Abstract
This study investigated the frequency of allelic imbalance (AI) in particular loci in conventional renal cell carcinoma tissue and premalignant lesions of the kidney. DNA from the tumor tissue, premalignant lesions and normal kidney tissue of radical nephrectomy specimens from 33 patients was obtained. It was amplified with a set of eight microsatellite markers, which are located on chromosomes 2, 3, 5, 8, 9, 11, 16, 17. AI in DNA samples was determined by analysis of the alteration in (CA)n repeats. The rates of AI in tumor tissue were found to be between 22.2% and 53.3% and in premalignant lesions between 11.1% and 40.0%. Premalignant lesions and tumor tissues in conventional renal cell carcinoma have the same genotypic changes in 50.0–87.8% (informative cases). These results suggest that the progressive accumulation of AI in areas of premalignant lesions may contribute to the development of renal cell carcinoma, representing an important molecular event in the multistep renal carcinogenesis cascade.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The etiology of renal cell carcinoma (RCC) is unclear. Furthermore, conventional renal cell carcinoma (cRCC) has an extremely variable clinical course [9]. The pathogenesis of cRCC is controversial, and premalignant alterations in the kidney have been described, however, very little is known about the molecular biology of these precancerous changes [11, 16, 20]. A consensus committee of the World Health Organisation suggested naming these lesions renal intratubular neoplasia (RIN), and considered them to be the precancerous abnormalities which might be involved, at intermediate stages, in the sequence of changes by which RCC develops from normal kidney tubules [11].
Microsatellites have been increasingly used to study the genetic structure of human populations and are valuable tools in the characterization of human genetic individuality for population genetics, forensic and clinical purposes [4, 7]. They also play a crucial part in unravelling the genetic basis of tumor formation and progression. Microsatellites have been used to identify genes for susceptibility to colon cancer, retinoblastoma and Wilm’s tumor [13]. Premalignant lesions of the kidney show similar immunocytochemical profiles toRCC, but the genetic changes are poorly understood [11, 20].
The aim of this study was to determine the allelic imbalance (AI) in particular loci in RIN and in tumor tissue to assess whether these have the same genotypic changes as cRCC.
Patients and methods
Specimen collection
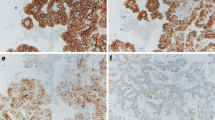
Tissue samples from 33 patients with cRCC who underwent radical nephrectomy were studied (mean age 62 years, male to female ratio 2.3:1). The tumors were histologically classified according to the criteria described by Störkel et al. [14]. The 1997 TNM classification was used for tumor staging, and tumors were graded according to the Fuhrman nuclear grading system [5, 12]. One section from each centimeter of the tumor, one section including both the tumor and adjacent normal paranchyma, at least two sections from normal renal tissue and sections from the normal-renal capsule, tumor-Gerota’s facia, tumor-renal pelvis, and renal vessels were blocked, cut at 5 μm thickness and stained with hematoxylin-eosin (HE). The number of slides per case ranged from 10 to 20, with an average of 14. The sections stained with HE were reviewed and re-examined by the same pathologist (M.K.) (Fig. 1). RIN was selected according to criteria described previously [11, 16, 20]. DNA was isolated from the tissue specimens of the tumor, RIN and normal tubules [6]. Serial 5 μm thick sections of selected tissue blocks were obtained on glass slides, and the areas of interest were microdissected after matching with an adjacent section stained with HE. For tumor and RIN samples, tissue was selected that contained more than 90% of the representative tumor and more than 70% of the RIN on the tissue section, respectively [10]. Corresponding samples of normal tubules that were free of tumor and RIN were obtained from each case as normal tissue. To eliminate cross-contamination, disposable microtome blades were used [10].
PCR and AI analysis
A total of 99 DNA samples were studied. These were isolated from the paraffin blocks in which normal, RIN and tumor tissues were found and were analyzed by using eight dinucleotide repeats which were located on different chromosomes [4, 6].
The method utilized for the characterization of microsatellite alterations was based on PCR amplification. Eight sets of primers were selected, based on chromosome localization and rate of polymorphism [3, 4]. Genomic DNA was amplified by PCR in a total of 25 μl of reaction mixture, which included 100 nm primers, 100 μM each of dNTP, 1× PCR buffer, 1.5 mM magnesium chloride and 2 units of Tag DNA polymerase (Fermentase, USA). Aliquots (5 μl) of the PCR products were added to 5 μl of loading buffer (95% formamide, 20 mM EDTA, 0.05% bromophenol blue and 0.05% xylene cyanol) and the entire sample was denatured at 94°C for 2 min and placed on ice. Aliquots (8 μl) were electrophoresed on 6% polyacrylamide gels and silver nitrate was used for the detection of bands [4]. Microsatellite alterations (microsatellite instability: MI, loss of heterozygosity: LOH) were identified as either a deletion or a band shift. A band shift was defined as an abnormal and reproducible pattern which revealed expansion, contraction or rearranged bands [10]. All altered cases were analyzed at least twice by an additional PCR and an electrophoretic run to confirm the results.
Results
Our findings are summarized in Table 1. Of the 33 tumors and RIN, 53.3% and 40% showed AI (MI and/or LOH), respectively, (informative cases). An example of microsatellite analysis is shown in Fig. 2a. The percentage of AI obtained from each locus in tumor tissue and RIN is summarized in Fig. 2b. When normal, RIN and tumor tissues were compared, the same alterations were found both in 50–87.8% of samples (Fig. 2b). The polymorphism information content (PIC) of the eight microsatellite DNA region in our findings were 0.88, 0.76, 0.91, 0.88, 0.82, 0.94, 0.82 and 0.88, respectively.
Discussion
This is the first study to examine the frequency and patterns of AI in RIN. If AI plays an early and significant role in renal carcinogenesis, one might expect to find it in RIN before the development of cancer. In this study, a high level of AI was detected in 53.3% of tumors and 40% of RIN tissues examined.
It is now widely accepted that human neoplasms are the end result of a multistep process in which the accumulation of several genetic alterations play a major role. Tumor suppressor genes, oncogenes and mismatch repair genes are associated with a variety of human cancers. The genetic instability is thought to increase the normal mutation rate and to cause multiple mutations in oncogenes and tumor suppressor genes. In certain precursor lesions of cancers, these genetic changes are poorly understood [15, 18]. AI is a distinct form of genomic instability linked to defects in DNA mismatch characterized by alterations in the length of simple homopolymeric sequences that are ubiquitous in the genome [1]. It is believed to be associated with genetic defects that promote tumorigenesis when present at multiple loci. The presence of AI in tumor tissue has been associated with unique clinical and pathological characteristics [5, 12, 18].
The rates of AI in RIN and tumor tissue at the D17S1807 locus were determined as 31% and 25%, respectively. Hypermethylated in cancer 1 gene (HIC1), replication protein A1 gene (RPA1), tumor suppressor gene (p53), tumor necrosis factor ligand superfamily-member 12 and MMR gene are localized in this region (17q13) [17]. The rates of AI in tumor tissue were higher than RIN in other loci (D2S301, D3S1613, D5S679, D8S514, D9S157, D11S1356, D16S3018). The AI at the D17S1807 locus might be an early event in cRCC carcinogenesis.
In the flanking region to the microsatellite loci that we studied, DNA repair genes, genomic instability genes, oncogenes, apoptosis inhibitory gene, cyclin dependent kinase, interferon alpha gene, HIC1, recombination or tumor suppressor genes are localized. These results suggest that DNA repair genes and/or other genes may be involved in cRCC. Diakoumis et al. and Von Knobloch et al. detected similar results using microsatellite markers on chromosomes 2, 3,5, 8, 9, 17 [2, 17].
Some studies have reported that laser microdissection is more sensitive in detecting LOH compared to manual microdissection due to a lower contamination by normal cells [19]. We used manual microdissection and although it may be inferior to laser capture microdissection others have reported good results with this technique [10, 15].
The events that take place in the development of cRCC are not well-known. RIN have been found in 23% of normal kidney tissues of patients with RCC and are now accepted as a precursor of RCC [20]. However, the sequence of genetic events that accompany this process are not known. The low local recurrence rate after nephron sparing surgery for small RCC suggests that these lesions may regress. Probably this event can be explained by molecular mechanisms.
The present study exhibited MI that has also been demonstrated to be defective in mismatch repair. The genetic instability associated with a mismatch repair defect occurs in tumors and premalignant lesions [1, 10, 15]. AI, reflected by changes in microsatellite repeat number or LOH, is thought to play an important role in carcinogenesis [12, 17, 18]. Acquisition of AI in premalignant lesions of the kidney such as RIN suggests that this is an early event in the genesis of sporadic carcinomas.
References
Cahill DP, Kinzler KW, Vogelstein B, Lengauer C (1999) Genetic instability and Darwinian selection in tumors. Trends Cell Biol 9:M57-M60
Diakoumis E, Sourvinos G, Kiaris H, Delakos D, Cranidis A, Spandidos DA (1998) Genetic instability in renal cell carcinoma. Eur Urol 33:227
Dib C, Faure S, Fizames C, Samson D, Drout N, Vignal A, Millasseau P, Marc S, Hazan J, Seboun E, Lathrop M, Gyapay G, Morissette J, Weissenbach J (1996) The Genethon human genetic linkage map. Nature 380:A-21, A-25, A-63, A-65, A-79, A-101, A-105
Erdem H, Pehlivan S, Topaloğlu H, Togan I, Ozguc M (1997) Allele distribution of D5S125, MAP1B5′ and D5S679 microsatellite markers in turkish spinal muscular atrophy families. Turk J Pediatr 39:447
Fuhrman S, Lasky L, Lumas C (1982) Prognostic significance of morphologic parameters in renal cell carcinoma. Am J Pathol 6:655
Goelz SE (1985) Purification of DNA from formaldehyde fixed and paraffin embedded human tissue. Biochem Biophys Res Commun 130:118
Ingvarsson S, Finndottir V, Sigurdsson A, Geirsson G (2000) Population studies and validation of paternity determinants by six microsatellite loci. Forensic Sci 45:692
Jiang F, Desper R, Papadimitriou CH, Schaffer AA, Kallioniemi OP, Richter J, Schraml P, Sauter G, Mihatsch MJ, Moch H (2000) Construction of evolutionary tree models for renal cell carcinoma from comparative genomic hybridization data. Cancer Res 60:6503
Kirkali Z, Lekili M (2003) Renal cell carcinoma: new prognostic factors? Curr Opin Urol 13:433
Leung WK, Kim JJ, Kim JG, Graham DY, Sepulveda AR (2000) Microsatellite instability in gastric intestinal metaplasia in patients with and without gastric cancer. Am J Pathol 156:537
Mourad WA, Nestok BR, Saleh GY, Solez K, Power RF, Jewell LD (1994) Dysplastic tubular epithelium in “normal” kidney associated with renal cell carcinoma. Am J Surg Pathol 18:1117
Sobin LH, Wittekind CH (1997) For the International Union Against Cancer (IUCC); TNM Classification of Malignant Tumors, 5th edn. Wiley-Liss, New York
Steiner G, Sidransky D (1996) Molecular differential diagnosis of renal cell carcinoma: from microscopes to microsatellites. Am J Pathol 149:2081
Störkel S, Eble JN, Adlakha K, Amin M, Blute ML, Bostwick DG, Darson M, Delahunt B, Iczkowski K (1997) Classification of renal cell carcinoma. Workgroup No I. Cancer 80:987
Van Dekken H, Alers JC, Riegman PHJ, Rosenberg C, Tilanus HW, Vissers K (2001) Molecular cytogenetic evaluation of gastric cardia adenocarcinoma and precursor lesions. Am J Pathol 158:1961
Van Poppel H, Nilsson S, Algaba F, Bergerheim U, DalCin P, Fleming S, Hellsten S, Kirkali Z, Klotz L, Lindblad P, Ljungberg B, Mulders P, Roskams T, Ross RK, Walker C, Wersall D (2000) Precancerous lesions in the kidney. Scand J Urol Nephrol Suppl 34:136
Von Knobloch R, Hegele A, Brandt H, Varga Z, Wille S, Kalble T, Heidenreich A, Hoffmann R (2002) High frequency of serum DNA alterations in renal cell carcinoma detected by fluorescent microsatellite analysis. Int J Cancer 98:889
Weber RG, Scheer M, Born IA, Joos S, Cobbers JM, Hofele C, Reifenberger G, Zoller JE, Lichter P (1998) Recurrent chromosomal imbalances detected in biopsy material from oral premalignant and malignant lesions by combined tissue microdissection, universal DNA amplification, and comparative genomic hybridization. Am J Pathol 153:295
Wild P, Knuechel R, Dietmaier W, Hofstaedter F, Hartmann A (2000) Laser microdissection and microsatellite analyses of breast cancer reveal a high degree of tumor heterogeneity. Pathobiology 68:180
Yörükoğlu K, Aktaş S, Mungan MU, Kirkali Z (1999) Tubular dysplasia and carcinoma in situ: precursors of renal cell carcinoma. Urology 53:684
Acknowledgement
This work was supported by the Turkish Scientific and Technical Research Council (TUBITAK) Research Fund no:SBAG-1940 and Ege University Research Fund no: 98/BIL/020.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pehlivan, S., Koyuncuoglu, M., Pehlivan, M. et al. Premalignant lesions of the kidney share the same genetics changes as conventional renal cell carcinoma. World J Urol 22, 120–123 (2004). https://doi.org/10.1007/s00345-003-0384-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00345-003-0384-6