Abstract
We summarize recent advances in the clinical genetics of hypercholesterolemia, hypertrophic cardiomyopathy (HCM), and lethal arrhythmia, all of which are monogenic cardiovascular diseases being essential to understanding the heart and circulatory pathophysiology. Among the issues of hypercholesterolemia which play a pivotal role in development of vascular damages, familial hypercholesterolemia is the common genetic cardiovascular disease; in addition to identifying the gene mutation coding low-density lipoprotein receptor, lipid kinetics in autosomal recessive hypercholesterolemia as well as in proprotein convertase subtilisin/kexin 9 gene mutation were recently demonstrated. As for HCM, some gene mutations were identified to correlate with clinical manifestations. Additionally, a gene polymorphism of the renin–angiotensin system in development of heart failure was identified as a modifier gene. The lethal arrhythmias such as sudden death syndromes, QT prolongation, and Brugada syndrome were found to exhibit gene mutation coding potassium and/or sodium ion channels. Interestingly, functional analysis of these gene mutations helped to identify the role of each gene mutation in developing these cardiovascular disorders. We suggest considering the genetic mechanisms of cardiovascular diseases associated with hyperlipidemia, myocardial hypertrophy, or lethal arrhythmia in terms of not only clinical diagnosis but also understanding pathophysiology of each disease with therapeutic aspects.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Recent advances in genetic analysis have made it possible to determine gene mutations for almost all hereditary diseases, irrespective of the coded gene region [1]. Therefore, the recognition, diagnosis, and suitable treatment of atherosclerosis, heart failure, and arrhythmias have become important targets for clinical genetics research in the field of cardiovascular medicine [2]. Definitive diagnosis of inheritable diseases by genetic methods could lead to tailor-made therapies using conventional medicine as well as gene silencing or targeting therapy.
Atherosclerosis develops in the presence of a lipid disorder typically represented by familial hypercholesterolemia (FH) [3]. Left ventricular hypertrophy is an independent risk factor for heart failure, and hypertrophic cardiomyopathy (HCM) is the most common genetic cause of left ventricular hypertrophy [4]. Life-threatening arrhythmias and sudden death frequently occur in inherited cardiac arrhythmia syndromes such as congenital long-QT syndrome (LQTS) [5]. In this article, we review current perspectives in genetic-related cardiovascular diseases from recent reports including our ones and provide insights into these disorders.
Advances in hypercholesterolemia research
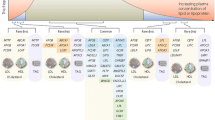
Large epidemiological studies have shown that coronary risk factors including hyperlipidemia, hypertension, impaired glucose tolerance, male sex, and smoking habits, are associated with the development of atherosclerosis [6]. Most coronary risk factors are associated with endothelial dysfunction, which is known to initiate atherosclerosis. Although menopausal hormone replacement therapy failed to decrease cardiovascular disease in older women [7], epidemiological data suggested that endogenous estrogen is antiatherogenic. Because aged vessels show characteristics of atherosclerosis, such as reduced number of vascular smooth muscle cells, increased collagen deposition, and fractured elastin, aging per se is a risk factor for atherosclerosis. Under these conditions, low-density lipoprotein (LDL) cholesterol, which directly accumulates in arterial walls, presents the most prominent coronary risk factor for atherosclerosis. Homozygous FH, usually defined as FH with homozygous mutation, and heterozygous FH are typical clinical models of atherosclerotic cardiovascular disease leading to premature coronary artery disease due to extreme hyper-LDL cholesterolemia. There exits the homozygous FH due to LDL receptor and other different gene mutations as double heterozygous FH. However, the most common genetic cause of FH, even in homo- and hetero-zygotes, is a mutation or defect in the LDL receptor gene associated with intracellular trafficking of LDL cholesterol (Fig. 1). Thus, homozygous FH, lacking the LDL receptor, is highly resistant to cholesterol-lowering medical therapy including HMGCoA reductase inhibitors (statins), although heterozygous FH was observed partially responding to statins [3]. A mutation in the apolipoprotein B-100 gene is also known to be a cause of FH. However, few FH patients with the apolipoprotein B-100 gene mutation had been detected in Japan [8, 9].
Autosomal recessive hypercholesterolemia (ARH) is a rare disorder; the reported number of patients with clinical manifestations resembling homozygous FH is approximately 70 individuals worldwide. The first ARH patient was reported in Japan [10], and we recently found the second ARH family [11]. The cause of ARH is a mutation of LDL receptor adaptor protein-1 (LDLRAP1) (Fig. 1). The clinical manifestations and severity of ARH differs from those in homozygous and heterozygous FH. Lipid reduction therapy including the use of statins is at least partially effective [12], because intracellular trafficking of very LDL via the LDL receptor without LDLRAP1 is preserved [13]. We performed a lipoprotein kinetic study in a patient with ARH and concluded that the fractional catabolic rate of LDL apolipoprotein B-100 was significantly lower and the direct removal of the very LDL remnant was significantly greater than that seen in normal controls [12, 14]. This suggests the presence of an alternate pathway of LDL cholesterol in ARH, which may be a new target of lipid-lowering therapy.
Recently, proprotein convertase subtilisin/kexin 9 (PCSK9), associated with intracellular degeneration of LDL receptor, was found as a causal gene for FH [15] (Fig. 1). We identified an E32 K mutation in the PCSK9 gene in patients with the milder FH phenotype. The frequency of PCSK9 E32 K carriers is 1.7 % in the general population; the LDL cholesterol serum level of PCSK9 E32 K carriers is significantly higher than that seen in the general population, but significantly lower than seen in individuals with the heterozygous LDL receptor gene mutation [16]. Plasma levels of PCSK9 increased following cholesterol-lowering medical therapy such as statins or fibrates, and may cancel out at least a part of the LDL cholesterol-lowering effect [17, 18], although there was no difference in increased PCSK9 levels between different statins such as pitavastatin and pravastatin [19]. Recently, treatment with anti-PCSK9 antibody was demonstrated to reduce cholesterol levels in animal as well as human trials, and further clinical trials will demonstrate its efficacy [20–28]. The S447X variant of the lipoprotein lipase gene is inversely associated with severity of coronary artery disease [29], probably due to functional loss of lipoprotein lipase associated with hyperlipoproteinemia.
Intensive genetic diagnosis sometimes reveals misdiagnosis of homozygous FH as heterozygous FH in patients with severe hypercholesterolemia without consanguinity or family history who respond to medical cholesterol-lowering therapy. For instance, an additional mutation in LDLRAP1 gene worsens the clinical manifestation such as increases in LDL cholesterol levels and xanthomas in FH with a single LDL receptor gene mutation [30]. Interestingly, genetic diagnosis of FH reveals that the frequency of FH is higher than previously estimated [3]. Because the available intensive LDL cholesterol-lowering regimens can reduce serum levels of LDL cholesterol < 100 mg/dl [17], accurate diagnosis of FH based on genetic methods should be considered before starting therapy. Plasma LDL apheresis instead of conventional statin therapy in FH could result in regression of coronary plaques [31].
Previous studies showed that high-density lipoprotein (HDL) cholesterol levels are associated with decreased frequencies of cardiovascular events [32]. Reverse cholesterol transport as well as anti-oxidant, anti-inflammatory, and anti-thrombotic effects are known to be antiatherogenic function of HDL. Statin therapy effectively increases HDL cholesterol levels and seems to be associated with regression of coronary atheroma [33, 34]. The study of genetic disorders associated with HDL, including cholesteryl ester transfer protein deficiency, first reported by our research group [35], has led to several developments in terms of genetic and functional analyses [36–38], as well as possible therapeutic implications [39, 40]. The reverse cholesterol transport-promoting concept has been a promising alternative to simple HDL-increasing therapy [41, 42], although some clinical trials have failed to demonstrate their efficacy [43, 44]. Additionally, genetic mechanisms that raise plasma HDL cholesterol do not seem to lower the risk of myocardial infarction [45]. These data challenge the concept that raising plasma HDL cholesterol will or will not uniformly translate into reductions of risk for myocardial infarction.
Hypertrophic cardiomyopathy
Hypertrophic cardiomyopathy is a heritable myocardial disorder characterized by increased ventricular wall thickness in the absence of other loading conditions, such as aortic stenosis or hypertension. Histopathologic abnormalities of HCM include myocyte hypertrophy, myocyte disarray, and increased interstitial and/or focal fibrosis. These anatomic and histopathologic findings contribute to diastolic dysfunction with impaired filling due to increased chamber stiffness, leading to elevated left ventricular end-diastolic and left atrial pressures. This condition is a slowly progressive disorder that manifests remarkably variable clinical courses from near-normal life expectancy with no symptoms to sudden cardiac death (SCD) in youth [4, 46].
The estimated prevalence of HCM in the general population is approximately 1 in 500 [47]; approximately half of the cases have a family history of HCM and the mode of inheritance is autosomal dominant. Since 1990, when a missense mutation in the β-myosin heavy chain (MYH7) gene was identified in patients with HCM [48], more than 17 disease-causing genes and several hundred mutations have been discovered [47] (Table 1). Mutations in the MYH7 and MYBPC3 genes are responsible for HCM in approximately 50 % of patients. Mutations in other sarcomere genes (TNNT2, TNNI3, TPM1, and ACTC1) account for about 10–15 % of HCM cases. Mutations in the genes that encode Z-disk-related proteins (including the genes MYOZ2, TCAP, and ANKRD1) are uncommon causes of HCM [4, 47, 49]. Collectively, the known disease-causing genes account for about two-thirds of HCM cases.
The majority of mutations are specific to a patient and the patient’s family members, and different mutations are usually identified in unrelated HCM families [47]. Therefore, identifying the mutation in HCM patients requires screening for all disease-causing genes, which also requires extensive time and cost. With recently developed high-throughput DNA sequencing technologies, comprehensive genetic diagnosis in HCM may now be available [50].
Genetic testing provides opportunities to assess the relevance of the genotype in the phenotype of HCM, and numerous genotype–phenotype correlations have been identified from family studies. Clinical manifestations of HCM caused by TNNT2 gene mutations often begin near adolescence, while MYBPC3 gene mutations typically trigger HCM in middle age [47, 51]. We recently demonstrated that MYBPC3 mutation carriers developed left ventricular systolic dysfunction less frequently than non-MYBPC3 mutation carriers [52].
Several studies have indicated that genetic modifiers such as polymorphisms in the renin–angiotensin–aldosterone system genes may influence the extent of left ventricular mass or left ventricular function, which could in part explain a wide spectrum of clinical expression in HCM [53]. Furthermore, HCM patients who carry more than one independent mutation may be at a higher risk for adverse clinical courses, which include development of advanced heart failure with left ventricular systolic dysfunction [54–56] (Fig. 2). Additionally, gender may have some influence on the development of ventricular hypertrophy in HCM. Because HCM is inherited in an autosomal dominant pattern, an equal ratio of males to females would be anticipated. However, previous reports of HCM cohorts have demonstrated a male predominance, with approximately 55–60 % of HCM probands [57]. It has been postulated that chromosomal or hormonal-specific factors and gender-specific single-nucleotide polymorphisms may influence the pathogenesis of hypertrophy development, which was observed in animal models [57, 58]. Although genotype–phenotype correlations may not be applied to therapy for HCM, one clinical advantage of genetic testing is gene-based preclinical diagnosis of the condition [59]. Prior to the development of genetic testing, clinicians were not able to predict whether young family members without ventricular hypertrophy would develop HCM later in life. With genetic testing, preclinical mutation carriers and non-carriers can be distinguished, which provides a relief for non-carriers and may contribute to better clinical management for the carriers [60–62].
Modifying factors of hypertrophic cardiomyopathy (HCM) phenotypes. Certain sarcomere gene mutations may be associated with malignant clinical courses. Polymorphisms in the renin–angiotensin–aldosterone system genes may influence HCM phenotypes. Furthermore, HCM patients who carry more than one independent mutation may exhibit severe phenotypes
Identification of disease-causing mutations enables researchers to generate genetically engineered animal models that harbor human HCM mutations [4, 63, 64]. From the studies of HCM animal as well as cellular models [65], molecular pathways such as transforming growth factor-beta1/Smads signals that trigger ventricular hypertrophy, fibrosis, and remodeling have been revealed [66], which could provide future therapeutic targets. There exists indirect evidence that the nonpeptide AVE0991 might attenuate myocardial hypertrophy as induced by angiotensin II through down-regulation of transforming growth factor-beta1/Smad2 expression [67]. Thus, genetic testing in HCM may contribute to gene-based accurate diagnosis, identifying “malignant” mutations in specific patients, and preclinical diagnosis in subjects who do not exhibit ventricular hypertrophy, although the clinical course of each HCM patient may not be predicted based solely on genotype.
Inherited cardiac arrhythmia syndrome
A line of evidence demonstrated that inherited tachycardia or bradycardia diseases are associated with monogenic gene mutations [68, 69]. LQTS is characterized by prolonged ventricular repolarization and a propensity for life-threatening ventricular tachyarrhythmias, typically torsades de pointes, resulting in syncopal attacks and sudden death [70]. Before puberty, boys with LQTS are at three to fourfold higher risk for cardiac events including syncope, aborted cardiac arrest (ACA), or SCD compared with respective females [71, 72]. In contrast, adult women in the range of 18–40 years have a significant 2.7-fold increase in the risk of ACA or SCD as compared with men [73]. LQTS can be subclassified into congenital and acquired forms. Genetic testing can identify a mutation in 50–75 % of clinically affected patients with congenital LQTS (cLQTS) [74–76], as well as some patients with acquired LQTS (aLQTS) [77–80]. Thirteen genetic forms of LQTS have been described, and the most prevalent forms are LQT1 and LQT2 associated with mutations in potassium channels, and LQT3 with a sodium channel mutation [68, 81–83] (Table 2).
The KCNQ1 and KCNE1, which are also expressed in the experimental animal such as a swine [84] genes encode the α and the β subunit, respectively, of the potassium channel conducting the IKs current underlying the LQT1 and LQT5 forms. The AKAP-9 encoding Yotiao, which assembles KCNQ1, was reported to be linked to the LQT11 form. The KCNH2 and the KCNE2 genes encode the α and the β subunit, respectively, of the potassium channel conducting the IKr current underlying the LQT2 and LQT6 forms [85]. Interestingly, there exists LQTS caused by a KCNQ1 mutation associated with left ventricular noncompaction [86].
The KCNJ2 gene encodes the inward rectifier potassium current (IK1) underlying Andersen’s syndrome (LQT7), with associated QT prolongation and ventricular arrhythmias. A mutation in KCNJ5 encoding the α-subunit of the acetylcholine-sensitive potassium current (IK-ACh) channel was reported to be responsible for LQT13.
The SCN5A gene encodes the protein of the cardiac sodium channel underlying the LQT3 form [87] (Fig. 3). The CAV3 encoding caveolin-3 and SCN4B encoding Navβ4, an auxiliary subunit of the cardiac sodium channel, are also reported to be associated with the LQT9 and LQT10 forms. The SNTA1 encoding a cytoskeletal protein syntrophin-α1 interacts with the cardiac sodium channel underlying the LQT12 form. A mutation in the CACNA1C gene results in an increase in Cav1.2 current, QT prolongation, and a phenotype that is characterized by syndactyly in both hands and feet and multiorgan dysfunction (Timothy’s syndrome) (Fig. 3). LQT4 is caused by mutations in the ankyrin B gene that produces a protein that functions as a membrane of versatile membrane adapters.
Life-threatening cardiac events tend to occur under specific circumstances in a gene-specific manner. Syncope or sudden death in LQT1 patients is triggered by emotional or physical stress such as diving and swimming. LQT1 patients tend to have an optimal response to β-blockers compared to another forms of LQTS. Interestingly, C-loop missense mutations in the KCNQ1 channel present a high risk for life-threatening events and have a pronounced benefit of treatment with β-blockades [88]. It is known that effectiveness is not equal for different β-blockers. A recent study shows that propranolol has a significantly better QTc-shortening effect compared to metoprolol and nadolol [89]. In addition, symptomatic LQTS patients treated with metoprolol are four times more likely to have breakthrough cardiac events than those treated with propranolol and nadolol [89]. Emotion and noise have been noted to cause cardiac symptoms more frequently in LQT2. LQT2 patients are particularly sensitive to sudden noise, such as a telephone or alarm clock ring, but β-blockers have been found to reduce cardiac events in patients with LQT2. Increased extracellular potassium concentration has been reported to shorten the QT interval in LQT2 patients. Patients with LQT3 have the highest risk of events when at rest or asleep and a relatively low risk of events during arousal. Sodium-channel blockers such as mexiletine may normalize the QTc interval in patients with LQT3.
Disease penetrance is low (= 25 %) in some families with cLQTS [90], and QT duration appears normal in about 30 % of LQTS mutation carriers [91]. Electrophysiological studies to date show that channel function varies with different mutations and mutation sites. The pore region of the hERG channel, which was encoded by the KCNH2 gene, provides an aqueous pathway for potassium ions. Most mutations involving this region are missense mutations with dominant-negative effects on IKr, and subjects with pore mutations exhibit a severe clinical course. In contrast, most mutations in the non-pore region of the KCNH2 gene are known to not exhibit dominant-negative effects, resulting in mild clinical phenotypes [92]. However, we recently identified a novel mutation of KCNH2 T473P in the non-pore region in patients with the congenital or acquired form of LQTS, which result in protein trafficking defects and exhibit dominant negative effects with severe phenotype [93]. These mutations seem to be concentrated in the region between S2 and S2-S3 linker of the hERG channel and may cause severe clinical course in affected patients [94]. In patients with LQTS caused by a mutation leading to mild dysfunction of a cardiac ion channel, marked QT prolongation or torsade de pointes may appear only upon the addition of a stressor such as a drug, or under conditions such as hypokalemia or bradycardia [95]. Up to 40 % of aLQTS patients with torsades de pointes carry mutations in KCNQ1, KCNH2, KCNE1, KCNE2, KCNE3, and SCN5A [77–80].
Previous study reported that the K+ channel regulator 1 (KCR1) protected KCNH2 current from drug block [96, 97]. Another study showed that the I447 V variant of KCR1 occurred less frequently in patients with drug-induced torsade de pointes compared to controls, and this variant was more effective at protecting KCNH2 against dofetilide inhibition in a heterologous expression study [98]. We identified a KCR1 genetic variant (E33D) in a patient who suffered ventricular fibrillation and QT prolongation; this variant diminished the ability of KCR1 to protect KCNH2 from inhibition by commonly used therapeutic agents and constitutes a risk factor for aLQTS [99] (Fig. 3).
Congenital short QT syndrome (SQT) is a hereditary disease characterized by a shorter than normal QT interval (< 350 ms), ventricular and atrial arrhythmias, and SCD [100, 101]. Some patients with SQTS develop lethal arrhythmias within the first year of life, and this syndrome is thought to be one of the causes of sudden infant death syndrome. Five ion channel genes were reported to be responsible for SQTS [102] (Table 2). In heterologous expression systems, mutations in KCNH2 (SQT1), KCNQ1 (SQT2), and KCNJ2 (SQT3) genes showed an increase in the potassium currents involved, resulting in short QT intervals. In contrast, mutations in the CACNA1C (SQT4) and CACNB2b (SQT5) showed loss of function of Calcium channels [102].
Brugada syndrome (BrS) is a channelopathy associated with conduction delay and ST-segment elevation of the right precordial leads with or without right bundle branch block in the absence of structural disease, characterized by syncope and premature sudden death due to ventricular fibrillation [103]. The ECG abnormality can be manifested by administration of channel blocker or by environmental influences, including fever or large meals. BrS affects men more commonly and severely than women. Previous study for a series of 384 patients with BrS showed that men were predominant (70.8 %) and had experienced syncope or aborted SCD more frequently than woman, as well as a greater event rate in follow-up [104].
At least 15 genes were reported to be linked to BrS [81, 105–113] (Table 2). There is no consensus on the relation between BrS types 8 and higher and responsible genes, probably because some associated mutations are not universally considered pathogenic. Mutations of SCN5A are reported to account for 18–30 % of clinically diagnosed BrS patients at present, and these mutations cause loss of function in the sodium current (BrS1). Less commonly, BrS may result from mutations in CACNA1C (BrS2) and CACNB2b (BrS3) through a reduction in the L-type calcium current, or from GPD1-L (BrS4), SCN1B (BrS5), or SCN3B (BrS7) causing a loss of sodium channel function. The KCNE3 mutation identified in the BrS family reduces the inhibitory effect of KCNE3 on the Kv4.3 channel, resulting in an increase in transient outward current (BrS6). Symptomatic patients with VF or syncope had higher recurrence rates of cardiac events compared to asymptomatic patients [114, 115], and therefore the symptomatic patients should be considered to receive an ICD for secondary and primary prevention of SCD. Risk stratification in asymptomatic patients is still controversial. Recent studies showed that a family history of SCD and the presence of early repolarization were predictors of a poor outcome in this patient group [115, 116].
Catecholaminergic polymorphic ventricular tachycardia (CPVT) is a familial disease characterized by stress-induced ventricular arrhythmias that result in syncope and sudden death in children or young adults [117]. The resting ECG is unremarkable: therefore, the diagnosis is mostly based on symptoms and on the detection of stress-induced arrhythmias during exercise stress test or continuous ECG recording. Three genes were reported to be responsible for CPVT. Mutations in the RYR2 gene, encoding the cardiac ryanodine receptor, account for autosomal dominant disease [118, 119]. In addition, mutations in the cardiac calsequestrin gene CASQ2 account for a small number of autosomal recessive cases [120]. In addition, mutations in KCNJ2 associated with Anderson syndrome, TRDN encoding triadin, which links RYR2 with calsequestrin, or CALM1, encoding the calcium-modulated protein calmodulin 1 may underlie the minority of cases [121–123] (Table 2). Under these conditions, calcium channel inhibition by verapamil rectified abnormal calcium handling in CPVT myocytes and prevented ventricular arrhythmias [124], however, the effect of verapamil is still controversial. Recent studies showed that flecainide reduced exercise-induced ventricular arrhythmias in patients with CPVT not controlled by conventional drug therapy [125, 126]. Therefore, flecainide in addition to β-blocker therapy including carvedilol should be considered for these CPVT patients.
Recently, fibroblasts derived from patients with LQTS were reprogrammed and the induced pluripotent stem (iPS) cells were directed to differentiate into cardiac myocytes. The patient-derived cells showed the electrophysiological properties of the disorder [127, 128]. Application of iPS cell-derived cardiomyocytes to study inherited cardiovascular diseases will enable us to understand the molecular mechanisms of human disease in more detail, although it is still controversial whether iPS cell-derived cells can represent mature ones in terms of channel expression [129].
Conclusions
We reviewed the current perspectives in cardiovascular genetic diseases such as FH, HCM, and hereditary arrhythmic syndrome, all of which are major fields of cardiovascular research. Functional analysis of disease-causing gene mutations will give us new insight into the mechanism of each disease, while development of newer technology for gene analysis may identify additional novel gene mutations.
References
Nabel EG (2012) Molecular biology and genetics. Principles of cardiovascular molecular biology and genetics. In: Bonow RO, Mann DL, Zipes DP, Lippy P (eds) A textbook of cardiovascular medicine, 9th edn. WB Saunders, New York, pp 57–69
Yamagishi M, Hayashi K, Konno T, Kawashiri MA, Ino H, Fujino N (2012) Current perspectives of genetic cardiovascular disease. J Cardiol Jpn Ed 6:105–114
Mabuchi H, Nohara A, Noguchi T, Kobayashi J, Kawashiri MA, Tada H, Nakanishi C, Mori M, Yamagishi M, Inazu A, Koizumi J (2011) Molecular genetic epidemiology of homozygous familial hypercholesterolemia in the Hokuriku district of Japan. Atherosclerosis 214:404–407
Konno T, Chang S, Seidman JG, Seidman CE (2010) Genetics of hypertrophic cardiomyopathy. Curr Opin Cardiol 25:205–209
Nannenberg EA, Sijbrands EJ, Dijksman LM, Alders M, van Tintelen JP, Birnie M, van Langen IM, Wilde AA (2012) Mortality of inherited arrhythmia syndromes: insight into their natural history. Circ Cardiovasc Genet 5:183–189
Kannel WB, McGee D, Gordon T (1976) A general cardiovascular risk profile: the Framingham Study. Am J Cardiol 38:46–51
Hulley S, Grady D, Bush T, Furberg C, Herrington D, Riggs B, Vittinghoff E (1998) Randomized trial of estrogen plus progestin for secondary prevention of coronary heart disease in postmenopausal women. Heart and Estrogen/progestin Replacement Study (HERS) Research Group. JAMA 280:605–613
Hansen PS, Rüdiger N, Tybjaerg-Hansen A, Faergeman O, Gregersen N (1991) Detection of the apoB-3500 mutation (glutamine for arginine) by gene amplification and cleavage with MspI. J Lipid Res 32:1229–1233
Nohara A, Yagi K, Inazu A, Kajinami K, Koizumi J, Mabuchi H (1995) Absence of familial defective apolipoprotein B-100 in Japanese patients with familial hypercholesterolaemia. Lancet 345:1438
Harada-Shiba M, Tajima S, Yokoyama S, Miyake Y, Kojima S, Tsushima M, Kawakami M, Yamamoto A (1992) Siblings with normal LDL receptor activity and severe hypercholesterolemia. Arterioscler Thromb 12:1071–1078
Tada H, Kawashiri M, Noguchi T, Nakanishi C, Tsuchida M, Takata M, Nohara A, Inazu A, Kobayashi J, Mabuchi H, Yamagishi M (2011) A novel type of homozygous familial hypercholesterolemia: double heterozygous mutations in LDL receptor and LDL receptor adaptor protein 1 gene (abstr). Circulation 124:A8511
Tada H, Kawashiri M, Ikewaki K, Terao Y, Noguchi T, Nakanishi C, Tsuchida M, Takata M, Miwa K, Konno T, Hayashi K, Nohara A, Inazu A, Kobayashi J, Mabuchi H, Yamagishi M (2012) Altered metabolism of low-density lipoprotein and very-low-density lipoprotein remnant in autosomal recessive hypercholesterolemia: results from stable isotope kinetic study in vivo. Circ Cardiovasc Genet 5:35–41
Jones C, Garuti R, Michaely P, Li WP, Maeda N, Cohen JC, Herz J, Hobbs HH (2007) Disruption of LDL but not VLDL clearance in autosomal recessive hypercholesterolemia. J Clin Investig 117:165–174
Tada H, Kawashiri MA, Tanaka A, Nakano T, Nakajima K, Inoue T, Noguchi T, Nakanishi C, Konno T, Hayashi K, Nohara A, Inazu A, Kobayashi J, Mabuchi H, Yamagishi M (2012) Post-prandial remnant lipoprotein metabolism in autosomal recessive hypercholesterolaemia. Eur J Clin Investig 42:1094–1099
Abifadel M, Varret M, Rabes JP, Allard D, Ouguerram K, Devillers M, Cruaud C, Benjannet S, Wickham L, Erlich D, Derre A, Villeger L, Farnier M, Beucler I, Bruckert E, Chambaz J, Chanu B, Lecerf JM, Luc G, Moulin P, Weissenbach J, Prat A, Krempf M, Junien C, Seidah NG, Boileau C (2003) Mutations in PCSK9 cause autosomal dominant hypercholesterolemia. Nat Genet 34:154–156
Noguchi T, Katsuda S, Kawashiri MA, Tada H, Nohara A, Inazu A, Yamagishi M, Kobayashi J, Mabuchi H (2010) The E32 K variant of PCSK9 exacerbates the phenotype of familial hypercholesterolaemia by increasing PCSK9 function and concentration in the circulation. Atherosclerosis 210:166–172
Kawashiri MA, Nohara A, Noguchi T, Tada H, Nakanishi C, Mori M, Konno T, Hayashi K, Fujino N, Inazu A, Kobayashi J, Mabuchi H, Yamagishi M (2012) Efficacy and safety of coadministration of rosuvastatin, ezetimibe, and colestimide in heterozygous familial hypercholesterolemia. Am J Cardiol 109:364–369
Noguchi T, Kobayashi J, Yagi K, Nohara A, Yamaaki N, Sugihara M, Ito N, Oka R, Kawashiri MA, Tada H, Takata M, Inazu A, Yamagishi M, Mabuchi H (2011) Comparison of effects of bezafibrate and fenofibrate on circulating proprotein convertase subtilisin/kexin type 9 and adipocytokine levels in dyslipidemic subjects with impaired glucose tolerance or type 2 diabetes mellitus: results from a crossover study. Atherosclerosis 217:165–170
Nozue T, Hattori H, Ishihara M, Iwasaki T, Hirano T, Kawashiri MA, Yamagishi M, Michishita I (2013) Comparison of effects of pitavastatin versus pravastatin on serum proprotein convertase subtilisin/kexin type 9 levels in statin-naive patients with coronary artery disease. Am J Cardiol 111:1415–1419. doi:10.1016/j.amjcard.2013.01.289
Chaparro-Riggers J, Liang H, DeVay RM, Bai L, Sutton JE, Chen W, Geng T, Lindquist K, Casas MG, Boustany LM, Brown CL, Chabot J, Gomes B, Garzone P, Rossi A, Strop P, Shelton D, Pons J, Rajpal A (2012) Increasing serum half-life and extending cholesterol lowering in vivo by engineering antibody with pH-sensitive binding to PCSK9. J Biol Chem 287:11090–11097
Dias CS, Shaywitz AJ, Wasserman SM, Smith BP, Gao B, Stolman DS, Crispino CP, Smirnakis KV, Emery MG, Colbert A, Gibbs JP, Retter MW, Cooke BP, Uy ST, Matson M, Stein EA (2012) Effects of AMG 145 on low-density lipoprotein cholesterol levels: results from 2 randomized, double-blind, placebo-controlled, ascending-dose phase 1 studies in healthy volunteers and hypercholesterolemic subjects on statins. J Am Coll Cardiol 60:1888–1898
Stein EA, Mellis S, Yancopoulos GD, Stahl N, Logan D, Smith WB, Lisbon E, Gutierrez M, Webb C, Wu R, Du Y, Kranz T, Gasparino E, Swergold GD (2012) Effect of a monoclonal antibody to PCSK9 on LDL cholesterol. N Engl J Med 366:1108–1118
McKenney JM, Koren MJ, Kereiakes DJ, Hanotin C, Ferrand AC, Stein EA (2012) Safety and efficacy of a monoclonal antibody to proprotein convertase subtilisin/kexin type 9 serine protease, SAR236553/REGN727, in patients with primary hypercholesterolemia receiving ongoing stable atorvastatin therapy. J Am Coll Cardiol 59:2344–2353
Roth EM, McKenney JM, Hanotin C, Asset G, Stein EA (2012) Atorvastatin with or without an antibody to PCSK9 in primary hypercholesterolemia. N Engl J Med 367:1891–1900
Sullivan D, Olsson AG, Scott R, Kim JB, Xue A, Gebski V, Wasserman SM, Stein EA (2012) Effect of a monoclonal antibody to PCSK9 on low-density lipoprotein cholesterol levels in statin-intolerant patients: the GAUSS randomized trial. JAMA 308:2497–2506
Raal F, Scott R, Somaratne R, Bridges I, Li G, Wasserman SM, Stein EA (2012) Low-density lipoprotein cholesterol-lowering effects of AMG 145, a monoclonal antibody to proprotein convertase subtilisin/kexin type 9 serine protease in patients with heterozygous familial hypercholesterolemia: the Reduction of LDL-C with PCSK9 Inhibition in Heterozygous Familial Hypercholesterolemia Disorder (RUTHERFORD) randomized trial. Circulation 126:2408–2417
Koren MJ, Scott R, Kim JB, Knusel B, Liu T, Lei L, Bolognese M, Wasserman SM (2012) Efficacy, safety, and tolerability of a monoclonal antibody to proprotein convertase subtilisin/kexin type 9 as monotherapy in patients with hypercholesterolaemia (MENDEL): a randomised, double-blind, placebo-controlled, phase 2 study. Lancet 380:1995–2006
Giugliano RP, Desai NR, Kohli P, Rogers WJ, Somaratne R, Huang F, Liu T, Mohanavelu S, Hoffman EB, McDonald ST, Abrahamsen TE, Wasserman SM, Scott R, Sabatine MS, LAPLACE-TIMI 57 Investigators (2012) Efficacy, safety, and tolerability of a monoclonal antibody to proprotein convertase subtilisin/kexin type 9 in combination with a statin in patients with hypercholesterolaemia (LAPLACE-TIMI 57): a randomised, placebo-controlled, dose-ranging, phase 2 study. Lancet 380:2007–2017
Agirbasli M, Sumerkan MC, Eren F, Agirbasli D (2011) The S447X variant of lipoprotein lipase gene is inversely associated with severity of coronary artery disease. Heart Vessels 26:457–463
Tada H, Kawashiri MA, Ohtani R, Noguchi T, Nakanishi C, Konno T, Hayashi K, Nohara A, Inazu A, Kobayashi J, Mabuchi H, Yamagishi M (2011) A novel type of familial hypercholesterolemia: double heterozygous mutations in LDL receptor and LDL receptor adaptor protein 1 gene. Atherosclerosis 219:663–666
Matsuzaki M, Hiramori K, Imaizumi T, Kitabatake A, Hishida H, Nomura M, Fujii T, Sakuma I, Fukami K, Honda T, Ogawa H, Yamagishi M (2002) Intravascular ultrasound evaluation of coronary plaque regression by low density lipoprotein-apheresis in familial hypercholesterolemia: the Low Density Lipoprotein-Apheresis Coronary Morphology and Reserve Trial (LACMART). J Am Coll Cardiol 40:220–227
Gordon T, Castelli WP, Hjortland MC, Kannel WB, Dawber TR (1977) High density lipoprotein as a protective factor against coronary heart disease. The Framingham Study. Am J Med 62:707–714
Takayama T, Hiro T, Yamagishi M, Daida H, Hirayama A, Saito S, Yamaguchi T, Matsuzaki M, COSMOS Investigators (2009) Effect of rosuvastatin on coronary atheroma in stable coronary artery disease: multicenter coronary atherosclerosis study measuring effects of rosuvastatin using intravascular ultrasound in Japanese subjects (COSMOS). Circ J 73:2110–2117
Kawashiri M, Yamagishi M, Sakamoto T, Takayama T, Hiro T, Daida H, Hirayama A, Saito S, Yamaguchi T, Matsuzaki M, The COSMOS Investigators (2013) Impact of intensive lipid lowering on lipid profiles over time and tolerability in stable coronary artery disease: insights from a subanalysis of the coronary atherosclerosis study measuring effects of rosuvastatin using intravascular ultrasound in Japanese subjects (COSMOS). Cardiovasc Ther (Epub ahead of print)
Inazu A, Brown ML, Hesler CB, Agellon LB, Koizumi J, Takata K, Maruhama Y, Mabuchi H, Tall AR (1990) Increased high-density lipoprotein levels caused by a common cholesteryl-ester transfer protein gene mutation. N Engl J Med 323:1234–1238
Inazu A, Nakajima K, Nakano T, Niimi M, Kawashiri MA, Nohara A, Kobayashi J, Mabuchi H (2008) Decreased post-prandial triglyceride response and diminished remnant lipoprotein formation in cholesteryl ester transfer protein (CETP) deficiency. Atherosclerosis 196:953–957
Miwa K, Inazu A, Kawashiri M, Nohara A, Higashikata T, Kobayashi J, Koizumi J, Nakajima K, Nakano T, Niimi M, Mabuchi H, Yamagishi M (2009) Cholesterol efflux from J774 macrophages and Fu5AH hepatoma cells to serum is preserved in CETP-deficient patients. Clin Chim Acta 402:19–24
Ohtani R, Inazu A, Noji Y, Wakasugi T, Miwa K, Tada H, Kawashiri MA, Noguchi T, Nohara A, Kobayashi J, Koizumi J, Yamagishi M, Mabuchi H (2012) Novel mutations of cholesteryl ester transfer protein (CETP) gene in Japanese hyperalphalipoproteinemic subjects. Clin Chim Acta 413:537–543
Cannon CP, Shah S, Dansky HM, Davidson M, Brinton EA, Gotto AM, Stepanavage M, Liu SX, Gibbons P, Ashraf TB, Zafarino J, Mitchel Y, Barter P, Determining the Efficacy and Tolerability Investigators (2010) Safety of anacetrapib in patients with or at high risk for coronary heart disease. N Engl J Med 363:2406–2415
Nicholls SJ, Brewer HB, Kastelein JJ, Krueger KA, Wang MD, Shao M, Hu B, McErlean E, Nissen SE (2011) Effects of the CETP inhibitor evacetrapib administered as monotherapy or in combination with statins on HDL and LDL cholesterol: a randomized controlled trial. JAMA 306:2099–2109
Kawashiri MA, Zhang Y, Puré E, Rader DJ (2002) Combined effects of cholesterol reduction and apolipoprotein A-I expression on atherosclerosis in LDL receptor deficient mice. Atherosclerosis 165:15–22
Moore RE, Kawashiri MA, Kitajima K, Secreto A, Millar JS, Pratico D, Rader DJ (2003) Apolipoprotein A-I deficiency results in markedly increased atherosclerosis in mice lacking the LDL receptor. Arterioscler Thromb Vasc Biol 23:1914–1920
Nissen SE, Tardif JC, Nicholls SJ, Revkin JH, Shear CL, Duggan WT, Ruzyllo W, Bachinsky WB, Lasala GP, Tuzcu EM, ILLUSTRATE Investigators (2007) Effect of torcetrapib on the progression of coronary atherosclerosis. N Engl J Med 356:1304–1316
Schwartz GG, Olsson AG, Abt M, Ballantyne CM, Barter PJ, Brumm J, Chaitman BR, Holme IM, Kallend D, Leiter LA, Leitersdorf E, McMurray JJ, Mundl H, Nicholls SJ, Shah PK, Tardif JC, Wright RS, dal-OUTCOMES Investigators (2012) Effects of dalcetrapib in patients with a recent acute coronary syndrome. N Engl J Med 367:2089–2099
Voight BF, Peloso GM, Orho-Melander M, Frikke-Schmidt R, Barbalic M, Jensen MK, Hindy G, Hólm H, Ding EL, Johnson T, Schunkert H, Samani NJ, Clarke R, Hopewell JC, Thompson JF, Li M, Thorleifsson G, Newton-Cheh C, Musunuru K, Pirruccello JP, Saleheen D, Chen L, Stewart A, Schillert A, Thorsteinsdottir U, Thorgeirsson G, Anand S, Engert JC, Morgan T, Spertus J, Stoll M, Berger K, Martinelli N, Girelli D, McKeown PP, Patterson CC, Epstein SE, Devaney J, Burnett MS, Mooser V, Ripatti S, Surakka I, Nieminen MS, Sinisalo J, Lokki ML, Perola M, Havulinna A, de Faire U, Gigante B, Ingelsson E, Zeller T, Wild P, de Bakker PI, Klungel OH, Maitland-van der Zee AH, Peters BJ, de Boer A, Grobbee DE, Kamphuisen PW, Deneer VH, Elbers CC, Onland-Moret NC, Hofker MH, Wijmenga C, Verschuren WM, Boer JM, van der Schouw YT, Rasheed A, Frossard P, Demissie S, Willer C, Do R, Ordovas JM, Abecasis GR, Boehnke M, Mohlke KL, Daly MJ, Guiducci C, Burtt NP, Surti A, Gonzalez E, Purcell S, Gabriel S, Marrugat J, Peden J, Erdmann J, Diemert P, Willenborg C, König IR, Fischer M, Hengstenberg C, Ziegler A, Buysschaert I, Lambrechts D, Van de Werf F, Fox KA, El Mokhtari NE, Rubin D, Schrezenmeir J, Schreiber S, Schäfer A, Danesh J, Blankenberg S, Roberts R, McPherson R, Watkins H, Hall AS, Overvad K, Rimm E, Boerwinkle E, Tybjaerg-Hansen A, Cupples LA, Reilly MP, Melander O, Mannucci PM, Ardissino D, Siscovick D, Elosua R, Stefansson K, O'Donnell CJ, Salomaa V, Rader DJ, Peltonen L, Schwartz SM, Altshuler D, Kathiresan S (2012) Plasma HDL cholesterol and risk of myocardial infarction: a mendelian randomisation study. Lancet 380:572–580 (Erratum in: Lancet. 2012;380:564)
Konno T, Shimizu M, Ino H, Matsuyama T, Yamaguchi M, Terai H, Hayashi K, Mabuchi T, Kiyama M, Sakata K, Hayashi T, Inoue M, Kaneda T, Mabuchi H (2003) A novel missense mutation in the myosin binding protein-C gene is responsible for hypertrophic cardiomyopathy with left ventricular dysfunction and dilation in elderly patients. J Am Coll Cardiol 41:781–786
Wang L, Seidman JG, Seidman CE (2010) Narrative review: harnessing molecular genetics for the diagnosis and management of hypertrophic cardiomyopathy. Ann Intern Med 152:513–520
Geisterfer-Lowrance AA, Kass S, Tanigawa G, Vosberg HP, McKenna W, Seidman CE, Seidman JG (1990) A molecular basis for familial hypertrophic cardiomyopathy: a beta cardiac myosin heavy chain gene missense mutation. Cell 62:999–1006
Arimura T, Bos JM, Sato A, Kubo T, Okamoto H, Nishi H, Harada H, Koga Y, Moulik M, Doi YL, Towbin JA, Ackerman MJ, Kimura A (2009) Cardiac ankyrin repeat protein gene (ANKRD1) mutations in hypertrophic cardiomyopathy. J Am Coll Cardiol 54:334–342
Lopes LR, Zekavati A, Syrris P, Hubank M, Giambartolomei C, Dalageorgou C, Jenkins S, McKenna W, Uk10 k Consortium, Plagnol V, Elliott PM (2013) Genetic complexity in hypertrophic cardiomyopathy revealed by high-throughput sequencing. J Med Genet 50:228–239
Anan R, Niimura H, Takenaka T, Hamasaki S, Tei C (2007) Mutations in the genes for sarcomeric proteins in Japanese patients with onset sporadic hypertrophic cardiomyopathy after age 40 years. Am J Cardiol 99:1750–1754
Fujino N, Konno T, Hayashi K, Hodatsu A, Fujita T, Tsuda T, Nagata Y, Kawashiri MA, Ino H, Yamagishi M (2013) Impact of systolic dysfunction in genotyped hypertrophic cardiomyopathy. Clin Cardiol 36:160–165
Funada A, Konno T, Fujino N, Muramoto A, Hayashi K, Tsubokawa T, Sakata K, Kawashiri MA, Takeda Y, Ino H, Yamagishi M (2010) Impact of renin–angiotensin system polymorphisms on development of systolic dysfunction in hypertrophic cardiomyopathy. Circ J 74:2674–2680
Maron BJ, Maron MS, Semsarian C (2012) Double or compound sarcomere mutations in hypertrophic cardiomyopathy: a potential link to sudden death in the absence of conventional risk factors. Heart Rhythm 9:57–63
Hodatsu A, Konno T, Funada A, Fujita T, Hayashi K, Fujino N, Yamagishi M (2012) Compound heterozygosity of cardiac myosin biding protein C gene mutation worsens clinical outcome of hypertrophic cardiomyopathy: evidence from comparison with simple founder mutation carriers (abstr). Circulation 126:A17669
Kubo T, Kitaoka H, Okawa M, Baba Y, Hirota T, Hayato K, Yamasaki N, Matsumura Y, Otsuka H, Arimura T, Kimura A, Doi YL (2011) Genetic screening and double mutation in Japanese patients with hypertrophic cardiomyopathy. Circ J 75:2654–2659
Page SP, Kounas S, Syrris P, Christiansen M, Frank-Hansen R, Andersen PS, Elliott PM, McKenna WJ (2012) Cardiac myosin binding protein-C mutations in families with hypertrophic cardiomyopathy: disease expression in relation to age, gender, and long term outcome. Circ Cardiovasc Genet 5:156–166
Geisterfer-Lowrance AA, Christe M, Conner DA, Ingwall JS, Schoen FJ, Seidman CE, Seidman JG (1996) A mouse model of familial hypertrophic cardiomyopathy. Science 272:731–734
Charron P, Arad M, Arbustini E, Basso C, Bilinska Z, Elliott P, Helio T, Keren A, McKenna WJ, Monserrat L, Pankuweit S, Perrot A, Rapezzi C, Ristic A, Seggewiss H, van Langen I, Tavazzi L, European Society of Cardiology Working Group on Myocardial and Pericardial Diseases (2010) Genetic counselling and testing in cardiomyopathies: a position statement of the European Society of Cardiology Working Group on Myocardial and Pericardial Diseases. Eur Heart J 31:2715–2726
Uchiyama K, Hayashi K, Fujino N, Konno T, Sakamoto Y, Sakata K, Kawashiri MA, Ino H, Yamagishi M (2009) Impact of QT variables on clinical outcome of genotyped hypertrophic cardiomyopathy. Ann Noninvasive Electrocardiol 14:65–71
Konno T, Shimizu M, Ino H, Yamaguchi M, Terai H, Uchiyama K, Oe K, Mabuchi T, Kaneda T, Mabuchi H (2004) Diagnostic value of abnormal Q waves for identification of preclinical carriers of hypertrophic cardiomyopathy based on a molecular genetic diagnosis. Eur Heart J 25:246–251
Konno T, Shimizu M, Ino H, Fujino N, Hayashi K, Uchiyama K, Kaneda T, Inoue M, Fujita T, Masuta E, Funada A, Mabuchi H (2005) Differences in diagnostic value of four electrocardiographic voltage criteria for hypertrophic cardiomyopathy in a genotyped population. Am J Cardiol 96:1308–1312
Kaneda T, Naruse C, Kawashima A, Fujino N, Oshima T, Namura M, Nunoda S, Mori S, Konno T, Ino H, Yamagishi M, Asano M (2008) A novel beta-myosin heavy chain gene mutation, p.Met531Arg, identified in isolated left ventricular non-compaction in humans, results in left ventricular hypertrophy that progresses to dilation in a mouse model. Clin Sci (Lond) 114:431–440
Konno T, Chen D, Wang L, Wakimoto H, Teekakirikul P, Nayor M, Kawana M, Eminaga S, Gorham JM, Pandya K, Smithies O, Naya FJ, Olson EN, Seidman JG, Seidman CE (2010) Heterogeneous myocyte enhancer factor-2 (Mef2) activation in myocytes predicts focal scarring in hypertrophic cardiomyopathy. Proc Natl Acad Sci USA 107:18097–18102
Lan F, Lee AS, Liang P, Sanchez-Freire V, Nguyen PK, Wang L, Han L, Yen M, Wang Y, Sun N, Abilez OJ, Hu S, Ebert AD, Navarrete EG, Simmons CS, Wheeler M, Pruitt B, Lewis R, Yamaguchi Y, Ashley EA, Bers DM, Robbins RC, Longaker MT, Wu JC (2013) Abnormal calcium handling properties underlie familial hypertrophic cardiomyopathy pathology in patient-specific induced pluripotent stem cells. Cell Stem Cell 12:101–113
Teekakirikul P, Eminaga S, Toka O, Alcalai R, Wang L, Wakimoto H, Nayor M, Konno T, Gorham JM, Wolf CM, Kim JB, Schmitt JP, Molkentin JD, Norris RA, Tager AM, Hoffman SR, Markwald RR, Seidman CE, Seidman JG (2010) Cardiac fibrosis in mice with hypertrophic cardiomyopathy is mediated by non-myocyte proliferation and requires Tgf-β. J Clin Invest 120:3520–3529
He JG, Chen SL, Huang YY, Chen YL, Dong YG, Ma H (2010) The nonpeptide AVE0991 attenuates myocardial hypertrophy as induced by angiotensin II through downregulation of transforming growth factor-beta1/Smad2 expression. Heart Vessels 25:438–443
Shimizu W, Horie M (2011) Phenotypic manifestations of mutations in genes encoding subunits of cardiac potassium channels. Circ Res 109:97–109
Makita N, Seki A, Sumitomo N, Chkourko H, Fukuhara S, Watanabe H, Shimizu W, Bezzina CR, Hasdemir C, Mugishima H, Makiyama T, Baruteau A, Baron E, Horie M, Hagiwara N, Wilde AA, Probst V, Le Marec H, Roden DM, Mochizuki N, Schott JJ, Delmar M (2012) A connexin40 mutation associated with a malignant variant of progressive familial heart block type I. Circ Arrhythm Electrophysiol 5:163–172
Moss AJ, Schwartz PJ, Crampton RS, Tzivoni D, Locati EH, MacCluer J, Hall WJ, Weitkamp L, Vincent GM, Garson A Jr (1991) The long QT syndrome. Prospective longitudinal study of 328 families. Circulation 84:1136–1144
Goldenberg I, Moss AJ, Peterson DR, McNitt S, Zareba W, Andrews ML, Robinson JL, Locati EH, Ackerman MJ, Benhorin J, Kaufman ES, Napolitano C, Priori SG, Qi M, Schwartz PJ, Towbin JA, Vincent GM, Zhang L (2008) Risk factors for aborted cardiac arrest and sudden cardiac death in children with the congenital long-QT syndrome. Circulation 117:2184–2191
Hobbs JB, Peterson DR, Moss AJ, McNitt S, Zareba W, Goldenberg I, Qi M, Robinson JL, Sauer AJ, Ackerman MJ, Benhorin J, Kaufman ES, Locati EH, Napolitano C, Priori SG, Towbin JA, Vincent GM, Zhang L (2006) Risk of aborted cardiac arrest or sudden cardiac death during adolescence in the long-QT syndrome. JAMA 296:1249–1254
Sauer AJ, Moss AJ, McNitt S, Peterson DR, Zareba W, Robinson JL, Qi M, Goldenberg I, Hobbs JB, Ackerman MJ, Benhorin J, Hall WJ, Kaufman ES, Locati EH, Napolitano C, Priori SG, Schwartz PJ, Towbin JA, Vincent GM, Zhang L (2007) Long QT syndrome in adults. J Am Coll Cardiol 49:329–337
Roden DM (2008) Clinical practice. Long-QT syndrome. N Engl J Med 358:169–176
Tester DJ, Will ML, Haglund CM, Ackerman MJ (2006) Effect of clinical phenotype on yield of long QT syndrome genetic testing. J Am Coll Cardiol 47:764–768
Itoh H, Shimizu W, Hayashi K, Yamagata K, Sakaguchi T, Ohno S, Makiyama T, Akao M, Ai T, Noda T, Miyazaki A, Miyamoto Y, Yamagishi M, Kamakura S, Horie M (2010) Long QT syndrome with compound mutations is associated with a more severe phenotype: a Japanese multicenter study. Heart Rhythm 7:1411–1418
Itoh H, Sakaguchi T, Ding WG, Watanabe E, Watanabe I, Nishio Y, Makiyama T, Ohno S, Akao M, Higashi Y, Zenda N, Kubota T, Mori C, Okajima K, Haruna T, Miyamoto A, Kawamura M, Ishida K, Nagaoka I, Oka Y, Nakazawa Y, Yao T, Jo H, Sugimoto Y, Ashihara T, Hayashi H, Ito M, Imoto K, Matsuura H, Horie M (2009) Latent genetic backgrounds and molecular pathogenesis in drug-induced long-QT syndrome. Circ Arrhythm Electrophysiol 2:511–523
Ohno S, Toyoda F, Zankov DP, Yoshida H, Makiyama T, Tsuji K, Honda T, Obayashi K, Ueyama H, Shimizu W, Miyamoto Y, Kamakura S, Matsuura H, Kita T, Horie M (2009) Novel KCNE3 mutation reduces repolarizing potassium current and associated with long QT syndrome. Hum Mutat 30:557–563
Paulussen AD, Gilissen RA, Armstrong M, Doevendans PA, Verhasselt P, Smeets HJ, Schulze-Bahr E, Haverkamp W, Breithardt G, Cohen N, Aerssens J (2004) Genetic variations of KCNQ1, KCNH2, SCN5A, KCNE1, and KCNE2 in drug-induced long QT syndrome patients. J Mol Med 82:182–188
Hayashi K, Shimizu M, Ino H, Yamaguchi M, Terai H, Hoshi N, Higashida H, Terashima N, Uno Y, Kanaya H, Mabuchi H (2004) Probucol aggravates long QT syndrome associated with a novel missense mutation M124T in the N-terminus of HERG. Clin Sci (Lond) 107:175–182
Shimizu W (2008) Clinical impact of genetic studies in lethal inherited cardiac arrhythmias. Circ J 72:1926–1936
Yang Y, Yang Y, Liang B, Liu J, Li J, Grunnet M, Olesen SP, Rasmussen HB, Ellinor PT, Gao L, Lin X, Li L, Wang L, Xiao J, Liu Y, Zhang S, Liang D, Peng L, Jespersen T, Chen YH (2010) Identification of a Kir3.4 mutation in congenital long QT syndrome. Am J Hum Genet 86:872–880
Hayashi K, Fujino N, Uchiyama K, Ino H, Sakata K, Konno T, Masuta E, Funada A, Sakamoto Y, Tsubokawa T, Nakashima K, Liu L, Higashida H, Hiramaru Y, Shimizu M, Yamagishi M (2009) Long QT syndrome and associated gene mutation carriers in Japanese children: results from ECG screening examinations. Clin Sci (Lond) 117:415–424
Soma K, Nagaoka K, Kuwahara M, Tsubone H, Ito K (2011) Abundant expression of KCNE1 in the left ventricle of the miniature pig. Heart Vessels 26:353–356
Hayashi K, Shuai W, Sakamoto Y, Higashida H, Yamagishi M, Kupershmidt S (2010) Trafficking-competent KCNQ1 variably influences the function of HERG long QT alleles. Heart Rhythm 7:973–980
Nakashima K, Kusakawa I, Yamamoto T, Hirabayashi S, Hosoya R, Shimizu W, Sumitomo N (2013) A left ventricular noncompaction in a patient with long QT syndrome caused by a KCNQ1 mutation: a case report. Heart Vessels 28:126–129
Makita N, Behr E, Shimizu W, Horie M, Sunami A, Crotti L, Schulze-Bahr E, Fukuhara S, Mochizuki N, Makiyama T, Itoh H, Christiansen M, McKeown P, Miyamoto K, Kamakura S, Tsutsui H, Schwartz PJ, George AL Jr, Roden DM (2008) The E1784 K mutation in SCN5A is associated with mixed clinical phenotype of type 3 long QT syndrome. J Clin Invest 118:2219–2229
Barsheshet A, Goldenberg I, O-Uchi J, Moss AJ, Jons C, Shimizu W, Wilde AA, McNitt S, Peterson DR, Zareba W, Robinson JL, Ackerman MJ, Cypress M, Gray DA, Hofman N, Kanters JK, Kaufman ES, Platonov PG, Qi M, Towbin JA, Vincent GM, Lopes CM (2012) Mutations in cytoplasmic loops of the KCNQ1 channel and the risk of life-threatening events: implications for mutation-specific response to β-blocker therapy in type 1 long-QT syndrome. Circulation 125:1988–1996
Chockalingam P, Crotti L, Girardengo G, Johnson JN, Harris KM, van der Heijden JF, Hauer RN, Beckmann BM, Spazzolini C, Rordorf R, Rydberg A, Clur SA, Fischer M, van den Heuvel F, Kääb S, Blom NA, Ackerman MJ, Schwartz PJ, Wilde AA (2012) Not all beta-blockers are equal in the management of long QT syndrome types 1 and 2: higher recurrence of events under metoprolol. J Am Coll Cardiol 60:2092–2099
Priori SG, Napolitano C, Schwartz PJ (1999) Low penetrance in the long-QT syndrome: clinical impact. Circulation 99:529–533
Napolitano C, Priori SG, Schwartz PJ, Bloise R, Ronchetti E, Nastoli J, Bottelli G, Cerrone M, Leonardi S (2005) Genetic testing in the long QT syndrome: development and validation of an efficient approach to genotyping in clinical practice. JAMA 294:2975–2980
Shimizu W, Moss AJ, Wilde AA, Towbin JA, Ackerman MJ, January CT, Tester DJ, Zareba W, Robinson JL, Qi M, Vincent GM, Kaufman ES, Hofman N, Noda T, Kamakura S, Miyamoto Y, Shah S, Amin V, Goldenberg I, Andrews ML, McNitt S (2009) Genotype–phenotype aspects of type 2 long QT syndrome. J Am Coll Cardiol 54:2052–2062
Liu L, Hayashi K, Kaneda T, Ino H, Fujino N, Uchiyama K, Konno T, Tsuda T, Kawashiri MA, Ueda K, Higashikata T, Shuai W, Kupershmidt S, Higashida H, Yamagishi M (2013) A novel mutation in the transmembrane non-pore region of the KCNH2 gene causes severe clinical manifestations of long QT syndrome. Heart Rhythm 10:61–67
Anderson CL, Delisle BP, Anson BD, Kilby JA, Will ML, Tester DJ, Gong Q, Zhou Z, Ackerman MJ, January CT (2006) Most LQT2 mutations reduce Kv11.1 (hERG) current by a class 2 (trafficking-deficient) mechanism. Circulation 113:365–373
Oka Y, Itoh H, Ding WG, Shimizu W, Makiyama T, Ohno S, Nishio Y, Sakaguchi T, Miyamoto A, Kawamura M, Matsuura H, Horie M (2010) Atrioventricular block-induced torsades de pointes with clinical and molecular backgrounds similar to congenital long QT syndrome. Circ J 74:2562–2571
Kupershmidt S, Yang IC, Hayashi K, Wei J, Chanthaphaychith S, Petersen CI, Johns DC, George AL Jr, Roden DM, Balser JR (2003) The IKr drug response is modulated by KCR1 in transfected cardiac and noncardiac cell lines. FASEB J 17:2263–2265
Nakajima T, Hayashi K, Viswanathan PC, Kim MY, Anghelescu M, Barksdale KA, Shuai W, Balser JR, Kupershmidt S (2007) HERG is protected from pharmacological block by alpha-1,2-glucosyltransferase function. J Biol Chem 282:5506–5513
Petersen CI, McFarland TR, Stepanovic SZ, Yang P, Reiner DJ, Hayashi K, George AL, Roden DM, Thomas JH, Balser JR (2004) In vivo identification of genes that modify ether-a-go–go-related gene activity in Caenorhabditis elegans may also affect human cardiac arrhythmia. Proc Natl Acad Sci USA 101:11773–11778
Hayashi K, Fujino N, Ino H, Uchiyama K, Sakata K, Konno T, Masuta E, Funada A, Sakamoto Y, Tsubokawa T, Hodatsu A, Yasuda T, Kanaya H, Kim MY, Kupershmidt S, Higashida H, Yamagishi M (2011) A KCR1 variant implicated in susceptibility to the long QT syndrome. J Mol Cell Cardiol 50:50–57
Gussak I, Brugada P, Brugada J, Wright RS, Kopecky SL, Chaitman BR, Bjerregaard P (2000) Idiopathic short QT interval: a new clinical syndrome? Cardiology 94:99–102
Funada A, Hayashi K, Ino H, Fujino N, Uchiyama K, Sakata K, Masuta E, Sakamoto Y, Tsubokawa T, Yamagishi M (2008) Assessment of QT intervals and prevalence of short QT syndrome in Japan. Clin Cardiol 31:270–274
Patel C, Yan GX, Antzelevitch C (2010) Short QT syndrome: from bench to bedside. Circ Arrhythm Electrophysiol 3:401–408
Chen Q, Kirsch GE, Zhang D, Brugada R, Brugada J, Brugada P, Potenza D, Moya A, Borggrefe M, Breithardt G, Ortiz-Lopez R, Wang Z, Antzelevitch C, O’Brien RE, Schulze-Bahr E, Keating MT, Towbin JA, Wang Q (1998) Genetic basis and molecular mechanism for idiopathic ventricular fibrillation. Nature 392:293–296
Benito B, Sarkozy A, Mont L, Henkens S, Berruezo A, Tamborero D, Arzamendi D, Berne P, Brugada R, Brugada P, Brugada J (2008) Gender differences in clinical manifestations of Brugada syndrome. J Am Coll Cardiol 52:1567–1573
Hu D, Barajas-Martinez H, Burashnikov E, Springer M, Wu Y, Varro A, Pfeiffer R, Koopmann TT, Cordeiro JM, Guerchicoff A, Pollevick GD, Antzelevitch C (2009) A mutation in the beta 3 subunit of the cardiac sodium channel associated with Brugada ECG phenotype. Circ Cardiovasc Genet 2:270–278
Ueda K, Hirano Y, Higashiuesato Y, Aizawa Y, Hayashi T, Inagaki N, Tana T, Ohya Y, Takishita S, Muratani H, Hiraoka M, Kimura A (2009) Role of HCN4 channel in preventing ventricular arrhythmia. J Hum Genet 54:115–121
Burashnikov E, Pfeiffer R, Barajas-Martinez H, Delpón E, Hu D, Desai M, Borggrefe M, Häissaguerre M, Kanter R, Pollevick GD, Guerchicoff A, Laiño R, Marieb M, Nademanee K, Nam GB, Robles R, Schimpf R, Stapleton DD, Viskin S, Winters S, Wolpert C, Zimmern S, Veltmann C, Antzelevitch C (2010) Mutations in the cardiac L-type calcium channel associated with inherited J-wave syndromes and sudden cardiac death. Heart Rhythm 7:1872–1882
Kattygnarath D, Maugenre S, Neyroud N, Balse E, Ichai C, Denjoy I, Dilanian G, Martins RP, Fressart V, Berthet M, Schott JJ, Leenhardt A, Probst V, Le Marec H, Hainque B, Coulombe A, Hatem SN, Guicheney P (2011) MOG1: a new susceptibility gene for Brugada syndrome. Circ Cardiovasc Genet 4:261–268
Giudicessi JR, Ye D, Tester DJ, Crotti L, Mugione A, Nesterenko VV, Albertson RM, Antzelevitch C, Schwartz PJ, Ackerman MJ (2011) Transient outward current (I(to)) gain-of-function mutations in the KCND3-encoded Kv4.3 potassium channel and Brugada syndrome. Heart Rhythm 8:1024–1032
Ohno S, Zankov DP, Ding WG, Itoh H, Makiyama T, Doi T, Shizuta S, Hattori T, Miyamoto A, Naiki N, Hancox JC, Matsuura H, Horie M (2011) KCNE5 (KCNE1L) variants are novel modulators of Brugada syndrome and idiopathic ventricular fibrillation. Circ Arrhythm Electrophysiol 4:352–361
Medeiros-Domingo A, Tan BH, Crotti L, Tester DJ, Eckhardt L, Cuoretti A, Kroboth SL, Song C, Zhou Q, Kopp D, Schwartz PJ, Makielski JC, Ackerman MJ (2010) Gain-of-function mutation S422L in the KCNJ8-encoded cardiac K (ATP) channel Kir6.1 as a pathogenic substrate for J-wave syndromes. Heart Rhythm 7:1466–1471
Hu D, Barajas-Martínez H, Medeiros-Domingo A, Crotti L, Veltmann C, Schimpf R, Urrutia J, Alday A, Casis O, Pfeiffer R, Burashnikov E, Caceres G, Tester DJ, Wolpert C, Borggrefe M, Schwartz P, Ackerman MJ, Antzelevitch C (2012) A novel rare variant in SCN1Bb linked to Brugada syndrome and SIDS by combined modulation of Na(v)1.5 and K(v)4.3 channel currents. Heart Rhythm 9:760–769
Ishikawa T, Sato A, Marcou CA, Tester DJ, Ackerman MJ, Crotti L, Schwartz PJ, On YK, Park JE, Nakamura K, Hiraoka M, Nakazawa K, Sakurada H, Arimura T, Makita N, Kimura A (2012) A novel disease gene for Brugada syndrome: sarcolemmal membrane-associated protein gene mutations impair intracellular trafficking of hNav1.5. Circ Arrhythm Electrophysiol 5:1098–1107
Probst V, Veltmann C, Eckardt L, Meregalli PG, Gaita F, Tan HL, Babuty D, Sacher F, Giustetto C, Schulze-Bahr E, Borggrefe M, Haissaguerre M, Mabo P, Le Marec H, Wolpert C, Wilde AA (2010) Long-term prognosis of patients diagnosed with Brugada syndrome: results from the FINGER Brugada Syndrome Registry. Circulation 121:635–643
Takagi M, Aonuma K, Sekiguchi Y, Yokoyama Y, Aihara N, Hiraoka M, Japan Idiopathic Ventricular Fibrillation Study (J-IVFS) Investigators (2013) The prognostic value of early repolarization (J wave) and ST-segment morphology after J wave in Brugada syndrome: multicenter study in Japan. Heart Rhythm 10:533–539
Kamakura S, Ohe T, Nakazawa K, Aizawa Y, Shimizu A, Horie M, Ogawa S, Okumura K, Tsuchihashi K, Sugi K, Makita N, Hagiwara N, Inoue H, Atarashi H, Aihara N, Shimizu W, Kurita T, Suyama K, Noda T, Satomi K, Okamura H, Tomoike H, Brugada Syndrome Investigators in Japan (2009) Long-term prognosis of probands with Brugada-pattern ST-elevation in leads V1-V3. Circ Arrhythm Electrophysiol 2:495–503
Priori SG, Napolitano C, Memmi M, Colombi B, Drago F, Gasparini M, DeSimone L, Coltorti F, Bloise R, Keegan R, Cruz Filho FE, Vignati G, Benatar A, DeLogu A (2002) Clinical and molecular characterization of patients with catecholaminergic polymorphic ventricular tachycardia. Circulation 106:69–74
Priori SG, Napolitano C, Tiso N, Memmi M, Vignati G, Bloise R, Sorrentino V, Danieli GA (2001) Mutations in the cardiac ryanodine receptor gene (hRyR2) underlie catecholaminergic polymorphic ventricular tachycardia. Circulation 103:196–200
Laitinen PJ, Brown KM, Piippo K, Swan H, Devaney JM, Brahmbhatt B, Donarum EA, Marino M, Tiso N, Viitasalo M, Toivonen L, Stephan DA, Kontula K (2001) Mutations of the cardiac ryanodine receptor (RyR2) gene in familial polymorphic ventricular tachycardia. Circulation 103:485–490
Lahat H, Pras E, Olender T, Avidan N, Ben-Asher E, Man O, Levy-Nissenbaum E, Khoury A, Lorber A, Goldman B, Lancet D, Eldar M (2001) A missense mutation in a highly conserved region of CASQ2 is associated with autosomal recessive catecholamine-induced polymorphic ventricular tachycardia in Bedouin families from Israel. Am J Hum Genet 69:1378–1384
Vega AL, Tester DJ, Ackerman MJ, Makielski JC (2009) Protein kinase A-dependent biophysical phenotype for V227F-KCNJ2 mutation in catecholaminergic polymorphic ventricular tachycardia. Circ Arrhythm Electrophysiol 2:540–547
Roux-Buisson N, Cacheux M, Fourest-Lieuvin A, Fauconnier J, Brocard J, Denjoy I, Durand P, Guicheney P, Kyndt F, Leenhardt A, Le Marec H, Lucet V, Mabo P, Probst V, Monnier N, Ray PF, Santoni E, Trémeaux P, Lacampagne A, Fauré J, Lunardi J, Marty I (2012) Absence of triadin, a protein of the calcium release complex, is responsible for cardiac arrhythmia with sudden death in human. Hum Mol Genet 21:2759–2767
Nyegaard M, Overgaard MT, Søndergaard MT, Vranas M, Behr ER, Hildebrandt LL, Lund J, Hedley PL, Camm AJ, Wettrell G, Fosdal I, Christiansen M, Børglum AD (2012) Mutations in calmodulin cause ventricular tachycardia and sudden cardiac death. Am J Hum Genet 91:703–712
Alcalai R, Wakimoto H, Arad M, Planer D, Konno T, Wang L, Seidman JG, Seidman CE, Berul CI (2011) Prevention of ventricular arrhythmia and calcium dysregulation in a catecholaminergic polymorphic ventricular tachycardia mouse model carrying calsequestrin-2 mutation. J Cardiovasc Electrophysiol 22:316–324
Watanabe H, Chopra N, Laver D, Hwang HS, Davies SS, Roach DE, Duff HJ, Roden DM, Wilde AA, Knollmann BC (2009) Flecainide prevents catecholaminergic polymorphic ventricular tachycardia in mice and humans. Nat Med 15:380–383
van der Werf C, Kannankeril PJ, Sacher F, Krahn AD, Viskin S, Leenhardt A, Shimizu W, Sumitomo N, Fish FA, Bhuiyan ZA, Willems AR, van der Veen MJ, Watanabe H, Laborderie J, Haïssaguerre M, Knollmann BC, Wilde AA (2011) Flecainide therapy reduces exercise-induced ventricular arrhythmias in patients with catecholaminergic polymorphic ventricular tachycardia. J Am Coll Cardiol 57:2244–2254
Moretti A, Bellin M, Welling A, Jung CB, Lam JT, Bott-Flügel L, Dorn T, Goedel A, Höhnke C, Hofmann F, Seyfarth M, Sinnecker D, Schömig A, Laugwitz KL (2010) Patient-specific induced pluripotent stem-cell models for long-QT syndrome. N Engl J Med 363:1397–1409
Egashira T, Yuasa S, Suzuki T, Aizawa Y, Yamakawa H, Matsuhashi T, Ohno Y, Tohyama S, Okata S, Seki T, Kuroda Y, Yae K, Hashimoto H, Tanaka T, Hattori F, Sato T, Miyoshi S, Takatsuki S, Murata M, Kurokawa J, Furukawa T, Makita N, Aiba T, Shimizu W, Horie M, Kamiya K, Kodama I, Ogawa S, Fukuda K (2012) Disease characterization using LQTS-specific induced pluripotent stem cells. Cardiovasc Res 95:419–429
Knollmann BC (2013) Induced pluripotent stem cell-derived cardiomyocytes. Boutique science or valuable arrhythmia model? Circ Res 112:969–976
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Additional information
M. Kawashiri, K. Hayashi, and T. Konno contributed equally to establishing this article.
A part of this article was reproduced from our article as listed in reference 2.
Rights and permissions
About this article
Cite this article
Kawashiri, Ma., Hayashi, K., Konno, T. et al. Current perspectives in genetic cardiovascular disorders: from basic to clinical aspects. Heart Vessels 29, 129–141 (2014). https://doi.org/10.1007/s00380-013-0391-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00380-013-0391-5