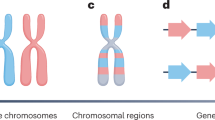
gene trees
and species trees. We construct different lineage histories for different genes, in spite of the fact that intragenic recombination ensures that building a gene tree can become an exercise in averaging over disparate (and reticulating) segmental phylogenies. Combining data across disparate gene trees leads to an average species tree, but whether that represents anything real is dubious. Another ploy is to study mitochondrial and/or chloroplast genomes, confidently asserted to be inherited in strictly lineal fashion, without recombination. Evidence is mounting, however, that even these organellar elements have recombination and that their phylogenies are reticulate. Given the generally reticulate process of evolution at the subspecific level, we should model the collection of relationships more as a redundant and multiply connected network than as a strictly radiating phylogeny.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Smouse, P. Reticulation inside the Species Boundary. J. of Classification 17, 165–173 (2000). https://doi.org/10.1007/s003570000015
Issue Date:
DOI: https://doi.org/10.1007/s003570000015