Abstract
Epigenetic patterns on the level of DNA methylation have already been shown to separate clinically relevant subgroups of meningiomas. We here set out to identify potential prognostic implications of epigenetic modification on the level of histones with focus on H3K27 trimethylation (H3K27me3). H3K27me3 was assessed by immunohistochemistry on 232 meningiomas from 232 patients. In 194 cases, trimethylation was detected in tumor cells. In 25 cases, staining was limited to vessels while all tumor cells were negative. Finally, 13 cases yielded equivocal staining patterns. Reduced abundance of H3K27me3 in cases with staining limited to vessels was confirmed by mass spectrometry on a subset of cases. Lack of staining for H3K27me3 in all tumor cells was significantly associated with more rapid progression (p = 0.009). In line, H3K27me3-negative cases were associated with a DNA methylation pattern of the more aggressive types among the recently introduced DNA methylation groups. Also, NF2 and SUFU mutations were enriched among cases with complete lack of H3K27me3 staining in tumor cells (p < 0.0001 and p = 0.029, respectively). H3K27me3 staining pattern added significant prognostic insight into WHO grade II cases and in the compound subset of WHO grade I and II cases (p = 0.04 and p = 0.007, respectively). However, it did not further stratify within WHO grade III cases. Collectively, these data indicate that epigenetic modifications beyond DNA methylation are involved in the aggressiveness of meningioma. It also suggests that H3K27me3 immunohistochemistry might be a useful adjunct in meningioma diagnostics, particularly for cases with WHO grade II histology or at the borderline between WHO grade I and II.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Meningiomas are the most frequent primary intracranial and spinal tumors. They are classified into WHO grades I to III. Grading is based on histological features [18]. This histological system, as per se all classificatory approaches, is imperfect; particularly, WHO grade I meningiomas can recur and infiltrate locally and, conversely, higher grade meningiomas can have an indolent clinical course [10].
Despite its limitations, the WHO grading system is the best currently available algorithm for risk stratification of meningiomas. Thus, it defines clinical management, with clear guidelines for grades I and III, recommending observation and adjuvant radiation, respectively. Grade II meningiomas, however, comprise a histologically heterogeneous group of tumors with a behavior that is more challenging to predict, leaving treatment decisions to be determined by institutional multidisciplinary consensus rather than formalized guidelines [10]. Treatment determination hinges on an imprecise balance between likelihood of recurrence and potentially latent treatment-induced morbidity. Thus, further improvement of the classification is needed, potentially employing novel, more reliable biomarkers.
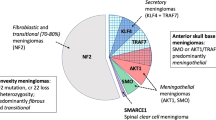
Besides the previously known mutations in NF2, genome wide and subsequent targeted sequencing studies of meningiomas have identified recurrent mutations in AKT1/TRAF7, KLF4/TRAF7, SMO, and PIK3CA which are strongly associated with histological features and location [2, 3, 5, 26, 33, 38]. Additionally, mutations in the TERT promoter are associated with higher risk of recurrence and shorter progression-free survival [11, 23, 29].
Epigenetic modifications may also indicate the risk of recurrence in meningiomas [5, 7, 8, 12, 16, 35]. Consequently, recent reports propose risk stratification schemes based on DNA methylation subgroups [21, 30].
Besides DNA methylation, a major epigenetic determinant of gene expression and cellular differentiation is the modification of histones, primarily by methylation and acetylation. Particularly modifications of lysine 27 (K27) of histone H3 play a crucial role in tumorigenesis [37]. Methylation of H3K27 is regulated by the EZH2 subunit of the PRC2 complex [4, 20, 36] and trimethylated H3K27 (H3K27me3) is associated with silencing of genes in the accompanying region [17]. Dysregulation of H3K27 methylation has been identified in several different cancers, including breast, prostate, colon, ovarian cancers, and malignant peripheral nerve sheath tumors [6, 15, 24, 31, 37, 39, 40]. As a result, assessment of H3K27 methylation status, particularly trimethylation (H3K27me3), has entered diagnostic practice as an immunohistochemical tool for several entities [1, 22].
We here tested for H3K27me3 staining patterns in meningioma to detect potentially clinically relevant subgroups identified by this marker.
Materials and methods
Case cohort
We assessed formalin-fixed paraffin-embedded tissue of 232 meningiomas of 232 patients. Since meningiomas at the interface of the common grade I and the less frequent grade II are most challenging to predict in terms of clinical course, the study was intentionally enriched for WHO grade II cases compared to epidemiological distribution (Table 1). Of note, all but two cases of WHO grade II were diagnosed based on mitotic count, the other two based on brain invasion. Of cases with recurrences, only material from the primary lesion was assessed. Based on information from available clinical records, no patient had prior radiotherapy or known neurofibromatosis type 2. Tumor size was estimated by measuring the largest diameter of contrast-enhancing tumor lesions in one plane on available imaging scans (CT or MRI). The cases and clinical data were provided by the Department of Neurology Zurich (Switzerland), the Department of Neuropathology Frankfurt (Germany), the Department of Pathology at the NYU Langone Medical Center (USA), and the Departments of Neurosurgery, Neurology and Neuropathology Heidelberg (Germany). The tissues from Frankfurt were assessed on tissue microarrays with two cores of each 2 mm diameter from each case. All other cases were analyzed as whole sections. Research use of tissue and clinical data were in accordance with local ethical regulations. Diagnoses were based on the WHO classification of brain tumors 2016. Cases initially diagnosed based on previous versions of the classification were reviewed (Zurich and Heidelberg samples in Heidelberg, other cases at the respective local institutions).
Immunohistochemistry and molecular analysis
Immunohistochemistry was performed on 4-µm-thick formalin-fixed, paraffin-embedded (FFPE) tissue sections. Tissues were pre-treated for 10 min at 121 °C in an autoclave at 210 kPa, subsequently further incubated with Ventana Cell Conditioner 1 immunostainer (Ventana Medical Systems, Tucson, AZ, USA) for 1 h. This pre-treatment was followed by incubation with rabbit monoclonal H3K27me3 antibody C36B11 (1:100, Cell Signaling, Danvers, MA, USA) on a Ventana BenchMark Ultra automated stainer for 2 h. Standard Ventana signal amplification was used including OptiView Amplifier Multimer and incubation with hematoxylin and bluing reagent for 4 min each. Intratumoral vessels served as positive controls. DNA methylation and panel sequencing data were obtained from previous analyses [30].
Statistics
Fisher’s exact test was used to compare categorical factors between H3K27 groups. Mann–Whitney test was used to compare quantitative parameters between H3K27 groups. Time to progression (TTP) was defined as time from initial surgery to first recurrence as determined by imaging. Patients without recurrence during follow-up were censored at last follow-up. Kaplan–Meier estimates and log-rank test were used to estimate and compare distribution of TTP between groups. Cox regression was used to assess the impact of factors on TTP. Interaction between WHO grade and H3K27 was tested in Cox regression to identify subgroup effects. Cox regression with Firth correction [13] was used in case of complete separation. For multivariable Cox regression model, multiple imputations of missing values with 100 imputations were performed using the chained equations (mice) algorithm [34]. Associations of staining patterns with mutations were analyzed with Fisher’s exact test. p values below 0.05 were considered statistically significant. Analysis was performed with statistical software R 3.4 (https://www.R-project.org/).
Mass spectrometry for histone modification
Nine frozen meningioma samples (from Zurich and Heidelberg) were available for mass spectrometry analysis. Histones were acid extracted, derivatized via propionylation, digested with trypsin, newly formed N-termini were propionylated as previously described [9] at ActiveMotif (Carlsbad, CA, USA), and then measured three separate times using the Thermo Scientific TSQ Quantum Ultra mass spectrometer coupled with an UltiMate 3000 Dionex nano-liquid chromatography system. The data were quantified using Skyline [19].
Results
Of 232 assessed cases, 194 showed positive staining for H3K27me3 in vessels and tumor cells, indicating trimethylation at H3K27 in the majority of meningiomas (Table 1). In 25 cases, however, H3K27me3 staining was limited to vessels while tumor cells were negative for this marker, pointing towards a loss of trimethylation (Fig. 1a, b). Cases with positive staining for H3K27me3 were subsequently tagged “retained” while cases without H3K27me3 in tumor cells were designated as “loss”. Of note, 13 cases had retained trimethylation with intermingled areas of negative staining. In these cases with ambiguous staining pattern, the vessels were also faintly stained or negative for H3K27me3 in the areas with negative tumor cells (Fig. 1c, d). Thus, this pattern of partial loss is more likely an artifact than due to a sub-clonal event. Consequently, these cases were grouped with the “retained” cases.
Cases with complete loss of trimethylation showed significantly less favorable outcome and more rapid progression (p = 0.009, Fig. 2a). This also held true when limiting the analysis to cases with clearly positive or negative H3K27me3 staining in tumor cells, excluding the cases with ambiguous pattern (Suppl. Fig. 1, p = 0.01). While this survival analysis was applied to the entire un-stratified cohort, for potential application in diagnostic routine, however, the added value on top of the current grading is more relevant. Interestingly, when further dissecting this overall discriminatory effect by the WHO grades, it was actually limited to WHO grade I and II (Fig. 2b, c). All WHO grade II cases with complete loss were diagnosed as atypical based on mitotic count. In contrast, histologically clearly anaplastic cases could not be further sub-divided for prognostic subgroups by H3K27me3 staining pattern (Suppl. Fig. 2) and showed in fact a significantly different prognostic impact of H3K27me3 than WHO I/II cases (interaction test p = 0.02).
Kaplan–Meier curves showing time-to-recurrence for all analyzed meningiomas (a) and restricted to cases of WHO grade I/II (b), stratified for H3K27me3 staining, with number of patients/events given in parenthesis. Hazard ratio for H3K27me (c): the first five lines (I, II, I/II, III, all) are based on univariable Cox regression models for H3K27me in the respective WHO grade subgroup. Line 6 (I/II adjusted) is based on the multivariable Cox regression model in the subgroup of WHO grade I/II patients (Table 3). Wald test p values are given
H3K27me3 staining, WHO grade, extent of resection (STR vs. GTR and Simpson grade) were all significantly associated with outcome in a univariable analysis of WHO grade I and II cases (Table 2). H3K27me3 staining pattern remained prognostically relevant when adjusting this subset for WHO grade and extent of resection in a multivariable analysis (Table 3).
Mutational status and DNA methylation subgroups
Mutational data were available for 98 cases. Among the most frequently mutated genes in meningioma, encompassing AKT1, KLF4/TRAF7, NF2, PIK3CA, SMO, SUFU and the TERT promoter, only mutations of NF2 and SUFU were significantly more frequent in cases without H3K27 trimethylation (p < 0.001 and p = 0.029, respectively, Suppl. Table 1). Other recurrent mutations, including aberrations of genes coding for histones that can also be associated with loss of trimethylation, were not detected.
Also, the DNA methylation status of 87 samples was analyzed in context of the H3K27me3 staining pattern. Case numbers with complete loss of trimethylation were too small to assess association with the previously introduced six individual DNA methylation subgroups [30]. Thus, an analysis for association with the two overarching DNA methylation groups “A” (comprising the three benign subgroups and subgroup intermediate A) and “B” (comprising subgroups intermediate B and malignant) was performed. Therein, complete loss of trimethylation was significantly associated with a DNA methylation pattern of the “group B”, comprising the subgroups “MC malignant” and “MC intermediate B” (p = 0.0046, Fisher’s exact test, Suppl. Table 2, Suppl. Fig. 3).
Mass spectrometry screen for histone modification
To assess the broader landscape of epigenetic regulation by histone modification, we performed a mass spectrometry-based screen for > 80 histone modifications. Availability of sufficient frozen tissue and cost per analysis restricted the case selection and sample size. The analysis included two samples with complete loss of H3K27me3, assigned to group B by DNA methylation analysis (subgroups MC malignant and MC intermediate B) and seven cases with retained trimethylation from group A (three each from MC benign-2 and benign-1 and one from MC intermediate A). Unsupervised clustering of these data yielded a pattern that exactly recapitulated the groups assigned by DNA methylation analysis (Fig. 3). Trimethylation of H3K27me (H3.1 and H3.3) was significantly lower in group B cases (p = 0.003 and p = 0.04, respectively).
Discussion
Associations between epigenetic modification and aggressiveness in meningioma have so far mostly been assessed on the level of DNA methylation. Thereby, DNA methylation analysis has evolved as promising candidate to add a molecular layer to upcoming WHO classifications, along with risk-related genetic aberrations like TERT promoter or BAP1 mutations [11, 14, 23, 29, 32].
However, the major alternative epigenetic modifier, histone methylation and acetylation, has not been further evaluated for prognostic potential. Yet, associations of H3K27-related regulation of expression and molecular subgroups of meningioma have already been reported. On the basis of chromatin immunoprecipitation sequencing (ChIP-Seq) for the regulatory mark H3K27Ac, the chief competing modification to trimethylation, differences between meningioma with AKT1 vs. NF2 mutations have been reported [5].
In contrast to ChIP-Seq and DNA methylation analysis, H3K27me3 immunohistochemistry is already implemented in many laboratories and can be readily applied. Its diagnostic potential has already been demonstrated for malignant peripheral nerve sheath tumors (MPNST) and ependymomas [22, 27]. In both entities, MPNST and posterior fossa ependymoma, the H3K27me3 staining pattern parallels a distinct DNA methylation. Also in our cohort, the H3K27me3 staining was associated with the previously introduced DNA methylation subgroups. In MPNST, the loss of trimethylation is mechanistically attributable to the perturbed PRC2 complex as a result of the EED/SUZ12 alterations [25]. For ependymomas and meningiomas, the functional background is not yet fully deciphered.
Although a merely descriptive finding, our data show that immunohistochemistry for H3K27me3 on meningioma samples can provide a useful tool in neuropathology practice. Complete loss of H3K27me3 staining predicts increased risk of recurrence in meningiomas, for the group of WHO grade I/II cases even independent of histological grade or extent of resection. While complete loss of trimethylation also occurred in WHO grade III cases, the staining pattern did not further stratify for risk-related subgroups among them. This might be due to other factors driving malignancy irrespective of H3K27 status. Also, an effect might be obscured by the study design that did not stratify for treatment. The majority of the high-grade cases might have received adjuvant therapy. However, only limited information on this was available for the present study which prevents definitive assessment of H3K27 trimethylation as a biomarker in WHO grade III meningiomas. Further, the fact that complete loss of trimethylation is associated with worse outcome within the entire cohort but not an obligatory prerequisite for high-grade meningiomas may also explain why a previous study could detect higher H3K27 trimethylation in a subset of WHO grade II cases compared to low-grade meningioma [12].
Of note, a challenge in application of H3K27me3 staining remains that different specificities have been reported for the various available antibodies [28]. More advanced proteomic methods including mass spectrometry will potentially elucidate which specific methylation status is detected at which specific sites and by which clones. Importantly, these studies may identify whether there is actually a functional background and relevance of these discrepant antibody specificities. By now, incorporation of this immunohistochemical biomarker as outlined here has the potential to predict which meningiomas are more likely to recur, helping to identify those patients that may benefit from adjuvant radiation or a more stringent clinical and radiological follow-up. Future larger and prospective studies stratifying patients based on H3K27me3 status are warranted to further validate its use in diagnostic routine and its correlation with mutations and DNA methylation.
References
Bender S, Tang Y, Lindroth AM et al (2013) Reduced H3K27me3 and DNA hypomethylation are major drivers of gene expression in K27M mutant pediatric high-grade gliomas. Cancer Cell 24:660–672
Bi WL, Greenwald NF, Abedalthagafi M et al (2017) Genomic landscape of high-grade meningiomas. NPJ Genom Med 2. https://doi.org/10.1038/s41525-017-0014-7
Brastianos PK, Horowitz PM, Santagata S et al (2013) Genomic sequencing of meningiomas identifies oncogenic SMO and AKT1 mutations. Nat Genet 45:285–289
Cao R, Wang L, Wang H et al (2002) Role of histone H3 lysine 27 methylation in Polycomb-group silencing. Science 298:1039–1043
Clark VE, Erson-Omay EZ, Serin A et al (2013) Genomic analysis of non-NF2 meningiomas reveals mutations in TRAF7, KLF4, AKT1, and SMO. Science 339:1077–1080
Cleven AH, Al Sannaa GA, Briaire-de Bruijn I et al (2016) Loss of H3K27 tri-methylation is a diagnostic marker for malignant peripheral nerve sheath tumors and an indicator for an inferior survival. Mod Pathol 29:582–590
Di Vinci A, Brigati C, Casciano I et al (2012) HOXA7, 9, and 10 are methylation targets associated with aggressive behavior in meningiomas. Transl Res 160:355–362
Gao F, Shi L, Russin J et al (2013) DNA methylation in the malignant transformation of meningiomas. PLoS ONE 8:e54114
Garcia BA, Mollah S, Ueberheide BM et al (2007) Chemical derivatization of histones for facilitated analysis by mass spectrometry. Nat Protoc 2:933–938
Goldbrunner R, Minniti G, Preusser M et al (2016) EANO guidelines for the diagnosis and treatment of meningiomas. Lancet Oncol 17:e383–e391
Goutagny S, Nault JC, Mallet M, Henin D, Rossi JZ, Kalamarides M (2014) High incidence of activating TERT promoter mutations in meningiomas undergoing malignant progression. Brain Pathol 24:184–189
Harmanci AS, Youngblood MW, Clark VE et al (2017) Integrated genomic analyses of de novo pathways underlying atypical meningiomas. Nat Commun 8:14433
Heinze G, Schemper M (2001) A solution to the problem of monotone likelihood in Cox regression. Biometrics 57:114–119
Juratli TA, Thiede C, Koerner MVA et al (2017) Intratumoral heterogeneity and TERT promoter mutations in progressive/higher-grade meningiomas. Oncotarget 8:109228–109237
Karczmarski J, Rubel T, Paziewska A et al (2014) Histone H3 lysine 27 acetylation is altered in colon cancer. Clin Proteom 11:24
Kishida Y, Natsume A, Kondo Y et al (2012) Epigenetic subclassification of meningiomas based on genome-wide DNA methylation analyses. Carcinogenesis 33:436–441
Kondo Y, Shen L, Cheng AS et al (2008) Gene silencing in cancer by histone H3 lysine 27 trimethylation independent of promoter DNA methylation. Nat Genet 40:741–750
Louis DN, Perry A, Reifenberger G et al (2016) The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131:803–820
MacLean B, Tomazela DM, Shulman N et al (2010) Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26:966–968
Nekrasov M, Klymenko T, Fraterman S et al (2007) Pcl-PRC2 is needed to generate high levels of H3–K27 trimethylation at Polycomb target genes. EMBO J 26:4078–4088
Olar A, Wani KM, Wilson CD et al (2017) Global epigenetic profiling identifies methylation subgroups associated with recurrence-free survival in meningioma. Acta Neuropathol 133:431–444
Panwalkar P, Clark J, Ramaswamy V et al (2017) Immunohistochemical analysis of H3K27me3 demonstrates global reduction in group-A childhood posterior fossa ependymoma and is a powerful predictor of outcome. Acta Neuropathol 134:705–714
Peyre M, Gauchotte G, Giry M et al (2017) De novo and secondary anaplastic meningiomas: a study of clinical and histomolecular prognostic factors. Neuro Oncol. https://doi.org/10.1093/neuonc/nox231
Puppe J, Drost R, Liu X et al (2009) BRCA1-deficient mammary tumor cells are dependent on EZH2 expression and sensitive to Polycomb Repressive Complex 2-inhibitor 3-deazaneplanocin A. Breast Cancer Res 11:R63
Reuss DE, Habel A, Hagenlocher C et al (2014) Neurofibromin specific antibody differentiates malignant peripheral nerve sheath tumors (MPNST) from other spindle cell neoplasms. Acta Neuropathol 127:565–572
Reuss DE, Piro RM, Jones DT et al (2013) Secretory meningiomas are defined by combined KLF4 K409Q and TRAF7 mutations. Acta Neuropathol 125:351–358
Rohrich M, Koelsche C, Schrimpf D et al (2016) Methylation-based classification of benign and malignant peripheral nerve sheath tumors. Acta Neuropathol 131:877–887
Rothbart SB, Dickson BM, Raab JR et al (2015) An interactive database for the assessment of histone antibody specificity. Mol Cell 59:502–511
Sahm F, Schrimpf D, Olar A et al (2016) TERT promoter mutations and risk of recurrence in meningioma. J Natl Cancer Inst 108(5):djv377. https://doi.org/10.1093/jnci/djv377
Sahm F, Schrimpf D, Stichel D et al (2017) DNA methylation-based classification and grading system for meningioma: a multicentre, retrospective analysis. Lancet Oncol 18:682–694
Schlesinger Y, Straussman R, Keshet I et al (2007) Polycomb-mediated methylation on Lys27 of histone H3 pre-marks genes for de novo methylation in cancer. Nat Genet 39:232–236
Shankar GM, Abedalthagafi M, Vaubel RA et al (2017) Germline and somatic BAP1 mutations in high-grade rhabdoid meningiomas. Neuro Oncol 19:535–545
Strickland MR, Gill CM, Nayyar N et al (2017) Targeted sequencing of SMO and AKT1 in anterior skull base meningiomas. J Neurosurg 127:438–444
Van Buuren S, Groothuis-Oudshoorn K (2011) mice: Multivariate imputation by chained equations in R. J Stat Softw 45(3):1–67
Vengoechea J, Sloan AE, Chen Y et al (2013) Methylation markers of malignant potential in meningiomas. J Neurosurg 119:899–906
Viré E, Brenner C, Deplus R et al (2006) The Polycomb group protein EZH2 directly controls DNA methylation. Nature 439:871–874
Wei Y, Xia W, Zhang Z et al (2008) Loss of trimethylation at lysine 27 of histone H3 is a predictor of poor outcome in breast, ovarian, and pancreatic cancers. Mol Carcinog 47:701–706
Yesiloz U, Kirches E, Hartmann C et al (2017) Frequent AKT1E17K mutations in skull base meningiomas are associated with mTOR and ERK1/2 activation and reduced time to tumor recurrence. Neuro Oncol 19:1088–1096
Yoo KH, Hennighausen L (2012) EZH2 methyltransferase and H3K27 methylation in breast cancer. Int J Biol Sci 8:59–65
Yu J, Yu J, Rhodes DR et al (2007) A polycomb repression signature in metastatic prostate cancer predicts cancer outcome. Can Res 67:10657–10663
Acknowledgements
The study was supported by grants of the German Cancer Aid (110983, 110670) and the “Else Kröner-Fresenius Stiftung” (2015_A60). We thank Laura Dörner, Antje Habel, Lisa Kreinbihl, and Hai Yen Nguyen for skillful technical assistance.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Katz, L.M., Hielscher, T., Liechty, B. et al. Loss of histone H3K27me3 identifies a subset of meningiomas with increased risk of recurrence. Acta Neuropathol 135, 955–963 (2018). https://doi.org/10.1007/s00401-018-1844-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-018-1844-9