Abstract
The extraradical hyphae-associated microbiome of arbuscular mycorrhizal fungi (AMF), the “hyphosphere microbiome,” harbors a diverse reservoir of microbes. The biological interactions in the AMF hyphosphere have major implications for soil carbon and nutrient cycling, soil food web dynamics, and plant nutrition and health. Hyphosphere microbial communities are thought to assist AMF in accessing organic nutrients by degrading complex organic compounds that AMF are unable to do by themselves. The AMF, in return, provide an energy-rich microhabitat supplied with hyphal exudates that facilitates microbial growth and mobility in the hyphosphere. However, our current knowledge of hyphosphere entities, their trophic interactions and functional roles, and the underlying mechanisms facilitating microbial co-occurrence and co-operation is largely incomplete. Here, we review the current state of knowledge on the identity and putative roles of AMF hyphae-associated microbes, with a specific focus on prokaryotes, and potential drivers of such microbial communities in the hyphosphere. Moreover, we discuss the knowledge gaps and open challenges that should be addressed and prioritized in future studies on the AMF microbiomes. We also provide an appraisal of available and emerging tools and technologies and highlight the need for innovative approaches to disentangle AMF hyphosphere processes and answer the many unresolved questions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Arbuscular mycorrhizal fungi (AMF), from the subphylum Glomeromycotina, are the most widespread plant root symbiotic partners in a wide range of terrestrial ecosystems, interconnecting the plant root system to the soil environment (Brundrett and Tedersoo 2018; Spatafora et al. 2016). The mutualistic symbiotic association between AMF and plants was established more than 400 million years ago and currently exists in more than 70% of all vascular plant species (Brundrett and Tedersoo 2018; Remy et al. 1994; van der Heijden et al. 2015). The fungal partner contributes to plant mineral nutrient uptake, and resistance against multiple biotic (e.g., pathogens) and abiotic (e.g., salinity, drought, heavy metals) stresses (Faghihinia et al. 2020; Gao et al. 2020; Kikuchi et al. 2016; Smith and Smith 2011; Zai et al. 2021). As obligate plant symbionts, AMF fully depend upon photosynthate carbon (C) supply from their host plants to thrive and propagate (Smith and Read 2008). In consequence, AMF also contribute to soil C fluxes and stabilization in terrestrial ecosystems by facilitating the transfer of atmospheric C fixed by the plants into the soil (Jansa and Treseder 2017; van der Heijden et al. 2015).
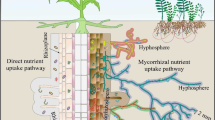
Of the total AMF biomass, extraradical fungal hyphae represent a significant component, and these mycorrhizal hyphal networks are largely responsible for the lateral C fluxes in soil (Godbold et al. 2006; Kaiser et al. 2015; Talbot et al. 2008; Treseder and Cross 2006). Extraradical AMF hyphae have enormous potential to exploit soil micropores beyond the rhizosphere zone to access macronutrients, particularly phosphorus (P) and nitrogen (N), and micronutrients such as zinc and copper (Adeyemi et al. 2021; Bukovská et al. 2018; Gao et al. 2021; Jiang et al. 2021; Tamayo et al. 2014; Thirkell et al. 2016), and possibly also facilitate soil–plant water fluxes (Kikuchi et al. 2016; Püschel et al. 2021). However, AMF are unable to efficiently cleave and utilize complex organic compounds due to their limited exo-enzymatic repertoire (but see Koide and Kabir (2000)), suggesting that AMF organic nutrient acquisition is facilitated by other soil microorganisms (Rozmoš et al. 2022; Tisserant et al. 2013). This could explain the existence of the microbial communities associated with the surface of extraradical fungal hyphae, termed the “hyphosphere microbiome” (Artursson et al. 2006; Bonfante and Anca 2009; Jansa and Hodge 2021). Indeed, AMF affect the soil immediately surrounding their hyphae, the hyphosphere, through the exudation of a range of compounds and signaling molecules by the extraradical hyphae (Jiang et al. 2021; Zhang et al. 2018a) and recruit specific microbiota through a mechanism called the “hyphosphere effect” (in parallel to the “rhizosphere effect”) (Bakker et al. 2013).
To date, only limited research efforts have been dedicated to characterizing the hyphosphere communities associated with different AMF species, using compartmentalized microcosms with separate root and hyphal compartments or ingrown mesh cores, and/or manipulating soil conditions (Emmett et al. 2021; Wang et al. 2016; Zhou et al. 2020) (see Table 1 and supplementary Table S1). In addition, the relationships between AMF and the hyphosphere microbiome members, and the outcome of their interactions on organic nutrient utilization and nutrient cycling, were only investigated in a few studies so far using in vitro experimental setups and single or multiple bacterial genotypes (Bukovská et al. 2018; Jiang et al. 2021) (see Table 2). Thus, our current knowledge of the functioning of the hyphospheric microorganisms and the underlying mechanisms of interactions between AMF and their associated microbiome is still largely incomplete (Jansa and Hodge 2021; Zhang et al. 2022).
The concept of the AMF microbiome offers a promising perspective to improve our understanding of microbe-AMF interactions, which subsequently influence plant nutrition and health (Artursson et al. 2006; Pivato et al. 2009; Zhang et al. 2022). Moreover, uncovering the link between AMF and hyphosphere microorganisms has significant implications for our understanding of soil nutrient and C cycling, and soil food webs.
This review scrutinizes and critically appraises the current state of knowledge on AMF hyphosphere and provides possible directions for future research efforts. It also highlights the methodological approaches and emerging tools and techniques that have great potential to contribute to a better understanding of the hyphosphere microbiome functioning. Contrary to the recent review of the AMF microbiome by Zhang et al. (2022), here the emphasis is on the functional role of microbes in the hyphosphere from a system-level perspective as well as providing a summary of important considerations for experimental design/approaches when studying the AMF hyphosphere microbiome. Particularly, we pay attention to a whole range of multitrophic interactions within the hyphosphere microbiome, not only positive, but also negative (such as competition for ammonium ions between AM hyphae and nitrification bacteria); in addition to focusing on P and N, we further mention processes within the S and Si cycling, and the role of common mycorrhizal networks in soil–plant nutrient cycling, something overlooked in the previous review paper.
Hyphosphere and its functional role
The term “mycorrhizal hyphosphere” has been around for several decades as the zone of interface between extraradical hyphae of mycorrhizal fungi and the adjacent soil (Andrade et al. 1997; Linderman 1991; Marschner 1995). This zone can further be separated into endo-hyphosphere (inner hyphosphere, i.e., inside the hyphae) and exo-hyphosphere (outer hyphosphere or the hyphosphere corresponding to the definition above). The exo-hyphosphere can further be operationally divided into hyphoplane (hyphal surface) and ecto-hyphosphere (i.e., the hyphosphere soil), although it is difficult to strictly differentiate between them since it is a continuum without a clear demarcation separating them from each other (Fig. 1).
A schematic representation of the arbuscular mycorrhizal fungal (AMF) hyphosphere, which could operationally be separated into endo-hyphosphere (inner hyphosphere, intracellular space) and exo-hyphosphere (outer or extracellular hyphosphere). Exo-hyphosphere can further be divided into hyphoplane (hyphal surface) and ecto-hyphosphere (i.e., hyphosphere soil)
The AMF endo-hyphosphere can be colonized by obligate bacterial endosymbionts which are nutritionally fully dependent on their fungal host, which could confer specific functions such as providing essential nutritional/metabolic factors, and have been reviewed elsewhere (Bonfante et al. 1994; Desirò et al. 2014). In this review, we mainly focus on the zones at the surface or in the immediate vicinity of the AMF mycelium. These zones are characterized by a variety of interactions between hyphae, soil minerals, organic and inorganic nutrients, gases, soluble compounds, and active microbial communities that participate in various soil biogeochemical cycles. The AMF extraradical hyphae release exudates containing a variety of compounds, including sugars and amino acids, that can be sensed and/or used by soil microbes, mainly bacteria or archaea, and stimulate them to move towards the hyphosphere (Jiang et al. 2021; Luthfiana et al. 2021). Indeed, due to their superior capability to grow into soil micropores and forage for spatially and temporarily heterogeneous nutrient resources, while exuding a variety of compounds along the way, extraradical hyphae provide a high-energy microhabitat for microbes and can facilitate their dispersal throughout the soil (Jansa and Hodge 2021; Jiang et al. 2021).
Scanning electron microscope images and other earlier experimental efforts show that some bacterial groups are able to attach firmly to the surface of the AMF hyphae (Artursson and Jansson 2003; Bianciotto et al. 2001, 1996; Jansa and Hodge 2021; Toljander et al. 2006), and there is also direct observation that some groups of bacteria are able to migrate along the water film-coated hyphae (Jiang et al. 2021). The movement of bacteria along such a “fungal highway” is essential for their ecological competence (i.e., the capacity to fulfill a specific ecosystem function) and competitive success due to many factors that limit their mobility and dispersal in the soil (Jiang et al. 2021; Junier et al. 2021; Otto et al. 2017). The role of fungal highways on bacterial dispersion in air-filled soil micropores have previously been highlighted for other groups of fungi (Deveau et al. 2018; Kohlmeier et al. 2005; Nazir et al. 2014; Otto et al. 2017; Wick et al. 2007). The fungal highway also facilitates contact between prey (e.g., bacteria) and their predators (e.g., protists), shaping the microbial communities and thus also soil food webs (Junier et al. 2021; Otto et al. 2017). These observations were made in some groups of fungi using novel tools such as 3D-printed microfluidic devices or controlled microcosm systems, but not yet in systems that included mycorrhizal fungi (Abeysinghe et al. 2020; Aleklett et al. 2018; Mafla-Endara et al. 2021).
AMF hyphae-associated microbes have been shown to enable or at least facilitate production of extracellular lytic enzymes (which could be regarded as “public goods”) that degrade soil organic matter to liberate nutrients that the AMF can then obtain to meet their nutritional needs (Rozmoš et al. 2022; Zhang et al. 2018a). Under in vitro culture conditions, Zhang et al. (2018a) observed significantly greater phosphatase excretion by hyphosphere bacteria supplied with AMF exudates. Furthermore, there was greater expression of the mycorrhizal phosphate transporter gene GintPT and polyP synthesis gene Vtc4p in Rhizophagus irregularis in the presence of Rahnella aquatilis (phosphate solubilizing bacterium, PSB) compared to R. irregularis not in association with R. aquatilis. In a series of well-designed experiment, Jiang et al. (2021) observed a significant role of R. aquatilis in organic P utilization by the AMF, and that R. aquatilis dispersion along the hyphal highway and enhanced metabolism is likely due, at least in part, to AMF hyphal exudates. Wang et al. (2016) found that combined inoculation with PSB (Pseudomonas alcaligenes M20, Bacillus megaterium C4, or Rahnella aquatilis HX2) and R. irregularis resulted in higher phytate-P mineralization and P contained in microbial biomass in the AMF hyphosphere compared to the treatments with R. irregularis or with the PSB alone. These observations suggest that AMF rely on their associated bacterial communities for the acquisition of P from organic sources in the soil.
There is also evidence for the potential role of some hyphosphere microbes in facilitating N acquisition from organic sources by the AMF hyphae, in addition to the effects of some soil bacteria on AMF germination and hyphal growth (Gryndler et al. 2000; Hildebrandt et al. 2002; Xavier and Germida 2003). For example, using quantitative real-time PCR, a significant positive correlation was observed between the hyphal proliferation of two AMF species, R. irregularis and Claroideoglomus claroideum, and the abundance of ammonia-oxidizing bacteria (including Nitrosospira sp.) in soil patches containing organic N (Bukovská et al. 2016). In contrast, however, a follow-up study with non-mycorrhizal controls showed that AMF hyphae actually suppressed the abundance of many soil microbes, including nitrification bacteria in a root-free soil that was supplied with organic nutrients as compared with to soil patches not amended with such nutrients (Bukovská et al. 2018). More recently, it has been demonstrated that a substantial amount of (otherwise unavailable) N supplied as chitin could be used up by the AMF hyphae in the presence of Paenibacillus sp. in root-free organic N patches (Rozmoš et al. 2022). These findings suggest that, in some cases, the AMF specifically recruit beneficial bacteria for their own nutritional needs, likely providing them with C resources in return, as well as providing them with a microhabitat for convenient movement throughout the soil. The apparent contradictions in results presented here are likely linked to a still far too superficial understanding on AMF-bacteria relationships and interactions, and specifically what cues might be used by the AMF to recruit beneficial bacteria (and/or archaea). Nonetheless, evidence does suggest that AMF can alter the microbe’s physicochemical environment via efficient acquisition and exporting of nutrients such as N and/or P from enriched patches and importing fresh C into the microbe’s microhabitats (Bukovská et al. 2018; Jiang et al. 2021; Nuccio et al. 2013; Wang et al. 2019; Zhang et al. 2014).
Overall, the AMF hyphosphere provides an energy-rich habitat for microbes, facilitates the dispersal of microbes through soil matrix via water film-coated hyphae, facilitates contact between prey and predators, selectively recruits beneficial bacteria (and/or archaea (Nuccio et al. 2022)), and can stimulate bacterial metabolic activities to degrade soil organic matter in order to liberate P and N that the AMF hyphae can then utilize. The hyphosphere and its microbial community would therefore be playing a greater role in soil processes than its physical size would indicate.
Hyphosphere residents
The hyphosphere microbiome differs from that of the mycorrhizosphere (soil zone under the influence of both roots and fungal components), the rhizosphere (soil zone under the influence of plant root components), and the bulk soil (traditionally defined as root-free soil) (Gahan and Schmalenberger 2015; Veresoglou et al. 2019; Zhang et al. 2018b; Zhou et al. 2020). These observations were made mainly by comparing soil microbial communities in root and hyphal compartments and also using extracted AMF hyphae, employing high-throughput amplicon sequencing (Emmett et al. 2021; Nuccio et al. 2013; Wang et al. 2016; Zhang et al. 2020; Zhou et al. 2020). To date, only a few studies have attempted to specifically characterize the hyphosphere microbial community (Table 1 and supplementary Table S1). Based on that research, it has been proposed that “core AMF hyphosphere microbiome” does exist (in spite of lacking consensus as to its exact definition) and that it is AMF species- and soil conditions-independent (Emmett et al. 2021). In addition, microbial community analyses reported so far have provided relative abundances of microbes in destructively obtained samples, which does not necessarily address the absolute abundances of the different microbes and the spatial and temporal complexity of such communities in the hyphosphere unless combined with complementary techniques such as qPCR (Table 3) (Alteio et al. 2021). When relying on amplicon sequencing, inclusion of spike-in standards has the potential to enable absolute quantification (Tourlousse et al. 2017), which is a strategy that should definitively be promoted in the future.
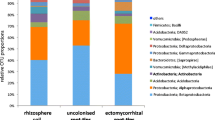
Recent compelling evidence suggests that bacteria are the dominant life form in the AMF hyphosphere microbiomes in terms of individuals and biomass (Bukovská et al. 2021). The results also indicate that despite possible differences in the relative abundance of bacteria at lower taxonomic levels (e.g., species), the composition of the bacterial community may not differ substantially with regard to the dominant phyla detected in the AMF hyphosphere. Despite differences in the relative abundances of the different species present, it appears that the microbiome of the AMF hyphosphere is dominated mainly by four bacterial phyla: Pseudomonadota, Actinomycetota, Gemmatimonadota, and Bacteroidota (Table 1 and supplementary Table S1). The substantial variation in taxonomic composition obvious at lower taxonomic ranks (genus and species levels) may be related to the particular environmental context or functional redundancy within the AMF-microbe interactions.
Although several high-throughput sequencing techniques have been used to identify the prokaryotic microbiota, mainly bacteria, in the AMF hyphosphere, we are not aware of any studies that similarly characterized the community of eukaryotes in the same zone, apart from indirect research comparing mycorrhizal and non-mycorrhizal pots (e.g., Gryndler et al. (2018)). Importantly, the significant role of bacterivores such as protists in the dynamics of bacterial populations should not be overlooked (Bukovská et al. 2018; Koller et al. 2013a; Mafla-Endara et al. 2021; Rozmoš et al. 2022). And of course, other organisms such as animals (e.g., Collembola and nematodes) or fungi (filamentous or not) may fulfil specific functions in the AMF hyphosphere, too (Poveda et al. 2019; Purin and Rillig 2008).
Factors shaping the hyphosphere microbiome
Our current knowledge suggests that interactions in the hyphosphere are regulated by a number of factors, contributed to by the AMF identity (Agnolucci et al. 2015; Bukovská et al. 2016; Emmett et al. 2021; Zhou et al. 2020), quality of soil organic matter, particularly organic P and bioavailable P levels (Gao et al. 2020; Jiang et al. 2021; Wang et al. 2019; Zhang et al. 2014, 2018b), and organic N quality and mineral N availability (Bukovská et al. 2018; Nuccio et al. 2013; Veresoglou et al. 2019), as well as soil physico-chemical properties (Emmett et al. 2021; Svenningsen et al. 2018).
By combining 13C-DNA stable isotope probing (13C-DNA-SIP) with MiSeq sequencing in compartmented mesocosms with split-root systems, Zhou et al. (2020) found distinct active microbial communities associated with different AMF species, Funneliformis mosseae, Gigaspora margarita, and R. intraradices, that had simultaneously colonized single cotton plant root system. Greater hyphal density, 13C abundance, and bacterial OTUs richness were observed in the hyphal compartments with F. mosseae or R. intraradices than in those with Gi. margarita (Zhou et al. 2020). The authors also found greater relative abundance of Streptomyces and Bacillus in the hyphosphere of R. intraradices and F. mosseae, and the greatest abundance of Pseudomonas in the hyphosphere of Gi. margarita (Zhou et al. 2020). Accordingly, Emmett et al. (2021), using 16S rRNA gene sequence analysis, compared the AMF hyphae-associated microbiome of two AMF species, R. irregularis and Glomus versiforme, and observed greater enrichment of Gammaproteobacteria and Alphaproteobacteria on extraradical mycelium of R. irregularis compared to those associated with Glomus versiforme. The evidence that AMF species can, to some extent, determine the composition of microbial communities in their hyphosphere is thus robust, albeit for a still relatively small number of examples. Furthermore, absolute quantification of microbial taxa within the communities (which are likely very relevant to ecosystem functions they confer) is still largely missing.
Identification of distinct microbial communities associated with hyphae of different AMF species might have been caused by differences in the composition of their hyphal exudates and/or by their different developmental and metabolic traits (Luthfiana et al. 2021). Notably, composition of hyphal exudates can change in response to nutrient availability in the vicinity of the hyphae, which likely influences recruitment of bacteria. Luthfiana et al. (2021) showed that the concentrations of 18 metabolites containing sugars, amino acids, and organic acids were significantly higher in hyphal exudates of R. clarus at low P than at high P supply. Conversely, the concentrations of 10 compounds in the hyphal exudates of R. irregularis were significantly lower under low P than under high P conditions (Luthfiana et al. 2021). Jiang et al. (2021) found a significant increase in bacterial (PSB) abundance and AMF (R. irregularis) hyphal biomass in the presence of organic P compared with treatments without organic P addition under in vitro culture condition. Thus, it appears plausible that organic P triggers changes in composition of hyphal exudates, hyphal growth/branching, and consequently the composition of the hyphosphere microbiome. The details behind such a process have yet to be revealed.
Furthermore, Nuccio et al. (2013) observed the influence of G. hoi on relative abundance of nearly 10% of all bacterial taxa inhabiting decomposing litter, suggesting that N acquisition by AMF is one possible mechanism by which AMF alter bacterial populations in an organic patch (i.e., experimentally created or naturally occurring organic-rich soil microsite) (Bunn et al. 2019). In a pot experiment consisting of spatially discrete (and root-free) organic patches containing different organic N forms and Andropogon gerardii as host plant, Bukovská et al. (2018) firmly established (using various qPCR assays) a significant suppression of the microbial abundances, particularly that of ammonia oxidizing bacteria, in the presence of R. irregularis hyphae networks. This suppression was attributed to the competition that occurs between AMF and microbes for free ammonium ions. Interestingly, Wang et al. (2019) reported significant changes in the alkaline phosphatase-harboring bacterial community associated with the F. mosseae hyphosphere attached to the leek root system in response to different forms of P (KH2PO4 or phytin) with a higher relative abundance of Pseudomonas in phytin treatments compared to KH2PO4 and the control treatments. It is worth noting that alkaline phosphatase activity has been shown to be positively associated with the relative abundance of rare microbial taxa (Liu et al. 2021; Wei et al. 2019). Overall, there is good evidence that both synergistic and antagonistic interactions between AMF and microbes are expected to be regulated to some extent by nutrient availability in the hyphosphere or by nutrient status of the AMF hyphae and/or the host plants.
The information discussed above would point to the existence of a degree of specificity between AMF and certain soil bacteria that is context-dependent (Artursson et al. 2005; Cruz-Paredes et al. 2021; de Boer 2017; Scheublin et al. 2010). A number of mutually interacting factors would therefore shape the structure and spatiotemporal dynamics of the hyphosphere microbiome and its functional role in the plant-AMF-soil continuum, particularly with respect to the C and nutrient cycling. Direct experimental evidence for this happening across environmental and temporal gradients at different scales, however, is still largely lacking.
Knowns and unknowns
To stimulate beneficial bacteria or other microbes, AMF are assumed to exude various compounds into the hyphosphere to act either as energy (C) resources or signals that facilitate microbial growth/migration and their metabolic activities (Jiang et al. 2021). Some microbes may intimately associate with hyphae by colonizing the AMF hyphal surface (Scheublin et al. 2010; Toljander et al. 2006), which could give them a competitive advantage for acquiring hyphal exudates. AMF hyphal colonization allows bacteria to take advantage of living in a nutrient- and water-rich environment, which clearly could facilitate their growth, proliferation, and metabolic activities, as well as mobility throughout the soil. Given the availability of resources, a high degree of competition among microorganisms for the available resources in this microhabitat is likely. Considering that AMF influence the hyphosphere microbial colonization particularly through modulating the composition of their hyphal exudates (Luthfiana et al. 2021), a degree of specialization would be expected to exist in the AMF microbiome. AMF may need to recruit different functional groups of microbes to exploit a wide range of organic compounds in the soil and this may have consequences in conferring specific functional services to the plants and other components of ecosystems. However, there is a scarcity of information about functional traits of different microbial taxa in the AMF hyphosphere and the processes that govern their interactions with the AMF. In other words, the fundamental question of “who does what?” remains largely unanswered.
On one hand, there appears to be a spectrum of associations between hyphae and bacteria ranging from tightly attached to non-attached bacteria (Toljander et al. 2006) and the tightness of association and the ability of attachment is different among various bacterial groups (Bonfante and Anca 2009; Scheublin et al. 2010). Furthermore, it remains unclear as to which taxa are just casual opportunists and which are mutualistically dependent on the specific niche provided by the AMF hyphae. On the other hand, at least some of the AMF hyphae-associated microbes have been demonstrated to provide specific services to the AMF, including enhancement of organic nutrient uptake (Rozmoš et al. 2022; Zhang et al. 2016). However, the underlying mechanisms by which AMF select their symbionts and balance their own need for essential nutrients such as N and P with those of their bacterial companions are not well understood. It also remains unclear whether and to what extent bacteria (or other hyphae-associated microbes) provide other benefits to the AMF such as boosting AMF defence mechanisms or inhibiting AMF pathogens.
Using a bacterium without flagella, Micrococcus luteus, Jiang et al. (2021) recently claimed that non-motile bacteria were unable (unlike the flagellated Rahnella aquatilis) to reach a distant organic P patch even in the presence of AMF hyphae, suggesting that bacterial motility (and particularly the presence of flagella) was required for their migration along the hyphae. However, scanning electron microscopy of AMF hyphae from an unsterile pot experiment revealed a plethora of microbes on the surface of the hyphae coated with liquid water film or mucilaginous substances (Bukovská et al. 2018; Holátko et al. 2021; Jansa and Hodge 2021). There appear to be some groups of bacteria that are immobile on the bumpy, rough surface of the mycorrhizal hyphae. It also is possible that the rough surface of the hyphae could provide various microhabitats that can be inhabited by functionally and/or structurally diverse microbes (Jansa and Hodge 2021). However, it has not yet been possible to unequivocally show that colonization of hyphal surfaces requires motility traits since bacteria are capable of various means (swimming, gliding, swarming) by which to move along a hydrated surface.
Water film thickness on the surface of the AMF hyphae may be a critically important factor contributing to bacterial movement along the hyphae (Jiang et al. 2021). Thus, the microbe migration and community composition along the hyphae could be influenced by the thickness and temporal dynamics of liquid water films. For example, it is plausible that the tortuosity of fungal highway increases dramatically in dry or water-unsaturated soils. The conditions in which bacteria are able to travel along the hyphae thus need to be explored from both AMF and bacterial perspectives.
In parallel to the interactions between AMF and bacteria, there are also interactions among bacteria themselves and with other soil microbes such as protists and/or saprotrophic fungi in the hyphosphere that may induce positive or negative feedbacks on the AMF microbiome. Zhang et al. (2014) indicated that organic P acquisition by AMF hyphae was influenced by the interactions among PSB. In addition, free-living bacterial grazers such as protists could alter bacterial community structure via selective feeding on bacteria or serving as their intermittent host and retaining them inside the protist transiently (Amaro and Martín-González 2021; Amaro et al. 2015; Šimek et al. 1997). Soil protists may also return some of the N they ingest (~ 30%) as free ammonium ions back to the soil (Bonkowski 2004), which can be taken up by other soil microorganisms, including AMF, and passed eventually onto plants (Bukovská et al. 2018; Koller et al. 2013b). Rozmoš et al. (2022) showed that the addition of an amoeboid protist, Polysphondylium pallidum, to AMF hyphosphere in the presence of Paenibacillus sp. significantly enhanced N uptake by AMF and their associated plant roots from an organic (chitin) source. These findings suggest that the interactions between AMF and their associated bacteria should not be addressed without considering the critical role of bacterivores involved in the complex soil food webs. It is also likely that certain hyphosphere interactions are synergistic, such as when the effect of a hyphosphere microorganism on AMF is enhanced by the presence of another organism from a different functional group (guild). Furthermore, it is possible that bacterial movement along hyphae could be aided by eukaryotes such as protists (Rubinstein et al. 2015), but this still remains to be directly demonstrated (Jansa and Hodge 2021). Uncovering the functional overlaps (redundancy) in the AMF hyphosphere microbiome by assembling communities based on their function (observed growth promotion, enhanced nutrient acquisition, etc.) also deserves further investigation. We have summarized known and hypothesized interactions between AMF hyphae and other microorganisms in the hyphosphere in Fig. 2.
Known and hypothesized interactions between arbuscular mycorrhizal fungal (AMF) hyphae and other microorganisms in their hyphosphere. Numbers refer to different processes taking place within AMF hyphosphere microbiome: (1) AMF hyphae are the main and rapid pathway for the transfer of plant carbon (C) to soil microbes (Kaiser et al. 2015). AMF stimulate a range of beneficial soil microbiota (mainly bacteria) through their hyphal exudation of a variety of molecules such as fructose, glucose, and trehalose (Jiang et al. 2021; Wang et al. 2016, 2022). Therefore, different AMF genotypes are thought to associate with different microbial communities due to inherent differences in hyphal exudation patterns. (2) Plant-derived C released by AMF leads to selection of certain bacteria, e.g., Bradyrhizobium, Pseudomonas, and Burkholderia (Drigo et al. 2010). Transmission of organic C from the AMF to surrounding bacteria is likely to be selective (e.g., AMF-mediated C transfer preferentially to Opitutus spp. Mucilanginibacter, Ohktaekwangia, and Massilia spp. (Hünninghaus et al. 2019)). (3) Possible role of some bacteria in promoting AMF spore germination and mycelial growth, e.g., Rhizobiales, Bacillales, and Pseudomonadales (Agnolucci et al. 2015). (4) Protists (e.g., Polysphondylium pallidum) increase N availability in the hyphosphere by releasing N from consumed bacterial biomass (in a process termed soil microbial loop) (Bonkowski 2004; Henkes et al. 2018; Rozmoš et al. 2022), supporting the hypothesis that interactions between AMF and protists could further alter prokaryote community composition/activity/abundance/growth pattern in the hyphosphere. AMF hyphal networks could also provide a prey-rich microhabitat for the protists or other (prokaryotic and eukaryotic) grazers. (5) Movement of motile bacteria toward the resource-rich patches along the fungal highway (Jiang et al. 2021). (6) Positive responses of some bacteria to the presence of AMF hyphae, e.g., Pseudomonas, Burkholderia, Streptomyces, Rhodococcus, and Myxococcales (see Table 1). These bacteria can bind tightly to hyphae or loosely swim in the water film surrounding the hyphosphere (Bonfante and Anca 2009; Jansa and Hodge 2021). They are likely to provide services to AMF such as enhancing nutrient availability or promoting AMF resistance to pathogens. (7) Cooperative interactions between AMF and phosphate-solubilizing bacteria (PSB) in the presence of silicon increase P availability in the hyphosphere and enhance P uptake by AMF (Etesami et al. 2021). (8) Organic N acquisition by AMF hyphae with microbial support (e.g., Paenibacillus sp.) (Nuccio et al. 2013; Rozmoš et al. 2022). Cooperative or antagonistic interactions between AMF and microbes can be determined based on nutrient patch quality (e.g., suppression of some ammonia oxidizing bacteria by AMF due to competition for free ammonia as a shared resource (Bukovská et al. 2018; Dudáš et al. 2022; Veresoglou et al. 2019)). (9) Cooperative interactions between AMF and PSB (e.g., Pseudomonas alcaligenes, Rahnella aquatilis) in the hyphosphere and enrichment of some alkaline phosphatase-harboring bacteria that enhance the mineralization of organic P and microbial P biomass (Jiang et al. 2021; Wang et al. 2016; Zhang et al. 2014). (10) Fructose exuded by the AMF, as a source of energy, stimulates P release by PSB and acts as a signal molecule to trigger P mineralization by bacteria (Gao et al. 2020; Zhang et al. 2018a). (11) The AMF highway interconnecting different plants belonging to the same or different plant species (so called common mycorrhizal network) may facilitate the transfer of some bacteria between the plants (e.g., the migration of Bradyrhizobium diazoefficiens, a N-fixing rhizobial strain, into legumes via the rhizodermis cracks along the AMF hyphae (de Novais et al. 2020)). (12) Mobilization of organically bound sulfur (S) can be facilitated by some hyphosphere bacteria such as Gammaproteobacteria (including Pseudomonas and Stenotrophomonas) and Actinobacteria, which enhances S uptake by AMF and plants (Gahan and Schmalenberger 2015)
Directions for future research
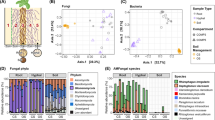
Recent experiments in characterizing the AMF microbiome using high-throughput sequencing techniques has led to the identification of a diversity of bacteria in the hyphosphere (Table 1 and supplementary Table S1). These studies have begun to shed light on the (possible) interactions between individuals or communities of AMF and various bacterial and archaeal taxa and have paved the way for more detailed functional studies of various combinations of AMF and microbes (Table 2).
To date, AMF microbiome research has focused almost exclusively on specific (dual) AMF-bacterial interactions, and less effort has been made to understand AMF microbiome functioning at the community level. Although focusing on bipartite interactions within a single functional type (e.g., AMF-PSB) could provide insights into the level of mutualism within such combinations of microbes, the importance of mutualism at the community level is underestimated because synergism or complementarity usually occurs among several and functionally distinct partners within the same guild. Moreover, a network of trophic interactions binds microorganisms together and cross-feeding cannot simply be ignored. Viewing the AMF microbiome from a systems-level perspective requires that we examine and interpret interactions not only between AMF and microbes, but also between different phylogenetic groups and/or functional guilds of microbes (e.g., bacteria, protists, archaea, and fungi). With the exception of the recent study by Rozmoš et al. (2022) on the interactions between bacteria and protists, we are not aware of any research that addresses specifically the interactions between different life forms in the AMF microbiome.
It is also important to acknowledge that capturing the whole system, with its dynamics and stability, resistance, and resilience to environmental perturbations, is experimentally challenging mainly due to a plethora of simultaneous interactions. While expecting the occurrence of such interactions in the AMF hyphosphere, it still remains challenging to identify key species or genes and their metabolic interactions due to spatial and temporal (micro)scales and due to significant diversity of the relevant microbiomes. To decipher the complexity of players and interactions, novel in vitro and mesocosm experimental setups using state-of-the-art approaches are required to adjust and alter specific microbial strain/guild-related factors such as absence/presence/abundance, and environmental factors including pH and/or nutrient availability.
Despite widely criticized artificiality of the approach, several in vitro culture systems have been used in a few recent studies. For example, Jiang et al. (2021) designed a two-compartment Petri plate system to test whether the PSB bacterium, R. aquatilis HX2, enhanced the uptake of organic P by R. irregularis by migrating along the hyphae to obtain nutrients. In an innovative way, they created an air gap by cutting the solid medium to halt the unspecific migration of bacteria through the medium (Fig. 3a). The same principle was used to design a three-compartment Petri dish system to test the effects of different levels of organic P on bacterial movement along the hyphae (see Jiang et al. (2021)). However, it is still extraordinarily challenging to identify bacteria that actually use the fungal highway to reach nutrient-rich microsites in natural soil.
Four examples of well-designed in vitro and mesocosm experiments aimed at characterizing the microbial community and deciphering biological interactions in the arbuscular mycorrhizal fungal hyphosphere. RC and HC stand for plant root and hyphal compartments, respectively. MSR refers to modified Strullu-Romand medium (Fortin et al. 2002)
Rozmoš et al. (2022) investigated the recycling of organic N via a microbial loop by applying a synthetic approach using 15 N-labelled chitin as an organic N source in a compartmented in vitro culture system (Fig. 3b). This study is unique in that it is the first attempt to unravel the biological interactions between AMF, bacteria, and protists under controlled settings, as this previously was assumed to have significant implications in nutrient cycling in soil (Koller et al. 2013a). Although these kinds of simplified and artificial experimental setups with low complexity may overlook the heterogeneity of soil microbes and soil microsites, such studies are of particular interest as they contribute to a better mechanistic understanding of the biological interaction occurring in soil, especially when combined with isotope labelling approaches. In addition, a major strength of the above study is the possibility to establish bacteria-free controls. It is worth mentioning that each inoculum and each open pot culture, including the non-mycorrhizal (mock) inoculants, contain specific microbiomes that cannot be precisely reconstructed with soil washes (Gryndler et al. 2018). This means that full replicability of microbiome manipulation experiments under open pot settings may be difficult to achieve, unless novel revolutionary methods such as molecular editing tools are applied to genetically manipulate complex microbiomes (Rubin et al. 2022; Tringe 2022).
In spite of that, there have been also some extraordinarily well-designed mesocosm experiments using root-free compartments, delimited from the root-zones by meshes of various sizes (from 25 to 50 µm), aiming to characterize the AMF hyphosphere-associated microbiome and the factors that shape such a community. For instance, in a series of mesocosm experiments, Emmett et al. (2021) examined the effects of different soils, AMF species, and timing on the bacterial community in the AMF hyphosphere. The novelty of their experimental setting was that they exposed root-associated AMF networks grown from microbiome-free (or nearly free) inoculum to a complex bacterial community in nonsterile field soils (Fig. 3c). By doing so, they gave the AMF hyphae access to a natural pool of different bacterial species/genotypes and allowed the AMF to preferentially recruit bacteria from such a complex pool. Thus, using field soil as a microbial starter for the AMF hyphae may reflect the complexity of biotic interactions that take place in natural environments. It would also allow us to ensure that all mycorrhizal and non-mycorrhizal treatments are exposed to an identical and relevant microbial input (rather than sieve-washing and filtrating unsterilized soil inoculum which lead to significant losses and could potentially narrow down the diversity of the microbiome (Ehlers et al. 2008)). In fact, inoculation with open pot-produced AMF inoculant will introduce a lot of microbes with the inoculant itself (e.g., Zhou et al. (2020)) and cannot really be corrected by applying soil washes to the non-mycorrhizal pots (Gryndler et al. 2018).
Another advantage of the study by Emmett et al. (2021) is that not only mycorrhizal and non-mycorrhizal root-free soils were collected and analyzed, but also the AMF hyphal samples. In fact, many preceding studies (Nuccio et al. 2013; Wang et al. 2016; Zhou et al. 2020) have characterized the microbial community of the soil sampled from the hyphal compartment but not from the hyphae itself (see supplementary Table S1 for details). The community in the soil of the hyphal compartment may not well reflect the community of bacteria that are tightly attached to the hyphal surface. The reason is that the hyphal exudates spread out only a few micrometers, so that the soil from the hyphal compartment as a whole is only slightly affected by the hyphae (Wang et al. 2022). However, collecting undisturbed hyphal samples is challenging, and we are not yet fully aware of the effects of washing, agitation, and centrifugation on the recovery and intactness of the hyphosphere microbial community. Although it may introduce some confounding effects and may not fully mimic natural soil, a possible and practical approach is to mix the soil with autoclaved sand and/or glass beads (Emmett et al. 2021; Wang et al. 2019; Zhang et al. 2018b; Zhou et al. 2020) to better be able to collect the fragile and highly dispersed AMF hyphae.
Another innovative mesocosm experiment conducted previously by Wang et al. (2016) consisted of hyphal compartments inoculated with various PSB strains (Fig. 3d, supplementary Table S1). The originality of that study was based on the combination of complementary techniques including 16S rRNA gene amplicon sequencing, terminal restriction fragment length polymorphism (T-RFLP) analysis, and 13CO2 pulse labeling. T-RFLP fingerprinting and DNA fractionation into different buoyant density fractions led to recognition of the hyphal associated PSB that was actively involved in translocation to/utilization of recent (13C-labeled) photosynthates in the soil. Indeed, stable-isotope probing (SIP) approaches combined with high-throughput sequencing and/or multiple omics approaches would be a powerful means to identify microbes that are metabolically associated with/dependent on the living AMF hyphae, and not blur the observations with microbes that consume dead hyphal walls/biomass (Dumont and Hernández García 2019; Radajewski et al. 2000) (supplementary Table S2). The SIP approaches, which are independent of cultivation, have been extensively applied in various systems, including diverse microbial communities in marine environments (Mayali and Weber 2018), soil (Liu et al. 2018; Schwarz et al. 2018), and rhizosphere (Pett-Ridge and Firestone 2017). New developments and refinements in these techniques (Nuccio et al. 2022) could be applied to the AMF microbiome to characterize hyphosphere communities, identify isotopically (13C, 15 N, or 18O) labeled, partially labeled, or unlabeled microbial guilds, estimate their population sizes, analyze their gene expression and metabolic networks, and decipher their functional roles and their positions in food webs, besides providing insights into element fluxes and in microbial cross-feeding in the hyphosphere. We have summarized some of the potential applications of SIP-omics approaches in AMF microbiome research in supplementary Table S2. We have also provided a summary of possible experimental design considerations in future AMF microbiome research in Table 3.
The microbes on the surface of the AMF hyphae can also be screened by nanometer-scale secondary ion mass spectrometry (NanoSIMS) to trace fluxes of stable (C, N, or O) isotopes at the microbial cell level. This technology has been applied to visualize and quantify 15 N and 13C in AMF hyphae (Nuccio et al. 2013) and to visualize C and N utilization on the hyphal surfaces of ectomycorrhizal fungi (Gorka et al. 2019; Kaiser et al. 2015; Mayerhofer et al. 2021). Application of NanoSIMS coupled with SIP-Raman microspectroscopy in AMF microbiome research could also help linking microbial species identities with their function in the hyphosphere at the single-cell level. Moreover, a combination of NanoSIMS with fluorescent in situ hybridization (FISH) approaches (NanoSIMS-FISH) or with gold nanoparticle DNA-hybridization (Kubota et al. 2014) could also provide as yet unexplored options to link identities of hyphosphere microbes with their functions (Musat et al. 2016).
In addition, novel microtechnologies such as microfluidic soil chips mimicking soil environment combined with advanced microspectroscopy techniques such as vibrational-infrared absorption, Raman scattering, and synchrotron radiation-based X-ray microspectroscopy offer opportunities to overcome common obstacles in the study of soil microbiomes in physically and chemically controlled microenvironments in real time (Aleklett et al. 2018; Arellano-Caicedo et al. 2021; Mafla-Endara et al. 2021; Pucetaite et al. 2021). These technologies have been recently applied for the studies of microbial interactions in the rhizosphere (Massalha et al. 2017; Noirot-Gros et al. 2020), formation of soil biogeochemical interfaces (Huang et al. 2017), soil microbial dispersal and interactions (Mafla-Endara et al. 2021), and fungal growth and foraging behavior at the single hyphal scale (Aleklett et al. 2021). In AMF microbiome research, the dynamics of in situ hyphosphere-bacteria interactions at the cellular and subcellular resolution could be visualized and monitored by designing a microfluidic imaging platform where microbe(s) are introduced to the hyphosphere and the microbe’s behavior or chemical responses to the defined environmental condition (e.g., nutrient supply, signaling gradients) monitored with minimal system disturbance.
Finally, a key element to consider in future research is how the presence of specific AMF-bacteria associations in the hyphosphere might drive fitness outcomes in the different AMF and plant species. Deciphering these complex interactions could eventually lead to the development of novel strategies and technologies to harness the full potential of beneficial bacteria or specific inter-organismal interactions. Such knowledge could be utilized to develop more profitable microbial inoculants or combinations of microbes for maintaining plant and soil health, improving agricultural sustainability and increasing crop yields particularly in low-input systems (Bonfante et al. 2019; Jiang et al. 2020; Messa and Savioli 2021; Ray et al. 2020). Enhancing phytoremediation of polluted or contaminated soils could also be envisaged as an additional benefit of improved knowledge of AMF hyphosphere communities. Yet, more experimental studies are needed to validate the interactions which so far have been postulated only theoretically or based on extremely simplified models. Besides, it needs a dedicated research to specifically address the importance of these fine-scale interactions at a full plant or plant community levels, taking into account plant C expenditure to AMF (and its associated microbes) and redistribution of symbiotic benefits and costs among plant individuals involved in common mycorrhizal networks (Walder et al. 2012; Weremijewicz and Janos 2013). It also needs more attention to broaden the scope to other symbiotic fungi, namely the fine root endophytes recruiting from Mucoromycotina, which occupy a very similar ecological niche as the Glomeromycotina (Orchard et al. 2017; Sinanaj et al. 2021) and possibly also to other documented plant-fungal symbioses (Hestrin et al. 2022).
Concluding remarks
The AMF provide both nutritional and non-nutritional benefits to various microorganisms in their hyphosphere, as they represent a microhabitat coated with a water film, rich in carbon/energy sources to facilitate bacterial growth and translocation over long distances. In return, bacteria enhance the nutrient uptake capacity of AMF by breaking down organic nutrients, which AMF only have a limited or no capability to do themselves.
The AMF and microbes could engage in both synergistic and antagonistic interactions. Synergistic interactions may include nutrient resource interdependence, facilitation of movement along hyphae, production of quorum sensing or other signaling molecules or extracellular polymeric matrices that support biofilm formation, and possibly production of volatile organic compounds that regulate microbial coexistence. Antagonistic interactions may include competition for available resources and/or production of suppressive (biocidal) compounds. Synergistic and antagonistic interactions may change with AMF identity and substrate (soil) properties such as nutrient and water availability.
The results of next generation sequencing of 16S rRNA gene from samples of AMF hyphae extracted from pots have suggested that the hyphosphere microbiome is possibly structured by a selection at higher taxonomic ranks. However, the relative abundance of various microbes at different taxonomic levels may simply reflect differences in their environment and fungal host. Thus, hyphosphere microbial communities should be studied not only at a broad phylum resolution, but more revealingly also at lower taxonomic levels where the full suite of ecological functions should become more apparent. This may eventually reveal that the current notion of high-rank selection of bacteria within AMF microbiome is merely based on our limited understanding of functional diversity within those high ranks.
The AMF microbiome research can be advanced through utilizing carefully designed experimental setups, by collecting actively growing hyphae, providing identical microbial inputs to both mycorrhizal and non-mycorrhizal treatments, using only axenically produced AMF inoculants, providing field soil as a microbial starter for the AMF hyphae, considering the temporal dynamic of hyphosphere microbial community by collecting hyphal samples at multiple time points and viewing the AMF microbiome from a systems-level perspective through studying both eukaryotes and prokaryotes in the hyphosphere. In addition, integrating a range of complementary approaches including omics-based technologies, SIP, and other isotope-enabled imaging tools could provide exciting perspectives to elucidate the roles of individuals, populations, genes, proteins, and metabolites in the AMF microbiome. This would also advance our understanding of the fate of C in AMF hyphal exudates and other elemental fluxes in the hyphosphere.
A key element of future research in the AMF microbiome should be a specific focus on better understanding of the functional processes underlying microbial interactions in the hyphosphere in order to facilitate development of optimal combinations of microorganisms that could be used as efficient and competent soil inoculants in sustainable agriculture. The AMF-bacteria interactions appear to be of paramount importance in low-input agricultural systems where biological mechanisms rather than chemical fertilizers are sustaining soil quality and plant production. Thus, increasing our understanding of fundamental aspects of AMF hyphosphere ecology and underlying mechanisms appears critical to securing sustainable food production for the future.
References
Abeysinghe G, Kuchira M, Kudo G, Masuo S, Ninomiya A, Takahashi K, Utada AS, Hagiwara D, Nomura N, Takaya N, Obana N, Takeshita N (2020) Fungal mycelia and bacterial thiamine establish a mutualistic growth mechanism. Life Sci Alliance 3:e202000878. https://doi.org/10.26508/lsa.202000878
Adeyemi NO, Atayese MO, Sakariyawo OS, Azeez JO, AbayomiSobowale SP, Olubode A, Mudathir R, Adebayo R, Adeoye S (2021) Alleviation of heavy metal stress by arbuscular mycorrhizal symbiosis in Glycine max (L.) grown in copper, lead and zinc contaminated soils. Rhizosphere 18:100325. https://doi.org/10.1016/j.rhisph.2021.100325
Agnolucci M, Battini F, Cristani C, Giovannetti M (2015) Diverse bacterial communities are recruited on spores of different arbuscular mycorrhizal fungal isolates. Biol Fertil Soils 51:379–389. https://doi.org/10.1007/s00374-014-0989-5
Aleklett K, Kiers ET, Ohlsson P, Shimizu TS, Caldas VEA, Hammer EC (2018) Build your own soil: exploring microfluidics to create microbial habitat structures. Isme J 12:312–319. https://doi.org/10.1038/ismej.2017.184
Aleklett K, Ohlsson P, Bengtsson M, Hammer EC (2021) Fungal foraging behaviour and hyphal space exploration in micro-structured Soil Chips. Isme J 15:1782–1793. https://doi.org/10.1038/s41396-020-00886-7
Alteio LV, Séneca J, Canarini A, Angel R, Jansa J, Guseva K, Kaiser C, Richter A, Schmidt H (2021) A critical perspective on interpreting amplicon sequencing data in soil ecological research. Soil Biol Biochem 160:108357. https://doi.org/10.1016/j.soilbio.2021.108357
Amaro F, Martín-González A (2021) Microbial warfare in the wild—the impact of protists on the evolution and virulence of bacterial pathogens. Int Microbiol 24:559–571. https://doi.org/10.1007/s10123-021-00192-y
Amaro F, Wang W, Gilbert JA, Roger Anderson O, Shuman HA (2015) Diverse protist grazers select for virulence-related traits in Legionella. Isme J 9:1607–1618. https://doi.org/10.1038/ismej.2014.248
Andrade G, Mihara KL, Linderman RG, Bethlenfalvay GJ (1997) Bacteria from rhizosphere and hyphosphere soils of different arbuscular-mycorrhizal fungi. Plant Soil 192:71–79. https://doi.org/10.1023/a:1004249629643
Arellano-Caicedo C, Ohlsson P, Bengtsson M, Beech JP, Hammer EC (2021) Habitat geometry in artificial microstructure affects bacterial and fungal growth, interactions, and substrate degradation. Commun Biol 4:1226. https://doi.org/10.1038/s42003-021-02736-4
Artursson V, Jansson JK (2003) Use of bromodeoxyuridine immunocapture to identify active bacteria associated with arbuscular mycorrhizal hyphae. Appl Environ Microbiol 69:6208–6215. https://doi.org/10.1128/AEM.69.10.6208-6215.2003
Artursson V, Finlay RD, Jansson JK (2005) Combined bromodeoxyuridine immunocapture and terminal-restriction fragment length polymorphism analysis highlights differences in the active soil bacterial metagenome due to Glomus mosseae inoculation or plant species. Environ Microbiol 7:1952–1966. https://doi.org/10.1111/j.1462-2920.2005.00868.x
Artursson V, Finlay RD, Jansson JK (2006) Interactions between arbuscular mycorrhizal fungi and bacteria and their potential for stimulating plant growth. Environ Microbiol 8:1–10. https://doi.org/10.1111/j.1462-2920.2005.00942.x
Bakker P, Berendsen R, Doornbos R, Wintermans P, Pieterse C (2013) The rhizosphere revisited: root microbiomics. Front Plant Sci 4:165. https://doi.org/10.3389/fpls.2013.00165
Bianciotto V, Bandi C, Minerdi D, Sironi M, Tichy HV, Bonfante P (1996) An obligately endosymbiotic mycorrhizal fungus itself harbors obligately intracellular bacteria. Appl Environ Microbiol 62:3005–3010. https://doi.org/10.1128/aem.62.8.3005-3010.1996
Bianciotto V, Andreotti S, Balestrini R, Bonfante P, Perotto S (2001) Mucoid mutants of the biocontrol strain Pseudomonas fluorescens CHA0 show increased ability in biofilm formation on mycorrhizal and nonmycorrhizal carrot roots. Mol Plant Microbe in 14:255–260. https://doi.org/10.1094/MPMI.2001.14.2.255
Bonfante P, Anca IA (2009) Plants, mycorrhizal fungi, and bacteria: a network of interactions. Annu Rev Microbiol 63:363–383. https://doi.org/10.1146/annurev.micro.091208.073504
Bonfante P, Balestrini R, Mend Gen K (1994) Storage and secretion processes in the spore of Gigaspora margarita Becker and Hall as revealed by high-pressure freezing and freeze substitution. New Phytol 128:93–101. https://doi.org/10.1111/j.1469-8137.1994.tb03991.x
Bonfante P, Venice F, Lanfranco L (2019) The mycobiota: fungi take their place between plants and bacteria. Curr Opin Microbiol 49:18–25. https://doi.org/10.1016/j.mib.2019.08.004
Bonkowski M (2004) Protozoa and plant growth: the microbial loop in soil revisited. New Phytol 162:617–631. https://doi.org/10.1111/j.1469-8137.2004.01066.x
Brundrett MC, Tedersoo L (2018) Evolutionary history of mycorrhizal symbioses and global host plant diversity. New Phytol 220:1108–1115. https://doi.org/10.1111/nph.14976
Bukovská P, Gryndler M, Gryndlerová H, Püschel D, Jansa J (2016) Organic nitrogen-driven stimulation of arbuscular mycorrhizal fungal hyphae correlates with abundance of ammonia oxidizers. Front Microbiol 7:711. https://doi.org/10.3389/fmicb.2016.00711
Bukovská P, Bonkowski M, Konvalinková T, Beskid O, Hujslová M, Püschel D, Řezáčová V, Gutiérrez-Núñez MS, Gryndler M, Jansa J (2018) Utilization of organic nitrogen by arbuscular mycorrhizal fungi-is there a specific role for protists and ammonia oxidizers? Mycorrhiza 28:465–465. https://doi.org/10.1007/s00572-018-0851-y
Bukovská P, Rozmoš M, Kotianová M, Gančarčíková K, Dudáš M, Hršelová H, Jansa J (2021) Arbuscular mycorrhiza mediates efficient recycling from soil to plants of nitrogen bound in chitin. Front Microbiol 12:325. https://doi.org/10.3389/fmicb.2021.574060
Bunn RA, Simpson DT, Bullington LS, Lekberg Y, Janos DP (2019) Revisiting the ‘direct mineral cycling’ hypothesis: arbuscular mycorrhizal fungi colonize leaf litter, but why? Isme J 13:1891–1898. https://doi.org/10.1038/s41396-019-0403-2
Cruz-Paredes C, Diera T, Davey M, Rieckmann MM, Christensen P, Dela Cruz M, Laursen KH, Joner EJ, Christensen JH, Nybroe O, Jakobsen I (2021) Disentangling the abiotic and biotic components of AMF suppressive soils. Soil Biol Biochem 159:108305. https://doi.org/10.1016/j.soilbio.2021.108305
de Boer W (2017) Upscaling of fungal–bacterial interactions: from the lab to the field. Curr Opin Microbiol 37:35–41. https://doi.org/10.1016/j.mib.2017.03.007
de Novais CB, Sbrana C, da Conceição JE, Rouws LFM, Giovannetti M, Avio L, Siqueira JO, Saggin Júnior OJ, da Silva EMR, de Faria SM (2020) Mycorrhizal networks facilitate the colonization of legume roots by a symbiotic nitrogen-fixing bacterium. Mycorrhiza 30:389–396. https://doi.org/10.1007/s00572-020-00948-w
Desirò A, Salvioli A, Ngonkeu EL, Mondo SJ, Epis S, Faccio A, Kaech A, Pawlowska TE, Bonfante P (2014) Detection of a novel intracellular microbiome hosted in arbuscular mycorrhizal fungi. Isme J 8:257–270. https://doi.org/10.1038/ismej.2013.151
Deveau A, Bonito G, Uehling J, Paoletti M, Becker M, Bindschedler S, Hacquard S, Hervé V, Labbé J, Lastovetsky OA, Mieszkin S, Millet LJ, Vajna B, Junier P, Bonfante P, Krom BP, Olsson S, van Elsas JD, Wick LY (2018) Bacterial–fungal interactions: ecology, mechanisms and challenges. FEMS Microbiol Rev 42:335–352. https://doi.org/10.1093/femsre/fuy008
Drigo B, Pijl AS, Duyts H, Kielak AM, Gamper HA, Houtekamer MJ, Boschker HT, Bodelier PL, Whiteley AS, Van Veen JA (2010) Shifting carbon flow from roots into associated microbial communities in response to elevated atmospheric CO2. P Natl Acad Sci USA 107:10938–10942. https://doi.org/10.1073/pnas.0912421107
Dudáš M, Pjevac P, Kotianová M, Gančarčíková K, Rozmoš M, Hršelová H, Bukovská P, Jansa J (2022) Arbuscular mycorrhiza and nitrification: disentangling processes and players through using synthetic nitrification inhibitors. Appl Environ Microbiol 88:20. https://doi.org/10.1128/aem.01369-22
Dumont MG, Hernández García M (2019) Stable isotope probing, methods and protocols Humana Press, New York, USA. 247
Ehlers K, Bünemann EK, Oberson A, Frossard E, Frostegård Å, Yuejian M, Bakken LR (2008) Extraction of soil bacteria from a Ferralsol. Soil Biol Biochem 40:1940–1946. https://doi.org/10.1016/j.soilbio.2008.04.005
Emmett BD, Lévesque-Tremblay V, Harrison MJ (2021) Conserved and reproducible bacterial communities associate with extraradical hyphae of arbuscular mycorrhizal fungi. Isme J 15:2276–2288. https://doi.org/10.1038/s41396-021-00920-2
Etesami H, Jeong BR, Glick BR (2021) Contribution of arbuscular mycorrhizal fungi, phosphate–solubilizing bacteria, and silicon to P uptake by plant. Front Plant Sci 12:1355. https://doi.org/10.3389/fpls.2021.69961
Faghihinia M, Zou Y, Chen Z, Bai Y, Li W, Marrs R, Staddon PL (2020) The response of grassland mycorrhizal fungal abundance to a range of long-term grazing intensities. Rhizosphere 13:100178. https://doi.org/10.1016/j.rhisph.2019.100178
Fortin JA, Bécard G, Declerck S, Dalpé Y, St-Arnaud M, Coughlan AP, Piché Y (2002) Arbuscular mycorrhiza on root-organ cultures. Can J Bot 80:1–20. https://doi.org/10.1139/b01-139
Gahan J, Schmalenberger A (2015) Arbuscular mycorrhizal hyphae in grassland select for a diverse and abundant hyphospheric bacterial community involved in sulfonate desulfurization. Appl Soil Ecol 89:113–121. https://doi.org/10.1016/j.apsoil.2014.12.008
Gao X, Guo H, Zhang Q, Guo H, Zhang L, Zhang C, Gou Z, Liu Y, Wei J, Chen A, Chu Z, Zeng F (2020) Arbuscular mycorrhizal fungi (AMF) enhanced the growth, yield, fiber quality and phosphorus regulation in upland cotton (Gossypium hirsutum L.). Sci Rep 10:2084. https://doi.org/10.1038/s41598-020-59180-3
Gao D, Pan X, Khashi u Rahman M, Zhou X, Wu F (2021) Common mycorrhizal networks benefit to the asymmetric interspecific facilitation via K exchange in an agricultural intercropping system. Biol Fertil Soils 57:959–971. https://doi.org/10.1007/s00374-021-01561-5
Godbold DL, Hoosbeek MR, Lukac M, Cotrufo MF, Janssens IA, Ceulemans R, Polle A, Velthorst EJ, Scarascia-Mugnozza G, De Angelis P, Miglietta F, Peressotti A (2006) Mycorrhizal hyphal turnover as a dominant process for carbon input into soil organic matter. Plant Soil 281:15–24. https://doi.org/10.1007/s11104-005-3701-6
Gorka S, Dietrich M, Mayerhofer W, Gabriel R, Wiesenbauer J, Martin V, Zheng Q, Imai B, Prommer J, Weidinger M, Schweiger P, Eichorst SA, Wagner M, Richter A, Schintlmeister A, Woebken D, Kaiser C (2019) Rapid transfer of plant photosynthates to soil bacteria via ectomycorrhizal hyphae and its interaction with nitrogen availability. Front Microbiol 10:168. https://doi.org/10.3389/fmicb.2019.00168
Gryndler M, Hršelová H, Stříteská D (2000) Effect of soil bacteria on hyphal growth of the arbuscular mycorrhizal fungus Glomus claroideum. Folia Microbiol 45:545–551. https://doi.org/10.1007/BF02818724
Gryndler M, Šmilauer P, Püschel D, Bukovská P, Hršelová H, Hujslová M, Gryndlerová H, Beskid O, Konvalinková T, Jansa J (2018) Appropriate nonmycorrhizal controls in arbuscular mycorrhiza research: a microbiome perspective. Mycorrhiza 28:435–450. https://doi.org/10.1007/s00572-018-0844-x
Henkes GJ, Kandeler E, Marhan S, Scheu S, Bonkowski M (2018) Interactions of mycorrhiza and protists in the rhizosphere systemically alter microbial community composition, plant shoot-to-root ratio and within-root system nitrogen allocation. Front Environ Sci 6:117. https://doi.org/10.3389/fenvs.2018.00117
Hestrin R, Kan MG, Lafler M, Wollard J, Kimbrel JA, Ray P, Blazewicz SJ, Stuart R, Craven K, Firestone M, Nuccio EE, Pett-Ridge J (2022) Plant-associated fungi support bacterial resilience following water limitation. Isme J Press. https://doi.org/10.1038/s41396-022-01308-6
Hildebrandt U, Janetta K, Bothe H (2002) Towards growth of arbuscular mycorrhizal fungi independent of a plant host. Appl Environ Microbiol 68:1919–1924. https://doi.org/10.1128/AEM.68.4.1919-1924.2002
Holátko J, Brtnický M, Kučerík J, Kotianová M, Elbl J, Kintl A, Kynický J, Benada O, Datta R, Jansa J (2021) Glomalin – truths, myths, and the future of this elusive soil glycoprotein. Soil Biol Biochem 153:108116. https://doi.org/10.1016/j.soilbio.2020.108116
Huang X, Li Y, Liu B, Guggenberger G, Shibistova O, Zhu Z, Ge T, Tan W, Wu J (2017) SoilChip-XPS integrated technique to study formation of soil biogeochemical interfaces. Soil Biol Biochem 113:71–79. https://doi.org/10.1016/j.soilbio.2017.05.021
Hünninghaus M, Dibbern D, Kramer S, Koller R, Pausch J, Schloter-Hai B, Urich T, Kandeler E, Bonkowski M, Lueders T (2019) Disentangling carbon flow across microbial kingdoms in the rhizosphere of maize. Soil Biol Biochem 134:122–130. https://doi.org/10.1016/j.soilbio.2019.03.007
Jansa J, Hodge A (2021) Swimming, gliding, or hyphal riding? On microbial migration along the arbuscular mycorrhizal hyphal highway and functional consequences thereof. New Phytol 230:14–16. https://doi.org/10.1111/nph.17244
Jansa J, Treseder K (2017) Introduction: Mycorrhizas and the carbon cycle. In: Johnson NC, Gehring C, Jansa J (eds) Mycorrhizal mediation of soil: fertility, structure and carbon storage. Elsevier, Amsterdam, pp 343–355
Jiang Y, Luan L, Hu K, Liu M, Chen Z, Geisen S, Chen X, Li H, Xu Q, Bonkowski M, Sun B (2020) Trophic interactions as determinants of the arbuscular mycorrhizal fungal community with cascading plant-promoting consequences. Microbiome 8:142. https://doi.org/10.1186/s40168-020-00918-6
Jiang F, Zhang L, Zhou JC, George TS, Feng G (2021) Arbuscular mycorrhizal fungi enhance mineralisation of organic phosphorus by carrying bacteria along their extraradical hyphae. New Phytol 230:304–315. https://doi.org/10.1111/nph.17081
Junier P, Cailleau G, Palmieri I, Vallotton C, Trautschold OC, Junier T, Paul C, Bregnard D, Palmieri F, Estoppey A, Buffi M, Lohberger A, Robinson A, Kelliher JM, Davenport K, House GL, Morales D, Gallegos-Graves LV, Dichosa AEK, Lupini S, Nguyen HN, Young JD, Rodrigues DF, Parra-Vasquez ANG, Bindschedler S, Chain PSG (2021) Democratization of fungal highway columns as a tool to investigate bacteria associated with soil fungi. FEMS Microbiol Rev 97:fiab003. https://doi.org/10.1093/femsec/fiab003
Kaiser C, Kilburn MR, Clode PL, Fuchslueger L, Koranda M, Cliff JB, Solaiman ZM, Murphy DV (2015) Exploring the transfer of recent plant photosynthates to soil microbes: mycorrhizal pathway vs direct root exudation. New Phytol 205:1537–1551. https://doi.org/10.1111/nph.13138
Kikuchi Y, Hijikata N, Ohtomo R, Handa Y, Kawaguchi M, Saito K, Masuta C, Ezawa T (2016) Aquaporin-mediated long-distance polyphosphate translocation directed towards the host in arbuscular mycorrhizal symbiosis: application of virus-induced gene silencing. New Phytol 211:1202–1208. https://doi.org/10.1111/nph.14016
Kohlmeier S, Smits TH, Ford RM, Keel C, Harms H, Wick LY, technology (2005) Taking the fungal highway: mobilization of pollutant-degrading bacteria by fungi. Environ Sci 39:4640–4646. https://doi.org/10.1021/es047979z
Koide RT, Kabir Z (2000) Extraradical hyphae of the mycorrhizal fungus Glomus intraradices can hydrolyse organic phosphate. New Phytol 148:511–517. https://doi.org/10.1046/j.1469-8137.2000.00776.x
Koller R, Rodriguez A, Robin C, Scheu S, Bonkowski M (2013) Protozoa enhance foraging efficiency of arbuscular mycorrhizal fungi for mineral nitrogen from organic matter in soil to the benefit of host plants. New Phytol 199:203–211. https://doi.org/10.1111/nph.12249
Koller R, Scheu S, Bonkowski M, Robin C (2013) Protozoa stimulate N uptake and growth of arbuscular mycorrhizal plants. Soil Biol Biochem 65:204–210. https://doi.org/10.1016/j.soilbio.2013.05.020
Kubota K, Morono Y, Ito M, Terada T, Itezono S, Harada H, Inagaki F (2014) Gold-ISH: a nano-size gold particle-based phylogenetic identification compatible with NanoSIMS. Syst Appl Microbiol 37:261–266. https://doi.org/10.1016/j.syapm.2014.02.003
Linderman R (1991) The rhizosphere and plant growth. In: Keister D.L., P.B. C (eds), Beltsville Symposia in Agricultural Research, p. 343–348. Springer, Dordrecht, The Netherlands
Liu P, Pommerenke B, Conrad R (2018) Identification of Syntrophobacteraceae as major acetate-degrading sulfate reducing bacteria in Italian paddy soil. Environ Microbiol 20:337–354. https://doi.org/10.1111/1462-2920.14001
Liu S, Zhang X, Dungait JAJ, Quine TA, Razavi BS (2021) Rare microbial taxa rather than phoD gene abundance determine hotspots of alkaline phosphomonoesterase activity in the karst rhizosphere soil. Biol Fertil Soils 57:257–268. https://doi.org/10.1007/s00374-020-01522-4
Luthfiana N, Inamura N, Tantriani ST, Saito K, Oikawa A, Chen W, Tawaraya K (2021) Metabolite profiling of the hyphal exudates of Rhizophagus clarus and Rhizophagus irregularis under phosphorus deficiency. Mycorrhiza 31:403–412. https://doi.org/10.1007/s00572-020-01016-z
Mafla-Endara PM, Arellano-Caicedo C, Aleklett K, Pucetaite M, Ohlsson P, Hammer EC (2021) Microfluidic chips provide visual access to in situ soil ecology. Commun Biol 4:1–12. https://doi.org/10.1038/s42003-021-02379-5
Marschner H (1995) Mineral nutrition of higher plants, 2nd ed. Academic Press London. 889
Massalha H, Korenblum E, Malitsky S, Shapiro OH, Aharoni A (2017) Live imaging of root–bacteria interactions in a microfluidics setup. P Natl Acad Sci USA 114:4549–4554. https://doi.org/10.1073/pnas.1618584114
Mayali X, Weber PK (2018) Quantitative isotope incorporation reveals substrate partitioning in a coastal microbial community. FEMS Microbiol Ecol 94 https://doi.org/10.1093/femsec/fiy047
Mayerhofer W, Schintlmeister A, Dietrich M, Gorka S, Wiesenbauer J, Martin V, Gabriel R, Reipert S, Weidinger M, Clode P, Wagner M, Woebken D, Richter A, Kaiser C (2021) Recently photoassimilated carbon and fungus-delivered nitrogen are spatially correlated in the ectomycorrhizal tissue of Fagus sylvatica. New Phytol 232:2457–2474. https://doi.org/10.1111/nph.17591
Messa VR, Savioli MR (2021) Improving sustainable agriculture with arbuscular mycorrhizae. Rhizosphere 19:100412. https://doi.org/10.1016/j.rhisph.2021.100412
Musat N, Musat F, Weber PK, Pett-Ridge J (2016) Tracking microbial interactions with NanoSIMS. Curr Opin Microbiol 41:114–121. https://doi.org/10.1016/j.copbio.2016.06.007
Nazir R, Tazetdinova DI, van Elsas JD (2014) Burkholderia terrae BS001 migrates proficiently with diverse fungal hosts through soil and provides protection from antifungal agents. Front Microbiol 5:598. https://doi.org/10.3389/fmicb.2014.00598
Noirot-Gros M-F, Shinde SV, Akins C, Johnson JL, Zerbs S, Wilton R, Kemner KM, Noirot P, Babnigg G (2020) Functional imaging of microbial interactions with tree roots using a microfluidics setup. Front Plant Sci 11:408. https://doi.org/10.3389/fpls.2020.00408
Nuccio EE, Hodge A, Pett-Ridge J, Herman DJ, Weber PK, Firestone MK (2013) An arbuscular mycorrhizal fungus significantly modifies the soil bacterial community and nitrogen cycling during litter decomposition. Environ Microbiol 15:1870–1881. https://doi.org/10.1111/1462-2920.12081
Nuccio EE, Blazewicz SJ, Lafler M, Campbell AN, Kakouridis A, Kimbrel JA, Wollard J, Vyshenska D, Riley R, Tomatsu A, Hestrin R, Malmstrom RR, Firestone M, Pett-Ridge J (2022) HT-SIP: a semi-automated Stable Isotope Probing pipeline identifies interactions in the hyphosphere of arbuscular mycorrhizal fungi. BioRxiv preprint. https://doi.org/10.1101/2022.07.01.498377
Orchard S, Standish RJ, Dickie IA, Renton M, Walker C, Moot D, Ryan MH (2017) Fine root endophytes under scrutiny: a review of the literature on arbuscule-producing fungi recently suggested to belong to the Mucoromycotina. Mycorrhiza 27:619–638. https://doi.org/10.1007/s00572-017-0782-z
Otto S, Bruni EP, Harms H, Wick LY (2017) Catch me if you can: dispersal and foraging of Bdellovibrio bacteriovorus 109J along mycelia. Isme J 11:386–393. https://doi.org/10.1038/ismej.2016.135
Pett-Ridge J, Firestone M (2017) Using stable isotopes to explore root-microbe-mineral interactions in soil. Rhizosphere 3:244–253. https://doi.org/10.1016/j.rhisph.2017.04.016
Pivato B, Offre P, Marchelli S, Barbonaglia B, Mougel C, Lemanceau P, Berta G (2009) Bacterial effects on arbuscular mycorrhizal fungi and mycorrhiza development as influenced by the bacteria, fungi, and host plant. Mycorrhiza 19:81–90. https://doi.org/10.1007/s00572-008-0205-2
Poveda J, Hermosa R, Monte E, Nicolás C (2019) Trichoderma harzianum favours the access of arbuscular mycorrhizal fungi to non-host Brassicaceae roots and increases plant productivity. Sci Rep 9:11650. https://doi.org/10.1038/s41598-019-48269-z
Pucetaite M, Ohlsson P, Persson P, Hammer E (2021) Shining new light into soil systems: spectroscopy in microfluidic soil chips reveals microbial biogeochemistry. Soil Biol Biochem 153:108078. https://doi.org/10.1016/j.soilbio.2020.108078
Purin S, Rillig MC (2008) Parasitism of arbuscular mycorrhizal fungi: reviewing the evidence. FEMS Microbiol Lett 279:8–14. https://doi.org/10.1111/j.1574-6968.2007.01007.x
Püschel D, Bitterlich M, Rydlová J, Jansa J (2021) Drought accentuates the role of mycorrhiza in phosphorus uptake. Soil Biol Biochem 157:108243. https://doi.org/10.1016/j.soilbio.2021.108243
Radajewski S, Ineson P, Parekh NR, Murrell JC (2000) Stable-isotope probing as a tool in microbial ecology. Nature 403:646–649. https://doi.org/10.1038/35001054
Ray P, Lakshmanan V, Labbé JL, Craven KD (2020) Microbe to microbiome: a paradigm shift in the application of microorganisms for sustainable agriculture. Front Microbiol 11:3323. https://doi.org/10.3389/fmicb.2020.622926
Remy W, Taylor TN, Hass H, Kerp H (1994) Four hundred-million-year-old vesicular arbuscular mycorrhizae. P Natl Acad Sci USA 91:11841–11843. https://doi.org/10.1073/pnas.91.25.11841
Rozmoš M, Bukovská P, Hršelová H, Kotianová M, Dudáš M, Gančarčíková K, Jansa J (2022) Organic nitrogen utilisation by an arbuscular mycorrhizal fungus is mediated by specific soil bacteria and a protist. Isme J 16:676–685. https://doi.org/10.1038/s41396-021-01112-8
Rubin BE, Diamond S, Cress BF, Crits-Christoph A, Lou YC, Borges AL, Shivram H, He C, Xu M, Zhou Z, Smith SJ, Rovinsky R, Smock DCJ, Tang K, Owens TK, Krishnappa N, Sachdeva R, Barrangou R, Deutschbauer AM, Banfield JF, Doudna JA (2022) Species- and site-specific genome editing in complex bacterial communities. Nat Microbiol 7:34–47. https://doi.org/10.1038/s41564-021-01014-7
Rubinstein RL, Kadilak AL, Cousens VC, Gage DJ, Shor LM (2015) Protist-facilitated particle transport using emulated soil micromodels. Environ Sci Technol 49:1384–1391. https://doi.org/10.1021/es503424z
Scheublin TR, Sanders IR, Keel C, van der Meer JR (2010) Characterisation of microbial communities colonising the hyphal surfaces of arbuscular mycorrhizal fungi. Isme J 4:752–763. https://doi.org/10.1038/ismej.2010.5
Schwarz A, Adetutu EM, Juhasz AL, Aburto-Medina A, Ball AS, Shahsavari E (2018) Microbial degradation of phenanthrene in pristine and contaminated sandy soils. Microb Ecol 75:888–902. https://doi.org/10.1007/s00248-017-1094-8
Šimek K, Vrba J, Pernthaler J, Posch T, Hartman P, Nedoma J, Psenner R (1997) Morphological and compositional shifts in an experimental bacterial community influenced by protists with contrasting feeding modes. Appl Environ Microbiol 63:587–595. https://doi.org/10.1128/aem.63.2.587-595.1997
Sinanaj B, Hoysted GA, Pressel S, Bidartondo MI, Field KJ (2021) Critical research challenges facing Mucoromycotina ‘fine root endophytes.’ New Phytol 232:1528–1534. https://doi.org/10.1111/nph.17684
Smith SE, Read DJ (2008) Mycorrhizal symbiosis, 3rd edn. Academic Press, San Diego CA, USA, p 787
Smith SE, Smith FA (2011) Roles of arbuscular mycorrhizas in plant nutrition and growth: new paradigms from cellular to ecosystem scales. Annu Rev Plant Biol 62:227–250. https://doi.org/10.1146/annurev-arplant-042110-103846
Spatafora JW, Chang Y, Benny GL, Lazarus K, Smith ME, Berbee ML, Bonito G, Corradi N, Grigoriev I, Gryganskyi A (2016) A phylum-level phylogenetic classification of zygomycete fungi based on genome-scale data. Mycologia 108:1028–1046. https://doi.org/10.3852/16-042
Svenningsen NB, Watts-Williams SJ, Joner EJ, Battini F, Efthymiou A, Cruz-Paredes C, Nybroe O, Jakobsen I (2018) Suppression of the activity of arbuscular mycorrhizal fungi by the soil microbiota. Isme J 12:1296–1307. https://doi.org/10.1038/s41396-018-0059-3
Talbot J, Allison S, Treseder K (2008) Decomposers in disguise: mycorrhizal fungi as regulators of soil C dynamics in ecosystems under global change. Funct Ecol 22:955–963. https://doi.org/10.1111/j.1365-2435.2008.01402.x
Tamayo E, Gómez-Gallego T, Azcón-Aguilar C, Ferrol N (2014) Genome-wide analysis of copper, iron and zinc transporters in the arbuscular mycorrhizal fungus Rhizophagus irregularis. Front Plant Sci 5:547. https://doi.org/10.3389/fpls.2014.00547
Thirkell TJ, Cameron DD, Hodge A (2016) Resolving the ‘nitrogen paradox’ of arbuscular mycorrhizas: fertilization with organic matter brings considerable benefits for plant nutrition and growth. Plant Cell Environ 39:1683–1690. https://doi.org/10.1111/pce.12667
Tisserant E, Malbreil M, Kuo A, Kohler A, Symeonidi A, Balestrini R, Charron P, Duensing N, dit Frey NF, Gianinazzi-Pearson V (2013) Genome of an arbuscular mycorrhizal fungus provides insight into the oldest plant symbiosis. P Natl Acad Sci USA 110:20117–20122. https://doi.org/10.1073/pnas.1313452110
Toljander JF, Artursson V, Paul LR, Jansson JK, Finlay RD (2006) Attachment of different soil bacteria to arbuscular mycorrhizal fungal extraradical hyphae is determined by hyphal vitality and fungal species. FEMS Microbiol Lett 254:34–40. https://doi.org/10.1111/j.1574-6968.2005.00003.x
Tourlousse DM, Yoshiike S, Ohashi A, Matsukura S, Noda N, Sekiguchi Y (2017) Synthetic spike-in standards for high-throughput 16S rRNA gene amplicon sequencing. Nucleic Acids Res 45:e23. https://doi.org/10.1093/nar/gkw984
Treseder KK, Cross A (2006) Global distributions of arbuscular mycorrhizal fungi. Ecosystems 9:305–316. https://doi.org/10.1007/s10021-005-0110-x
Tringe SG (2022) A toolkit for microbial community editing. Nat Rev Microbiol 20:383. https://doi.org/10.1038/s41579-022-00747-4
van der Heijden MGA, Martin FM, Selosse MA, Sanders IR (2015) Mycorrhizal ecology and evolution: the past, the present, and the future. New Phytol 205:1406–1423. https://doi.org/10.1111/nph.13288
Veresoglou SD, Verbruggen E, Makarova O, Mansour I, Sen R, Rillig MC (2019) Arbuscular mycorrhizal fungi alter the community structure of ammonia oxidizers at high fertility via competition for soil NH4. Microb Ecol 78:147–158. https://doi.org/10.1007/s00248-018-1281-2
Walder F, Niemann H, Natarajan M, Lehmann MF, Boller T, Wiemken A (2012) Mycorrhizal networks: common goods of plants shared under unequal terms of trade. Plant Physiol 159:789–797. https://doi.org/10.1104/pp.112.195727
Wang F, Shi N, Jiang RF, Zhang FS, Feng G (2016) In situ stable isotope probing of phosphate-solubilizing bacteria in the hyphosphere. J Exp Bot 67:1689–1701. https://doi.org/10.1093/jxb/erv561
Wang F, Kertesz MA, Feng G (2019) Phosphorus forms affect the hyphosphere bacterial community involved in soil organic phosphorus turnover. Mycorrhiza 29:351–362. https://doi.org/10.1007/s00572-019-00896-0
Wang F, Zhang L, Zhou J, Rengel Z, George TS, Feng G (2022) Exploring the secrets of hyphosphere of arbuscular mycorrhizal fungi: processes and ecological functions. Plant Soil Press. https://doi.org/10.1007/s11104-022-05621-z
Wei X, Hu Y, Razavi BS, Zhou J, Shen J, Nannipieri P, Wu J, Ge T (2019) Rare taxa of alkaline phosphomonoesterase-harboring microorganisms mediate soil phosphorus mineralization. Soil Biol Biochem 131:62–70. https://doi.org/10.1016/j.soilbio.2018.12.025
Weremijewicz J, Janos DP (2013) Common mycorrhizal networks amplify size inequality in Andropogon gerardii monocultures. New Phytol 198:203–213. https://doi.org/10.1111/nph.12125
Wick LY, Remer R, Würz B, Reichenbach J, Braun S, Schäfer F, Harms H (2007) Effect of fungal hyphae on the access of bacteria to phenanthrene in soil. Environ Sci 41:500–505. https://doi.org/10.1021/es061407s
Xavier LJC, Germida JJ (2003) Bacteria associated with Glomus clarum spores influence mycorrhizal activity. Soil Biol Biochem 35:471–478. https://doi.org/10.1016/S0038-0717(03)00003-8
Zai X-M, Fan J-J, Hao Z-P, Liu X-M, Zhang W-X (2021) Effect of co-inoculation with arbuscular mycorrhizal fungi and phosphate solubilizing fungi on nutrient uptake and photosynthesis of beach palm under salt stress environment. Sci Rep 11:5761. https://doi.org/10.1038/s41598-021-84284-9
Zhang L, Fan JQ, Ding XD, He XH, Zhang FS, Feng G (2014) Hyphosphere interactions between an arbuscular mycorrhizal fungus and a phosphate solubilizing bacterium promote phytate mineralization in soil. Soil Biol Biochem 74:177–183. https://doi.org/10.1016/j.soilbio.2014.03.004
Zhang L, Xu MG, Liu Y, Zhang FS, Hodge A, Feng G (2016) Carbon and phosphorus exchange may enable cooperation between an arbuscular mycorrhizal fungus and a phosphate-solubilizing bacterium. New Phytol 210:1022–1032. https://doi.org/10.1111/nph.13838
Zhang L, Feng G, Declerck S (2018) Signal beyond nutrient, fructose, exuded by an arbuscular mycorrhizal fungus triggers phytate mineralization by a phosphate solubilizing bacterium. Isme J 12:2339–2351. https://doi.org/10.1038/s41396-018-0171-4
Zhang L, Shi N, Fan JQ, Wang F, George TS, Feng G (2018) Arbuscular mycorrhizal fungi stimulate organic phosphate mobilization associated with changing bacterial community structure under field conditions. Environ Microbiol 20:2639–2651. https://doi.org/10.1111/1462-2920.14289
Zhang L, Peng Y, Zhou JC, George TS, Feng G (2020) Addition of fructose to the maize hyphosphere increases phosphatase activity by changing bacterial community structure. Soil Biol Biochem 142:107724. https://doi.org/10.1016/j.soilbio.2020.107724
Zhang L, Zhou J, George TS, Limpens E, Feng G (2022) Arbuscular mycorrhizal fungi conducting the hyphosphere bacterial orchestra. Trends Plant Sci 27:402–411. https://doi.org/10.1016/j.tplants.2021.10.008
Zhou JC, Chai XF, Zhang L, George TS, Wang F, Feng G (2020) Different arbuscular mycorrhizal fungi cocolonizing on a single plant root system recruit distinct microbiomes. mSystems 5:e00929-00920. https://doi.org/10.1128/mSystems.00929-20
Funding
This work was supported by The Ministry of Education, Youth and Sports of Czech Republic (CZ.02.2.69/0.0/0.0/18_053/0017705), Czech Academy of Sciences (RVO 61388971), and Grant Agency of the Czech Republic (21-07275S).
Author information
Authors and Affiliations
Contributions
Faghihinia and Jansa conceived and developed the review. Faghihinia drafted the first version of the manuscript. Faghihinia, Jansa, Halverson, and Staddon contributed to revisions and all approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Faghihinia, M., Jansa, J., Halverson, L.J. et al. Hyphosphere microbiome of arbuscular mycorrhizal fungi: a realm of unknowns. Biol Fertil Soils 59, 17–34 (2023). https://doi.org/10.1007/s00374-022-01683-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00374-022-01683-4