Abstract
Purpose
We evaluated the effect of low-calorie mediterranean (MD) and vegetarian (VD) diets on gut microbiome (GM) composition and short-chain-fatty acids (SCFA) production.
Methods
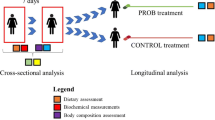
We performed next generation sequencing (NGS) of 16S rRNA and SCFA analysis on fecal samples of 23 overweight omnivores (16 F; 7 M) with low-to-moderate cardiovascular risk. They were randomly assigned to a VD or MD, each lasting 3 months, with a crossover study design.
Results
Dietary interventions did not produce significant diversity in the GM composition at higher ranks (family and above), neither between nor within MD and VD, but they did it at genus level. MD significantly changed the abundance of Enterorhabdus, Lachnoclostridium and Parabacteroides, while VD significantly affected the abundance of Anaerostipes, Streptococcus, Clostridium sensu stricto, and Odoribacter. Comparison of the mean variation of each SCFA between MD and VD showed an opposite and statistically significant trend for propionic acid (+ 10% vs − 28%, respectively, p = 0.034). In addition, variations of SCFA were negatively correlated with changes of some inflammatory cytokines such as VEGF, MCP-1, IL-17, IP-10 and IL-12, only after MD. Finally, correlation analyses showed a potential relationship—modulated by the two diets—between changes of genera and changes of clinical and biochemical parameters.
Conclusions
A short-term dietary intervention with MD or VD does not induce major change in the GM, suggesting that a diet should last longer than 3 months for scratching the microbial resilience. Changes in SCFA production support their role in modulating the inflammatory response, thus mediating the anti-inflammatory and protective properties of MD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The human gut microbiota (GM) is a complex biological network made up of hundreds of thousands of microorganisms. It is mainly composed of bacteria, but also includes archaea, viruses, fungi, and protozoa [1]. They tend to associate with the host in a symbiotic way: the host provides the nutrients to the GM while the GM makes it capable to digest certain foods which are indigestible for human enzymes, balances the immune and metabolic functions and provides protection against opportunistic pathogens. Furthermore, the bacterial fermentation of dietary fiber produces biologically active short chain fatty acids (SCFA), such as acetate, butyrate and propionate, which lead to favorable effects, ranging from glucose and lipid homeostasis to cancer protection [2].
A structurally and functionally balanced GM is necessary for optimal health status; therefore, this dynamic microbial community has fueled research investigating a potential impact on different aspects of human physiology and pathological disorders, such as obesity [3], inflammatory diseases [4] immune system [5], and susceptibility to disease [6, 7].
Among the various environmental factors, diet is a source of dominant variation of the entire intestinal microbial community [8]. At the same time, diet represents also one of the most important modifiable factors for the cardiovascular prevention. In this regard, Mediterranean diet (MD) and vegetarian diet (VD) are certainly two of the most studied dietary profiles for their potential beneficial effects on human health. MD is a dietary pattern typical of countries bordering the Mediterranean Sea, characterized by large quantities of fruit, vegetables, cereals, legumes, olive oil as the main source of fats, moderate quantities of fish, poultry and dairy products and low quantities of red meat and wine. VD is characterized by abstention from the consumption of meat and meat products, poultry, fish, seafood and flesh from any other animal and a large amount of plant-derived foods. These two dietary patterns have proved to be widely useful for patients with a well-established disease and for subjects with traditional risk factors for chronic diseases [9, 10].
Regarding the dietary impact on GM composition, to date, few studies have finely investigated the link between nutrition and GM composition. Wu and colleagues showed the association of different dietary profiles with three enterotypes [11]: a carbohydrate-dominated metabolism and the VD were associated with genus Prevotella; diets high in protein and of animal origin (mostly omnivorous) were associated with Bacteroides, and finally a westernized fat-based diet was related to Firmicutes [8]. De Filippis et al. reported that nutritional models rich in plant-based foods promote a more favorable GM profile based on the high amount of dietary fiber and SCFA [12]. In a cross-sectional study, the GM of two USA omnivorous and vegan cohorts was compared with negligible differences [13]. Recently, Shankar et al. explored the differences in the GM structure of two groups of adolescents (USA vs Egypt) based on their diets, documenting an enrichment of polysaccharide-degrading genera in the guts of Egyptian teenagers [14]. Given the information on the influence of the habitual diet on GM and its metabolism, the present paper aims to evaluate and compare GM composition and SCFA production in a group of subjects in primary prevention for cardiovascular disease that underwent two different nutritional interventions based on MD and VD.
Methods
Study participants and design
The study population consisted of 23 clinically healthy Caucasian subjects (16 women, 7 men; mean age 58.6 ± 9.8 years) enrolled in the CARDIVEG study, a dietary intervention study aimed at comparing the effect of MD and VD on cardiovascular risk factors of people with a low-to-moderate cardiovascular risk profile [15]. The study protocol has been previously described [16]. Briefly, the study participants were asked to follow a MD or VD for 3 months each and then they were crossed. Both diets were hypocaloric with respect to the energy needs of subjects, but they were isocaloric between them and consisted of about 50–55% of energy from carbohydrate, 25–30% from total fat and 15–20% of proteins. The MD was characterized by the consumption of all food groups, including meat and meat products, poultry and fish. The VD was characterized by abstinence to consume meat and meat products, poultry, fish, seafood and flesh from any other animal, but included eggs and dairy products. The data collection and follow-up measures were performed at the Clinical Nutrition Unit of the Careggi University Hospital (Florence, Italy). Body weight and body composition were measured at each clinical evaluation. Venous blood samples and fecal samples were collected at baseline and at the end of each intervention phase. Blood samples were used to assess a number of laboratory measurements (i.e., blood chemistry tests, plasma oxidative stress profile, and serum inflammatory cytokines) as previously reported [15, 16]. Stool sample collection kits, including containers, were provided for the participants. Fecal samples (four or five scoops totaling 4 g) were collected in sterile containers before and after each intervention phase—for a total of three fecal samples for each participant—and immediately frozen at − 20 °C, before being transferred to − 80 °C until analysis. Prior to enrollment, informed written consent was obtained from each participant. The study was approved by the Ethics Committee (SPE 15.054) of the Tuscany Region, Careggi University Hospital and registered at https://www.clinicaltrials.gov (Unique identifier: NCT02641834). The study was adhered to the principles of the Declaration of Helsinki and the Data Protection Act.
Laboratory analysis
Fecal Samples collection
Total DNA was extracted using the DNeasy PowerLyzer PowerSoil Kit (Qiagen, Hilden, Germany) from frozen (− 80 °C) fecal samples according to the manufacturer’s instructions [17]. In particular, 0.25 g of fecal samples were added to a bead beating tube and homogenized rapidly and thoroughly using a FastPrep instrument (Qiagen, Hilden, Germany). Cell lysis occurred by mechanical and chemical interaction. Total genomic DNA was captured on a silica membrane in a spin column format. DNA was washed and eluted from the membrane. The quality and quantity of extracted DNA was assessed using the NanoDrop ND-1000 (Thermo Fisher Scientific, Waltham, US) and the Qubit Fluorometer (Thermo Fisher Scientific), respectively.
Analysis of 16S rRNA Sequences and profiling of the gut microbiota
Amplicons of the V3–V4 region of the bacteria 16s rRNA were obtained and sequenced with a 2 × 300 bp paired-end approach on the Illumina MiSeq platform (IGA technology services, Udine, Italy), according to the Illumina 16S Metagenomic Sequencing Library Preparation protocol [17]. Demultiplexed sequence reads were merged using PEAR [18] and quality-checked using dedicated tool included in the QIIME suite (v. 1.9) [19]. Merged amplicons were then processed with USEARCH 6.1 [20] to detect potential chimera sequences and to generate cluster of operational taxonomic units (OTUs), with a minimum pair-wise identity threshold of 97%. A quality-filtering step was performed on the generated OTUs to filtered out those OTUs showing a relative abundance less than 0.005% [21]. The SILVA database (release 128) was employed for taxonomic classification [22]. Microbial alpha (Richness, Simpson’s, Gini-Simpson’s, inverse Simpson and Shannon’s indices, evenness and dominance) and beta (Weighted UniFrac and Bray–Curtis dissimilarity) diversity measures were assessed using QIIME (v. 1.9) [19]. To infer the functional contribution of microbial communities on 16S rDNA sequencing data, metagenomes prediction was carried out using PICRUSt (Phylogenetic Investigation of Communities by Reconstruction of Unobserved States) evaluating KEGG (Kyoto Encyclopedia of Genes and Genomes) pathways [23]. The presence of potential distinguishing biomarker among subjects’ categories was assessed through the Linear Discriminant Analysis (LDA Effect Size (LEfSe) analysis [24].
Short-chain fatty acids measurements
Methanol and tert-butyl methyl ether (Chromasolv grade), sodium bicarbonate and hydrochloric acid (Reagent grade), [2H3]acetic, [2H3]propionic, [2H7]isobutyric and [2H9]isovaleric (used as internal standards or ISTDs), acetic acid, propionic acid, butyric acid, isobutyric acid, valeric acid and isovaleric acid (analytical standards grade) were purchased by Sigma-Aldrich (Milan, Italy). MilliQ water 18 MΩ was obtained from Millipore’s Simplicity system (Milan, Italy). The short chain fatty acids analyses were performed by agilent gas chromatography–mass spectrometry system composed with 5971 single quadrupole mass spectrometer, 5890 gas chromatograph and 7673 autosampler. The SCFA in the samples were analyzed as free acid form using a Supelco Nukol column 30 m length, 0.25 mm internal diameter and 0.25 µm of film thickness with the temperatures program as follows: initial temperature of 40 °C was held for 1 min, then it was increased to 150 °C at 30 °C/min, finally grow up to 220 °C at 20 °C/min. A 1 µl aliquot of extracted sample was injected in spitless mode (spitless time 1 min) at 250 °C, while the transfer line temperature was 280 °C. The carrier flow rate was maintained at 1 mL/min.
Statistical analyses
Statistical analyses were performed using R for Statistical Computing 3.5.1. In detail, permutational analysis of variance (PERMANOVA) were performed using the package ‘vegan’ [25], based on 999 permutations. Categorical variables were presented in terms of frequencies and percentages. The dichotomous variables were analyzed by the Chi square test. Data were treated as paired samples from a crossover study: for each subject, they were clustered into four groups: before MD (MD0), after MD (MD1), before VD (VD0), and after VD (VD1). The two interventions were analyzed by combining the results obtained in the two phases of both groups. p values less than 0.05 were considered as statistically significant.
Data normalization was conducted to exclude the potential confounding effect of sequencing depth in differential abundance analysis.
Principal component analysis (PCA) and partial least squares discriminant analysis (PLSDA) were applied to bacterial relative abundances at each taxonomic level (Genus, Family, Order, Class and Phylum) to identify possible similarities of the samples within the four groups. Afterwards, PCA was integrated with co inertia analysis (CIA) [26] to identify potential shared biological trends between the four groups.
Principal coordinate analysis (PCoA) was applied to the Weighted Unifrac and the Bray–Curtis beta diversity metrics to identify potential patterns of the samples within the four groups. For each alpha diversity index, bacterial abundance and predicted KEGG pathway abundance, minimum, maximum and quartiles were reported for each group at each taxonomic level (Genus, Family, Order, Class and Phylum), and boxplots were used to represent the results graphically. The Wilcoxon signed-rank test was applied to assess the effect of each dietary intervention (MD0 versus MD1, and VD0 versus VD1). The Kruskal–Wallis test was applied to compare the effect of the two dietary interventions (ΔMD versus ΔVD).
Permutational analysis of variance (PERMANOVA) was applied with three objectives: to assess the effect of the dietary interventions on PCoA scores of beta diversity metrics; to assess the effect of the dietary interventions on the whole predicted KEGG pathways; to assess the effect of the dietary interventions and of several metadata (SCFA’ levels, nutrients’ intake, clinical and biochemical parameters) on the whole microbial composition. In the last chance, the analysis was performed at each taxonomic level (Genus, Family, Order, Class and Phylum), with a separated model for each metadata. Each analysis was performed using a model including both the main effects and the interaction of the two dietary interventions.
Spearman correlation coefficients between the changes in microbial composition, SCFA’ levels, nutrients’ intake and in clinical and biochemical parameters were computed and statistically tested for each dietary intervention. Results were visualized through heatmaps with hierarchical clustering based on the complete linkage algorithm.
Results
To investigate the effect of MD and VD on GM, we performed next generation sequencing (NGS) of 16S rRNA amplicons on fecal samples collected at three time points from 23 healthy subjects, 11 of which started to follow a MD and 12 a VD. The amount of genomic DNA extracted from the fecal sample of a subject after MD intervention was not sufficient to perform the NGS analysis. The baseline characteristics of the study population, according to the first dietary randomization, are reported in Table 1. No significant differences were observed in demographic, anthropometric and clinical parameters, as well as in dietary habits between the two groups. Overall, the OTU table generated with QIIME included 210 observations and 69 samples, with 7,590,334 total sequences and 110,004 mean sequences per sample.
Impact of the dietary interventions on microbiota composition
To obtain an overview of the GM composition, we first analyzed the microbiota profile in terms of taxonomic assignment and more abundant taxa. Analysis of the taxonomic composition of all the subjects’ GM at baseline revealed that over 98% of the collected sequences belonged to 4 phyla: Firmicutes (64.9%), Actinobacteria (14.8%), Bacteroidetes (13.6%), and Verrucomicrobia (4.7%). The bar plots of relative abundances at each taxonomic level are shown in the online Data Supplement (Supplemental Figure 1). Next, we investigated the richness and evenness of the GM in all samples. No significant differences were observed at all levels and between different time points in any groups. The comparison of the alpha diversity indices between and within the dietary interventions did not reveal significant changes (online Data Supplement, Supplemental Table 1). Similarly, the Firmicutes/Bacteroidetes ratio was not statistically different before and after each intervention phase (MD0 vs MD1: p = 0.545; VD0 vs VD1: p = 0.689) as well as between the two diets (ΔMD vs ΔVD: p = 0.756). We observed a similar situation for the Firmicutes/Proteobacteria ratio: no significant difference was found after each intervention phases (MD0 vs MD1: p = 0.082; VD0 vs VD1: p = 0.708) as well as between the two diets (ΔMD vs ΔVD: p = 0.385). Detailed results can be found in the online Data Supplement (Supplemental Table 2).
To explore the composition of the GM between and within dietary intervention, PCA, CIA and PLSDA were performed. The PCA with two components was applied to the abundances relative to the bacterial taxa to identify possible clustering between the four groups (MD0, MD1, VD0, VD1) (Supplemental Figure 2, online Data Supplement). Data obtained through PCA were integrated using the CIA to explore the biological trends shared between the two dietary interventions. In agreement with PCA, the CIA did not provide clearly separated clusters based on the type of diet (Supplemental Figure 3, online Data Supplement). To find a separation between the 4 groups, the PLSDA with two-variate classification rule was applied to the relative abundances of bacterial taxa. The variance explained by the PLSDA was almost 38% at the phylum level (Supplemental Figure 4, online Data Supplement). A trend of separation between groups was only collected at the genus level, explaining 9.3% of total variance.
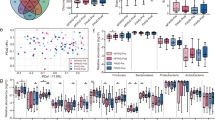
The PCoA on beta diversity computed using the Weighted Unifrac and the Bray–Curtis dissimilarity metrics did not provide clearly separated clusters based on the type of diet (Fig. 1). Similarly, the search for presumed distinctive microbial biomarkers, performed through the LEfSe tool, did not show significantly discriminating characteristics based on the type of diet (data not shown).
Principal Coordinate Analysis (PCoA). Scatterplot of principal coordinate scores obtained from beta diversity dissimilarities a Weighted Unifrac, b Bray–Curtis. Subjects are colored according to the group (MD0, MD1, VD0, VD1). Each axis label shows the percentage of explained variance of the corresponding component. The total percentage of variance explained by the two coordinates is indicated at the top of each scatterplot. MD0 before Mediterranean dietary intervention, MD1 after Mediterranean dietary intervention, n number of samples, VD0 before vegetarian dietary intervention, VD1 after vegetarian dietary intervention
We performed univariate analyses to identify the bacterial taxa influenced by MD and VD. Among the taxonomic ranks, no significant changes were observed except for genus level; in detail, 7 genera reported a statistically significant variation (p < 0.05) due to diets (Fig. 2a). In particular, MD resulted in a significant change in the abundance (median pre vs post) of Enterorhabdus (151 vs 240, p = 0.002), Lachnoclostridium (211 vs 417, p = 0.039), Parabacteroides (190 vs 40.5, p = 0.037), while VD determined a significant change in Anaerostipes (280 vs 502, p = 0.041), Clostridium sensu stricto (66 vs 26, p = 0.020), Odoribacter (9 vs 5, p = 0.025) and Streptococcus (78 vs 183, p = 0.038). Furthermore, to test the different impact of the two diets on the GM, the mean variation of each microorganism within any taxonomic rank (post–pre diet) between MD and VD was compared. We found an opposite and statistically significant effect of the two treatments concerning the following bacterial genera: Anaerostipes (ΔMD = − 64.5 vs ΔVD = 58, p = 0.048), Clostridium sensu stricto (ΔMD = 15 vs ΔVD = -18; p = 0.005), Enterorhabdus (ΔMD = 83, ΔVD = − 17; p = 0.003), Veillonella (ΔMD = 1.5 vs ΔVD = − 1; p = 0.029) (Fig. 2b). Detailed results can be found in the online Data Supplement Supplemental Table 3.
Effect of dietary intervention on fecal microbiota variation. a Boxplot showing the abundance of the genera associated with a statistically significant variation due to the Mediterranean dietary intervention (MD) or vegetarian dietary intervention (VD) assessed with Wilcoxon signed-rank test (p < 0.05). b Boxplot showing the abundance of genera associated with a statistically significant variation between Mediterranean and Vegetarian dietary intervention assessed with Kruskal–Wallis test (p < 0.05). MD0 before Mediterranean dietary intervention, MD1 after Mediterranean dietary intervention, n number of samples, VD0 before vegetarian dietary intervention, VD1 after vegetarian dietary intervention, n number of samples
To compare the entire bacterial composition within and between dietary interventions, PERMANOVA analysis was performed. The results obtained (online Data Supplement, Supplemental Table 4) did not show statistically significant difference in the entire microbiota composition due to treatments in any taxonomic rank.
Impact of the dietary interventions on fecal SCFA’ concentrations
Overall, the most representative SCFA at baseline were acetic acid [39.31 (4.42) µmol/g], butyric acid [17.77 (2.69) µmol/g], and propionic acid [14.46 (1.96) µmol/g]. The mean change in SCFA according to the prescribed diet is shown in Table 2. No statistically significant differences in the amount of SCFA for MD were observed. On the other hand, a marginal reduction in fecal propionate and an increase in both branched chain SCFA, isobutyric and isovaleric acid (p = 0.034) emerged for VD. Furthermore, the comparison of the mean variation of each SCFA (post–pre diet) between MD and VD showed an opposite and statistically significant trend for propionic acid (p = 0.034); in particular, MD led to an increase of 10%, while VD to a decrease of − 28% (Fig. 3). To examine the association between the taxonomic composition of GM at the genus level and the bacteria’s end-products within each dietary intervention, a Spearman correlation was conducted and a network diagram was constructed to show the correlations between bacterial genera and SCFA (Fig. 4a). In detail, focusing on the relationship between the SCFA and modified genera, within or between the dietary interventions, we observed a negative correlation between Veillonella and isobutyrate and isovalerate (p < 0.05) in subjects following MD intervention (Fig. 4b).
Correlations between fecal microbiota and SCFA. a Network diagram showing Spearman’s correlations between bacterial genera and SCFA according to Mediterranean diet and vegetarian diet. Genera, represented to circles, are colored according to phylum. SCFA are represented by squares. Edges size is proportional to the number of correlation. Blue edges-negative correlation, red edge-positive correlation. Only correlation with statistical significance is reported. b Heatmap of correlations between genera that showed significant differences after the two dietary intervention and changes of SCFA levels, according, respectively to Mediterranean or Vegetarian diet. Rows and columns are clustered by Euclidean distance with Ward linkage hierarchical clustering. The intensity of colors represents the degree of association between the changes in the abundance of genera and change in the abundance of SCFA due to each diet, as measured by the Spearman’s correlations. Asterisks indicates the statistically significant Spearman correlations (*p < 0.05, **p < 0.01, ***p < 0.001). MD Mediterranean dietary intervention, VD vegetarian dietary intervention
To find the association between nutrients and SCFA variations, Spearman’s correlation was calculated between nutrient intake (post–pre intervention) and SCFA (post–pre intervention) for each diet. By plotting the correlation between SCFA and dietary information, it was possible to detect the influence of the diet on the SCFA levels (Fig. 5a). The nutrients that seem to be most associated with the variation of SCFA are total fats, in particular saturated fats and polyunsaturated fatty acids omega-3, carbohydrates and fiber. Notably, SCFA that were negatively correlated with fat intake was acetic acid, propionic acid, valeric acid and isobutyric acid, whereas the carbohydrate was positively correlated with butyric acid in MD. In contrast, energy intake, carbohydrates and fiber seemed to be negatively associated with isobutyric and isovaleric acids in VD.
Correlation between fecal SCFA, dietary information and cytokines. a Heatmaps of correlations between changes in nutrients’ intake and changes in SCFA, according to Mediterranean and vegetarian diets. b Heatmaps of correlations between immunological parameters and changes of SCFA levels, according, respectively to Mediterranean or vegetarian diet. Rows and columns are clustered by Euclidean distance and Ward linkage hierarchical clustering. The intensity of colors represents the degree of association between the changes in the inflammatory profile (post–pre diet) and change in the levels of SCFA due to each diet, as measured by the Spearman correlations. Asterisks indicates the statistically significant Spearman correlations (*p < 0.05, **p < 0.01, ***p < 0.001). MD Mediterranean dietary intervention, VD vegetarian dietary intervention
Finally, correlation analyses were calculated between the changes in SCFA and the differences in the inflammatory cytokines evaluated after the two dietary interventions (Fig. 5b). Some registered correlations have suggested that different SCFA have a linear relationship with cytokines. During the MD period, the variations of the most representative SCFA—propionic, acetic and butyric acid—were negatively correlated with the variations of some inflammatory cytokines such as VEGF, MCP-1, IL-17, IP-10 and IL-12. In detail, propionate showed a negative correlation with IP-10, IL-12 (p < 0.05), and with VEGF (p < 0.01); acetic acid with IP-10, IL-10, Il-17 (p < 0.05), VEGF, and IL-12 (p < 0.01); butyric acid with VEGF, MCP-1 (p < 0.05), IL-12 and IL-17 (p < 0.01); isovalerate with IL-1RA (p < 0.05); isobutyric acid with IL-1RA and MCP-1(p < 0.05). On the other hand, only few modifications appeared after the VD period: isobutyric acid was negatively associated with both MCP-1 and IL-12 (p < 0.05), while isovaleric acid was negatively associated with MCP-1 (p < 0.05) (Fig. 5b).
Quantification of the dietary impact on microbiota, clinical and biochemical variables
PERMANOVA analysis was subsequently used to seek for possible associations between some biochemical variable and GM composition. Variables significantly associated (p < 0.05) with the microbial composition at any rank were: butyrate, isobutyrate, isovalerate, IL-10, white blood cell count, vitamin B12, AST. The complete associations, for each taxonomic rank are shown in online Data Supplement Supplemental Table 5.
Moreover, to find a potential association—modulated by the diet—between taxa and metadata (clinical and biochemical parameters) the Spearman correlation was conducted between variations of microorganisms that showed statistically significant changes after dietary interventions and variation of clinical parameters (Fig. 6). We found several statistically significant correlations (p < 0.05, excepting *: p < 0.01) for MD and VD. In particular for MD, the variation of Anaerostipes was positively correlated with LDL-cholesterol and total cholesterol.; changes in Clostridium sensu stricto was positively associated with serum ferritine. HDL-cholesterol and IFN-gamma* were negatively correlated with the variations of Enterorhabdus. The variation in Parabacteroidetes was positively correlated with MCP-1. Finally, the variation of Lachnoclostridium was related to negative variations of IL-6, AST*, ALT and vitamin B12. A greater number of associations emerged also for VD. The variation of Anaerostipes was found to be positively correlated with total cholesterol*, serum iron and IP-10, while AST showed a negative correlation. The change in the abundance of Clostridium sensu stricto was positively associated with IL-10 and LDL-cholesterol, while a negative correlation was observed with changes in insulin levels. A negative correlation between Enterorhabdus and the oxidative profile emerged, notably M-derived ROS*, L-derived ROS*, G-derived ROS and AST; variations in Veillonella were positively associated with VEGF*, serum urea and triglycerides, whereas on the contrary MCV and IL-6 were negatively associated. The variation in Odoribacter abundance showed a positive correlation with serum calcium* and glucose, and a negative correlation with IL-17*. Finally, Streptococcus was positively associated with serum uric acid, glucose and HOMA index, as well as eicosapentaenoic acid, docosahexaenoic acid and glucose from the diet.
Correlation between fecal microbiota and clinical and nutritional variables. Heatmap of correlations between changes in nutrients’ intake, clinical and biochemical parameters and genera that showed significant differences after the two dietary intervention, according to Mediterranean (a) or vegetarian (b) diet. Rows and columns are clustered by Euclidean distance with Ward linkage hierarchical custering. The intensity of colors represents the degree of association between the changes in the levels of nutrients’ intake, clinical and biochemical parameters and the changes in the abundance of genera due to each diet, as measured by the Spearman correlations. Asterisks indicates the statistically significant Spearman correlations (*p < 0.05, **p < 0.01, ***p < 0.001). MD Mediterranean dietary intervention, VD vegetarian dietary intervention
Finally to infer the functional contribution of microbial communities on the 16S rDNA sequencing data, metagenome prediction was performed using PICRUSt, which revealed some significant, but not relevant, differences in the main functional classes (KEGG), resulting from functional acquisitions associated with different dietary habits (online Data Supplement Supplemental Table 6).
Discussion
Increasing evidence suggests that diets with low animal protein and high vegetable and fiber intake are associated with the prevention of cardiovascular disease. A recent publication about the first results of the CARDIVEG study, documented that VD and MD are equally effective in reducing cardiovascular risk [15]. Current trend in nutritional therapy involves the beneficial use of diet to improve human health through GM performances [27], although the mechanisms underlying this association are not fully clarified. Research on the microbiome is expanding rapidly thanks to technological and computational advances that make DNA sequencing cheaper and data analysis easier.
To deepen our knowledge in this field, we used the fecal sample of a cohort of omnivorous with a low-to-moderate cardiovascular risk profile previously enrolled in the CARDIVEG study to evaluate, for the first time, the impact of short-term fully controlled low-calorie MD and VD on the functional composition of the fecal microbiota. The crossover design of the study, in which each subject has its own control, was selected to evaluate the quantification of the dietary impact on GM composition and on the anthropometric, clinical and nutritional parameters of the host. Although it is conceivable that a long-term dietary intake is able to change the GM structural composition [11, 12, 28,29,30,31], it is unclear how rapidly the human GM responds to short-term dietary changes; thus, the understanding of the GM resilience is crucial for determining the effectiveness of therapeutic diets.
In our study, a 3-month intervention period did not lead to marked changes in the composition and diversity of microbiota at higher ranks (family and above), either between the dietary interventions or within, suggesting a modest effect of a short-term Mediterranean or vegetarian profile in modeling gut microbial communities. In accordance with the previous results [28], the two short-term dietary interventions have produced small changes in the composition of the microbiota in our subjects. In fact, a slight separation between the groups was only collected at the genus level, explaining 9.3% of the total variance. Anyway, such variance is not negligible, given the high individual genetic, environmental and clinical variability of the subjects enrolled.
Interestingly, our findings align with other previous studies that have shown a stability of the general pattern of GM in response to short-term dietary changes [11, 28]. For example, diets high in fat/low in fiber or low in fat/high in fiber for 10 days [11] or 3 months in a vegetarian diet [28] were associated with small variations in species and/or genetic content when compared to baseline interpersonal changes.
In our report the overall GM composition of our omnivorous subjects was made up mostly by the phylum Firmicutes, often associated with a westernized fat-based diet in the literature [8, 29]. Many papers have identified a relatively low Firmicutes to Bacteroidetes (F/B) ratio in association with health and elevated ratios with metabolic disorders [29, 32]. However, we did not observe significant changes in the abundance of Firmicutes or in F/B abundance ratio after the dietary interventions, despite an improvement in the metabolic conditions previously reported in these subjects [15]. In any case, considerable changes in F/B ratio may be due to several factors [29, 32], so that it cannot be considered an absolute indicator of gut wellbeing and, therefore, of human health [33].
Looking at the changes produced by dietary interventions, the MD was associated with change in the abundance of three genera (Enterorhabdus, Lachnoclostridium and Parabacteroides; Fig. 1) whereas VD of 4 genera (Anaerostipes, Clostridium sensu stricto, Odoribacter and Streptococcus). In particular, Lachnoclostridium and Enterorhabdus abundance increased after MD intervention, whereas Parabacteroides decreased after MD intervention. Enterorhabdus belonging to Actinobacteria phylum was previously identified only in lean, but not obese or diabetic animals [34]. Interestingly, the enrichment of the genus Parabacteroides has recently been proposed as a microbial marker for hypertension [35], therefore, our findings may support the cardiovascular protection offered by the MD. On the other hand, the VD intervention produced an increase in the abundance of Streptococcus and Anaerostipes genera and the reduction of Clostridium sensu stricto and Odoribacter genera. It is believed that Anaerostipes promote the gut barrier integrity of the host through anti-inflammation, anti-tumorigenesis, and the exclusion of the pathogen in the colon [36] thus mediating a beneficial role of the VD. Regarding the increase in Streptococcus genus abundance, our result is in contrast to previous studies, where this type negatively correlates with the plant-based diet patterns [12]. It also seems beneficial to reduce the bacteria belonging to the genus Clostridium sensu stricto, which are generally perceived as pathogens [37] as well as an indicator of a less healthy microbiota [38]. To be noted, Clostridium spp. have been reported to be strong inducers of colonic T regulatory cell (Treg) accumulation [39], whereas Odoribacter are known for their ability to produce butyrate, which plays a key role in promoting the development of Treg and in the regulation of the Treg/Th17 balance [40]. Overall, the reduction of these two genera, together with the decrease in fecal butyrate observed in VD, could be associated with the promotion of a Th17 profile, consistent with the increase in the level of interleukin-17 previously reported in our subjects after VD intervention [15]. In conclusion, our results on the VD intervention are in agreement with two recent works that report a significant decrease of Clostridium, and an increase of the Lactobacillus genera, such as Streptococcus, in subjects consuming plant protein [41, 42].
To compare the effects of the proposed dietary pattern on the composition of the microbiota, we compared the GM taxonomic structure between the two diets, observing an opposite and statistically significant variation in 4 different genera. To deepen this aspect, we have analyzed the type and amount of fecal SCFA, because changes in the amount of fiber intake and the consequent microbiota modulation have an impact on the production of microbial metabolites. According to previous studies, in all the subjects enrolled, acetate was the predominant fecal SCFA, followed by propionate and butyrate.
In our study, the two dietary interventions appear to have opposite effects on the fecal concentrations of SCFA, with a not significant, but only suggestive increase after MD and a decrease after VD. In particular, in VD we observed a reduction trend in propionic acid, which, in turn, shows an increasing trend after MD. This is in line with the opposite variation of abundance of Veillonella in VD compared to MD, which is one of the leading propionate producers via the succinate pathway [43]. Furthermore, there was an opposite trend for acetic and butyric acid, although not statistically significant. In contrast, both dietary interventions are associated with increasing levels of isobutyric, isovaleric and valeric acid that have reached statistical significance for isobutyrate and isovalerate after VD. Isovalerate and isobutyrate are microbial metabolites named “Branched Chain Fatty Acids (BCFA)”, which are mainly produced in the distal colon. While saccharolytic fermentation is a major source of luminal SCFA, protein fermentation leads to the release of marked quantities of BCFA which may have multiple effects on the colonic epithelium, that have been poorly described. BCFA contribute very little (5%) to total SCFA production, but they are most often considered detrimental for gut integrity and metabolic health [44]. In vitro experiments showed that both isovalerate and isobutyrate were able to induce apoptosis [45]. In our study, the significant increment of isovalerate and isobutyrate suggests an increase in microbial fermentative activity after VD. Interestingly, this result is in contrast with the previous studies where the intake of dairy and processed meats, that provided cholesterol and a small amount of fiber, was associated with increased production of BCFA [46].
SCFA exerts an immunoregulatory effect and influences the metabolism of lipids, cholesterol and glucose, thus playing consequently a role in various disorders, such as metabolic and cardiovascular disease [47]. Hence, we correlated the changes in SCFA with the dissimilarity of the inflammatory cytokines after the two dietary interventions. Interestingly, changes in the amount of propionic, acetic, butyric, isovaleric and isobutyric acids after MD intervention were negatively associated with some pro-inflammatory cytokines, like VEGF, MCP-1, IL-17, IP-10, IL-12 MCP-1 and IL-1RA, which supports the anti-inflammatory properties of the MD [48]. Conversely, SCFA variations after VD intervention often showed an opposite correlation with the evaluated cytokines, thus suggesting a possible pro-inflammatory effect of this nutritional intervention.
Finally, our correlation analysis showed the influence of various metadata (clinical and biochemical parameters) on the composition of the microbiota and on the amount of fecal SCFA, thus confirming a complex interaction between diet and the gut microbes.
Our study presents main limitations related to the low number of subjects investigated and short duration of the dietary intervention, which may have influenced the statistical power of the study. To confirm these results, studies with a larger population and longer duration are needed. It is also important to consider that it is very difficult to perform a dietary intervention study that includes a change of nutritional habits such as that considered with VD in a group of omnivorous subjects. However, the study includes also several strengths, such as the rate of adherence, the crossover design, the comparability between the two diets in terms of total energy and contribution from macronutrients, the number of blood and fecal parameters analyzed in the same group at different time points.
In summary, our study indicates that a short-term Mediterranean or vegetarian dietary pattern does not induce major changes in the structural composition of GM, thus confirming that dietary interventions should last for longer periods to scratch the resilience of GM. Anyway, we have shown some differences in the abundance of specific bacterial genera, and in the production of SCFA. In particular, the negative association between SCFA and a number of inflammatory cytokines reported only after MD, seems to support the anti-inflammatory properties of the MD. Furthermore, several associations between certain bacterial groups, SCFA, clinical and nutritional parameters, let us hypothesized that the cardiovascular protection associated with the two diets could be due—at least in part—to a modulation of the GM. However, as previously stated, studies with larger number of subjects and longer periods of dietary intervention are needed to confirm the present results.
Data availability
Sequence data sets from 16S ribosomal RNA gene sequencing supporting the conclusion of this article are available in GenBank with study accession number: PRJNA510080
References
Sommer F, Bäckhed F (2013) The gut microbiota—masters of host development and physiology. Nat Rev Microbiol 11:227–238
Russell WR, Hoyles L, Flint HJ, Dumas ME (2013) Colonic bacterial metabolites and human health. Curr Opin Micr 16:246–254
Salonen A, Lahti L, Salojärvim J, Holtrop G, Korpelam K, Duncan SH, Date P, Farquharson F, Johnstone AJ, Lobley GE, Louis P, Flint HJ, de Vos WM (2014) Impact of diet and individual variation on intestinal microbiota composition and fermentation products in obese men. ISME J 8:2218–2230
Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER, Gordon JI (2006) An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444:1027–1031
Cerf-Bensussan N, Gaboriau-Routhiau V (2010) The immune system and the gut microbiota: friends or foes? Nat Rev Immunol 10:735–744
Burcelin R, Garidou L, Pomié C (2012) Immuno-microbiota cross and talk: the new paradigm of metabolic diseases. Sem Immunol 24:67–74
Everard A, Cani PD (2013) Diabetes, obesity and gut microbiota. Best Prac Res Clin Gastroenterol 27:73–83
De Filippo C, Cavalieri D, Di Paola M, Ramazzotti M, Poullet JB, Massart S, Collini S, Pieraccini G, Lionetti P (2010) Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc Natl Acad Sci USA 107:14691–14696
Dinu M, Abbate R, Gensini GF, Casini A, Sofi F (2017) Vegetarian, vegan diets and multiple health outcomes: a systematic review with meta-analysis of observational studies. Crit Rev Food Sci Nutr 57:3640–3649
Dinu M, Pagliai G, Casini A, Sofi F (2018) Mediterranean diet and multiple health outcomes: an umbrella review of meta-analyses of observational studies and randomised trials. Eur J Clin Nutr 72:30–43
Wu GD, Chen J, Hoffmann C, Bittinger K, Chen YY, Keilbaugh SA, Bewtra M, Knights D, Walters WA, Knight R, Sinha R, Gilroy E, Gupta K, Baldassano R, Nessel L, Li H, Bushman FD, Lewis JD (2011) Linking long-term dietary patterns with gut microbial enterotypes. Science 334:105–108
De Filippis F, Pellegrini N, Vannini L, Jeffery IB, La Storia A, Laghi L, Serrazanetti DI, Di Cagno R, Ferrocino I, Lazzi C, Turroni S, Cocolin L, Brigidi P, Neviani E, Gobbetti M, O’Toole PW, Ercolini D (2016) High-level adherence to a Mediterranean diet beneficially impacts the gut microbiota and associated metabolome. Gut 65:1812–1821
Wu GD, Compher C, Chen EZ, Smith SA, Shah RD, Bittinger K, Chehoud C, Albenberg LG, Nessel L, Gilroy E, Star J, Weljie AM, Flint HJ, Metz DC, Bennett MJ, Li H, Bushman FD, Lewis JD (2014) Comparative metabolomics in vegans and omnivores reveal constraints on diet-dependent gut microbiota metabolite production. Gut 65:63–72
Shankar V, Gouda M, Moncivaiz J, Gordon A, Reo NV, Hussein L, Paliy O (2017) Differences in gut metabolites and microbial composition and functions between Egyptian and U.S. children are consistent with their diets. Systems 2:e00169-16
Sofi F, Dinu M, Pagliai G, Cesari F, Gori AM, Sereni A, Becatti M, Fiorillo C, Marcucci R, Casini A (2018) Low-calorie vegetarian vs. Mediterranean diets for reducing body weight and improving cardiovascular risk profile: the CARDIVEG study. Circulation 137:1103–1113
Sofi F, Dinu M, Pagliai G, Cesari F, Marcucci R, Casini A (2016) Mediterranean vs. vegetarian diet for cardiovascular prevention (the CARDIVEG study): study protocol for a randomized controlled trial. Trials 17:233
Russo E, Bacci G, Chiellini C, Fagorzi C, Niccolai E, Taddei E, Ricci F, Ringressi MN, Borrelli R, Melli F, Miloeva M, Bechi P, Mengoni A, Fani R, Amedei A (2018) Preliminary comparison of oral and intestinal human microbiota in patients with colorectal cancer: a pilot study. Front Microbiol 8:2699
Zhang J, Kobert K, Flouri T, Stamatakis A (2014) PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30:614–620
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461
Bokulich NA, Subramanian S, Faith JJ, Gevers D, Gordon JI, Knight R, Mills DA, Caporaso JG (2013) Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat Methods 10:57–59
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:D590–D596
Langille MG, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA, Clemente JC, Burkepile DE, Vega Thurber RL, Knight R, Beiko RG, Huttenhower C (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31:814–821
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS, Huttenhower C (2011) Metagenomic biomarker discovery and explanation. Genome Biol 12:R60
Zapala MA, Schork NJ (2006) Multivariate regression analysis of distance matrices for testing associations between gene expression patterns and related variables. Proc Natl Acad Sci USA 103:19430–19435
Fagan A, Culhane AC, Higgins DG (2007) A multivariate analysis approach to the integration of proteomic and gene expression data. Proteomics 7:2162–2171
Cho I, Blaser MJ (2012) The human microbiome: at the interface of health and disease. Nat Rev Genet 13:260–270
Zhang C, Björkman A, Cai K, Liu G, Wang C, Li Y, Xia H, Sun L, Kristiansen K, Wang J, Han J, Hammarström L, Pan-Hammarström Q (2018) Impact of a 3-months vegetarian diet on the gut microbiota and immune repertoire. Front Immunol 9:908
Ley RE, Turnbaugh PJ, Klein S, Gordon JI (2006) Microbial ecology: human gut microbes associated with obesity. Nature 444:1022–1023
Duncan SH, Belenguer A, Holtrop G, Johnstone AM, Flint HJ, Lobley GE (2007) Reduced dietary intake of carbohydrates by obese subjects results in decreased concentrations of butyrate and butyrate-producing bacteria in feces. Appl Environ Microbiol 73:1073–1078
Walker AW, Ince J, Duncan SH, Webster LM, Holtrop G, Ze X, Brown D, Stares MD, Scott P, Bergerat A, Louis P, McIntosh F, Johnstone AM, Lobley GE, Parkhill J, Flint HJ (2011) Dominant and diet-responsive groups of bacteria within the human colonic microbiota. ISME J 5:220–230
Mariat D, Firmesse O, Levenez F, Guimarăes V, Sokol H, Doré J, Corthier G, Furet JP (2009) The firmicutes/bacteroidetes ratio of the human microbiota changes with age. BMC Microbiol 9:123
Hollister EB, Riehle K, Luna RA, Weidler EM, Rubio-Gonzales M, Mistretta TA, Doddapaneni HR, Metcalf GA, Muzny DM, Gibbs RA, Petrosino JF, Shulman RJ, Versalovic J (2015) Structure and function of the healthy pre-adolescent pediatric gut microbiome. Microbiome 3:36
Geurts L, Lazarevic V, Derrien M, Everard A, Van Roye M, Knauf C, Valet P, Girard M, Muccioli GG, François P, de Vos WM, Schrenzel J, Delzenne NM, Cani PD (2011) Altered gut microbiota and endocannabinoid system tone in obese and diabetic leptin-resistant mice: impact on apelin regulation in adipose tissue. Front Microbiol 2:149
Yan O, Gu Y, Li X, Yang W, Jia L, Chen C, Han X, Huang Y, Zhao L, Li P, Fang Z, Zhou J, Guan X, Ding Y, Wang S, Khan M, Xin Y, Li S, Ma Y (2017) Alterations of the Gut Microbiome in Hypertension. Front Cell Infect Microbiol 7:381
Velasquez-Manoff M (2015) Gut microbiome: the peacekeepers. Nature 518:S3–S11
Rajilić-Stojanović M, de Vos WM (2014) The first 1000 cultured species of the human gastrointestinal microbiota. FEMS Microbiol Rev 38:996–1047
Lakshminarayanan B, Harris HM, Coakley M, O’Sullivan O, Stanton C, Pruteanu M, Shanahan F, O’Toole PW, Ross RP, ELDERMET consortium (2013) Prevalence and characterization of Clostridium perfringens from the faecal microbiota of elderly Irish subjects. J Med Microbiol 62:457–466
Atarashi K, Tanoue T, Shima T, Imaoka A, Kuwahara T, Momose Y, Cheng G, Yamasaki S, Saito T, Ohba Y, Taniguchi T, Takeda K, Hori S, Ivanov II, Umesaki Y et al (2011) Induction of colonic regulatory T cells by indigenous Clostridium species. Science 331:337–341
Kosiewicz MM, Dryden GW, Chhabra A, Alard P (2014) Relationship between gut microbiota and development of T cell associated disease. FEBS Lett 588:4195–4206
Kim CH, Park J, Kim M (2014) Gut microbiota-derived short-chain fatty acids, T cells, and inflammation. Immune Net 14:277–288
Meddah AT, Yazourh A, Desmet I, Risbourg B, Verstraete W, Romond MB (2001) The regulatory effects of whey retentate from bifidobacteria fermented milk on the microbiota of the simulator of the human intestinal microbial ecosystem (SHIME). J Appl Microbiol 6:1110–1117
Scott KP, Martin JC, Campbell G, Mayer CD, Flint HJ (2006) Whole-genome transcription profiling reveals genes up-regulated by growth on fucose in the human gut bacterium “Roseburia inulinivorans”. J Bacteriol 188:4340434–4340439
Mortensen PB, Clausen MR (1996) Short-chain fatty acids in the human colon: relation to gastrointestinal health and disease. Scand J Gastroenterol 216(s):132–148
Sakurazawa T, Ohkusa T (2005) Cytotoxicity of organic acids produced by anaerobic intestinal bacteria on cultured epithelial cells. J Gastroenterol 40(6):600–609
Yang J, Rose DJ (2016) The impact of long-term dietary pattern of fecal donor on in vitro fecal fermentation properties of inulin. Food Funct 7(4):1805–1813
Richards LB, Li M, van Esch BCAM, Garssen J, Folkerts G (2016) The effects of short-chain fatty acids on the cardiovascular system. PharmaNutrition 4:68–111
Casas R, Urpi-Sardà M, Sacanella E, Arranz S, Corella D, Castañer O, Lamuela-Raventós RM, Salas-Salvadó J, Lapetra J, Portillo MP, Estruch R (2017) Anti-inflammatory effects of the mediterranean diet in the early and late stages of atheroma plaque development. Med Inflamm 1:1–12
Acknowledgements
The research was founded with a grant from the regional contribution of “The Programma Attuativo Regionale (Toscana) funded by FAS (now FSC), Grant number 4042.16092014.066000029, the Italian Ministry of University and Research (MIUR) and the Foundation ‘Ente Cassa di Risparmio di Firenze’.
Author information
Authors and Affiliations
Contributions
AA and FS conceived the study, participated in the design of the study, wrote the study protocol, and prepared the final version of the manuscript. FS has been responsible for recruitment and clinical evaluations. GP, ER and MD, participated in the design of the study. ER and SB conducted microbiota experiments. GB, SB and MM performed SCFA analysis. AM, EN and VP performed the statistical analysis. RM participated in the writing of the study protocol. AC, GMR and BG participated in the design of the study and critical revision of the manuscript for important intellectual content. EN, GP, ER wrote the manuscript. All authors read and approved the final manuscript. All authors read and commented on the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Pagliai, G., Russo, E., Niccolai, E. et al. Influence of a 3-month low-calorie Mediterranean diet compared to the vegetarian diet on human gut microbiota and SCFA: the CARDIVEG Study. Eur J Nutr 59, 2011–2024 (2020). https://doi.org/10.1007/s00394-019-02050-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00394-019-02050-0