Abstract
The use of an appropriate reference gene to ensure accurate normalisation is crucial for the correct quantification of gene expression using qPCR assays and RNA arrays. The main criterion for a gene to qualify as a reference gene is a stable expression across various cell types and experimental settings. Several reference genes are commonly in use but more and more evidence reveals variations in their expression due to the presence of on-going neuropathological disease processes, raising doubts concerning their use. We conducted an analysis of genome-wide changes of gene expression in the human central nervous system (CNS) covering several neurological disorders and regions, including the spinal cord, and were able to identify a number of novel stable reference genes. We tested the stability of expression of eight novel (ATP5E, AARS, GAPVD1, CSNK2B, XPNPEP1, OSBP, NAT5 and DCTN2) and four more commonly used (BECN1, GAPDH, QARS and TUBB) reference genes in a smaller cohort using RT-qPCR. The most stable genes out of the 12 reference genes were tested as normaliser to validate increased levels of a target gene in CNS disease. We found that in human post-mortem tissue the novel reference genes, XPNPEP1 and AARS, were efficient in replicating microarray target gene expression levels and that XPNPEP1 was more efficient as a normaliser than BECN1, which has been shown to change in expression as a consequence of neuronal cell loss. We provide herein one more suitable novel reference gene, XPNPEP1, with no current neuroinflammatory or neurodegenerative associations that can be used for gene quantitative gene expression studies with human CNS post-mortem tissue and also suggest a list of potential other candidates. These data also emphasise the importance of organ/tissue-specific stably expressed genes as reference genes for RNA studies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Reference or so-called “housekeeping” genes are widely used as internal standards in mRNA-based techniques such as reverse transcription quantitative real-time polymerase chain reaction (RT-qPCR) [9, 31] and microarray technologies [35]. Current guidelines recommend the use of reference genes over housekeeping genes to refer to internal controls [10]. A reference gene is one that is constitutively expressed in all nucleated cell types, is involved in basal cell metabolism or survival and is defined as a gene showing the most stable expression across various tissue types and experimental conditions. Quantitative mRNA studies are commonly conducted to compare expression levels of a target gene or gene of interest (GOI) between disease and non-disease and such data need to be normalised to a reference gene.
It is essential when conducting such studies to ensure that all the samples contain the same amount of RNA. Biological parameters such as tissue volume, mass, size or cell number were first employed as normalisation procedures (absolute quantification). However, variability in biological samples rendered this method impractical and unreliable [8]. When comparing the amount of RNA, it was found that normalisation with an internal reference gene was more appropriate (relative or comparative quantification), since it compensates for sample-to-sample variations in total RNA amount. Relative quantification compares changes of mRNA levels between two genes with one of them being the endogenous reference (the normaliser) and will determine the expression of a GOI relative to the reference gene. The most commonly used reference genes are GAPDH (glyceraldehyde-3-phosphate dehydrogenase), HPRT1 (hypoxanthine phosphoribosyltransferase 1), albumin, actins, tubulins, cyclophilin, microglobulins, beclin and ribosomal subunits [48, 49]. Although well established, data normalisation using an internal reference gene is subject to frequent criticism since the quality of normalised expression levels is entirely dependent on the quality of the normaliser [26]. Any variations in expression of the normaliser will result in obscuring real changes and will produce artifactual changes [7]. Hence, to carry out appropriate normalisation in RT-qPCR, suitable reference genes are vital to the generation of biologically relevant results [3, 17].
There are increasing concerns about some of the more conventional reference genes when investigating mRNA expression levels in the human brain [26]. For example, GAPDH is mostly known for its role in glycolysis, but recent reports have demonstrated increased GAPDH activity associated with cell fate and neurodegeneration [14, 43]. Furthermore, increased beclin expression has been associated with neurones and astrocytes at the site of traumatic brain injury [12]. A similar scenario is emerging for actin [40]. Some concerns have also been raised when normalising against 18S ribosomal RNA, which is much more abundant than other mRNA species and, therefore, may not be the best to compare with mRNA species of lower abundance [9, 44]. Moreover, we have demonstrated that not all mRNAs in the CNS, 18S ribosomal RNA being one of them, appear to deteriorate in a similar manner with increasing post-mortem delay based on RIN assessment [19]. These studies would suggest that the identification of a universal human reference gene might prove more challenging than initially thought and that, perhaps, novel organ and tissue-specific reference genes might be more appropriate and useful. Indeed, the literature shows that an ideal and universal reference gene remains to be discovered [6, 7, 42, 48]. There are very few investigative reports to date that have been published trying to establish and validate reference genes for quantitative gene expression analysis of human CNS post-mortem tissue [15, 27].
Several studies have been conducted to isolate stably expressed genes using microarray data. Large scale categorisation of reference genes based on microarray data has generated three lists of reference genes widely cited in the literature, Warrington et al. [51], Hsiao et al. [25], and Eisenberg et al. [22, 46]. Zhu et al. [54] generated an up-dated list of reference genes using more advanced arrays with an increased number of probes. Finally, de Jonge et al. [16] conducted a meta-analysis of 13,629 human gene array samples and found a selection of candidate genes showing a stable expression across various human cell types and experimental conditions. Very few studies to date have been conducted using qPCR to investigate stable gene expression in the CNS. Penna et al. [39] found that CYC1 and EIF4A2 showed stable expression in Alzheimer’s disease only over a small collection of pre-selected reference genes available within a commercially available kit.
We have recently carried out an extensive and unique analysis of genome-wide changes of gene expression in human brain tissue from Alzheimer’s disease (AD), amyotrophic lateral sclerosis (ALS), Huntington’s disease (HD), multiple sclerosis (MS), Parkinson’s disease (PD), schizophrenia (SZ) and their respective region-specific controls using the Illumina whole genome HumanRef8 v2 BeadChip (GEO accession number GSE26927). A total of 117 tissue samples were investigated representing various brain regions including the spinal cord. Since RNA samples were available, we were able to validate with RT-qPCR expression levels of candidate reference genes extracted from the microarray data. Herein, we report a validation exercise by identifying the most suitable reference genes that can be used as internal controls for RT-qPCR studies in human CNS post-mortem tissue.
Materials and methods
Tissue samples
Gene expression levels from a total of 117 samples (111 different cases with 6 replicates) were included in this study covering six neurological conditions and six different brain areas with their respective controls (Table 1). Basic clinical and demographic details can be found in supplemental material 1. RNA samples were kept at −80 °C. A subset was used for the quantification of mRNA expression by RT-qPCR. The subset cohort for RT-qPCR included four cases from each neurological condition and five controls (n = 29).
Microarray analysis and selection of reference genes
Gene expression analysis was performed on RNA extracted (RNeasy® tissue lipid mini kit; Qiagen Ltd, Crawley, UK) from snap frozen tissues [19] with the Illumina whole genome HumanRef8 v2 BeadChip (Illumina, London, UK) as described previously [21]. RNA concentration and purity was assessed by spectrophotometry (NanoDrop ND1000; NanoDrop Technologies, Delaware, USA). RNA integrity was further assessed using an Agilent 2100 Bioanalyzer and lab-on-a-chip platform technology (Agilent Technologies UK Ltd, West Lothian, UK). Data normalisation was conducted using the Rosetta error models available in the Rosetta Resolver® system v7.0. Gene expression data from Illumina BeadStudio version 3.0 were loaded into the Rosetta Resolver® Gene Expression Analysis System version 7.0 and an error model applied to each expression intensity from each sample. The basis of this is described in detail in Weng et al. [52], and consists of applying an error model to each expression intensity that provides a p value and error term of the conservative estimate of the gene-independent propagated error as well as the largely gene-dependent scattered error. Inter-array normalisation was achieved by scaling probe intensities by a scaling factor derived from the mean intensity of the array with 10 % outlier exclusion. Expression intensities from probes representing the same transcript were combined using an error weighted average. Annotations were up-dated with the Rosetta Resolver system.
To establish the most stable expressed genes across all samples, coefficients of variance (CV; CV = standard deviation divided by the mean and is expressed as a percentage) were calculated for all genes from the normalised log2-transformed expression data as described by de Jongue et al. [16]. Genes were ranked thereafter based on their CV. The maximum fold change (MFC, the ratio of the maximum and minimum) was also taken into account. In addition, genes showing a ceiling effect (saturated) with their expression level were identified when the mean was higher than the maximum minus 2 standard deviations and were consequently not considered. The genes of unknown function were also removed. A Pubmed search (July 2012) was carried out on a selection of candidate genes to verify their non-involvement in any abnormal neurological condition.
Quantification of mRNA expression by RT-qPCR
The two-step RT-qPCR was performed using the QuantiTect® reverse transcription kit, the QuantiTect® SYBR Green kit and with QuantiTect® primer assays (Qiagen) as previously described [19]. Briefly, RT-qPCR experiments were performed using the Mx3000P™ real-time PCR system with software version 4.01 (Stratagene, La Jolla, USA). The QuantiTect® primer assays are listed in Table 3. For each sample, reactions were set up in duplicate with the following cycling protocol, 95 °C for 15 min, 40 cycles with a 3-step program (94 °C for 15 s, 55 °C for 30 s, and 72 °C for 30 s) and a final melting curve analysis with a ramp from 55 to 95 °C. Samples with no reverse transcriptase reaction to test for contaminating DNA and a negative control containing no RNA template were included in each run. To control for variation in RNA levels due to manipulation across samples, the same cDNA stock was used to investigate expression levels of the 12 reference genes and all RT-qPCR assays for a particular gene were undertaken at the same time in a 96-well plate under identical conditions. Different cDNA stocks were used to assess expression levels of target gene with different normaliser.
To determine expression stability of the reference genes, the quantification cycle (Cq; aka threshold cycle), which is the PCR cycle at which an increase in reporter fluorescence from SYBR green dye can be first detected above noise signal, was determined for each sample. The same threshold level was used for all experiments. Mean Cq from duplicates generated by the software were used to calculate the CV for each gene investigated and then, the reference genes were ranked.
Data accession
All data are available online through Gene Expression Omnibus (GEO accession number GSE26927).
Statistical analysis
The following software packages were used; GraphPad Prism 5.01 (GraphPad Software Inc, La Jolla, CA, USA) and Microsoft® Office Excel® 2007 (Microsoft UK Headquarters, Reading, UK). A parametric t test was used to assess group difference and the Pearson correlation test for a relationship between two variables. To compare variance between groups, a Barlett’s test for equal variances test (multiple groups) and an F test (two groups) was used with p < 0.05 suggesting a significant difference in variances between groups. A paired non-parametric test was used to compare gene rankings. To test whether a group of samples came from a Gaussian distribution, a d’Agostino and Pearson omnibus normality test was conducted. For multiple repeated measures comparison we used the Friedman test with Dunn’s multiple comparison test. For a particular gene, gene expression intensity data and RT-qPCR relative quantity data were divided by the mean of the control group and presented as a ratio. This procedure scaled mean gene expression levels of the controls at one. The ratio of data is presented as mean ± standard error of means.
Results
Detection of constant genes using the microarray data
All gene transcripts from the Illumina whole genome HumanRef8 v2 BeadChip (20,589) were ranked according to their coefficient of variance (CV). The CV ranged from 1.11 % (least variance) to 105.78 % (most variance). The conventional reference genes ranked as follows in the microarray dataset: beclin-1 (rank: 361), GAPDH (408), QARS (1,708), actin-β (4,882), β2-microglobulin (5,334), tubulin-β (6,008), albumin (8,376), cyclophilin A (10,457) and HPRT1 (13,249). CYC1 and EIF4A2 [39] were ranked 259 and 938, respectively. An arbitrary cut-off of <2 % for the CV was used to select a set of the most stable genes. From the 170 genes selected, saturated genes and genes of unknown functions were removed, leaving a total of 131 genes, which can be found in supplementary material 2. We conducted a thorough search using PubMed on the genes at the top of the list to retain only those with no known reports (at the time of the search) of abnormal expression in any CNS disorder until we isolated a number of potential candidates. A total of eight candidate reference genes of various functions were retained. Post-mortem delay (PMD) and age at death were taken into consideration since reports suggest that these variables may impact on the expression of certain mRNA species despite extreme care given to convert the same amount of mRNA from all samples [19]. RIN was not considered since the samples in the present study were pre-selected for their “good” and homogeneous RNA quality, which is one of the prerequisites for conducting a microarray experiment. Consequently, they do not represent the full range of tissue quality available within tissue banks as tests showed that RIN values did not come from a Gaussian distribution, unlike PMD (p = 0.0123) and age at death (p = 0.0097). Despite small gene expression variances, significant shifts in expression due to disease were detected for some genes, which we believe may account for our results, especially for GAPVD1 and BECN1 (see later section; Table 2). Finally, variations in expression between brain regions, groups and disease type (Barlett’s tests for equal variances and F test) were considered. One of the homogeneity of variance violations found for the 12 genes as listed in Table 2 was for GAPDH showing a more stable expression in the superior frontal gyrus grey matter compared to other brain regions (Fig. 1 and supplemental material 3a). No consistent variance in expression between control and disease group across disease states were observed for those 12 genes (supplemental material 3b) and no significant changes were observed across diseases (supplemental material 3c).
Expression variation across brain regions for GAPDH. The Barlett’s test for equal variances was significant for GAPDH (p = 0.0022) meaning that there was a significant effect in expression variability across brain regions (control + disease samples). More precisely, GAPDH expression was more stable in the superior frontal gyrus grey matter (grey matter lesions from MS patients) than any other brain region (refer to Table 1 for the full brain region listing)
Comparison of expression between two hybridisation techniques
Expression stability of the eight novel reference genes, in addition to four more commonly used reference genes (GAPDH, BECN1, QARS and TUBB), was verified using RT-qPCR on a subset cohort (n = 29) from the microarray study. We first verified that selecting a smaller subset cohort of the microarray dataset did not significantly affect the ranking of the selected genes. We compared ranking based on CV. Only minor signal expression variations in ranking were detected between the entire and the subset cohort but were not found to be significantly different (Wilcoxon signed rank test). The four common reference genes that showed the least expression stability over 117 samples remained the least stable in the smaller cohort comprised 29 samples. Cq values from RT-qPCR experiment (n = 29) were also ranked based on CV. Finally, a final average rank was given and all rankings can be found in Table 3 and a summary in Table 4.
We then compared the stability in expression of those 12 genes between the two hybridisation techniques. Statistically, there was no difference between all 3 ranking orders of those 12 genes nor was any statistical difference in ranking found between the two hybridisation techniques (Wilcoxon matched pairs test). However, some changes in ranking order were noticed. The most striking disparity in expression between the two techniques was for BECN1 and ATP5E. Expression levels for BECN1 with RT-qPCR were very stable and placed BECN1 at position 1 compared to the microarray data, while ATP5E moved from position 2 in the microarray data to position 12 with RT-qPCR. In summary, the novel investigated genes XPNPEP1, OBSP and AARS showed an overall better or comparable expression stability than the commonly used reference genes.
Performance as normalisers of novel reference genes
Since, RT-qPCR represents the gold standard to validate gene expression changes from cDNA microarrays, we assessed the eight top reference genes from the overall ranking (XPNPEP1, AARS, GAPVD1, ATP5E, OSBP, CSNK2B, BECN1 and QARS; in ranking order; see Table 4) as normalisers for a GOI to determine whether they were able to replicate the increased microarray expression levels of the GOI. Transforming growth factor, beta receptor II (TGFBR2) was chosen as the test-gene of interest. This gene was up-regulated in Huntington’s disease by 1.74-fold by microarray analysis. Except for GAPVD1 and QARS, all normalisers were able to reproduce significantly increased levels of TGFBR2 in HD from the microarray, albeit with a different magnitude of change (Fig. 2a). A good correlation in expression levels between the microarray and the RT-qPCR data was found with the exception of GAPVD1 (Table 5). From the novel reference genes, XPNPEP1, AARS, OSBP and CSNK2B replicated most efficiently the fold change of the test-gene of interest. TGFRB2 expression in HD was 1.0-fold higher with BECN1 (mean = 3.22 ± 1.07) and reduced with ATPE5 (mean = 1.6 ± 2.03) as normalisers. The higher expression of TGFRB2 when using BECN1 as normaliser could be explained due to significant variances of mRNA BECN1 levels in the disease group compared to controls, similarly for GAPDV1 (Fig. 3). XPNPEP1 and AARS were retained as will be discussed thereafter. In addition, using another GOI, TNFRSF14 (tumour necrosis factor receptor superfamily, member 14), which was significantly up-regulated (microarray data: ×1.94; p < 0.001) in PD [21], expression levels were compared using BECN1 and XPNPEP1 as normalisers (Fig. 2b). The significantly increased expression levels of TNFRSF14 were more closely matched to the expression levels from the microarray output when XPNPEP1 was used as normaliser (×2.04; p < 0.01) compared to BECN1 (×1.61; p < 0.05).
qPCR data for TGFBR2 (a) and TNFRSF14 (b) using different normalisers. The black dotted lines represent the expression levels of controls. In the first panel a, except for GAPVD1 and QARS, all normalisers were able to replicate significantly (p < 0.05) increased levels of TGFBR2 in disease (×1.74—microarray data). In b, XPNPEP1 (×2.04; p < 0.01) as normaliser produces a closer match to microarray output (microarray data ×1.94; p < 0.001) than BECN1 (×1.61; p < 0.05). **p < 0.01 and *p < 0.05
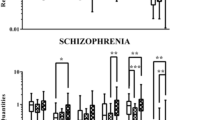
Expression levels of XPNPEP1 (a), GAPVD1 (b) and BECN1 (c). Despite a smaller coefficient of variance of expression for GAPVD1 (b; CV = 1.56) than XPNPEP1 (a; CV = 1.76) across our samples (control = 55 and disease = 62), there was a significant shift toward an increased expression in the disease group not observed with XPNPEP1 where expression was homogeneous irrespective of disease state. A significant shift towards a decrease in expression was found for BECN1 (c). ***p < 0.001 and **p < 0.01
Validation using independent datasets
In the first instance, we used two additional microarray datasets, available from our laboratory, to consolidate that XPNPEP1 is stably expressed across samples and compared its expression to more commonly references genes such as BECN1 and GAPDH. XPNPEP1 showed less variance in expression across samples in both datasets (see Table 6). Also, using a different RT-qPCR protocol and different samples, we found that XPNPEP1 expression was more stable measured by qPCR on Cq values (CV = 7.22; mean = 27.30; SD = 1.974) than GAPDH (CV = 10.60; mean = 21.85; SD = 2.317) in tissue from 10 control white matter samples and 12 MS white matter lesions [20]. We also used an external microarray dataset (GSE13162) from human tissue (frontotemporal cortex) samples (Affymetrix Human Genome U133A 2.0 Array) available from Gene Expression Omnibus (GEO) [13]. The CV of individual probes for XPNPEP1, ACTB, BECN1, GAPDH and HPRT1 show that XPNPEP1 expression was more stable than BECN1 but not GAPDH or ACTB (Table 7).
Finally, we investigated further whether the use of XPNPEP1 as a novel reference gene could also be applied to RT-qPCR mouse brain studies. From a listing available at the Allen Institute for Brain Science, we selected three mouse microarray datasets. The first dataset (GSE16496) was published by Kasukawa et al. [29] and used the Affymetrix platform (Affymetrix Mouse Genome 430 2.0 Array). The second study (GSE26024) used the Illumina MouseRef-8 v2.0 BeadChip on 21 samples [5] and the third (GSE9566) used again the Affymetrix platform as mentioned above, but, instead of whole tissue samples, gene expression changes in isolated CNS cells, i.e., neurones, astrocytes and oligodendrocytes were compared [11]. A summary of the results can be found in Table 8. Xpnpep1 was well within the lower CV range amongst the selected common reference genes, Actb, Becn1, Gapdh and Hprt (see supplementary material 4 for individual data), which placed Xpnpep1 as an equal or better reference gene than some common ones currently in use for RT-qPCR studies for human, mouse and single cell CNS samples.
Discussion
The pathogenetic mechanisms underlying the major neurodegenerative disorders are still poorly understood. Our recent microarray gene expression studies aimed to look for common pathways of degeneration underlying disease. In doing so, we wanted to overcome some of the shortcomings previously encountered when using microarray technology, such as the use of different microarray platforms, different analytical software tools or inadequate normalisation. By studying expression data from six different neurological conditions together with their respective controls, we have been able to generate a list of the most stably expressed genes within the CNS not affected by on-going disease processes that can now be used as reference genes. We propose herein XPNPEP1 as a more suitable reference gene for the purpose of normalisation for CNS gene expression studies using post-mortem tissues and also offer a list of potential alternatives.
The expression of the XPNPEP1 [X-prolyl aminopeptidase (aminopeptidase P) 1, soluble] gene showed excellent stability in our analysis. XPNPEP1 is localised on chromosome 10q25.3 and can be found in a wide variety of organisms including mammals, yeasts, and bacteria and a variety of tissues including lung, kidney, brain, adrenal medulla and cell types such platelets, lymphocytes, and astrocytes [36]. Northern blot analysis indicates that this gene product is ubiquitously expressed [50]. It shows amino acid homology with Escherichia coli methionine aminopeptidase, a prototypical member of the “pita bread-fold” family [4]. This novel class of proteases remove the N-terminal methionine from proteins and peptides, a crucial step in the maturation of many proteins towards initiating biological activity, adequate subcellular localisation and eventual degradation [37, 38]. The human XPNPEP1 consists of 623 amino acid residues with a calculated molecular mass of 69,886 Da, and has been shown to hydrolyse bradykinin—a small vasoactive peptide involved in a variety of biological processes [36]. Our findings suggest that increased expression levels of a GOI were better replicated with XPNPEP1 as normaliser than with BECN1. Beclin-1 plays a central role in autophagy [28] and has recently been shown to increase in expression in neurones at the site of traumatic brain injury [12]. Also, neurodegeneration was shown to induce up-regulation of beclin-1 [23]. In the mammalian adult brain beclin-1 is expressed in neurones and astrocytes, and up-regulation of beclin-1 has been reported in injured neurones [53] and in a subpopulation of astrocytes reacting to injured neuronal components [18]. Absolute mRNA expression of beclin-1 will be therefore subject to variances depending on total neuronal cell numbers and disease activity. Thus, caution is required in the use of this reference gene in the study of neurodegeneration.
AARS (alanyl-tRNA synthetase) belongs to a family of 20 cellular enzymes, the aminoacyl-tRNA synthetases, responsible for translating nucleotide triplets in genes and mRNAs. These enzymes are involved in a number of functions such as translational fidelity, tRNA processing, RNA splicing, RNA trafficking, apoptosis and transcriptional and translational regulation [41, 47]. No changes in expression of alanyl-tRNA synthetase (AlaRS) have been associated with any neurodegenerative disorders; however, mutation in the AlaRS protein was shown to lead to Purkinje cell degeneration in rodents [34], to peripheral axonal degeneration and cause autosomal dominant Charcot-Marie-Tooth disease [45]. Our preliminary evidence would suggest that AARS was more invariant than QARS (a common reference gene). We have investigated only one member (AARS) of this group of genes. However, other members may also be good candidates. OSBP (Oxysterol binding protein) is thought to act as a sensor of sterols [24] and overexpression of OSBP has been linked to increased cholesterol and sphingomyelin levels [32]. Although there is no known involvement of OSBP in any CNS disorders as yet, cholesterol metabolism alterations have nevertheless been shown to play a role in the pathophysiology of multiple CNS disorders [1, 30] and therefore would not be an ideal reference gene. Finally, the CSNK2B (Casein kinase 2, beta polypeptide) gene is located on chromosome 6p21, the human major histocompatibility complex [2]. Although not associated with any diseases as yet, it is well established that the human major histocompatibility complex is associated with many autoimmune/inflammatory diseases, and therefore was not retained.
In conclusion, the generation of expression profiles for several neurological disorders, representing various CNS regions allowed us to provide a selection of candidate invariant genes with potential to be useful as novel reference genes for appropriate normalisation of qPCR data from human CNS post-mortem tissue studies. By doing so, we overcame some of the shortcomings encountered with microarray studies. We isolated an initial 131 genes from the microarray data using the coefficient of variance over other methods available [49] and verified expression levels from a selection of genes in a sub-cohort of the microarray study using RT-qPCR. We were able to test several candidate genes as normalisers for a target GOI. XPNPEP1 and AARS performed best in replicating gene expression levels from the microarray data using RT-qPCR. XPNPEP1 performed better than BECN1 in being able to replicate microarray data. This impetus in the search for an appropriate reference gene has emerged due to rising concerns about the validity of the commonly used reference genes [8, 9, 33] and the importance of following stringent standards for molecular expression studies [10]. Although it is advisable to test for the most appropriate reference gene in each experimental setting and to use multiple reference genes, it is nevertheless very useful to have one established reliable tissue-specific reference gene, in particular, for single target gene expression investigations using RT-qPCR in a particular CNS disease. We provide herein one validated novel reference gene, XPNPEP1, useful for RT-qPCR studies of the human CNS and a list of other potential candidates.
References
Adibhatla RM, Hatcher JF (2008) Altered lipid metabolism in brain injury and disorders. Subcell Biochem 49:241–268
Albertella MR, Jones H, Thomson W, Olavesen MG, Neville M, Campbell RD (1996) Localisation of eight additional genes in the human major histocompatibility complex, including the gene encoding the casein kinase II beta subunit, and DNA sequence analysis of the class III region. DNA Seq J DNA Seq Mapp 7:9–12
Barrachina M, Castano E, Ferrer I (2006) TaqMan PCR assay in the control of RNA normalization in human post-mortem brain tissue. Neurochem Int 49:276–284
Bazan JF, Weaver LH, Roderick SL, Huber R, Matthews BW (1994) Sequence and structure comparison suggest that methionine aminopeptidase, prolidase, aminopeptidase P, and creatinase share a common fold. Proc Natl Acad Sci USA 91:2473–2477
Bottomly D, Walter NA, Hunter JE, Darakjian P, Kawane S, Buck KJ, Searles RP, Mooney M, McWeeney SK, Hitzemann R (2011) Evaluating gene expression in C57BL/6J and DBA/2J mouse striatum using RNA-Seq and microarrays. PLoS ONE 6:e17820
Bustin SA (2000) Absolute quantification of mRNA using real-time reverse transcription polymerase chain reaction assays. J Mol Endocrinol 25:169–193
Bustin SA (2002) Quantification of mRNA using real-time reverse transcription PCR (RT-PCR): trends and problems. J Mol Endocrinol 29:23–39
Bustin SA (2004) A–Z of quantitative PCR. International University Line, La Jolla
Bustin SA, Benes V, Nolan T, Pfaffl MW (2005) Quantitative real-time RT-PCR—a perspective. J Mol Endocrinol 34:597–601
Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, Mueller R, Nolan T, Pfaffl MW, Shipley GL, Vandesompele J, Wittwer CT (2009) The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55:611–622
Cahoy JD, Emery B, Kaushal A, Foo LC, Zamanian JL, Christopherson KS, Xing Y, Lubischer JL, Krieg PA, Krupenko SA, Thompson WJ, Barres BA (2008) A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. J Neurosci 28:264–278
Cao Y, Klionsky DJ (2007) Physiological functions of Atg6/Beclin 1: a unique autophagy-related protein. Cell Res 17:839–849
Chen-Plotkin AS, Geser F, Plotkin JB, Clark CM, Kwong LK, Yuan W, Grossman M, Van Deerlin VM, Trojanowski JQ, Lee VM (2008) Variations in the progranulin gene affect global gene expression in frontotemporal lobar degeneration. Hum Mol Genet 17:1349–1362
Colell A, Green DR, Ricci JE (2009) Novel roles for GAPDH in cell death and carcinogenesis. Cell Death Differ 16:1573–1581
Coulson DT, Brockbank S, Quinn JG, Murphy S, Ravid R, Irvine GB, Johnston JA (2008) Identification of valid reference genes for the normalization of RT qPCR gene expression data in human brain tissue. BMC Mol Biol 9:46
de Jonge HJ, Fehrmann RS, de Bont ES, Hofstra RM, Gerbens F, Kamps WA, de Vries EG, van der Zee AG, te Meerman GJ, ter Elst A (2007) Evidence based selection of housekeeping genes. PLoS ONE 2:e898
Dheda K, Huggett JF, Chang JS, Kim LU, Bustin SA, Johnson MA, Rook GA, Zumla A (2005) The implications of using an inappropriate reference gene for real-time reverse transcription PCR data normalization. Anal Biochem 344:141–143
Diskin T, Tal-Or P, Erlich S, Mizrachy L, Alexandrovich A, Shohami E, Pinkas-Kramarski R (2005) Closed head injury induces upregulation of Beclin 1 at the cortical site of injury. J Neurotrauma 22:750–762
Durrenberger PF, Fernando S, Kashefi SN, Ferrer I, Hauw JJ, Seilhean D, Smith C, Walker R, Al-Sarraj S, Troakes C, Palkovits M, Kasztner M, Huitinga I, Arzberger T, Dexter DT, Kretzschmar H, Reynolds R (2010) Effects of antemortem and postmortem variables on human brain mRNA quality: a BrainNet Europe study. J Neuropathol Exp Neurol 69:70–81
Durrenberger PF, Ettorre A, Kamel F, Webb LV, Sim M, Nicholas RS, Malik O, Reynolds R, Boyton RJ, Altmann DM (2012) Innate Immunity in multiple sclerosis white matter lesions: expression of natural cytotoxicity triggering receptor 1 (NCR1). J Neuroinflamm 9:1
Durrenberger PF, Grunblatt E, Fernando FS, Monoranu CM, Evans J, Riederer P, Reynolds R, Dexter DT (2012) Inflammatory pathways in Parkinson’s disease; A BNE Microarray Study. Parkinsons Dis 2012:214714
Eisenberg E, Levanon EY (2003) Human housekeeping genes are compact. Trends Genet 19:362–365
Erlich S, Shohami E, Pinkas-Kramarski R (2006) Neurodegeneration induces upregulation of Beclin 1. Autophagy 2:49–51
Fairn GD, McMaster CR (2008) Emerging roles of the oxysterol-binding protein family in metabolism, transport, and signaling. Cell Mol Life Sci 65:228–236
Hsiao LL, Dangond F, Yoshida T, Hong R, Jensen RV, Misra J, Dillon W, Lee KF, Clark KE, Haverty P, Weng Z, Mutter GL, Frosch MP, Macdonald ME, Milford EL, Crum CP, Bueno R, Pratt RE, Mahadevappa M, Warrington JA, Stephanopoulos G, Gullans SR (2001) A compendium of gene expression in normal human tissues. Physiol Genomics 7:97–104
Huggett J, Dheda K, Bustin S, Zumla A (2005) Real-time RT-PCR normalisation; strategies and considerations. Genes Immun 6:279–284
Johansson S, Fuchs A, Okvist A, Karimi M, Harper C, Garrick T, Sheedy D, Hurd Y, Bakalkin G, Ekstrom TJ (2007) Validation of endogenous controls for quantitative gene expression analysis: application on brain cortices of human chronic alcoholics. Brain Res 1132:20–28
Kang R, Zeh HJ, Lotze MT, Tang D (2011) The Beclin 1 network regulates autophagy and apoptosis. Cell Death Differ 18:571–580
Kasukawa T, Masumoto KH, Nikaido I, Nagano M, Uno KD, Tsujino K, Hanashima C, Shigeyoshi Y, Ueda HR (2011) Quantitative expression profile of distinct functional regions in the adult mouse brain. PLoS ONE 6:e23228
Korade Z, Kenworthy AK (2008) Lipid rafts, cholesterol, and the brain. Neuropharmacology 55:1265–1273
Kubista M, Andrade JM, Bengtsson M, Forootan A, Jonak J, Lind K, Sindelka R, Sjoback R, Sjogreen B, Strombom L, Stahlberg A, Zoric N (2006) The real-time polymerase chain reaction. Mol Aspects Med 27:95–125
Lagace TA, Byers DM, Cook HW, Ridgway ND (1999) Chinese hamster ovary cells overexpressing the oxysterol binding protein (OSBP) display enhanced synthesis of sphingomyelin in response to 25-hydroxycholesterol. J Lipid Res 40:109–116
Lanoix D, Lacasse AA, St-Pierre J, Taylor SC, Ethier-Chiasson M, Lafond J, Vaillancourt C (2012) Quantitative PCR pitfalls: the case of the human placenta. Mol Biotechnol
Lee JW, Beebe K, Nangle LA, Jang J, Longo-Guess CM, Cook SA, Davisson MT, Sundberg JP, Schimmel P, Ackerman SL (2006) Editing-defective tRNA synthetase causes protein misfolding and neurodegeneration. Nature 443:50–55
Lee S, Jo M, Lee J, Koh SS, Kim S (2007) Identification of novel universal housekeeping genes by statistical analysis of microarray data. J Biochem Mol Biol 40:226–231
Li X, Lou Z, Zhou W, Ma M, Cao Y, Geng Y, Bartlam M, Zhang XC, Rao Z (2008) Structure of human cytosolic X-prolyl aminopeptidase: a double Mn(II)-dependent dimeric enzyme with a novel three-domain subunit. J Biol Chem 283:22858–22866
Lowther WT, Matthews BW (2000) Structure and function of the methionine aminopeptidases. Biochim Biophys Acta 1477:157–167
Mogk A, Schmidt R, Bukau B (2007) The N-end rule pathway for regulated proteolysis: prokaryotic and eukaryotic strategies. Trends Cell Biol 17:165–172
Penna I, Vella S, Gigoni A, Russo C, Cancedda R, Pagano A (2011) Selection of candidate housekeeping genes for normalization in human postmortem brain samples. Int J Mol Sci 12:5461–5470
Ruan W, Lai M (2007) Actin, a reliable marker of internal control? Clin Chim Acta 385:1–5
Schimmel P (2008) Development of tRNA synthetases and connection to genetic code and disease. Protein Sci 17:1643–1652
Schmittgen TD, Zakrajsek BA (2000) Effect of experimental treatment on housekeeping gene expression: validation by real-time, quantitative RT-PCR. J Biochem Biophys Methods 46:69–81
Sirover MA (1999) New insights into an old protein: the functional diversity of mammalian glyceraldehyde-3-phosphate dehydrogenase. Biochim Biophys Acta 1432:159–184
Solanas M, Moral R, Escrich E (2001) Unsuitability of using ribosomal RNA as loading control for Northern blot analyses related to the imbalance between messenger and ribosomal RNA content in rat mammary tumors. Anal Biochem 288:99–102
Stum M, McLaughlin HM, Kleinbrink EL, Miers KE, Ackerman SL, Seburn KL, Antonellis A, Burgess RW (2011) An assessment of mechanisms underlying peripheral axonal degeneration caused by aminoacyl-tRNA synthetase mutations. Mol Cell Neurosci 46:432–443
Su AI, Cooke MP, Ching KA, Hakak Y, Walker JR, Wiltshire T, Orth AP, Vega RG, Sapinoso LM, Moqrich A, Patapoutian A, Hampton GM, Schultz PG, Hogenesch JB (2002) Large-scale analysis of the human and mouse transcriptomes. Proc Natl Acad Sci USA 99:4465–4470
Szymanski M, Deniziak M, Barciszewski J (2000) The new aspects of aminoacyl-tRNA synthetases. Acta Biochim Pol 47:821–834
Thellin O, Zorzi W, Lakaye B, De Borman B, Coumans B, Hennen G, Grisar T, Igout A, Heinen E (1999) Housekeeping genes as internal standards: use and limits. J Biotechnol 75:291–295
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A, Speleman F (2002) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3: RESEARCH0034
Vanhoof G, Goossens F, Juliano MA, Juliano L, De Meester I, Hendriks D, Schatteman K, Scharpe S (1997) Human lymphocyte X-prolyl aminopeptidase (aminopeptidase P)-like protein. A new member of the proline peptidase family? Adv Exp Med Biol 421:25–29
Warrington JA, Nair A, Mahadevappa M, Tsyganskaya M (2000) Comparison of human adult and fetal expression and identification of 535 housekeeping/maintenance genes. Physiol Genomics 2:143–147
Weng L, Dai H, Zhan Y, He Y, Stepaniants SB, Bassett DE (2006) Rosetta error model for gene expression analysis. Bioinformatics 22:1111–1121
Yue Z, Horton A, Bravin M, DeJager PL, Selimi F, Heintz N (2002) A novel protein complex linking the delta 2 glutamate receptor and autophagy: implications for neurodegeneration in lurcher mice. Neuron 35:921–933
Zhu J, He F, Song S, Wang J, Yu J (2008) How many human genes can be defined as housekeeping with current expression data? BMC Genomics 9:172
Acknowledgments
We would like to thank all the tissue donors and their families. Also we are grateful to Veronique Sazdovitch and Kasztner Magdolna for technical assistance. We would like also to thank Charles Mein at The Genome Centre (John Vane Science Centre, Queen Mary, University of London, Charterhouse Square, London EC1M 6BQ) for his assistance and expertise. This study was supported by the European Commission under the Sixth Framework Programme (BrainNet Europe II, LSHM-CT-2004-503039). The Multiple Sclerosis and Parkinson’s Disease Tissue Banks at Imperial were supported by the MS Society of Great Britain and Northern Ireland and the Parkinson’s UK respectively.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Durrenberger, P.F., Fernando, F.S., Magliozzi, R. et al. Selection of novel reference genes for use in the human central nervous system: a BrainNet Europe Study. Acta Neuropathol 124, 893–903 (2012). https://doi.org/10.1007/s00401-012-1027-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-012-1027-z