons, which are disturbed in mutant TDP-43 related to ALS [2, 21]. The amount of MAP1B mRNA is decreased in axons of Drosophila by TDP-43 aggregate formation in the cytoplasm of neurons [10]. TDP-43 also regulates translation of several mRNAs including Rac1 mRNA in concert with FMRP in dendrites of neurons [9, 31]. However, it is undetermined whether these phenomena are linked with the neurodegeneration seen in ALS and FTLD.
Here we found a subset of RP mRNAs as target mRNAs for axonal transport by TDP-43, and that the transported mRNAs serve to maintain ribosomal function in axons. Localization of ribosomes in neurites including axons and dendrites has previously been reported [46]. RP mRNAs have also been shown to be present in axons and growth cones, especially of immature neurons, suggesting that these mRNAs have a specific transport mechanism and a critical function at the local site during morphogenesis [56]. Here we demonstrated that RP mRNAs were decreased in axons of motor neurons from ALS patients that have the TDP-43 pathology. In mature neurons, although demand for protein synthesis in axons may not be as high as in immature neurons [39], local protein synthesis in axons is still crucial for maintaining neuronal morphological integrity in response to stress or forming new synapses during learning and memory. It is well presumed that degeneration progresses chronically according to the gradual deterioration of these functions in adult neurons that develop TDP-43 pathology in ALS and FTLD. Suppression of TDP-43 expression also impaired the processing of immature ribosomal RNA confined to neurites. Since RPs are known to regulate ribosomal RNA processing, it may reflect the functional defect of RPs derived from the deficit of RP mRNA transport by TDP-43 in axons [15].
TDP-43, especially ALS/FTLD-related mutant forms, is shown to be mis-localized and forms aggregates in axons of chick motor neurons over time, causing a reduction of the axonal outgrowth. This is conceivable because the aggregated TDP-43 in the cytoplasm or even in axons may deplete or disturb the functional, movable TDP-43 in axons, which is the same situation as TDP-43 knockdown at the site [45]. It is also possible that deposited TDP-43 itself is involved in neurotoxicity of the diseases by occupying the physical space in neuronal compartments or causing the exhaust of protein degrading systems, etc. Although the role of gain of function by TDP-43 on the pathogenesis of ALS and FTLD could not be examined in our study, it is also important to elucidate the significance.
We have revealed that the transported RP mRNAs are translated and incorporated into ribosomes in axons to promote axonal outgrowth. Interestingly, we found that axonal outgrowth inhibition by TDP-43 knock-down was counteracted by the expression of a single RP, with variable effects depending on which RP was chosen. We speculate that this becomes possible because ribosomal function in axons may be maintained by replacing specific deteriorated RPs with newly synthesized ones at the local site. It was reported that ribosomes bearing damaged RPs are repaired to extend their lifespan by exchanging proteins with a free cytoplasmic RP pool in mice [32], which supports this speculation. The varied effects by different RPs may reflect how easy or difficult it is to replace each RP locally. For instance, eukaryotic Rpl26 is located at the subunit interface of a ribosome [48]; whereas Rplp1 exists internally in the stalk of the large subunit between Rplp0 and Rplp2 [36], which may cause differences in accessibility to ribosomes and result in differential effects on axonal outgrowth by expressing Rpl26 and Rplp1. Recently, a subset of RP mRNAs has been shown to be translated locally and assembled into ribosomes in axons, which is consistent with our observations [38].
While we chose to examine genes down-regulated in neurites by TDP-43 knock-down, we found more than 2400 genes up-regulated (Fig. 1c). We found that the up-regulated genes were not classified to any particular group by gene ontology analysis. Glutathione peroxidase 4, the most highly up-regulated gene, may be related to the defense mechanism against oxidative stress caused by TDP-43 knock-down. Genes encoding transcription factors were also found among the up-regulated genes, which may reflect possible transcriptional changes caused by TDP-43 dysregulation, but the detailed mechanisms are yet to be determined. Among the genes down-regulated in neurites by TDP-43 knock-down, there were some genes which were unrelated to protein translation, and none of them formed any particular group by gene ontology analysis either. One of such genes was tubulin, whose subtype was previously shown to be mutated in familial ALS cases [40]. It is interesting that TDP-43 may be involved in regulating localization of tubulin.
We found mRNAs encoding ribosomal proteins as the only group of genes whose expressions are significantly altered by TDP-43 down-regulation. There were also a number of other genes dysregulated in axons by TDP-43 down-regulation, but those were not identified as constituents of functional group of genes by our gene ontology analysis. This was unexpected, and the reasons for this are unclear. It is possible that re-analysis of the dataset by different algorithms may result in identification of other gene groups related to particular signaling pathways. Future re-analysis may also be important, since previously unidentified gene functions are constantly added. Another possibility may be that TDP-43 has more specialized functions in the axon than in the nucleus or cell body, which may be relevant to the neuron-specific phenotype with TDP-43 proteinopathy. Future analysis may be required to fully elucidate the mechanism of TDP-43-dependent regulation of axonal transport of mRNAs.
In conclusion, we identified RP mRNAs as target mRNAs transported to axons by TDP-43. TDP-43 plays a role not only in the axonal transport of RP mRNAs, but also in maintaining functionality of axonal ribosomes, which is required for local protein synthesis in response to stimulation and stress to axons. Furthermore, impairment of the TDP-43 function disturbs axonal maintenance in neurons, which is reminiscent of the pathology in ALS and FTLD. Our results elucidate a novel mechanism of maintaining morphological integrity of neuronal processes, and prove the influence of its transport mechanism for neurodegeneration in ALS and FTLD associated with TDP-43 proteinopathy. Our findings may also have implications for developing new treatment strategies against these incurable diseases by promoting the expression or transport of RP mRNAs in axons.
References
Akashi K, Kakizaki T, Kamiya H, Fukaya M, Yamasaki M, Abe M et al (2009) NMDA receptor GluN2B (GluR epsilon 2/NR2B) subunit is crucial for channel function, postsynaptic macromolecular organization, and actin cytoskeleton at hippocampal CA3 synapses. J Neurosci 29:10869–10882
Alami NH, Smith RB, Carrasco MA, Williams LA, Winborn CS, Han SS et al (2014) Axonal transport of TDP-43 mRNA granules is impaired by ALS-causing mutations. Neuron 81:536–543
Arai T, Hasegawa M, Akiyama H, Ikeda K, Nonaka T, Mori H et al (2006) TDP-43 is a component of ubiquitin-positive tau-negative inclusions in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Biochem Biophys Res Commun 351:602–611
Araki T, Milbrandt J (2003) ZNRF proteins constitute a family of presynaptic E3 ubiquitin ligases. J Neurosci 23:9385–9394
Araki T, Sasaki Y (2004) Milbrandt, J (2004) Increased nuclear NAD biosynthesis and Sir2 activation prevent axonal degeneration. Science 305:1010–1013
Bannai H, Fukatsu K, Mizutani A, Natsume T, Iemura S, Ikegami T et al (2004) An RNA-interacting protein, SYNCRIP (heterogeneous nuclear ribonuclear protein Q1/NSAP1) is a component of mRNA granule transported with inositol 1,4,5-trisphosphate receptor type 1 mRNA in neuronal dendrites. J Biol Chem 279:53427–53434
Bertrand E, Chartrand P, Schaefer M, Shenoy SM, Singer RH, Long RM (1998) Localization of ASH1 mRNA particles in living yeast. Mol Cell 2:437–445
Caldarola S, De Stefano MC, Amaldi F, Loreni F (2009) Synthesis and function of ribosomal proteins–fading models and new perspectives. Febs J 276:3199–3210
Chu J, Majumder P, Chatterjee B, Huang S, Shen CJ (2019) TDP-43 regulates coupled dendritic mRNA transport-translation processes in co-operation with FMRP and staufen1. Cell Rep 29:3118–3133
Coyne AN, Siddegowda BB, Estes PS, Johannesmeyer J, Kovalik T, Daniel SG et al (2014) Futsch/MAP1B mRNA is a translational target of TDP-43 and is neuroprotective in a Drosophila model of amyotrophic lateral sclerosis. J Neurosci 34:15962–15974
Damgaard CK, Lykke-Andersen J (2011) Translational coregulation of 5'TOP mRNAs by TIA-1 and TIAR. Genes Dev 25:2057–2068
Dunn KW, Kamocka MM, McDonald JH (2011) A practical guide to evaluating colocalization in biological microscopy. Am J Physiol Cell Physiol 300:C723–742
Fallini C, Bassell GJ, Rossoll W (2012) The ALS disease protein TDP-43 is actively transported in motor neuron axons and regulates axon outgrowth. Hum Mol Genet 21:3703–3718
Fallini C, Donlin-Asp PG, Rouanet JP, Bassell GJ, Rossoll W (2016) Deficiency of the survival of motor neuron protein impairs mRNA localization and local translation in the growth cone of motor neurons. J Neurosci 36:3811–3820
Gazda HT, Preti M, Sheen MR, O'Donohue MF, Vlachos A, Davies SM et al (2012) Frameshift mutation in p53 regulator RPL26 is associated with multiple physical abnormalities and a specific pre-ribosomal RNA processing defect in diamond-blackfan anemia. Hum Mutat 33:1037–1044
Habuchi S, Tsutsui H, Kochaniak AB, Miyawaki A, van Oijen AM (2008) mKikGR, a monomeric photoswitchable fluorescent protein. PLoS ONE 3:e3944
Hamilton TL, Stoneley M, Spriggs KA, Bushell M (2006) TOPs and their regulation. Biochem Soc Trans 34:12–16
Igaz LM, Kwong LK, Lee EB, Chen-Plotkin A, Swanson E, Unger T et al (2011) Dysregulation of the ALS-associated gene TDP-43 leads to neuronal death and degeneration in mice. J Clin Invest 121:726–738
Iguchi Y, Katsuno M, Niwa J, Takagi S, Ishigaki S, Ikenaka K et al (2013) Loss of TDP-43 causes age-dependent progressive motor neuron degeneration. Brain 136:1371–1382
Intine RV, Tenenbaum SA, Sakulich AL, Keene JD, Maraia RJ (2003) Differential phosphorylation and subcellular localization of La RNPs associated with precursor tRNAs and translation-related mRNAs. Mol Cell 12:1301–1307
Ishiguro A, Kimura N, Watanabe Y, Watanabe S, Ishihama A (2016) TDP-43 binds and transports G-quadruplex-containing mRNAs into neurites for local translation. Genes Cells 21:466–481
Kabashi E, Lin L, Tradewell ML, Dion PA, Bercier V, Bourgouin P et al (2010) Gain and loss of function of ALS-related mutations of TARDBP (TDP-43) cause motor deficits in vivo. Hum Mol Genet 19:671–683
Kabashi E, Valdmanis PN, Dion P, Spiegelman D, McConkey BJ, Vande Velde C et al (2008) TARDBP mutations in individuals with sporadic and familial amyotrophic lateral sclerosis. Nat Genet 40:572–574
Kakegawa T, Ohuchi N, Hayakawa A, Hirata S, Matsuda M, Kogure K et al (2007) Identification of AUF1 as a rapamycin-responsive binding protein to the 5'-terminal oligopyrimidine element of mRNAs. Arch Biochem Biophys 465:274–281
Kawahara Y, Mieda-Sato A (2012) TDP-43 promotes microRNA biogenesis as a component of the Drosha and Dicer complexes. Proc Natl Acad Sci U S A 109:3347–3352
Kelleher RJ 3rd, Govindarajan A, Tonegawa S (2004) Translational regulatory mechanisms in persistent forms of synaptic plasticity. Neuron 44:59–73
Krichevsky AM, Kosik KS (2001) Neuronal RNA granules: a link between RNA localization and stimulation-dependent translation. Neuron 32:683–696
Lagier-Tourenne C, Polymenidou M, Hutt KR, Vu AQ, Baughn M, Huelga SC et al (2012) Divergent roles of ALS-linked proteins FUS/TLS and TDP-43 intersect in processing long pre-mRNAs. Nat Neurosci 15:1488–1497
Ling SC, Polymenidou M, Cleveland DW (2013) Converging mechanisms in ALS and FTD: disrupted RNA and protein homeostasis. Neuron 79:416–438
Liu-Yesucevitz L, Bassell GJ, Gitler AD, Hart AC, Klann E, Richter JD et al (2011) Local RNA translation at the synapse and in disease. J Neurosci 31:16086–16093
Majumder P, Chu JF, Chatterjee B, Swamy KB, Shen CJ (2016) Co-regulation of mRNA translation by TDP-43 and Fragile X syndrome protein FMRP. Acta Neuropathol 132:721–738
Mathis AD, Naylor BC, Carson RH, Evans E, Harwell J, Knecht J et al (2017) Mechanisms of in vivo ribosome maintenance change in response to nutrient signals. Mol Cell Proteom 16:243–254
Meyuhas O (2000) Synthesis of the translational apparatus is regulated at the translational level. Eur J Biochem 267:6321–6330
Neumann M, Sampathu DM, Kwong LK, Truax AC, Micsenyi MC, Chou TT et al (2006) Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 314:130–133
Polymenidou M, Lagier-Tourenne C, Hutt KR, Huelga SC, Moran J, Liang TY et al (2011) Long pre-mRNA depletion and RNA missplicing contribute to neuronal vulnerability from loss of TDP-43. Nat Neurosci 14:459–468
Qiu D, Parada P, Marcos AG, Cardenas D, Remacha M, Ballesta JP (2006) Different roles of P1 and P2 Saccharomyces cerevisiae ribosomal stalk proteins revealed by cross-linking. Mol Microbiol 62:1191–1202
Ratti A, Buratti E (2016) Physiological functions and pathobiology of TDP-43 and FUS/TLS proteins. J Neurochem 138(Suppl 1):95–111
Shigeoka T, Koppers M, Wong HHW, Lin JQ, Cagnetta R, Dwivedy A et al (2019) On-site ribosome remodeling by locally synthesized ribosomal protein in axons. Cell Rep 29:3605–3619
Slomnicki LP, Pietrzak M, Vashishta A, Jones J, Lynch N, Elliot S et al (2016) Requirement of neuronal ribosome synthesis for growth and maintenance of the dendritic tree. J Biol Chem 291:5721–5739
Smith BN, Ticozzi N, Fallini C, Gkazi AS, Topp S, Kenna KP et al (2014) Exome-wide rare variant analysis identifies TUBA4A mutations associated with familial ALS. Neuron 84:324–331
Sreedharan J, Blair IP, Tripathi VB, Hu X, Vance C, Rogelj B et al (2008) TDP-43 mutations in familial and sporadic amyotrophic lateral sclerosis. Science 319:1668–1672
Swarup V, Phaneuf D, Bareil C, Robertson J, Rouleau GA, Kriz J et al (2011) Pathological hallmarks of amyotrophic lateral sclerosis/frontotemporal lobar degeneration in transgenic mice produced with TDP-43 genomic fragments. Brain 134:2610–2626
Tollervey JR, Curk T, Rogelj B, Briese M, Cereda M, Kayikci M et al (2011) Characterizing the RNA targets and position-dependent splicing regulation by TDP-43. Nat Neurosci 14:452–458
Tomono T, Hirai Y, Okada H, Adachi K, Ishii A, Shimada T et al (2016) Ultracentrifugation-free chromatography-mediated large-scale purification of recombinant adeno-associated virus serotype 1 (rAAV1). Mol Ther Methods Clin Dev 3:15058
Tripathi VB, Baskaran P, Shaw CE, Guthrie S (2014) Tar DNA-binding protein-43 (TDP-43) regulates axon growth in vitro and in vivo. Neurobiol Dis 65:25–34
Twiss JL, Fainzilber M (2009) Ribosomes in axons–scrounging from the neighbors? Trends Cell Biol 19:236–243
van Niekerk EA, Willis DE, Chang JH, Reumann K, Heise T, Twiss JL (2007) Sumoylation in axons triggers retrograde transport of the RNA-binding protein La. Proc Natl Acad Sci U S A 104:12913–12918
Villarreal J Jr, Lee JC (1998) Yeast ribosomal protein L26 is located at the ribosomal subunit interface as determined by chemical cross-linking. Biochimie 80:321–324
Wakatsuki S, Furuno A, Ohshima M, Araki T (2015) Oxidative stress-dependent phosphorylation activates ZNRF1 to induce neuronal/axonal degeneration. J Cell Biol 211:881–896
Wegorzewska I, Bell S, Cairns NJ, Miller TM, Baloh RH (2009) TDP-43 mutant transgenic mice develop features of ALS and frontotemporal lobar degeneration. Proc Natl Acad Sci U S A 106:18809–18814
Wils H, Kleinberger G, Janssens J, Pereson S, Joris G, Cuijt I et al (2010) TDP-43 transgenic mice develop spastic paralysis and neuronal inclusions characteristic of ALS and frontotemporal lobar degeneration. Proc Natl Acad Sci U S A 107:3858–3863
Wu LS, Cheng WC, Shen CK (2012) Targeted depletion of TDP-43 expression in the spinal cord motor neurons leads to the development of amyotrophic lateral sclerosis-like phenotypes in mice. J Biol Chem 287:27335–27344
Xu YF, Gendron TF, Zhang YJ, Lin WL, D'Alton S, Sheng H et al (2010) Wild-type human TDP-43 expression causes TDP-43 phosphorylation, mitochondrial aggregation, motor deficits, and early mortality in transgenic mice. J Neurosci 30:10851–10859
Yokoseki A, Shiga A, Tan CF, Tagawa A, Kaneko H, Koyama A et al (2008) TDP-43 mutation in familial amyotrophic lateral sclerosis. Ann Neurol 63:538–542
Yoon BC, Jung H, Dwivedy A, O'Hare CM, Zivraj KH, Holt CE (2012) Local translation of extranuclear lamin B promotes axon maintenance. Cell 148:752–764
Zivraj KH, Tung YC, Piper M, Gumy L, Fawcett JW, Yeo GS et al (2010) Subcellular profiling reveals distinct and developmentally regulated repertoire of growth cone mRNAs. J Neurosci 30:15464–15478
Acknowledgements
We thank Dr. Robert H. Singer (Albert Einstein College of Medicine) and Dr. Katsuhiko Mikoshiba (RIKEN) for providing NLS-MS2-Venus and IP3R 3′UTR-MS2bs plasmids, respectively.
Funding
This work was supported by the Grant-in-Aid for Scientific Research (C) (22590932, 25461302 and 16K09690 to S.N.), the Grant-in-Aid for Scientific Research on Innovative Areas (16H06277 to S.N.), AMED (JP20lm0203007 and JP20ek0109320 to S.N., JP18dm0107103 to Y.S.), grants from Japan Foundation for Neuroscience and Mental Health and Strategic Research Program for Brain Sciences (to S.N.), Intramural Research Grant for Neurological and Psychiatric Disorders of NCNP (24-9, 27-7, and 27-9 to S.N. and T.A.), Astellas Research Support, Pfizer Academic Contributions, and Takeda Research Support (T.A.).
Author information
Authors and Affiliations
Contributions
SN and TA designed the study. SN, JJ, RFA, YJ, MS, SW, SH, MN, KS, OO, HO, TO SW and HM performed experiments. YS, JTF and SM collected and evaluated human samples. SN and TA wrote the manuscript. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
Authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Movie 1. Walking of TARDBP Flox/Flox, Eno2-Cre(-), Flox/Flox, Eno2-Cre Tg and Flox/+, Eno2-Cre Tg mice at 10 days of age. TARDBP Flox/Flox, Eno2-Cre Tg mice exhibited tremors and a waddling gait. (MP4 11681 kb)
Supplementary file2 (MP4 15348 kb)
Supplementary file3 (MP4 334 kb)
Supplementary file4 (MP4 365 kb)
Movie 2. Movement of Rpl41 mRNA (green) and mCherry-TDP-43 (red) in the axon of live cortical neurons. Both signals in the same granule move simultaneously along the axon (MP4 184 kb)
Supplementary file6 (MP4 11196 kb)
401_2020_2205_MOESM7_ESM.tif
Supplementary Figure 1. The phenotype of neuron-specific TARDBP deficient mice is unrelated to neuronal death. a: Representative photomicrographs showing NeuN immunostaining of cerebral cortices in TARDBP Flox/Flox, Eno2-Cre(-), Flox/Flox, Eno2-Cre Tg and Flox/+, Eno2-Cre Tg mice at 21 days of age. Scale bar, 100 μm. b: The number of NeuN-immunostained neurons in mice from each genotype (n = 3 in each genotype). No significant difference was observed between the different genotypes by one-way ANOVA test. c: The amount of TDP-43 mRNA in the white matter of the mice from each genotype (n = 3 in each genotype). The expression level of TDP-43 mRNA normalized to that of β-actin mRNA was quantified at 21 days of age and values relative to that in TARDBP Flox/Flox, Eno2-Cre(-) mice are shown. The value in TARDBP Flox/Flox, Eno2-Cre Tg mice was significantly lower than that in TARDBP Flox/Flox, Eno2-Cre(-) mice. *P < 0.05 compared with the value in TARDBP Flox/Flox, Eno2-Cre(-) mice by unpaired t-test (TIF 845 kb)
401_2020_2205_MOESM8_ESM.tif
Supplementary Figure 2. Co-localization of RP mRNAs with TDP-43. Double staining of Rplp1 or Rps7 mRNA (green) and TDP-43 protein (red) in neuronal axons by in situ hybridization and immunostaining, respectively. Similar to Rpl41 mRNA, both mRNAs co-localized with TDP-43 in a granular pattern in control axons. Both signals were attenuated in TDP-43 down-regulated axons. Scale bar, 10 μm. (TIF 1399 kb)
401_2020_2205_MOESM9_ESM.tif
Supplementary Figure 3. RP mRNAs are transported into axons as neuronal RNA granules by TDP-43. a: Representative photomicrographs of differential interference contrast (DIC) and tau immunostaining images of neurons cultured in the microfluidic device. Neurites on the right side were regarded as axons and harvested from the area more than 450 μm apart from cell bodies on the left side for the analysis of the expression of RPs and TDP-43. b: Pearson’s R values of co-localization of each RP or GAPDH mRNA by in situ hybridization with antisense or sense probe with TDP-43 protein by immunostaining (n = 17-23 in each set from 3 independent experiments). Note that the value of the RP mRNA by antisense probe was significantly higher than the respective one by sense probe or that of GAPDH mRNA by antisense probe. *P < 1.0 x 10-8 compared with the respective value by sense probe by unpaired t-test, and #P < 0.0001 compared with the value of GAPDH by one-way ANOVA test. c: Percentage of TDP-43 immunoreactivity co-localized with indicated RP mRNA visualized by in situ hybridization in primary cultured mouse cortical neurons. d, e: Quantified fluorescent intensities of TDP-43 (d) and each RP mRNA (e) by immunostaining and in situ hybridization, respectively, in axons of control or TDP-43 shRNA-transduced neurons (n=20-22 in each group from 3 independent experiments). The value of each intensity was normalized to the value of respective protein/mRNA in control shRNA-transduced neurons. *P < 1.0 x 10-6 and **P < 1.0 x 10-8 compared with the value in control shRNA-transduced neurons by unpaired t-test. (TIF 937 kb)
401_2020_2205_MOESM10_ESM.tif
Supplementary Figure 4. The expression of Rpl41 and EGFP-TDP-43-Myc constructs in Neuro2a cells used in the experiment shown in Figure 2b. a: Representative immunoblot analysis showing expression of GFP and GAPDH in Neuro2a cells transfected with mock (control), IP3R (negative control) or Rpl41 full length or Δ5’/3’UTR together with EGFP-TDP-43-Myc. IB - immunoblot. b: The expression level of endogenous GAPDH mRNA and exogenous Rpl41 mRNA sequence in each group (n = 3 in each group from 3 independent experiments). The expression level of each mRNA was quantified and shown normalized to that of GAPDH mRNA relative to that in Neuro2a cells transfected with Rpl41 full length and EGFP-TDP-43-Myc. (TIF 321 kb)
401_2020_2205_MOESM11_ESM.tif
Supplementary Figure 5. Detection of RP mRNA localization by the MS2-MS2 binding RNA sequence system. a: Representative photomicrographs showing co-localization of RP mRNA signals detected by the Venus-labeled MS2-MS2 binding RNA sequence system and in situ hybridization in cell bodies and axons of cortical neurons. Scale bar, 20 μm. b: The number of RP mRNA granules along neurites measured on images by the MS2-MS2 binding RNA sequence system (n = 30-36 in each group from 3 independent experiments). The number of granules of Rpl41 and Rplp1 full length mRNA was significantly more than that of the mock construct or respective mRNA without 5’UTR. *P < 0.0005 compared with the number of the mock construct and #P < 0.05 compared with that of respective mRNA without 5’UTR by Welch’s ANOVA test. c: The number of RP mRNA granules along neurites by the MS2-MS2 binding RNA sequence system with control or TDP-43 shRNA-transduced neurons (n = 35-43 in each group from 3 independent experiments). The number of granules of Rpl41 and Rplp1 full length mRNA in TDP-43 shRNA-transduced neurons was significantly less than that of respective mRNA in control shRNA-transduced neurons. *P < 0.0001 and **P < 0.0005 compared with the number of respective mRNA in control shRNA-transduced neurons by unpaired t-test. d: Pearson’s R values of co-localization of MS2-Venus signals of Rpl41 full length or IP3R mRNA and mCherry-TDP-43 signals in neurites of cortical neurons (n = 23-25 in each group from 3 independent experiments). Co-localization of TDP-43 was higher with Rpl41 mRNA than with IP3R mRNA. *P < 5.0 x 10-8 compared with the value with IP3R mRNA by unpaired t-test. e: Pearson’s R values of co-localization of Rpl41 full length mRNA signals by the MS2-MS2 binding RNA sequence system and FLAG-La, FLAG-TIA-1 or FLAG-AUF1 signals by FLAG immunostaining in neurites of cortical neurons (n = 22-29 in each group from 3 independent experiments). Co-localization of Rpl41 full length mRNA was significantly the highest with FLAG-La among the groups. *P < 0.0001 compared with the value with FLAG-TIA-1 and FLAG-AUF1 by Welch’s ANOVA test. f: Representative photomicrographs showing localization of Rpl41 mRNA and La in the axons of cultured cortical neurons using MS2-MS2 binding RNA sequence system are shown. Cortical neurons were transfected with expression constructs for Rpl41 with 5’UTR bearing MS2 binding RNA sequence, NLS-MS2-Venus, and FLAG-La. Tau is used as an axonal marker. Scale bar, 20 μm. g, h: The median transported distances from the nucleus and the number of RP mRNA granules along neurites by the MS2-MS2 binding RNA sequence system with control or TDP-43 shRNA-transduced neurons (n = 15 in each group). Note that no significant difference of the median transported distance and the number of granules of Rpl41 and Rplp1 mRNA without 5’UTR was observed between the groups by unpaired t-test. (TIF 2033 kb)
401_2020_2205_MOESM12_ESM.tif
Supplementary Figure 6. La enhances expression of RPs. a: Representative photomicrographs showing co-localization of RP mRNAs by in situ hybridization and La by immunocytochemistry in axons of cortical neurons. Scale bar, 10 μm. b: Representative immunoblots showing expression of Rpl26 and Rps6 in the neurite fraction of cortical neurons overexpressing EGFP (control) or La on the chamber insert. c: La expression significantly increased the amount of Rps6, and caused a trend to increase the amount of Rpl26 (n = 3 in each group from 3 independent experiments). The expression level of each RP against that of GAPDH was normalized relative to control neurites. *P < 0.05 compared with the value in control neurites by unpaired t-test. d: The levels of Rpl41 mRNA co-immunoprecipitated with TDP-43. TDP-43 and Rpl41 were expressed together with La in Neuro2a cells and the amount of Rpl41 mRNA co-precipitated with TDP-43 was measured by quantitative PCR. The levels are shown as a value relative to the level of mRNA precipitated with control IgG (n = 3 for each group). *P < 0.001 compared with the negative control (IP3R) by one-way ANOVA test. There was no significant change of binding between Rpl41 mRNA and TDP-43 by La expression. e: The expression levels of indicated genes in neurites of cortical neurons transduced with rAAV1-CAG-FLAG-La relative to the control rAAV1-transduced neurons determined by quantitative RT-PCR (n = 3 in each group). No difference was seen compared with the value of respective mRNA in the control rAAV1-transduced neurons by multiple t-test. (TIF 824 kb)
401_2020_2205_MOESM13_ESM.tif
Supplementary Figure 7. Local translation of RP mRNAs in axons. a: Representative immunoblots of La in control and BDNF-stimulated neurites of cortical neurons on the chamber insert. b: The amount of La was not altered by stimulation with BDNF (n = 3 in each group from 3 independent experiments). The expression level of La was quantified and shown normalized to that of GAPDH relative to the level in control neurites. c: Translation of RPs is enhanced by BDNF in axons independently of that in cell bodies. Expression of Rpl26 detected by immunocytochemistry was quantified in neurites of mouse cortical neurons cultured in the microfluidic device after BDNF application only to the axon compartment, with or without CHX treatment of the cell bodies. Rpl26 expression level in each condition normalized to tau expression relative to the level in untreated axons (control + control) is shown (n = 30 in each group from 3 independent experiments). *P < 0.0001 compared with the value in control + control axons by Welch’s ANOVA test. Note that the expression of Rpl26 was increased significantly by stimulation with BDNF even when cell bodies were treated with CHX. d: The expression levels of indicated genes in neurites of cortical neurons treated with BDNF relative to the control neurons determined by quantitative RT-PCR (n = 3 in each group). No significant difference was observed compared with the value of respective mRNA in the control neurons by multiple t-test. e: Representative immunoblots of TDP-43 and RPs in control and TDP-43-knockdown neurites of cortical neurons on the chamber insert. f: The levels of Rpl26 and Rps6 in neurites were significantly decreased by TDP-43 knockdown (n = 3 in each group from 3 independent experiments). The expression level of each indicated protein normalized to that of GAPDH relative to the level in control neurites is shown. *P < 0.05 compared to the value in control neurites by unpaired t-test. (TIF 323 kb)
401_2020_2205_MOESM14_ESM.tif
Supplementary Figure 8. Expression of HA-tagged RPs in neurons. Representative photomicrographs of immunostaining of cell bodies and axons of cultured cortical neurons transfected with RP-HA including 5’UTR or pcDNA using indicated primary antibodies. Note that proteins translated from RP expression constructs including 5’UTR sequences are detectable in axons. Scale bar, 20 μm. (TIF 877 kb)
401_2020_2205_MOESM15_ESM.xlsx
Supplementary Files 1 and 2 Raw data with graphs of microarray analysis described in Figure 1. Data from experiments with TDP-43 shRNA#1 and #2 are shown in file 1 and 2, respectively. In each experiment, Cy5 and Cy3 labeling show control and shRNA-mediated knock-down condition, respectively. (XLSX 7263 kb)
401_2020_2205_MOESM18_ESM.docx
Supplementary File 3 Summary of all the annotated transcripts down-regulated or up-regulated by TDP-43 knock-down identified in the microarray analysis. Among the transcripts that were detected in controls and TDP-43-knock down conditions, the ones down-regulated to less than half or up-regulated to more than 2-fold by both TDP-43 shRNA constructs (#1 and #2) are listed. (DOCX 35 kb)
Rights and permissions
About this article
Cite this article
Nagano, S., Jinno, J., Abdelhamid, R.F. et al. TDP-43 transports ribosomal protein mRNA to regulate axonal local translation in neuronal axons. Acta Neuropathol 140, 695–713 (2020). https://doi.org/10.1007/s00401-020-02205-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-020-02205-y