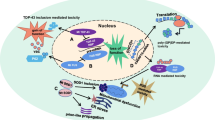
Abstract
Amyotrophic lateral sclerosis (ALS) is a progressive neurodegenerative disease that primarily involves the motor neuron system. The author initially summarizes the principal features of human ALS neuropathology, and subsequently describes in detail ALS animal models mainly from the viewpoint of pathological similarities and differences. ALS animal models in this review include strains of rodents that are transgenic for superoxide dismutase 1 (SOD1), ALS2 knockout mice, and mice that are transgenic for cytoskeletal abnormalities. Although the neuropathological results obtained from human ALS autopsy cases are valuable and important, almost all of such cases represent only the terminal stage. This makes it difficult to clarify how and why ALS motor neurons are impaired at each clinical stage from disease onset to death, and as a consequence, human autopsy cases alone yield little insight into potential therapies for ALS. Although ALS animal models cannot replicate human ALS, in order to compensate for the shortcomings of studies using human ALS autopsy samples, researchers must inevitably rely on ALS animal models that can yield very important information for clarifying the pathogenesis of ALS in humans and for the establishment of reliable therapy. Of course, human ALS and all ALS animal models share one most important similarity in that both exhibit motor neuron degeneration/death. This important point of similarity has shed much light on the pathomechanisms of the motor neuron degeneration/death at the cellular and molecular levels that would not have been appreciated if only human ALS autopsy samples had been available. On the basis of the aspects covered in this review, it can be concluded that ALS animal models can yield very important information for clarifying the pathogenesis of ALS in humans and for the establishment of reliable therapy only in combination with detailed neuropathological data obtained from human ALS autopsy cases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Amyotrophic lateral sclerosis (ALS) in humans is a progressive disease characterized by degeneration of both upper and lower motor neurons. Upper motor neurons are located mainly in layer V of the motor cortex, and are known as Betz cells, which are giant cells approximately 60–120 μm in diameter. Cytoarchitecturally, the axons of the upper motor neurons connect directly with the lower motor neurons located in the motor nuclei of the brainstem and the anterior horn of the spinal cord. The axons of the lower motor neurons project mainly to skeletal muscles. Although an essential pathological feature of ALS is motor neuron loss, affected motor neurons often contain characteristic inclusions in the perikarya, dendrites and axons. The concept of “ALS” has been widely debated for over 130 years since Charcot first introduced the term “la sclérose latérale amyotrophique” in 1874 [15]. Currently, in the era of advanced genetics, the molecular bases and mechanisms of ALS are now known to be very complex, thus explaining the different phenotypes of ALS. Therefore, ALS is considered to be a type of syndrome rather than a single disease entity.
Human pathologists are entirely dependent on human ALS autopsy samples in order to acquire a more definitive understanding of the etiology and pathogenesis of ALS motor neuron death. As almost all ALS autopsy samples are obtained from patients at the terminal stage, it is difficult to clarify how and why ALS motor neurons are impaired at each clinical stage from disease onset to death. Therefore, analyses of ALS autopsy samples alone is not sufficient to lead to possible therapies for ALS. Under these circumstances, many researchers including pathologists have come to rely on animal models of ALS in order to gain insights into both the mechanisms involved in motor neuron death and possible therapeutic approaches. Inevitably, these ALS animal models have different characteristics, because even human ALS, which is the prototype for these models, is not a single disease. On the basis of the aspects covered in this review, however, the author wish to emphasize that ALS animal models can yield very important information for clarifying the pathogenesis of ALS in humans only in combination with detailed neuropathological data obtained from human ALS autopsy cases.
Human ALS neuropathology
Sporadic ALS (SALS)
Human ALS is classified into two major subtypes: sporadic ALS (SALS) and familial ALS (FALS). In SALS, the degenerative change is mainly restricted to the motor neuron system, except for the oculomotor, trochlear and abducens nuclei controlling eye movements, as well as Onufrowicz’s nucleus which functions in fecal and urinary continence. Upper motor neurons such as the Betz cells in the motor cortex are also affected. Degeneration of the corticospinal tracts in the anterior and lateral columns of the spinal cord are particularly evident. Especially, the corticospinal tract degeneration is most evident in the lower spinal cord segment, supporting the hypothesis of a dying back degeneration of axons. In the degenerated primary motor tracts, there is loss of large myelinated fibers in association with variable astrocytic gliosis. Destruction of these fibers is usually associated with the appearance of lipid-laden macrophages.
An essential histopathological feature of SALS is loss of both upper and lower motor neurons, especially large anterior horn cells throughout the length of the spinal cord. In addition, striated muscles demonstrate denervation atrophy, i.e., neurogenic muscle atrophy. The surviving motor neurons often show histopathological features including cytoplasmic shrinkage and lipofuscin granules. The cytopathology of the affected motor neurons in SALS is characterized by the following two important intracytoplasmic inclusions.
Bunina bodies
Bunina bodies are small eosinophilic granular inclusions (1–3 μm in diameter) in the anterior horn cells, appearing either singly or in a group, and sometimes arranged in small beaded chains. These Bunina bodies are observed within the cytoplasm or dendrites (Fig. 1a), but they have not been found within the axoplasm. Bunina bodies stain bright red with hematoxylin and eosin (H&E) staining, deep blue with phosphotungstic acid hematoxylin, and blue with Luxol fast blue. Immunohistochemical studies have revealed that they express only cystatin C (Fig. 1b), and do not have a variety of other immunoreactive epitopes such as ubiquitin. Ultrastructurally, Bunina bodies consist of electron-dense amorphous material that contains tubules or vesicular structures. The amorphous material frequently includes a cytoplasmic island containing neurofilaments and other micro-organelles [58].
Neuropathological findings of human ALS. a, b Light micrographs of Bunina bodies. a Bunina bodies in an anterior horn cell of an SALS patient. Bunina bodies are small eosinophilic inclusions (approximately 1–3 μm in diameter) and staining bright red with H&E. Bunina bodies are observed within the cytoplasm (double arrows), being arranged in a chain-like formation. In addition, Bunina bodies are also seen within the dendrites (arrow), appearing as a cluster. H&E. b Cystatin C immunostaining. Bunina bodies are positive for cystatin C (double arrows). A Bunina body (indicated by the center arrow) appears as a round structure with central-lucent core, which corresponds to a cytoplasmic island containing neurofilaments and other microorgans at the ultrastructural level. c–e Light micrographs of neuronal Lewy-body-like hyaline inclusions (LBHIs). c A round LBHI (arrow) is observed in the cytoplasm of the anterior horn cell. This round LBHI is composed of eosinophilic core with paler peripheral halo. A small ill-defined LBHI (arrowhead) is also seen in the cytoplasm of the anterior horn cell, and consists of obscure slightly eosinophilic materials. H&E. d, e Serial sections of a neuronal LBHI in a spinal anterior horn cell immunostained with the antibodies against SOD1 (d) and ubiquitin (e). An intraneuronal LBHI is clearly labeled by the antibodies to SOD1 and ubiquitin. The immunoreactivity for both SOD1 and ubiquitin is almost restricted to the halo of the LBHI. f Light micrograph of an astrocytic hyaline inclusion (Ast-HI) in the spinal cord anterior horn. The Ast-HI (arrow) is round and eosinophilic. The astrocyte-bearing the Ast-HI is morphologically different from the adjacent neuron. The nucleus of the cell bearing the Ast-HI resembles that of a reactive astrocyte (double arrowheads) and not the nucleus of an oligodendrocyte (arrowhead). H&E. g A neurofilamentous conglomerate inclusion showing intense immunohistochemical positivity for phosphorylated neurofilament protein. Scale bar a (also for b–e, g) 30 μm, Scale bar f 10 μm
These specific bodies were originally described by a Russian pathologist Bunina [12] in 1962 in two FALS patients belonging to two different families. After Bunina’s report, Hirano immediately confirmed that these inclusion bodies were present in a number of ALS patients, including those with SALS and FALS neuropathologically identical to SALS, as well as Guamanian ALS [41]. Although not all ALS patients necessarily have Bunina bodies histologically, Bunina bodies themselves are currently considered a specific histopathological hallmark of ALS. Broadly speaking, Bunina bodies can be observed in SALS, frontotemporal lobar degeneration with motor neuron disease (FTLD-MND; terminologically identical to ALS with dementia, ALSD), mutant SOD1-unlinked FALS that is neuropathologically identical to SALS, and Guamanian ALS. However, there have been no reports that Bunina bodies are present in SOD1-mutated FALS (Table 1) or transgenic rodents with mutant SOD1.
Skein-like inclusions (SLIs) and round hyaline inclusions (RHIs)
The SLIs are intracytoplasmic filamentous structures [58] that are frequently encountered in preparations immunostained for ubiquitin, although in H&E preparations they are hardly visible or sometimes detected as faintly eosinophilic structures. SLIs form essential aggregates of thread-like structures. In more aggregated forms, SLIs show dense collections of filaments, in which the SLIs appear as spherical structures. RHIs are pale eosinophilic inclusions with halos in the remaining anterior horn cells in H&E preparations [57]. Sometimes, they lack halos and have irregular margins associated with filamentous structures similar to SLIs. Immunohistochemically, SLIs and RHIs are positive for ubiquitin, but negative for phosphorylated (p) neurofilament protein (NFP; pNFP) and SOD1 [58]. Ultrastructurally, the essential abnormal filaments of the SLIs range in width from approximately 15 nm in the naked parts without granules to about 20 nm in parts with fuzzy granules (i.e., fibrils with granules); these fibrils with granules often form bundles. In more aggregated forms, SLIs that appear to be spherical structures are composed of many bundles of these fibrils with granules [58]. Ultrastructurally, the essential abnormal filaments of the RHIs range in width from approximately 15 nm in the naked parts without granules to about 20 nm in the parts with fuzzy granules (fibrils with granules), being very similar to the fibrils with granules in SLIs [58]. As a whole, RHIs form spherical aggregates without a limiting membrane, and a hypothesis that SLIs evolve to RHIs has even been proposed [58]. Unlike Lewy body-like hyaline inclusions (LBHIs) in mutant SOD1-linked motor neurons, although RHIs are similar to LBHIs in H&E preparations, RHIs are negative for SOD1 [58]. Simply and clearly to state, SLIs/RHIs are ultrastructurally composed of 15–20-nm fibrils with granules, and LBHIs comprised of 15–25-nm granule-coated fibrils with SOD1 epitope [58] (see “Neuronal LBHIs and Ast-HIs”).
Familial ALS (FALS)
The FALS accounts for approximately 5–10% of all cases of ALS, and is histopathologically subclassified into two types. One type of FALS is neuropathologically identical to SALS, and frequently contains Bunina bodies. On the basis of the author’s observations of two mutant SOD1-unlinked siblings with FALS, this mutant SOD1-non-linked FALS type has similar neuropathological features including Bunina bodies and SLIs/RHIs (Table 1). It is likely that the inclusion bodies originally described by Bunina (Bunina bodies) would have been present in this FALS type. The other form of FALS is that showing posterior column involvement. In addition to the pathological features of SALS, this form also shows degeneration of the middle zone of the posterior column, Clarke nuclei and posterior spinocerebellar tracts. In 1967, Hirano et al. [42] reported the presence of Lewy body-like hyaline inclusions (LBHIs) in the anterior horn cells throughout the spinal cord in this type of FALS (Fig. 1c). This led to the establishment of this entity as FALS with posterior column involvement. In H&E preparations, neuronal LBHIs show an eosinophilic core with a paler peripheral halo, and their name is derived from their H&E staining features, which resemble those of Lewy bodies in patients with Parkinson’s disease.
To date, more than 100 different mutations within all exons of the SOD1 gene and its introns have been identified as being involved in the development of chromosome 21q-linked FALS. These SOD1 gene mutations are present in about 20% of FALS patients. SOD1-mutated FALS shows a variety of clinical phenotypes according to which type of SOD1 gene mutation is responsible: the disease onset and duration are reported to be closely linked to the type of the SOD1 mutation (Fig. 2), i.e., as Fig. 2 demonstrates, FALS with mutant G93A SOD1 shows rapid disease progression with severe neurological symptoms (about 1 year to at most 4 years), while FALS with mutant H46R SOD1 has a very long disease duration with mild neurological signs (11–24 years). From the viewpoint of only the copy number of the mutant SOD1 gene, almost all FALS patients with heterozygosity for the mutant SOD1 gene carry only a single copy of mutant SOD1 gene, i.e., most patients with SOD1-mutated FALS have the same transgene copy number. Based on this notion, it might be expected that FALS patients with heterozygosity for the mutant SOD1 gene would exhibit a similar expression level of mutant SOD1 protein. However, the protein expression level of mutant SOD1 in FALS patients differs due to the difference in stability of the mutant SOD1 protein itself; FALS patients with unstable-type mutant SOD1 show a lower protein expression level than FALS patients with stable-type mutant SOD1 [86].
Schema showing relationship between the main characteristics of transgenic rodents with mutant SOD1 and the main mutation of the human SOD1 gene. The main characteristics of transgenic rodents and human SOD1 gene structure are shown diagrammatically. With respect to transgenic rodents, the main characteristics of copy number, SOD1 protein level (SOD1 Pr), and SOD1 activity (SOD1 Act) in the G93A mouse (lines = G1H, G1L, G5, G5/G5, G12, G20), G37R mouse (lines = 42, 9, 106, 29), G85R mouse (line = 148), G86R mouse (line = M1), L126dellTT mouse (lines = D, DF), G127insTGGG mouse, H46R rat (line = 4), and G93A rat (lines = 39, 26H) are shown within each rodent body; exact values for each unit are described in the text (See “Main characteristics of animal models based on SOD1”). As for the human SOD1 gene structure, large blue areas (E1–E5) = exon 1 to 5; small open areas = intron; small dark gray area = 3 prime untranslated region. Large black areas (beta-sheets) = beta-sheet 1–8; large open areas (beta-loops) = beta-loops I to VII [IV, VII = active site; III, VI = Greek key]. Superscript Cu = copper binding site; superscript Zn = zinc binding site; superscript dc = dimer contact. Amino acid or nucleotide substitutions are indicated at appropriate codons according to the accepted international nomenclature. Even SALS has an SOD1 abnormality; each encircled superscript S indicates the SALS. Each encircled superscript N indicates ALS without a familial history. Disease durations (years) are shown in parentheses. FALS shows various disease durations according to the type of SOD1 gene mutation responsible; enzyme activities (% value relative to normal control erythrocytes) are shown in brackets. Mouse SOD1 gene structure of exon 4 (E4) is shown in yellow
Neuropathologically, it is of note that many SOD1-mutated FALS cases are of the posterior column involvement type with neuronal LBHIs (Table 1). In addition to neuronal LBHIs, certain long-surviving FALS patients with SOD1 gene mutations show astrocytic hyaline inclusions (Ast-HIs), which were first reported by the author in 1996 [53] (Fig. 1f). Neuronal LBHIs and Ast-HIs are characteristic intracytoplasmic structures in SOD1-mutated cells. By marked contrast, Bunina bodies have not been described in mutant SOD1-linked FALS unlike mutant SOD1-unlinked FALS with Bunina bodies (Table 1).
Neuronal LBHIs and Ast-HIs
Neuronal LBHIs and Ast-HIs in SOD1-mutated FALS have strong SOD1 immunoreactivity (Fig. 1d), and both type of inclusion are strongly positive for ubiquitin (Fig. 1e). Interestingly, LBHIs/Ast-HIs contain both wild-type and mutant SOD1 protein [11]. Ultrastructurally, neuronal LBHIs, which consist of filaments and granular materials, exhibit dense cores with rough peripheral halos and lacking a limiting membrane. The filaments of these inclusions are composed of approximately 15–25-nm granule-coated fibrils in association with normal 10-nm neurofilaments [53–57]. The neurofilaments are located mainly in the periphery and rarely in the central portion. Ast-HIs appear as globular structures that are well demarcated from other cytoplasmic structures, and have no limiting membrane. Ast-HIs are composed of about 15–25-nm granule-coated fibrils with granular materials, sometimes surrounded by normal glial filaments [53–57]. The essential constituents common to both neuronal LBHIs and Ast-HIs are granule-coated fibrils, and the indirect immunogold technique shows labeling with colloidal gold particles for wild-type and mutant SOD1 only on the surface of the granule-coated fibrils: the essential common constituents of LBHIs/Ast-HIs are SOD1-positive granule-coated fibrils [57].
Another characteristic pathological feature of patients with SOD1-mutated FALS is slight or mild corticospinal tract involvement, in contrast to severe degeneration of the lower motor neurons (Table 1). In patients with FALS lacking LBHI with the I113T mutation, neurofilament pathology is an almost universal feature. Frequently, neurofilamentous conglomerate inclusions (NFCIs) are evident, while they are less common in patients with SALS (less than 5%). Although NFCIs are recognized as homogeneous, faintly eosinophilic, oval or multi-lobulated inclusions in H&E preparations, they are intensely positive for pNFP immunohistochemically (Fig. 1g). Two FALS siblings with wild-type SOD1 in the bottom row of Table 1 show Bunina bodies and SLIs/RHIs that are usually observed in SALS, and neuropathologically they showed histology that was almost identical to SALS. These two mutant SOD1-unlinked FALS siblings did not exhibit LBHIs or posterior column degeneration. Although SLIs are seen in SALS and mutant SOD1-unlinked FALS as well as FALS with H48Q and E100G, many cases of SOD1-mutated FALS do not show SLIs (Table 1).
Transgenic animal models linked to superoxide dismutase 1 (SOD1)
Main characteristics of animal models based on SOD1
It is possible to create transgenic animals (rodents) expressing the human wild-type and mutated SOD1 gene. Currently, transgenic rodents expressing human mutant SOD1 are thought to provide the most suitable animal model of human ALS. There are several lines of transgenic rodents expressing human mutant SOD1, and their main characteristics are summarized in this paragraph in detail and shown schematically in Fig. 2. There are six major strains of ALS transgenic rodents carrying human mutant SOD1: G93A mice (lines: G1, G5, G12 and G20) [19–21, 36], G37R mice (lines: 42, 9, 106 and 29) [98], G85R mice (lines: 221, 164, 103, 148, 74, 46, 87 and 124) [10], H46R/H48Q mice (lines:139, 73, 67, and 58) [96], H46R rats (line: 4) [2, 72], and G93A rats (lines: 39) [2, 72] and 26H [43]). In addition, seven types of transgenic mice bearing human mutant SOD1 have been produced: G93R mice [31], D90A mice [49], H46R mice [1], L84V mice [1], L126delTT mice [97], L126Z (Z = stop) mice [23], and G127insTGGG mice [48]. There is a major line of transgenic mice carrying mouse G86R SOD1 (lines: M1, M2 and M3) [81].
With respect to G93A mice [19–21, 36], the original mouse strain G1-G93A carries 18.0 ± 2.6 transgene copy numbers expressing 4.1 ± 0.54 ng human SOD1 protein per μg of total protein (ng human SOD1) with SOD1 activity of 42.6 ± 2.1 U per μg of total protein (U). G5-G93A mice with 4.0 ± 0.6 copies show 1.3 ± 0.21 ng human SOD1 with 27.0 ± 2.9 U SOD1 activity. G12-G93A mice with 2.2 ± 0.8 copies exhibit 1.1 ± 0.22 ng human SOD1 with 19.5 ± 0.8 U SOD1 activity, and G20-G93A mice with 1.7 ± 0.6 copies express 0.7 ± 0.06 ng human SOD1 with 16.9 ± 0.4 U SOD1 activity. The original G1 line differentiates into two sub-lines derived from the same founder. These two sub-lines, named G1H and G1L, express different transgene copy numbers: G1H-G93A mice carrying 25 transgene copy numbers express 3.36 ± 0.84 ng SOD1 protein per μg soluble brain protein (ng SOD/sbp) with SOD1 activity of 3.04 ± 0.71 U per μg sbp (U/sbp), and G1L-G93A mice expressing 18 copies show 2.20 ± 0.9 ng SOD1/sbp with 2.56 ± 0.23 U/sbp SOD1 activity. G5/G5-G93A mice have been newly generated from G5-G93A mice: G5/G5-G93A mice with 10 copies express 1.28 ± 0.44 ng SOD1/sbp with 1.61 ± 0.14 U/sbp SOD1 activity.
With respect to G37R mice [98], the spinal cord tissues of G37R-42 mice express the human SOD1 protein at a level 12.3 times that of the endogenous mouse SOD1 protein with SOD1 activity 14.5 times higher than the control, G37R-9 mice show a 6.2-fold SOD1 protein level in spinal cord tissues and 9.0-fold SOD1 activity relative to the control, G37R-106 mice show a 5.3-fold SOD1 protein level and 7.2-fold SOD1 activity, and G37R-29 mice express a 5.0-fold SOD1 protein level and 7.0-fold SOD1 activity. There is no detailed description of transgene copy numbers in the original literature.
With regard to G85R mice [10], eight lines have been established with transgene copy numbers of 2–15, and lines 221, 164, 103, 148, 74, 87, 124 show a clinical phenotype. The SOD1 protein level (human/mouse) in these lines is 0.5, 0.8, 0.6, 1.0, 0.2, 0.6 and 0.6, respectively. Among seven lines in G85R mice, G85R-148 mice carrying 15 transgene copies numbers express the same SOD1 activity comparable to the control.
As for G86R mice [81], although there were three original lines of M1, M2 and M3 in G86R mice, only line M3 has not been perpetuated. M1 and M3 show a clinical phenotype, whereas all M2 mice appear phenotypically normal. The M1 line exhibits an equal amount of total SOD1 protein to the control and show almost identical SOD1 activity. Since G86R mice express murine mutant G86R SOD1, they are advantageous for researchers in that the mouse genomic sequence can be used to reproduce the mutation in SOD1.
With regard to transgenic rats with human mutant SOD1, H46R-4 rats carrying 25 transgene copy numbers express human SOD1 protein at 6 times the level of endogenous rat SOD1 and show SOD1 activity in the spinal cord tissues that is 0.21 times the control activity [2, 72]. G93A-39 rats expressing 10 transgene copies express 2.5 times the human SOD1 protein level and show SOD1 activity in spinal cord tissues that is 3 times the control activity [2, 72]. G93A-26H rats with 64 copies express 8.6 times the human SOD1 protein level, and their founder rats (G93A-26HL) with 72 copies show 10.4 times the human SOD1 level [43].
Neurology and neuropathology of animal models based on SOD1
The first report of transgenic mice expressing human mutant SOD1 was published by Gurney et al. [36]. Therefore, these mice—known as Gurney mice—are the most well known and have been used extensively by many researchers. Among these Gurney mice, the G1H- and G1L-G93A strains are the most widely studied [20, 36]. The first clinical symptom of G1H-G93A mice is reportedly a fine tremor in one or more limbs, which appears at approximately 90 to 100 days of age [20]. On the basis of the author’s experimental experience with G1H- and G1L-G93A mice, the term “jittering” seems to be a more appropriate descriptor for the early clinical symptoms they show, rather than “tremor”. Although spontaneous jittering/tremor as a single sign is not specific as the initial symptom, it is frequently observed in one or both hind legs when motor work loading such as the extension reflex is applied at a very early clinical stage. Many G1H-G93A mice at approximately 100 days of age exhibit clinical symptoms such as slow walking without agility due to muscle weakness, limp tail, jittering/tremor sometimes associated with motor work loading, and incomplete paresis of a single hind limb. These clinical symptoms can be regarded as significant signs of limb paresis and/or muscle weakness. From the author’s experience [52, 59], clinical onset in almost all G1H-G93A mice (B6SJL-TgN[SOD1-G93A]1Gur, JR2726) originating from Jackson Laboratory (Bar Harbor, ME, USA) occurs at about 100 days after birth, and death occurs at approximately 120 days. Neurologically, almost all G1H-G93A mice show a uniform clinical course, beginning with muscle weakness and/or paresis in the hind limbs, followed by ascent of paresis or paralysis to the forelimbs, until by the end stage the mice show severe quadriplegia, lying sideways or in a moribund state.
The essential cytopathological features of transgenic rodents overexpressing human mutant SOD1 are motor neuron loss with astrocytosis, the presence of SOD1-positive inclusions including LBHIs/Ast-HIs, and vacuole formation. Among these three major pathologic features, motor neuron loss with gliosis is the most essential, and is shared between human ALS and human mutant SOD1 transgenic rodents. In the G1H-G93A spinal cord [52, 59], the number of anterior horn cells at 90 days of age is not decreased significantly in comparison with age-matched littermates, whereas neuropil and/or neuronal intracytoplasmic vacuolation is already evident (Fig. 3a). The numbers of anterior horn cells in G1H-G93A mice at 100 days of age are slightly decreased, with abundant vacuoles and few inclusions (Fig. 3b). At 110 days, the mice demonstrate loss of anterior horn cells, some inclusions and vacuole formation. At 120 days, the mice exhibit quadriplegia and/or a moribund state, with severe loss of anterior horn cells and prominent inclusion pathology, although vacuolation pathology is less marked than that at disease onset (Fig. 3c). Although the author has utilized over 200 G1H-G93A mice hitherto and observed that the numbers of anterior horn cells in G1H-G93A mice at 100 days are slightly decreased, a review paper from Bendotti and Carri has indicated that there is 44% loss of motor neurons at 105 days [8]. This difference might be based on variations in the methods used to perpetuate the original G1H-G93A mice among researchers’ institutions, the presence or absence of backcross breeding of B6 mice × original G1H-G93A mice, and differences in quantitation methodologies among researchers. Therefore, researchers need to be sufficiently aware of the differences in the genetic background of G1H-G93A mice and research methodologies before drawing conclusions. Clinical onset in most G1L-G93A mice (B6SJL-TgN[SOD1-G93A]1 Gurdl, JR2300) from Jackson Laboratory occurs at about 185 days after birth, and death occurs after 250 days. G1L-G93A mice examined at 90, 100, 120, and 150 days of age show only vacuolation pathology without significant neuronal loss or inclusion pathology, whereas those aged 180 days reveal slightly decreased numbers of neurons with many vacuoles and a few inclusions. At the terminal stage after 250 days of age, there is significant loss of anterior horn cells, with both inclusion and vacuolation pathologies [52, 59].
Neuropathological findings of G1H-G93A mice as a gold-standard ALS model. a The number of anterior horn cells at 90 days of age is not decreased significantly; approximately 10 anterior horn cells can are observed, although an anterior horn cell with intracytoplasmic vacuolation is evident (arrow). b The numbers of anterior horn cells at 100 days of age are slightly decreased; about eight anterior horn cells can be seen, whereas abundant vacuoles are observed. c At 120 days, when the mouse is quadriplegic and moribund, there is severe loss of anterior horn cells and prominent inclusion pathology (arrowheads), although the vacuolation pathology is less marked than that at disease onset (in Fig. b). There is an inverse correlation between the number of vacuoles and the number of inclusions. a–c H&E. Scale bar a (also for b, c) 200 μm. d Light micrograph of a round neuronal LBHI (arrow) in a spinal anterior horn cell of a G1H-G93A mouse. This LBHI is observed in the cytoplasm of an anterior horn cell, and is composed of eosinophilic core with paler peripheral halo. Vacuolation pathology is evident in the neuropil. H&E. Scale bar d 20 μm. e-h Immunostaining with antibody against human SOD1 in the spinal cord anterior horn of the G1H-G93A mouse. This antibody (MBL, Nagoya, Japan) recognizes only human SOD1, i.e., mutant human G93A-SOD1. e Littermate mouse, showing no expression of G93A-SOD1. f In a 90-day-old G1H-G93A mouse, G93A-SOD1 is expressed mainly in the neuropil, and sometimes expressed intensely within the rims of vacuoles in the neuropil and motor neuron cytoplasm (arrows). However, it is not expressed in the motor neurons. g In a 110-day-old G1H-G93A mouse, G93A-SOD1 is expressed within the motor neuron cytoplasm in addition to the vacuole rims in the neuropil (arrows). h: In a 120-day-old G1H-G93A mouse, LBHIs are strongly positive for G93A-SOD1 (arrows). Scale bar e (also for f–h) 10 μm
Among transgenic rats with human mutant SOD1 [2, 52, 59, 72], H46R line-4 rats develop motor deficits at approximately 145 days of age, whereas G93A line-39 rats show clinical signs at around 125 days. The number of anterior horn cells in H46R rats at 110 days is not significantly decreased in comparison with age-matched littermates. The anterior horn cells of H46R rats at 135 days are slightly decreased in numbers and show inclusions, whereas at 160, 170 and over 180 days, the anterior horn cells are markedly decreased, and show severe inclusion pathology, featuring neuronal LBHIs and Ast-HIs. With respect to G93A rats, the number of anterior horn cells at 70, 90 and 110 days of age is almost the same as that in age-matched littermates, although at 90 and 110 days of age there is marked vacuolation pathology. At 130, 150 and over 180 days of age, there is marked loss of anterior horn cells, along with both inclusion and vacuolation pathology.
Invariably, the core pathology of rodents carrying mutant SOD1 is the lower motor neuron degeneration/death. As far as can be clarified from the original literature, however, the pathological lesions of G93A mice [19–21, 36] are distributed mainly in the spinal cord anterior horn, brainstem motor nuclei, brainstem reticular formation, dorsal motor nucleus of the vagus nerve, red nucleus, interpeduncular nucleus, and substantia nigra. In G37R mice [98], the areas of degeneration are mainly the spinal cord anterior horn, brainstem motor nuclei, and ventral and lateral white matter of the spinal cord. Vacuole formation is also evident in the olfactory bulb, pyriform cortex, striatum, thalamus, hypothalamus, and choroid plexus. G85R mice show severe neuronal abnormalities in the spinal cord ventral motor neurons, including small neurons near the central canal, and rare interneurons of the dorsal horns, as well as brainstem neurons, especially in the pons [10]. G86R mice show a pronounced motor neuron loss within the spinal cord ventral horns [81]. In the brainstem motor neurons, G86R mice show severe motor neuron depletion in the facial nuclei, while the oculomotor and hypoglossal nuclei show less extreme involvement [75]. From a neurochemical viewpoint, G86R mice exhibit vulnerability of the spinal cord motor neurons, which are positive for pNFP, CAT and calretinin proteins [68]. In H46R-4 and G93A-39 rats [2, 72], the central nervous system lesions are located in the spinal cord ventral horns and brainstem motor nuclei, but no pathology is evident in the cerebral cortex and cerebellum. G93A-26H rats are pathologically similar to G93A-39 rats, showing motor neuron loss with prominent vacuolation [43].
Since the pathology of ALS rodents bearing mutant SOD1 is expressed primarily as lower motor neuron degeneration/death, the number of reports describing upper motor neuron pathology in these models is limited in comparison with more frequent focusing on lower motor neuron pathology. This can be explained mainly in terms of the anatomical difference in the corticospinal tract system between humans and rodents: the main corticospinal tracts in the human spinal cord are the lateral and anterior columns, while the main spinal cord pyramidal tracts in rodents are the dorsal columns. Another major consideration is that even at necropsy with end-stage pathology, SOD1-mutated FALS, which is the prototype for these models, demonstrates only slight or mild corticospinal tract degeneration. Although there are a relatively few reports, some are pertinent. One group has reported that 110-day-old G1H-G93A mice demonstrate degeneration of the corticospinal and bulbospinal systems, in which 53% of corticospinal, 41% of bulbospinal and 43% of rubospinal neurons are lost (the bulbospinal neuron system in mice comprises three systems: rubospinal, vestibulospinal and reticulospinal neurons) [103]. Another group has reported that G85R mice at the end stage show progressive axonal degeneration of corticospinal tracts in the dorsal and lateral columns of the spinal cord [101].
Inclusion pathology
Ever since Hirano et al. [42] emphasized the presence of neuronal LBHIs in the anterior horn cells of FALS patients with posterior column involvement in 1967, and the author discovered Ast-HIs in 1996 [53], LBHI/Ast-HI have been considered pathognomonic features of mutant SOD1-linked FALS with posterior column involvement, which is the prototype form of human mutant SOD1 transgenic rodents, and in these transgenic rodents neuronal LBHIs are frequently observed in the soma (Fig. 3d) and neurites, although rarely in axons. Although Ast-HIs are sometimes found in only long-surviving patients with SOD1-mutated FALS [53, 54], they are frequently seen in G85R mice and H46R rats as well as in both G1H-/G1L-G93A mice and G93A rats at terminal stage [56, 59].
The author has examined six different lines of G1H-/G1L-G93A mice, G85R-148 mice, L84V mice, and H46R-4 and G93A-39 rats by electron microscopy, and all show almost identical ultrastructural features of neuronal LBHIs and Ast-HIs. Interestingly, neuronal LBHIs observed in both SOD1-mutated rodents and humans have a similar ultrastructure, being composed mainly of randomly oriented granule-coated fibrils approximately15–25 nm in diameter and granular materials. Ast-HIs seen in both SOD1-mutated rodents and humans also have the same ultrastructure. Therefore, the essential ultrastructural common components of neuronal LBHIs and Ast-HIs in SOD1-mutated rodents and humans are granule-coated fibrils about 15–25 nm in diameter and granular materials. As the inclusions develop, the granule-coated fibrillar component increases and the amount of granular material decreases, suggesting that the former might be derived from the latter [57].
It is of considerable interest that LBHIs/Ast-HIs observed in SOD1-mutated rodents are light- and electron-microscopically identical to those in patients with SOD1-mutated FALS. The presence of LBHI/Ast-HI is a morphological hallmark of cells affected by mutant SOD1 [58], and the formation of LBHI/Ast-HI is reported to be correlated with disease severity and progression [90]. However, the mechanism by which SOD1 mutation in vivo leads to the formation of 15–25-nm granule-coated fibrils as an essential component of LBHI/Ast-HI remains poorly defined. An important clue for explaining the formation of the granule-coated fibrils as an ultrastructural hallmark of mutant SOD1 has been reported: the same granule-coated fibrils as those in SOD1-mutated cells in vivo are induced by endoplasmic reticulum (ER) stress in vitro using neuroblastoma cells overexpressing human mutant L84V SOD1 [100]. Transgenic mice with L84V SOD1 show aberrant aggregation of the ER in association with early-stage neuronal LBHIs, suggesting that the LBHIs might arise as a result of ER dysfunction [100]. Collectively, the presence of LBHI/Ast-HI is a light-microscopical hallmark of SOD1-mutated cells, and the 15–25-nm granule-coated fibrils as an essential component of the LBHI/Ast-HI provide ultrastructural authentication of SOD1-mutated cells. In marked contrast, Bunina bodies are not found in the transgenic rodents bearing mutant SOD1.
Vacuolation pathology
With regard to vacuolation pathology, although transgenic rodents expressing mutant SOD1 exhibit vacuoles of various sizes in neurons and neuropil, similar features are not evident in autopsy cases of mutant SOD1-related FALS, which is the prototype of the mutant SOD1 transgenic rodent. This vacuolation pathology is also undetectable in mutant SOD1-unrelated FALS and SALS. Ultrastructurally, the vacuolation is evident in somata, dendrites and axons of motor neurons. At an early stage, these alterations occur in the rough ER and mitochondria. In particular, perinuclear vacuoles in somata at the early stage are derived from dilated ER cisternae. As the disease progresses, the number of mitochondria-derived vacuoles increases, while the number of vacuoles originating from the ER decreases. Mitochondria-derived vacuoles originate through expansion of the mitochondrial intermembrane space and extension of the outer mitochondrial membrane [40]. These vacuoles are apparently more abundant at disease onset, and decline thereafter; vacuole formation itself is reported to be related to disease onset rather than disease progression [61]. There are abundant vacuoles and few LBHIs in the early course of degeneration in G1L-G93A and G1H-G93A mice (Fig. 3b), and mice at the terminal stage show less abundant vacuoles and many LBHIs (Fig. 3c), i.e., there is a significant inverse correlation between the numbers of vacuoles and LBHIs [91]. Another important finding in G1H- and G1L-G93A mice at the presymptomatic stage is fragmentation of the Golgi apparatus in the spinal cord anterior horn cells [69].
Contribution of mutant SOD1 in each cell type
Transgenic rodents overexpress human mutant G93A SOD1 in all cells because the transgene is driven by a non-cell-specific endogenous promoter. On the other hand, there are transgenic mice in which mutant SOD1 expression is driven by a neuron-specific promoter such as the neurofilament light chain. In transgenic mice whose anterior horn cells specifically overexpress mutant SOD1, neither motor neuron impairment nor degeneration is evident [80]. Transgenic mice that overexpress the mutant SOD1 transgene in neurons after birth also do not show motor neuron pathology [65]. Some other types of transgenic mice overexpress mutant G86R SOD1 only in astrocytes under control of the GFAP promoter. Despite the fact that these mice develop astrocytosis, they show no motor neuron degeneration and develop normally [34]. Although neurons or astrocytes play very important role in ALS pathogenesis, it is of considerable interest that mutant SOD1, when over expressed either in neurons or astrocytes, does not sufficiently contribute to the onset of ALS. In culture study, conversely, astrocytes expressing mutated SOD1 kill spinal primary and embryonic mouse stem cell-derived motor neurons [73]. In addition to neurons and astrocytes, microglia are closely related to neuron injury not only in ALS but also other neurodegenerative disorders. Approaches such as the use of a deletable mutant SOD1 transgene have demonstrated that diminishing mutant SOD1 within microglia has little effect on the early disease phase but sharply slows later disease progression: i.e., SOD1 mutated motor neurons are a determinant of onset and early disease, and mutant accumulation within microglia accelerates disease progression [9]. Interestingly, microglia themselves have a double-edged sword effect; wild-type microglia can extend the survival of G93A mice with PU.1 knockout mice (which are unable to develop myeloid and lymphoid cells) by using bone marrow transplantation [6]. Since retraction of motor axons from synaptic connections to muscle is among the earlier presymptomatic morphological findings in SOD1-mutated mice, muscle itself is also a likely primary source of mutant SOD1 toxicity. However, use of a deletable mutant gene to eliminate mutant SOD1 from muscle does not affect disease onset or survival: SOD1-mutant-mediated damage within muscles is not a significant contributor to non-cell-autonomous pathogenesis in ALS [66].
Relation between mutant SOD1 and disease progression
Unlike patients with SOD1-mutated FALS, transgenic rodents bear both human mutant SOD1 and native endogenous rodent SOD1. Native endogenous rodent SOD1 catalyzes the conversion of the superoxide radical to hydrogen peroxide and molecular oxygen. Even overinduced human mutant SOD1 also detoxifies the superoxide radical, which is a source of reactive oxygen species generated from aerobic organisms, and protects cells, including motor neurons, from oxidative injury. Based on the gain-of-function theory, human mutant SOD1 itself acts as a cytotoxic factor, and in G1H-G93A mice human mutant G93A-SOD1 shows cytotoxicity for motor neurons. In 90-day-old G1H-G93A mice that show no significant motor neuron loss and only slight vacuolation pathology, G93A-SOD1 is already present but its expression level is not marked, and immunohistochemically it is expressed mainly in the neuropil, sometimes being expressed intensely within the rims of vacuoles in the neuropil and motor neuron cytoplasm (Fig. 3e,f). In contrast, motor neurons do not express mutant SOD1 (Fig. 3e,f). In 100-day-old G1H-G93A mice that demonstrate a slightly decreased number of motor neurons and prominent vacuolation pathology, mutant G93A-SOD1 is highly expressed in comparison with the level at 90 days of age. At 110 days, the mice that show loss of anterior horn cells with some inclusions and vacuole formation also exhibit high expression of mutant G93A-SOD1, which morphologically is located within the motor neuron cytoplasm and the vacuole rims in the neuropil (Fig. 3e,g). End-stage G1H-G93A mice that show severe motor neuron loss as well as vacuolation and inclusion pathologies demonstrate high expression of mutant G93A-SOD1, and immunohistochemically mutant G93A-SOD1 is aggregated and sequestered into the LBHIs, which are strongly positive for mutant G93A-SOD1 (Fig. 3e, h). Considered in connection with the abundance of neuropil vacuoles and few LBHIs at the early stage, and the fact that mice at the terminal stage show many LBHIs and less abundant vacuoles, as well as the accumulation of mutant G93A-SOD1 in vacuoles at the early stage and marked aggregation of mutant G93A-SOD1 in LBHIs at the late stage, it is possible that cytotoxic mutant G93A-SOD1 within vacuoles at the early stage leaks into the neurons and then aggregates within neurons as LBHIs with disease progression. Along with disease progression, there is a breakdown of cytotoxic mutant SOD1 sequestration in vacuoles, and the mutant SOD1 aggregates in motor neurons, resulting in their degeneration/death.
Development of rats with human mutant SOD1
The ultimate aim of developing transgenic rodents expressing human mutant SOD1 are as follows: to gain an understanding of the mechanism of motor neuron death in the presence of mutant SOD1, and to test new ALS therapies. Freshly obtained mouse spinal cord including the nerve roots, cauda equina and filum terminale, weighs only about 110 mg and is approximately 4 cm in length. In order to perform more extensive analysis of the mechanism of motor neuron death and to devise new ALS therapies that are difficult or impossible to explore using the small spinal cord of the mouse, transgenic rats expressing human mutant SOD1 have been developed [2, 43, 72]. As rats are a larger species than mice, they are easier to use in studies involving manipulations of spinal fluid (e.g., implantation of intrathecal catheters for chronic therapeutic studies and CSF sampling) and the spinal cord (e.g., direct administration of viral and cell-mediated therapies).
Relationship between ALS and TDP-43
The 43-kDa TAR DNA-binding protein (TDP-43) is localized to the nucleus. Originally, TDP-43 was identified as a component of ubiquitinated inclusions in frontotemporal lobar degeneration with ubiquitinated inclusions (FTLD-U) and ALS [3, 74]. Analyses of TDP-43 immunohistochemistry in SALS (two patients), mutant SOD1-unrelated FALS (two patients) and ALSD (one patient) have shown TDP-43-immunoreactive inclusions such as SLI/RHI in the anterior horn cells of the spinal cord [95]; as mentioned above, SLI/RHI are characteristic morphological structures in ALS, and mutant SOD1-unrelated FALS is neuropathologically indistinguishable from SALS. TDP-43 immunoreactivity has also been detected in the motor neurons of the hypoglossal nucleus in 4 patients with FTLD-MND/ALSD and 11 patients with ALS; TDP-43-positive structures include SLI/RHI [26]. Although Bunina bodies are a pathognomonic structure in SALS, mutant SOD1-unrelated FALS and FTLD-MND/ALSD, Bunina bodies themselves are negative for TDP-43 [95]. With regard to mutant SOD1-related motor neuron death, LBHIs, which are characteristic structures in mutant SOD1-related FALS with A4T (one patient) and D101Y (one patient) reportedly do not express TDP-43 [95]. In SALS (one patient) as well as mutant SOD1-related FALS with A4T (one patient) and I113T (one patient), TDP-43 is mislocalized from the nucleus to the cytoplasm [83]. Especially, one case of SALS was reported to show ubiquitin-positive RHIs with TDP-43 staining pattern [83]. By marked contrast, human mutant G93A, G37R and G85R SOD1-transgenic mice do not show any TDP-43 abnormalities including either TDP-43-positive inclusions or TDP-43 mislocalization [83]. It could be stated that in general, TDP-43 contributes to mutant SOD1-unlinked motor neuron degeneration, whereas mutant SOD1-linked motor neuron degeneration may not be essentially related to TDP-43 abnormality; in particular, human mutant SOD1 transgenic mice do not show TDP-43 abnormality [83].
Mouse models of ALS2
As already mentioned, SOD1 mutations have been identified as a cause of autosomal dominant FALS [22, 84]. Mutation in a second ALS-related (ALS2) gene has also been identified as the cause of a rare autosomal recessive form of juvenile-onset ALS, also referred to as ALS2 [7, 37, 102], as well as juvenile-onset primary lateral sclerosis (PLS) [37], and infantile-onset ascending hereditary spastic paralysis (HSP) [24, 29, 35]. In humans, the ALS2 gene is located on chromosome 2 at position 33.2, and encodes a protein called ALS2 protein or alsin. ALS2 protein is produced in a wide range of normal tissues, with the highest amounts in the brain and spinal cord. ALS2 protein is composed of 1,657 amino acids with three predicted guanine nucleotide exchange factor (GEF)-like domains [37, 102]: an N-terminal regulator of chromatin condensation (RCC 1)-like domain (RLD) homologous to GEF for Ran GTPase [77], middle Db1 homology (DH) and pleckstrin homology (PH) (DH/PH)-like domains resembling GEF for Rho GTPase [87], and a C-terminal vacuolar protein sorting 9 (VPS9)-like domain similar to GEF for Rab5 GTPase [79]. This ALS2 protein is particularly abundant in motor neurons. ALS2 protein is preferentially associated with the cytoplasmic face of the endosomal membrane [79]. Although the function of ALS2 protein in motor neurons is unclear, it may play an important role in regulating cell membrane organization and the movement of molecules within motor neurons. Therefore, it would be expected to play a role in the development of axons and dendrites. It is unclear how and why loss of ALS2 protein function causes the ALS2-linked diseases: ALS2, juvenile-onset PLS, and infantile-onset ascending HSP. In order to gain insight into the physiological role of ALS2 protein and the pathogenesis of ALS2-linked diseases, four types of ALS2 knockout mice have been successfully developed.
The ALS2 knockout mice with disruption of exon 3 of the murine ALS2 gene reported by Cai et al. [13] show a higher anxiety response as well as an age-dependent deficit of motor coordination and learning. Histopathologically and biologically, ALS2 knockout mice are characterized by a lack of neuropathological abnormality, no alteration of peripheral nerve conduction or electromyography features, susceptibility to oxidative stress, and increased susceptibility to glutamate receptor-mediated excitotoxicity. In the ALS2 knockout mice reported by Hadano et al. [38], exon 3 of the murine ALS2 gene is disrupted by inserting a stop codon. These mice demonstrate no obvious developmental, reproductive or motor abnormalities. However, histopathologically and biologically, they are characterized by an age-dependent decrease in the size and number of ventral motor axons and cerebellar Purkinje cells, astrocytosis and microglial activation in the spinal cord and brain, motor unit remodeling and fiber redistribution in skeletal muscle, and slightly affected endosomal dynamics. ALS2 knockout mice with disruption of both exons 3 and 4 of the murine ALS2 gene reported by Devon et al. [25] show mild hypoactivity. Neuropathologically, at the age of 12 months, they show significantly smaller cortical motor neurons, and in addition, marked diminution of Rab5-dependent endosome fusion activity and disturbance in endosomal transport of the insulin-like growth factor 1 and BDNF receptors. ALS2 knockout mice with disruption of exon 4 of the murine ALS2 gene reported by Yamanaka et al. [101] demonstrate slowed movement without muscle weakness and progressive axonal degeneration in the lateral spinal cord. Significantly, all four of these ALS2 knockout murine models show no human ALS2-like symptoms and are not neurologically analogous to humans with ALS2.
Among previously reported human patients with ALS2-linked disease, the members of a Tunisian family with 138delA ALS2 gene mutation showed development of spasticity in all limbs between 3–10 years of age [7, 37, 102], and their clinical symptoms might be classifiable as part of a spectrum of HSP rather than typical ALS. The members of a Kuwaiti family with 1425_1426delAG ALS2 gene mutation showed infantile-onset spastic paralysis without lower motor neuron involvement at 1–2 years of age [37]. Members of a Saudi Arabian family with1867_1868delCT ALS2 gene mutation developed PLS between 1–2 years of age [33, 102]. Up to now, eight additional ALS2-linked diseases have been reported [24, 29, 30, 35, 62], and a major common characteristic is infantile-onset spastic paralysis, reflecting upper motor disturbance. Although lower motor neuron impairment has been reported in a limited number of patients with ALS2 gene mutation, the majority of ALS2 gene mutations appear to be linked to upper motor neuron diseases from the viewpoint of human clinical data of 11-type ALS2-linked diseases. Although to the author’s knowledge there has been no reported autopsy case of ALS2, detailed neuropathological data from human ALS2 autopsy cases would clarify this point. In this context, although the author is unable to address the similarities and differences between human ALS2 and ALS2 animal models from a neuropathological viewpoint, it might be concluded that data from ALS2 knockout mice and ALS2-linked diseases mentioned above would become more valuable for clarifying the pathogenesis of human ALS2 if detailed human ALS2 autopsy data were also available.
Animal models based on cytoskeletal abnormalities
Animal models based on neurofilament abnormalities
The neuron cytoskeleton consists of three major filaments: actin microfilaments, microtubules, and neurofilaments. Neurofilaments biochemically comprise three different isoforms known as neurofilament triplet proteins: light subunit (68 kDa), medium subunit (160 kDa), and heavy subunit (200 kDa). Ultrastructurally, neurofilaments are approximately 10 nm in diameter, but in cross-section they appear tubular in structure with a narrow central electron-lucent core, and have fine side arms. Their size places them in the so-called “intermediate filaments” morphologically.
Neurofilament-lacking mice
Mice lacking any of the neurofilament triplet protein genes show no developmental problems [82]. However, mice lacking the neurofilament light subunit show a lack of intermediate filament structure, axonal hypotrophy, and aggregation of neurofilament medium and heavy subunit in motor neurons, although significant motor neuron loss is not evident [5]. Mice lacking the neurofilament medium subunit show axonal atrophy without significant motor neuron loss, and reduce contents of neurofilament light subunit [28]. Mice without the neurofilament heavy subunit also exhibit axonal atrophy without significant motor neuron loss [47]. Therefore, model mice lacking any of the neurofilament triplet protein subunits are not compatible with human ALS patients, as no significant motor neuron loss is evident.
Transgenic mice expressing the human wild-type neurofilament gene
Transgenic mice expressing the human wild-type neurofilament heavy chain gene show defective axonal transport and axonal atrophy in association with ultrastructural diminution of cytoskeletal components, the smooth endoplasmic reticulum, and mitochondria [16]. In mice showing a high level of expression, neurofilament aggregation is observed in the cytoplasm of neurons and proximal axons [17]. However, this transgenic mouse model shows no significant motor neuron loss [16]. Like neurofilament heavy chain-type transgenic mice, the neurofilament light chain-type transgenic mice show accumulation of neurofilaments in the neurons and axonal degeneration without significant motor neuron loss [99]. Therefore, transgenic mouse models expressing the human wild-type neurofilament gene bear no histopathological resemblance to human ALS in terms of significant motor neuron loss.
Transgenic mice expressing the human mutant neurofilament light chain gene with the L394P
Transgenic mice expressing the human mutant neurofilament light chain gene with the L394P develop neurological symptoms of muscle weakness. Unlike mice expressing the human wild-type neurofilament gene, these mice show significant motor neuron loss [64]. From this viewpoint, this mouse model is closely similar to human ALS on the basis of neurofilament pathology, although in human ALS there is no mutation of the neurofilament light chain gene with L394P [27].
Transgenic mice expressing the peripherin gene
Peripherin is a 58-kDa type III intermediate filament protein, which has been reported to be a component of ubiquitinated inclusion bodies in motor neurons of ALS patients [39]. As the name “peripherin” indicates, the protein exists mainly in the peripheral nervous system, and only a small amount is expressed with a selective distribution in the central nervous system. Overexpression of peripherin in transgenic mice leads to loss of spinal cord anterior horn cells and formation of inclusions that are immunoreactive for peripherin [4].
Transgenic mice expressing the dynamitin gene
The dynein/dynactin-complex is a type of motor protein responsible for minus-end-directed movement along the microtubule and plays an important role in fast retrograde-related axonal transport. Supporting the hypothesis that impairment of retrograde axonal transport causes motor neuron death, point mutations of the p150 subunit of the dynactin gene have been reported in ALS patients [70]. Experimentally, on the basis of this retrograde axonal transport impairment theory, mice overexpressing dynamitin, which is a subunit of dynactin, have been produced, and these mice show disruption of the dynein/dynactin complex, leading to inhibition of retrograde axonal transport. Histologically, such dynamitin-overexpressing mice show motor neuron loss [63].
Concluding remarks
Human ALS pathology exhibits a variety of cytopathological features including Bunina bodies, SLIs/RHIs, LBHIs/Ast-HIs, and NFCIs in addition to motor neuron degeneration. Among various rodent models of ALS, rodents with mutant SOD1 recapitulate motor neuron degeneration and SOD1-immunoreactive LBHIs/Ast-HIs, both of them found in SOD1-mutated FALS patients. Even with these similarities, human ALS pathology is different from that of rodents carrying SOD1 mutation, because (1) ALS is not a single entity but rather a heterogeneous syndrome, (2) relevant anatomical structures are different between humans and rodents, and (3) human pathology generally deals with only the terminal stage of the disease. In spite of these differences, motor neuron degeneration in rodent models provides us with opportunities to analyze the motor neuron degeneration process in detail and even to test therapeutic attempts. It is necessary to be aware not only of the similarities but also of differences between these ALS models and human ALS, because they are complementary.
References
Aoki M (2004) Amyotrophic lateral sclerosis: recent insights from transgenic animal models with SOD1 mutations. Rinsho Shinkeigaku 44:778–791
Aoki M, Kato S, Nagai M, Itoyama Y (2005) Development of a rat model of amyotrophic lateral sclerosis expressing a human SOD1 transgene. Neuropathology 25:365–370
Arai T, Hasegawa M, Akiyama H, Ikeda K, Nonaka T, Mori H, Mann D, Tsuchiya K, Yoshida M, Hashizume Y, Oda T (2006) TDP-43 is a component of ubiquitin-positive inclusions in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Biochem Biophys Res Commun 351:602–611
Beaulieu JM, Nguyen MD, Julien JP (1999) Late onset death of motor neurons in mice over-expressing wild-type peripherin. J Cell Biol 147:531–544
Beaulieu JM, Jacomy H, Julien JP (2000) Formation of intermediate filament protein aggregates with disparate effects in two transgenic mouse models lacking the neurofilament light subunit. J Neurosci 20:5321–5328
Beers DR, Henkel JS, Xiao Q, Zhao W, Wang J, Yen AA, Siklos L, McKercher SR, Appel SH (2006) Wild-type microglia extend survival in PU.1 knockout mice with familial amyotrophic lateral sclerosis. Proc Natl Acad Sci USA 103:16021–16026
Ben Hamida M, Hentati F, Ben Hamida C (1990) Hereditary motor system diseases (chronic juvenile amyotrophic lateral sclerosis). Conditions combining a bilateral pyramidal syndrome with limb and bulbar amyotrophy. Brain 113:347–363
Bendotti C, Carrì MT (2004) Lessons from models of SOD1-linked familial ALS. Trends Mol Med 10:393–400
Boillée S, Yamanaka K, Lobsiger CS, Copeland NG, Jenkins NA, Kassiotis G, Kollias G, Cleveland DW (2006) Onset and progression in inherited ALS determined by motor neurons and microglia. Science 312:1389–1392
Bruijn LI, Becher MW, Lee MK, Anderson KL, Jenkins NA, Copeland NG, Sisodia SS, Rothstein JD, Borchelt DR, Price DL, Cleveland DW (1997) ALS-linked SOD1mutant G85R mediates damage to astrocytes and promotes rapidly progressive disease with SOD1-containing inclusions. Neuron 18:327–338
Bruijn LI, Houseweart MK, Kato S, Anderson KL, Anderson SD,Ohama E, Reaume AG, Scott RW, Cleveland DW (1998) Aggregation and motor neuron toxicity of an ALS-linked SOD1 mutant independent from wild-type SOD1. Science 281:1851–1854
Bunina TL (1962) On intracellular inclusions in familial amyotrophic lateral sclerosis. Zh Nevropathol Psikhiatr Im S S Korsakova 62:1293–1299
Cai H, Lin X, Xie C, Laird FM, Lai C, Wen H, Chiang HC, Shim H, Farah MH, Hoke A, Price DL, Wong PC (2005) Loss of ALS2 function is insufficient to trigger motor neuron degeneration in knock-out mice but predisposes neurons to oxidative stress. J Neurosci 25:7567–7574
Cervenakova L, Protas II, Hirano A, Votiakov VI, Nedzved MK, Kolomiets ND, Taller I, Park KY, Sambuughin N, Gajdusek DC, Brown P, Goldfarb LG (2000) Progressive muscular atrophy variant of familial amyotrophic lateral sclerosis (PMA/ALS). J Neurol Sci 177:124–130
Charcot JM (1874) De la sclérose latérale amyotrophique. Prog Méd 2: 325–327, 341–342, 453–455
Collard JF, Côté F, Julien JP (1995) Defective axonal transport in a transgenic mouse model of amyotrophic lateral sclerosis. Nature 375:61–64
Côté F, Collard JF, Juien JP (1993) Progressive neuronopahty in transgenic mice expressing the human neurofilament heavy gene: a mouse model of amyotrophic lateral sclerosis. Cell 73:35–46
Cudkowicz ME, McKenna-Yasek D, Chen C, Hedley-Whyte ET, Brown RH Jr (1998) Limited corticospinal tract involvement in amyotrophic lateral sclerosis subjects with the A4V mutation in the copper/zinc superoxide dismutase gene. Ann Neurol 43:703–710
Dal Canto MC, Gurney ME (1995) Neuropathological changes in two lines of mice carrying a transgene for mutant human Cu, Zn SOD, and in mice overexpressing wild type human SOD: a model of familial amyotrophic lateral sclerosis (FALS). Brain Res 676:25–40
Dal Canto MC, Mourelatos Z, Gonatas NK, Chiu A, Gurney ME (1996) Neuropathological changes depend on transgene copy numbers in transgenic mice for mutant human Cu, Zn superoxide dismutase (SOD). In: Nakano I, Hirano A (eds) Amyotrophic lateral sclerosis. progress and perspectives in basic research and clinical application. XIth TMIN international symposium, Tokyo, international congress series 1104. Elsevier, Tokyo, pp 331–338
Dal Canto MC, Gurney ME (1997) A low expressor line of transgenic mice carrying a mutant human Cu, Zn superoxide dismutase (SOD1) gene develops pathological changes that most closely resemble those in human amyotrophic lateral sclerosis. Acta Neuropathol (Berl) 93:537–550
Deng HX, Hentati A, Tainer JA, Iqbal Z, Cayabyab A, Hung WY, Getzoff ED, Hu P, Herzfeldt B, Roos RP, Warner C, Deng G, Soriano E, Smyth C, Parge HE, Ahmed A, Roses AD, Hallewell RA (1993) Amyotrophic lateral sclerosis and structural defects in Cu, Zn superoxide dismutase. Science 261:1047–1051
Deng HX, Shi Y, Furukawa Y, Zhai H, Fu R, Liu E, Gorrie GH, Khan MS, Hung WY, Bigio EH, Lukas T, Dal Canto MC, O’Halloran TV, Siddique T (2006) Conversion to the amyotrophic lateral sclerosis phenotype is associated with intermolecular linked insoluble aggregates of SOD1 in mitochondria. Proc Natl Acad Sci USA 103:7142–7147
Devon RS, Helm JR, Rouleau GA, Leitner Y, Lerman-Sagie T, Lev D, Hayden MR (2003) The first nonsense mutation in alsin results in a homogeneous phenotype of infantile-onset ascending spastic paralysis with bulbar involvement in two siblings. Clin Genet 64:210–215
Devon RS, Orban PC, Gerrow K, Barbieri MA, Schwab C, Cao LP, Helm JR, Bissada N, Cruz-Aguado R, Davidson TL, Witmer J, Metzler M, Lam CK, Tetzlaff W, Simpson EM, McCaffery JM, El-Husseini AE, Leavitt BR, Hayden MR (2006) Als2-deficient mice exhibit disturbances in endosome trafficking associated with motor behavioral abnormalities. Proc Natl Acad Sci USA 103:9595–9600
Dickson DW, Josephs KA, Amador-Ortiz C (2007) TDP-43 in differential diagnosis of motor neuron disorders. Acta Neuropathol 114:71–79
Doble A, Kennel P (2000) Animal models of amyotrophic lateral sclerosis. Amyotroph Lateral Scler Other Motor Neuron Disord 1:301–312
Elder GA, Friedrich VL Jr, Bosco P, Kang C, Gourov A, Tu PH, Lee VM, Lazzarini RA (1998) Absence of the mid-sized neurofilament subunit decreases axonal calibers, levels of light neurofilament (NF-L), and neurofilament content. J Cell Biol 141:727–739
Eymard-Pierre E, Lesca G, Dollet S, Santorelli FM, di Capua M, Bertini E, Boespflug-Tanguy O (2002) Infantile-onset ascending hereditary spastic paralysis is associated with mutations in the alsin gene. Am J Hum Genet 71: 518–527
Eymard-Pierre E, Yamanaka K, Haeussler M, Kress W, Gauthier-Barichard F, Combes P, Cleveland DW, Boespflug-Tanguy O (2006) Novel missense mutation in ALS2 gene results in infantile ascending hereditary spastic paralysis. Ann Neurol 59:976–980
Friedlander RM, Brown RH, Gagliardini V, Wang J, Yuan J (1997) Inhibition of ICE slows ALS in mice. Nature 388:31
Fujimura H, Sumi H, Fukada K, Takayasu S, Sakoda S, Nakanishi T, Shimuzu A, Kato S (2002) Stability of mutant superoxide dismutase-1 and incidence of Lewy body-like hyaline inclusions in familial amyotrophic lateral sclerosis (in Japanese with English abstract) Rinsho Shinkeigaku 42:1283
Gascon GG, Chavis P, Yaghmour A, Stigsby B, Shums A, Ozand P, Siddique T (1995) Familial childhood primary lateral sclerosis with associated gaze paresis. Neuropediatrics 26:313–319
Gong YH, Parsadanian AS, Andreeva A, Snider WD, Elliott JL (2000) Restricted expression of G86R Cu/Zn superoxide dismutase in astrocytes results in astrocytosis but does not cause motoneuron degeneration. J Neurosci 20:660–665
Gros-Louis F, Meijer IA, Hand CK, Dubé MP, MacGregor DL, Seni MH, Devon RS, Hayden MR, Andermann F, Andermann E, Rouleau GA (2003) An ALS2 gene mutation causes hereditary spastic paraplegia in a Pakistani kindred. Ann Neurol 53:144–145
Gurney ME, Pu H, Chiu AY, Dal Canto MC, Polchow CY, Alexander DD, Caliendo J, Hentati A, Kwon YW, Deng H-X, Chen W, Zhai P, Sufit RL, Siddique T (1994) Motor neuron degeneration in mice that express a human Cu, Zn superoxide dismutase mutation. Science 264:1772–1775
Hadano S, Hand CK, Osuga H, Yanagisawa Y, Otomo A, Devon RS, Miyamoto N, Showguchi-Miyata J, Okada Y, Singaraja R, Figlewicz DA, Kwiatkowski T, Hosler BA, Sagie T, Skaug J, Nasir J, Brown RH Jr, Scherer SW, Rouleau GA, Hayden MR, Ikeda JE (2001) A gene encoding a putative GTPase regulator is mutated in familial amyotrophic lateral sclerosis 2. Nat Genet 29:166–173
Hadano S, Benn SC, Kakuta S, Otomo A, Sudo K, Kunita R, Suzuki-Utsunomiya K, Mizumura H, Shefner JM, Cox GA, Iwakura Y, Brown RH Jr, Ikeda JE (2006) Mice deficient in the Rab5 guanine nucleotide exchange factor ALS2/alsin exhibit age-dependent neurological deficits and altered endosome trafficking. Hum Mol Genet 15:233–250
He CZ, Hays AP (2004) Expression of peripherin in ubiquitinated inclusions of amyotrophic lateral sclerosis. J Neurol Sci 217:47–54
Higgins CMJ, Jung C, Xu Z (2003) ALS-associated mutant SOD1G93A causes mitochondrial vacuolation by expansion of the intermembrane space and by involvement of SOD1 aggregation and peroxisomes. BMC Neurosci 4:16
Hirano A (1965) Pathology of amyotrophic lateral sclerosis. In: Gajdusek DC, Gibbs CJ Jr, Alpers MP (eds) Slow, latent, and temperate virus infections. NINDB Monograph No.2 Washington: U.S. National Institute of Neurological Diseases and Blindness, National Institute of Health (U.S.), pp 23–37
Hirano A, Kurland LT, Sayre GP (1967) Familial amyotrophic lateral sclerosis. A subgroup characterized by posterior and spinocerebellar tract involvement and hyaline inclusions in the anterior horn cells. Arch Neurol 16:232–243
Howland DS, Liu J, She Y, Goad B, Maragakis NJ, Kim B, Erickson J, Kulik J, DeVito L, Psaltis G, DeGennaro LJ, Cleveland DW, Rothstein JD (2002) Focal loss of the glutamate transporter EAAT2 in a transgenic rat model of SOD1 mutant-mediated amyotrophic lateral sclerosis (ALS). Proc Natl Acad Sci USA 99:1604–1609
Ince PG, Shaw PJ, Slade JY, Jones C, Hudgson P (1996) Familial amyotrophic lateral sclerosis with a mutation in exon 4 of the Cu/Zn superoxide dismutase gene: pathological and immunocytochemical changes. Acta Neuropathol 92:395–403
Ince PG, Tomkins J, Slade JY, Thatcher NM, Shaw PJ (1998) Amyotrophic lateral sclerosis associated with genetic abnormalities in the gene encoding Cu/Zn superoxide dismutase: molecular pathology of five new cases, and comparison with previous reports and 73 sporadic cases of ALS. J Neuropathol Exp Neurol 57:895–904
Inoue K, Fujimura H, Ogawa Y, Satoh T, Shimada K, Sakoda S (2002) Familial amyotrophic lateral sclerosis with a point mutation (G37R) of the superoxide dismutase 1 gene: a clinicopathological study. Amyotroph Lateral Scler Other Motor Neuron Disord 3:244–247
Jacomy H, Zhu Q, Couillard-Després S, Beaulieu JM, Julien JP (1999) Disruption of type IV intermediate filament network in mice lacking the neurofilament medium and heavy subunits. J Neurochem 73:972–984
Jonsson PA, Ernhill K, Andersen PM, Bergemalm D, Brännström T, Gredal O, Nilsson P, Marklund SL (2004) Minute quantities of misfolded mutant superoxide dismutase-1 cause amyotrophic lateral sclerosis. Brain 127:73–88
Jonsson PA, Graffmo KS, Brännström T, Nilsson P, Andersen PM, Marklund SL (2006) Motor neuron disease in mice expressing the wild type-like D90A mutant superoxide dismutase-1. J Neuropathol Exp Neurol 65:1126–1136
Kadekawa J, Fujimura H, Yanagihara T, Sakoda S (2001) A clinicopathological study of patient with familial amyotrophic lateral sclerosis associated with a two-base pair deletion in the copper/zinc superoxide dismutase (SOD1) gene. Acta Neuropathol 101:415
Katayama S, Watanabe C, Noda K, Ohishi H, Yamamura Y, Nishisaka T, Inai K, Asayama K, Murayama S, Nakamura S (1999) Numerous conglomerate inclusions in slowly progressive familial amyotrophic lateral sclerosis with posterior column involvement. J Neurol Sci 171:72–77
Kato M, Kato S, Abe Y, Nishino T, Ohama E, Aoki M, Itoyama Y (2006) Histological recovery of the hepatocytes is based on the redox system upregulation in the animal models of mutant superoxide dismutase (SOD)1-linked amyotrophic lateral sclerosis. Histol Histopathol 21:729–742
Kato S, Shimoda M, Watanabe Y, Nakashima K, Takahashi K, Ohama E (1996) Familial amyotrophic lateral sclerosis with a two base pair deletion in superoxide dismutase 1 gene: multisystem degeneration with intracytoplasmic hyaline inclusions in astrocytes. J Neuropathol Exp Neurol 55:1089–1101
Kato S, Hayashi H, Nakashima K Nanba E, Kato M, Hirano A, Nakano I, Asayama K, Ohama E (1997) Pathological characterization of astrocytic hyaline inclusions in familial amyotrophic lateral sclerosis. Am J Pathol 151:611–620
Kato S, Saito M, Hirano A, Ohama E (1999) Recent advances in research on neuropathological aspects of familial amyotrophic lateral sclerosis with superoxide dismutase 1 gene mutations: neuronal Lewy body-like hyaline inclusions and astrocytic hyaline inclusions. Histol Histopathol 14:973–989
Kato S, Horiuchi S, Liu J, Cleveland DW, Shibata N, Nakashima K, Nagai R, Hirano A, Takikawa M, Kato M, Nakano I, Ohama E (2000) Advanced glycation endproduct-modified superoxide dismutase-1 (SOD1)-positive inclusions are common to familial amyotrophic lateral sclerosis patients with SOD1 gene mutations and transgenic mice expressing human SOD1 with a G85R mutation. Acta Neuropathol 100:490–505
Kato S, Takikawa M, Nakashima K, Hirano A. Cleveland DW, Kusaka H, Shibata N, Kato M, Nakano I, Ohama E (2000) New consensus research on neuropathological aspects of familial amyotrophic lateral sclerosis with superoxide dismutase 1 (SOD1) gene mutations: Inclusions containing SOD1 in neurons and astrocytes. Amyotroph Lateral Scler Other Motor Neuron Disord 1:163–184
Kato S, Shaw P, Wood-Allum C, Leigh PN, Show C (2003) Amyotrophic lateral sclerosis. In: Dickson DW (ed) Neurodegeneration: the molecular pathology of dementia and movement disorders. ISN Neuropath, Basel, pp 350–368
Kato S, Kato M, Abe Y, Matsumura T, Nishino T, Aoki M, Itoyama Y, Asayama K, Awaya A, Hirano A, Ohama E (2005) Redox system expression in the motor neurons in amyotrophic lateral sclerosis (ALS): immunohistochemical studies on sporadic ALS, superoxide dismutase 1 (SOD1)-mutated familial ALS, and SOD1-mutated ALS animal models. Acta Neuropathol 110:101–112
Kokubo Y, Kuzuhara S, Narita Y, Kikugawa K, Nakano R, Inuzuka T, Tsuji S, Watanabe M, Miyazaki T, Murayama S, Ihara Y (1999) Accumulation of neurofilaments and SOD1-immunoreactive products in a patient with familial amyotrophic lateral sclerosis with I113T SOD1 mutation. Arch Neurol 56:1506–1508
Kong J, Xu Z (1998) Massive mitochondrial degeneration in motor neurons triggers the onset of amyotrophic lateral sclerosis in mice expressing a mutant SOD1. J Neurosci 18:3241–3250
Kress JA, Kühnlein P, Winter P, Ludolph AC, Kassubek J, Müller U, Sperfeld AD (2005) Novel mutation in the ALS2 gene in juvenile amyotrophic lateral sclerosis. Ann Neurol 58:800–803
LaMonte BH, Wallace KE, Holloway BA, Shelly SS, Ascaño J, Tokito M, Van Winkle T, Howland DS, Holzbaur EL (2002) Disruption of dynein/dynactin inhibits axonal transport in motor neurons causing late-onset progressive degeneration. Neuron 34:715–727
Lee MK, Marszalek JR, Cleveland DW (1994) A mutant neurofilament subunit causes massive, selective motor neuron death: implications for the pathogenesis of human motor neuron disease. Neuron 13:975–988
Lino MM, Schneider C, Caroni P (2002) Accumulation of SOD1 mutants in postnatal motoneurons does not cause motoneuron pathology or motoneuron disease. J Neurosci 22:4825–4832
Miller TM, Kim SH, Yamanaka K, Hester M, Umapathi P, Arnson H, Rizo L, Mendell JR, Gage FH, Cleveland DW, Kaspar BK (2006) Gene transfer demonstrates that muscle is not a primary target for non-cell-autonomous toxicity in familial amyotrophic lateral sclerosis. Proc Natl Acad Sci USA 103:19546–19551
Mochizuki Y, Mizutani T, Nakano R, Fukushima T, Honma T, Nemoto N, Takei K (2003) Clinical features and neuropathological findings of familial amyotrophic lateral sclerosis with an H43R mutation in Cu/Zn superoxide dismutase. Rinsho Shinkeigaku 43:491–449
Morrison BM, Gordon JW, Ripps ME, Morrison JH (1996) Quantitative immunocytochemical analysis of the spinal cord in G86R superoxide dismutase transgenic mice: neurochemical correlates of selective vulnerability. J Comp Neurol 373:619–631
Mourelatos Z, Gonatas NK, Stieber A, Gurney ME, Dal Canto MC (1996) The Golgi apparatus of spinal cord motor neurons in transgenic mice expressing mutant Cu, Zn superoxide dismutase becomes fragmented in early, preclinical stages of the disease. Proc Natl Acad Sci USA 93:5472–5477
Münch C, Sedlmeier R, Meyer T, Homberg V, Sperfeld AD, Kurt A, Prudlo J, Peraus G, Hanemann CO, Stumm G, Ludoph AC (2004) Point mutations of the p150 subunit of dynactin (DCTN1) gene in ALS. Neurology 63:724–726
Murayama S, Namba E, Nishiyama, Kitamura Y, Morita T, Nakashima K, Ishida T, Mizutani T, Kanazawa I (1997) Molecular pathological studies of familial amyotrophic lateral sclerosis. Neuropathology 17(Suppl):219
Nagai M, Aoki M, Miyoshi I, Kato M, Pasinelli P, Kasai N, Brown RH Jr, Itoyama Y (2001) Rats expressing human cytosolic copper-zinc superoxide dismutase transgenes with amyotrophic lateral sclerosis: associated mutations develop motor neuron disease. J Neurosci 21:9246–9254
Nagai M, Re DB, Nagata T, Chalazonitis A, Jessell TM, Wichterle H, Przedborski S (2007) Astrocytes expressing ALS-linked mutated SOD1 release factors selectively toxic to motor neurons. Nat Neurosci 10:615–622
Neumann M, Sampathu DM, Kwong LK, Truax A, Micsenyi MC, Chou TT, Bruce J, Schuck T, Grossman M, Clark CM, McCluskey LF, Miller BL, Masliah E, Mackenzie IR, Feldman H, Feiden W, Kretzschmar HA, Trojanowski JQ, Lee VM-Y (2006) Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 314:130–133
Nimchinsky EA, Young WG, Yeung G, Shah RA, Gordon JW, Bloom FE, Morrison JH, Hof PR (2000) Differential vulnerability of oculomotor, facial, and hypoglossal nuclei in G86R superoxide dismutase transgenic mice. J Comp Neurol 416:112–125
Ohi T, Nabeshima K, Kato S, Yazawa S, Takechi S (2004) Familial amyotrophic lateral sclerosis with His46Arg mutation in Cu/Zn superoxide dismutase presenting characteristic clinical features and Lewy body-like hyaline inclusions. J Neurol Sci 225: 19–25
Ohtsubo M, Kai R, Furuno N, Sekiguchi T, Sekiguchi M, Hayashida H, Kuma K, Miyata T, Fukushige S, Murotsu T (1987) Isolation and characterization of the active cDNA of the human cell cycle gene (RCC1) involved in the regulation of onset of chromosome condensation. Genes Dev 1:585–593
Orrell RW, King AW, Hilton DA, Campbell MJ, Lane RJM, de Belleroche JS (1995) Familial amyotrophic lateral sclerosis with a point mutation of SOD-1: intrafamilial heterogeneity of disease duration associated with neurofibrillary tangles. J Neurol Neurosurg Psychiatry 59:266–270
Otomo A, Hadano S, Okada T, Mizumura H, Kunita R, Nishijima H, Showguchi-Miyata J, Yanagisawa Y, Kohiki E, Suga E, Yasuda M, Osuga H, Nishimoto T, Narumiya S, Ikeda JE (2003) ALS2, a novel guanine nucleotide exchange factor for the small GTPase Rab5, is implicated in endosomal dynamics. Hum Mol Genet 12:1671–1687
Pramatarova A, Laganière J, Roussel J, Brisebois K, Rouleau GA (2001) Neuron-specific expression of mutant superoxide dismutase 1 in transgenic mice does not lead to motor impairment. J Neurosci 21:3369–3374
Ripps ME, Huntley GW, Hof PR, Morrison JH, Gordon JW (1995) Transgenic mice expressing an altered murine superoxide dismutase gene provide an animal model of amyotrophic lateral sclerosis. Proc Natl Acad Sci USA 92:689–693
Robertson J, Kriz J, Nguyen MD, Julien JP (2002) Pathways to motor neuron degeneration in transgenic mouse models. Biochimie 84:1151–1160
Robertson J, Sanelli T, Xiao S, Yang W, Horne P, Hammond R, Pioro EP, Strong MJ (2007) Lack of TDP-43 abnormalities in mutant SOD1 transgenic mice shows disparity with ALS. Neurosci Lett 420:128–132
Rosen DR, Siddique T, Patterson D, Figlewicz DA, Sapp P, Hentati A, Donaldson D, Goto J, O’Regan JP, Deng HX, Rahmani Z, Krizus A, McKenna-Yasek D, Cayabyab A, Gaston SM, Berger R, Tanzi RE, Halperin JJ, Herzfeldt B, Van den Bergh R, HungWY, Bird T, Deng G, Mulder DW, Smyth C, Laing NG, Soriano E, Pericak-Vance MA, Haines J, Rouleau GA, Gusella JS, Horvitz HR, Brown RH (1993) Mutations in Cu/Zn superoxide dismutase gene are associated with familial amyotrophic lateral sclerosis. Nature 362:59–62
Rouleau GA, Clark AW, Rooke K, Pramatarova A, Krizus A, Suchowersky O, Julien JP, Figlewicz D (1996) SOD1 mutation is associated with accumulation of neurofilaments in amyotrophic lateral sclerosis. Ann Neurol 39:128–131
Sato T, Nakanishi T, Yamamoto Y, Andersen PM, Ogawa Y, Fukada K, Zhou Z, Aoike F, Sugai F, Nagano S, Hirata S, Ogawa M, Nakano R, Ohi T, Kato T, Nakagawa M, Hamasaki T, Shimizu A, Sakoda S (2005) Rapid disease progression correlates with instability of mutant SOD1 in familial ALS. Neurology 65:1854–1860
Schmidt A, Hall A (2002) Guanine nucleotide exchange factors for Rho GTPases: turning on the switch. Genes Dev 16:1587–1609
Shaw CE, Enayat ZE, Powell JF, Anderson VER, Radunovic A, al-Sarraj S, Leigh PN (1997) Familial amyotrophic lateral sclerosis: molecular pathology of a patient with a SOD1 mutation. Neurology 49:1612–1616
Shibata N, Hirano A, Kobayashi M, Siddique T, Deng HX, Hung WY, Kato T, Asayama K (1996) Intense superoxide dismutase-1 immunoreactivity in intracytoplasmic hyaline inclusions of familial amyotrophic lateral sclerosis with posterior column involvement. J Neuropathol Exp Neurol 55:481–490
Stieber A, Gonatas JO, Gonatas NK (2000) Aggregation of ubiquitin and a mutant ALS-linked SOD1 protein correlate with disease progression and fragmentation of the Golgi apparatus. J Neurol Sci 173: 53–62
Sumi H, Nagano S, Fujimura H, Kato S, Sakoda S (2006) Inverse correlation between the formation of mitochondria-derived vacuoles and Lewy-body-like hyaline inclusions in G93A superoxide-dismutase-transgenic mice. Acta Neuropathol 112:52–63
Takahashi H, Makifuchi T, Nakano R, Sato S, Inuzuka T, Sakimura K, Mishina M, Honma Y, Tsuji S, Ikuta F (1994) Familial amyotrophic lateral sclerosis with a mutation in the Cu/Zn superoxide dismutase gene. Acta Neuropathol 88:185–188
Takehisa Y, Ujike H, Ishizu H, Terada S, Haraguchi T, Tanaka Y, Nishinaka T, Nobukuni K, Ihara Y, Namba R, Yasuda T, Nishibori M, Hayabara T, Kuroda S (2001) Familial amyotrophic lateral sclerosis with a novel Leu126Ser mutation in the copper/zinc superoxide dismutase gene showing mild clinical features and Lewy body-like hyaline inclusions. Arch Neurol 58:736–740
Tan CF, Piao YS, Hayashi S, Obata H, Umeda Y, Sato M, Fukushima T, Nakano R, Tsuji S, Takahashi H (2004) Familial amyotrophic lateral sclerosis with bulbar onset and a novel Asp101Tyr Cu/Zn superoxide dismutase gene mutation. Acta Neuropathol 108:332–336
Tan CF, Eguchi H, Tagawa A, Onodera O, Iwasaki T, Tsujino A, Nishizawa M, Kakita A, Takahashi H (2007) TDP-43 immunoreactivity in neuronal inclusions in familial amyotrophic lateral sclerosis with or without SOD1 gene mutation. Acta Neuropathol 113:535–542
Wang J, Xu G, Gonzales V, Coonfield M, Fromholt D, Copeland NG, Jenkins NA, Borchelt DR (2002) Fibrillar inclusions and motor neuron degeneration in transgenic mice expressing superoxide dismutase 1 with a disrupted copper-binding site. Neurobiol Dis 10:128–138
Watanabe Y, Yasui K, Nakano T, Doi K, Fukada Y, Kitayama M, Ishimoto M, Kurihara S, Kawashima M, Fukuda H, Adachi Y, Inoue T, Nakashima K (2005) Mouse motor neuron disease caused by truncated SOD1 with or without C-terminal modification. Mol Brain Res 135:12–20
Wong PC, Pardo CA, Borchelt DR, Lee MK, Copeland NG, Jenkins NA, Sisodia SS, Cleveland DW, Price DL (1995) An adverse property of a familial ALS-linked SOD1 mutation causes motor neuron disease characterized by vacuolar degeneration of mitochondria. Neuron 14:1105–1116
Xu Z, Cork LC, Griffin JW, Cleveland DW (1993) Increased expression of neurofilament subunit NF-L produces morphological alterations that resemble the pathology of human motor neuron disease. Cell 73:23–33
Yamagishi S, Koyama Y, Katayama T, Taniguchi M, Hitomi J, Kato M, Aoki M, Itoyama Y, Kato S, Tohyama M (2007) An in vitro model for Lewy body-like hyaline inclusion/astrocytic hyaline inclusion: induction by ER stress with an ALS-linked SOD1 mutation. PLoS One 2:e1030
Yamanaka K, Miller TM, McAlonis-Downes M, Chun SJ, Cleveland DW (2006) Progressive spinal axonal degeneration and slowness in ALS2-deficient mice. Ann Neurol 60:95–104
Yang Y, Hentati A, Deng HX, Dabbagh O, Sasaki T, Hirano M, Hung WY, Ouahchi K, Yan J, Azim AC, Cole N, Gascon G, Yagmour A, Ben-Hamida M, Pericak-Vance M, Hentati F, Siddique T (2001) The gene encoding alsin, a protein with three guanine-nucleotide exchange factor domains, is mutated in a form of recessive amyotrophic lateral sclerosis. Nat Genet 29:160–165
Zang DW, Cheema SS (2002) Degeneration of corticospinal and bulbospinal systems in the superoxide dismutase 1G93A G1H transgenic mouse model of familial amyotrophic lateral sclerosis. Neurosci Lett 332:99–102
Acknowledgments
This study was supported in part by a Grant-in-Aid for Scientific Research (c) from the Ministry of Education, Culture, Sports, Science and Technology of Japan (SK: 17500229), a Grant for Research on Psychiatric and Neurological Disease and Mental Health from the Ministry of Health, Labor and Welfare of Japan (SK), and a Grant from The Research Group on Development of Novel Therapeutics for ALS of the Ministry of Health, Labor and Welfare of Japan (SK).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kato, S. Amyotrophic lateral sclerosis models and human neuropathology: similarities and differences. Acta Neuropathol 115, 97–114 (2008). https://doi.org/10.1007/s00401-007-0308-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-007-0308-4