Abstract
Soil salinity is one of the major abiotic stresses restricting crop production. Mechanisms of salt response have been intensively studied in model plants such as Arabidopsis and rice, but are rarely known in sorghum. In this study, we compared the transcriptome profiles between two cultivars with different salt tolerance under salt treatment (0.8% NaCl) for 0, 48, and 72 h. On average, about 243.9 million clean reads, representing 32.4 thousand transcripts and 26.4 thousand unigenes with 829 new genes were detected in each library. Also, over 112,000 single nucleotide polymorphisms were identified, which may supply useful resources for marker development. In total, 5647 differentially expressed genes (DEGs) were identified from all of the comparisons. Functional annotation analysis indicated that expression of genes in transcriptional regulation, signal transduction, and secondary metabolism changed significantly between the two varieties under salt stress, and hundreds of genes involved in the salt stress response were differentially expressed, especially genes encoding receptors like kinases and transcription factors. Besides, qRT-PCR analysis of expression profiles of the selected DEGs was in keeping with the results from RNA-seq analysis. Based on the findings, we proposed several candidate genes that might be used to improve salt tolerance in sorghum. The transcriptional profiles presented here provide further understanding of the salt-tolerance mechanism in sorghum.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Salinity is a major environmental factor limiting plant growth and productivity. High salinity leads to ionic stress and water scarcity, which damages cellular ion homeostasis, membrane stability, enzyme activity, as well as biological processes including photosynthesis and respiration (Hasegawa and others 2000; Zhu 2001). A series of developmental processes such as seed germination, and vegetative and reproductive growth are adversely affected under high salt conditions, resulting in the decline of crop yield and quality (Monreal and others 2013). Therefore, it is very important to elucidate salt response mechanisms and discover key genes involved in the salt stress response to achieve efficient crop production. Many physiological and molecular biological studies have investigated mechanisms of salt tolerance in model plants such as Arabidopsis thaliana (Shi and others 2000, 2003; Yokoi and others 2002), Oryza sativa (Lin and others 2004; Hu and others 2006), Medicago truncatula (Zahaf and others 2012) and so on. Other noteworthy reports have provided further understanding of salt-tolerance mechanisms in recent decades (Munns and Tester 2008; Deinlein and others 2014; Munns and Gilliham 2015). However, many indistinct molecular mechanisms remain to be understood due to the complexity of the plant response to salt stress.
As one of the most important cereal crops, sorghum is one of the few resilient crops with good adaptive ability to changing climatic conditions, especially to increasingly serious phenomena such as salinity, drought, and high temperature (Carpita and McCann 2008). In addition, the genome of sorghum is small (∼ 730 Mb), thus making it an attractive model system in exploring the molecular mechanisms of adaptation to abiotic stresses (Paterson and others 2009; Calvino and Messing 2012; Mace and others 2013). It is, however, difficult to discover salt-tolerance mechanisms in sorghum because of lack of detailed genetic and sequence information. Fortunately, the development of next generation sequencing (NGS) technologies and the release of genome sequences of several sorghum lines (Paterson and others 2009; Mace and others 2013) provide favorable chances for investigating significant mechanisms in sorghum on the transcriptome level. So far, a series of studies about sorghum have been reported and genes were discovered in response to abiotic stress (Sui and others 2015; Surender Reddy and others 2015; Sayyad-Amin and others 2016). However, the integrated dissection of the molecular mechanism in salt response is still largely unknown.
In this study, to get a comprehensive understanding of genes and gene networks in regulating salt stress, we exploited RNA-seq technology to obtain a whole transcriptome and characterize it by employing the sequenced genomes of sorghum, Arabidopsis, and rice. We also explored orthologous genes with differential expression levels between two species at different time points after salt treatment.
Materials and Methods
Plant Culture and RNA Isolation
Seeds of sorghum cultivars, 623B and HN (Henong 16), were disinfected with 70% ethanol, soaked, and pre-germinated for 24 h at room temperature (25 °C ± 1 °C). Then the budding and neat seeds were cultured in sterilized quartz sand in a green house. At the three leaves stage, plants were treated with 0.8% NaCl for 0, 48, and 72 h. The NaCl solution was renewed every 24 h. Then the whole plants were collected and frozen in liquid nitrogen and stored at − 80 °C for mRNA isolation. Each treatment was repeated three times.
Total RNAs were extracted using the Trizol plus kit (ThermoFisher, USA), followed by DNA digestion with DNase I. RNA quality and integrity was measured using Nanodrop, Qubit 2.0 and Aglient 2100. Qualified RNA was used for subsequent RNA library construction.
Preparation of mRNA Library and Sequencing
The mRNA library was constructed and sequenced by the BMK cloud company. In brief, the mRNAs with a polyA tail were firstly collected and fragmented using fragmentation buffer. Then first-strand cDNA was synthesized using reverse transcriptase with hexamer primers. Subsequently, the second-strand cDNA was generated using RnaseH and DNA polymerase I, and double-stranded cDNA was purified by AMPure XP beads, and then end repaired. Adapter ligation was carried out after adding “A” tail, and templates with the desired size range were subjected to PCR. Then the mRNA libraries were enriched by PCR and tested with Qubit 2.0 and Agilent 2100, and finally sequenced using Illumina HiSeq 2500 method.
Sequencing Data Quality Control and Transcriptome Alignment
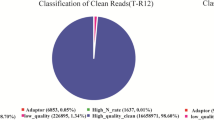
Raw data quality was controlled with criteria of less than 10% low-quality bases (Phred score < 20), followed by adapter sequences and removing of primers. Then the filtered reads (Clean Reads) were mapped to the reference genome using TopHat2 software.
Annotation and Function Analyses of Genes
To annotate the identified genes, the sequences were analyzed with various nucleotide and protein databases, including the non-redundant protein database (nr), SwissPort, Gene Ontology (GO), and Clusters of Orthologous Groups (COG) with criteria of E values less than 10−5. The Blast2GO program was used to classify GO terms, and pathway annotation was carried out by search against the KEGG database.
SNP Identification
SNPs were identified between the two cultivars and reference genome using SAMtools. SNP was screened by four factors, TopHat2 score ≥ 50, interval scale between two single mismatched base ≥ 5 bp, Variant Calling score ≥ 20 , and Sequencing Depth Between 5x and 100x.
Differentially Expressed Gene Identification
For the discovery of DEGs, FPKM (Fragments Per Kilobase of transcript per Million fragments mapped) and DESeq (Anders and Huber 2010) were employed to analyze gene expression levels, and the DEGs were screened with Fold Change ≥ 2 and FDR ≤ 0.01.
Results
Statistics of Transcriptome Sequencing Results
The Illumina platform, powerful in quantifying gene expression, is the most common approach to RNA-seq. To investigate molecular mechanisms of sorghum response to salt stress, 6 cDNA libraries from two cultivars, salt-sensitive HN and relatively salt-tolerant 623B were designed (623B-CK, 623B-48h, and 623B-72h; HN-CK, HN-48h, and HN-72h). 623B-CK, 623B-48h, and 623B-72h represented 623B libraries which were treated with 0.8% NaCl for 0, 48, and 72 h, respectively. HN-CK, HN-48h, and HN-72h represented HN libraries which were treated with 0.8% NaCl for 0, 48, and 72 h respectively. Seedling plants with three leaves were used for mRNA isolation, and mRNAs were isolated and subjected to paired-end sequencing using the Illumina HiSeqTM2500 method in the current study.
After quality control (trimming adapters and filtering out low-quality reads), 23,416,958–25,820,196 clean reads (the percentage of Q20 and GC being 92.86–93.25% and 52.68–53.93%) were obtained, which were blasted to reference genome using software TopHat2 (Kim and others 2013). Most reads (76.43–80.95%) could be mapped to the reference genome, among which 69.73–78.43% were unique mapped ones (Table 1).
On average, 32,465 transcripts with 2100 bp in mean length were obtained in every library (Table 2). Among the transcripts, more than 99.6% of them had lengths over 300 bp and about 88% over 1000 bp. The transcripts were mapped to 26,373 unigenes with 829 new genes on average (Table 2). Functions of the new genes from unannotated genomic regions were further systematically explored by searching other databases. Results showed that 466 new genes could be matched with an annotation in GO (Ashburner and others 2000), 104 in COG (Tatusov and others 2000), 23 in KEGG (Kanehisa and others 2004), 398 in Swiss-Prot/UniProt (Apweiler and others 2004), and 755 in NR (Pruitt and others 2005) (Table 3). On average, the length of the 99.74% of annotated new genes (756) was more than 300 bp, and that of 78.50% annotated new genes (595) was over 1000 bp.
SNP Identification in Sorghum
SNP is a useful molecular marker in the study of plant genetics and plant breeding. In this study, SAMtools (Li and others 2009) were used to screen SNPs based on the comparison of the mapping results of two sorghum cultivars and reference genome. More than 5000 SNPs in each 623B comparison and about 30,000 SNPs in every HN comparison including homozygous and heterozygous were identified. Among the SNPs, more than 90% were genic and about 60% were transitional. Besides, the heterozygosity of 623B sets was 85.74–88.65%, which is about 5 times higher than that of HN (15.65–17.74%), although the SNP number of 623B was much less than that of HN. Detailed information is listed in Supplemental File 1.
Exploration of DEGs in Response to Salt Stress
According to the design, gene responses to salt could be systematically explored in a time course manner, which is applicable to the two cultivars (623B-48h/623B-CK and 623B-72h/623B-CK; HN-48h/HN-CK and HN-72h/HN-CK) as well as genes involved in salt stress regulation (HN-CK/623B-CK, HN-48h/623B-48h, and HN-72h/623B-72h).
To identify genes with a significant change in expression, the FPKM method (Fragments Per Kilobase per Million reads) was used and differentially expressed genes (DEGs) were detected at a threshold of |log2ratio| ≥ 1 with p value ≤ 0.05 and FDR ≤ 0.01.
As listed in Table 4, a total of 5647 differentially expressed genes (DEGs) including 375 new genes were identified from all of the seven comparisons. In detail, 604 up-regulated genes and 431 down-regulated genes were detected in the 623B-48h/623B-CK set, whereas in the 623B-72h/623B-CK set, 711 up-regulated genes and 643 down-regulated genes were found. Meanwhile, 477 up-regulated genes and 418 down-regulated genes were identified in HN-48h/HN-CK, whereas in HN-72h/HN-CK, 853 up-regulated genes and 1240 down-regulated genes were picked.
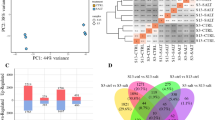
Venn diagram analysis showed that 278 up-regulated and 175 down-regulated genes overlapped between 623B-48h/623B-CK and 623B-72h/623B-CK, whereas 189 common genes were found to be up-regulated and 133 down-regulated between HN-48h/HN-CK and HN-72h/HN-CK (Fig. 1).
Furthermore, we compared the DEGs between HN and B23B under both normal and salt stress conditions (Table 4). In the absence of salt stress, 995 genes were up-regulated and 1119 genes were down-regulated in HN-CK/623B-CK. In contrast, under salt stress condition, 1035 genes were up-regulated and 1133 down-regulated in HN-48h/623B-48h, whereas 900 were up-regulated and 1378 genes were down-regulated in HN-72h/623B-72h. Venn diagram results showed that 305 common genes were up-regulated and 631 were down-regulated in these three comparisons (Fig. 2).
Functional Classification of Differentially Expressed Genes
To describe functional information of the DEGs, GO, COG, and KEGG enrichment analyses were carried out.
Figure 3 shows GO classification results of the DEGs, which were summarized into three categories: cellular component, molecular function, and biological process. Among these categories, “extracellular part and region,” “electron carrier activity,” “antioxidant activity,” “nutrient reservoir activity,” and “cell killing” were significantly enriched in all the four comparisons. However, in 623B-48h/623B-CK and 623B-72h/623B-CK, the percent of DEGs involved in “macromolecular complex,” “structural molecule activity,” and “protein binding transcription factor activity” significantly decreased, whereas DEGs with “nucleoid” or “extracellular matrix/part” function enriched. In HN-48h/HN-CK and HN-72h/HN-CK, genes with “enzyme regulator activity” and “receptor activity” were also enriched, whereas the percent of DEGs involved in “macromolecular complex” and “structural molecule activity” largely decreased only in HN-48h/HN-CK, and DEGs of “nucleoid” just enriched in HN-72h/HN-CK.
Besides, the category “response to salt stress (GO:0009651)” was also significantly enriched among all the comparisons. One hundred and three and 142 DEGs with 54 common genes in 623B-48h/623B-CK and 623B-72h/623B-CK, 111 and 215 DEGs with 55 common genes in HN-48h/HN-CK and HN-72h/HN-CK were classified into salt stress response (Fig. 4a, b). In HN-CK/623B-CK, HN-48h/623B-48h, and HN-72h/623B-72h, there were respectively 145, 144, and 159 DEGs which shared 43 genes in the GO term “response to salt stress” (Fig. 4c).
In addition, the DEGs of HN-48h/623B-48h and HN-72h/623B-72h were subjected to the Clusters of Orthologous Groups (COG) classification, 629 and 700 DEGs from which were classified into 22 COG categories, respectively (Fig. 5). Among the COG, the cluster for “general function prediction only” comprises the largest proportion, followed by “secondary metabolites biosynthesis, transport and catabolism,” “Signal transduction mechanisms,” “transcription,” “amino acid transport and metabolism,” “carbohydrate transport and metabolism,” and “posttranslational modification, protein turnover, chaperones.” The categories “cell motility,” “extracellular structure,” and “nuclear structure” had no corresponding genes.
Furthermore, to identify potential biological pathways represented in the transcriptome, the Kyoto Encyclopedia of Genes and Genomes database (KEGG) was used for further analysis. Results showed that 173 DEGs in HN-72h/623B-72h were assigned to 87 KEGG pathways, and “Flavonoid biosynthesis” [PATHWAY: ko00941], “Biosynthesis of unsaturated fatty acids” [PATHWAY: ko01040], “Photosynthesis” [PATHWAY: ko00195], “Glycolysis/Gluconeogenesis” [PATHWAY: ko00010], and “Nitrogen metabolism” [PATHWAY: ko00910] were significantly enriched (Supplemental File 1). However, in both 623B-72h/623B-CK and HN-72h/HN-CK, KEGGs such as “Protein processing in endoplasmic reticulum,” “Plant hormone signal transduction,” and “Starch and sucrose metabolism” were highly enriched (Supplemental File 2 and 3).
Identification of Functional Genes Related to Salt Tolerance
Protein Kinases
Protein kinases play important roles in plant growth and add to tolerance to unfavorable environments (Xiong and others 2002). Among all protein kinases, receptor-like kinases (RLKs) play critical parts in perceiving external signals and activating a series of intracellular reactions under salt and other stress (Lease and others 1998; Ouyang and others 2010; Osakabe and others 2013; Sun and others 2013a, b; Park and others 2014). In our results, 57 and 54 DEGs respectively from 623B-48h/623B-CK and 623B-72h/623B-CK were predicted to encode protein kinases, among which more than 81% were receptor kinases (47 genes in 623B-48h/623B-CK, 82.5%; and 44 genes in 623B-72h/623B-CK, 81.5%). Besides, there were 19 common protein kinases identified among them, 18 of which were receptor kinases. Meanwhile, there were 40 and 94 protein kinases detected in HN-48h/HN-CK and HN-72h/HN-CK, and more than 80% of them were receptor kinases (34 genes, 85.0% in HN-48h/HN-CK; and 76 genes, 80.9% in HN-72h/HN-CK). In addition, there were 14 receptor kinases among all the 15 common protein kinases between the two comparisons. All of the comparisons shared 3 receptor kinase genes with similar expression levels, two down-regulated genes (Sobic.002G326400 and Sobic.006G005500) and one up-regulated gene Sobic.008G175000 (Supplemental File 4). Therefore, these genes might play important roles in salt tolerance.
Transcription Factors
Transcription factors (TFs) extensively participated in various biological processes and play significant roles in regulating the expression of specific downstream genes in response to abiotic stress (Golldack and others 2011). In this investigation, classification for differentially expressed transcripts identified 28 transcription factors belonging to 12 families from 623B-48h/623B-CK, and ethylene-responsive transcription factors were significantly enriched. In 623B-72h/623B-CK, 67 transcription factors from DEGs were divided into 17 clusters, among which the most distinguished groups are ethylene-responsive transcription factors, MYB and bHLH transcription factors. For all the genes, there were 20 common transcription factors in 623B-48h/623B-CK and 623B-72h/623B-CK. In addition, in HN-48h/HN-CK and HN-72h/HN-CK, there were 32 and 84 differentially expressed transcription factors belonging to more than 7 families, among which ERF, MYB, bHLH, and heat stress transcription factor families were dramatically differentially expressed and they had 16 genes in common (Supplemental File 5). The results indicated that these TFs may elevate salt tolerance in sorghum.
Quantitative Real-Time PCR Validation of DEGs
To validate the transcriptome sequencing results, 9 genes were selected for qRT-PCR analysis, and each gene contained at least one differentially expressed result. Detailed information of these genes and primers are listed in Supplemental File 6 and qRT-PCR results are presented in Fig. 6. In total, 8 of the 9 genes showed different expression patterns between HN and 623B under salt treatment. Expression of Sobic.001G001200, Sobic.001G044600, Sobic.002G013000, and Sobic.003G413600 displayed significant change in HN comparisons, but kept relatively steady in 623B comparisons. Expression of Sobic.001G066900, Sobic.005G041000, and Sobic.001G008600 increased in HN sets but decreased in 623B sets, whereas the expression of Sobic.002G020000 displayed opposite results. Only Sobic.002G012800 showed a similar expression trend between HN and 623B. Overall, 5 and 7 genes showed similar expression patterns in HN and 623B RNA-seq data compared with qRT-PCR results. For example, Sobic.003G413600, which may encode an HAK5-like potassium transporter, was down-regulated in HN, but not differentially expressed in 623B, and Sobic.002G012800, a NADP-linked oxidoreductase superfamily gene, dramatically increased in both of HN and 623B sets.
Discussion
The development of high-throughput sequencing technologies brings great benefits to uncover the molecular basis of stress response, and assists in the understanding of genome-wide expression patterns of genes in sorghum under abiotic stress (Zhang and others 2010; Dugas and others 2011; Gelli and others 2014; Johnson and others 2014; Chopra and others 2015; Yang and others 2015). Expression profiling using RNA-seq is an effective way for discovering genes and pathways associated with the salinity response in sorghum.
Here, we performed transcriptome analysis of two cultivars under salt conditions and compared the expression profiles between control and salt-treated plants. Data of transcriptome sequencing results such as clean reads, mapped reads, as well as transcripts and unigenes were very similar between the two cultivars, which indicated that the sequencing results were credible and suitable for further analysis. In this study, over 110,000 SNPs were identified in total and about 91.8% of them were genic SNPs. Interestingly, the SNP number was 4–5 times more in the salt-sensitive cultivar HN than in the salt-resistant cultivar 623B, whereas the heterozygosity of HN was about 1/5 compared to that of 623B. These results strongly indicated that these SNPs may play important roles in the salt response and could be very valuable markers for salt resistance breeding.
As is well known, a stress signal transduction pathway contains a series of extracellular and intracellular reactions, including outside signal perception, second messenger generation, and protein phosphorylation cascade activation. Then target genes are activated to strengthen the cell resistance or protect cellular activities (Xiong and others 2002). In this study, KEGG pathways such as signal transduction, protein process, and carbohydrate metabolism were highly enriched in both of the two cultivars under salt treatment, which may reflect their similar response to stress. Notably, genes involved in resistance metabolism, photosynthesis, and nutrition processes as well as many newly discovered genes showed significantly different expression profiles between HN and 623B, and genes with “enzyme regulator activity” and “receptor activity” were enriched in HN rather than 623B. These results may contribute to explaining the reasons why HN is more tolerant than 623B under salt treatment, and exploring the mechanisms of salt resistance.
Previous studies have indicated that SOS1, NHX1, and HKT1 are key Na+ transporters involved in plant salt tolerance (Apse and others 1999; Shi and others 2000; Platten and others 2006). In our RNA-seq data, the expression of NHX-like and CHX-like genes remained relatively steady after salt treatment, whereas 2 of the 3 HKT1-like genes displayed dramatically higher expression levels in HN than those in 623B (Supplemental File 7). A recent study in Puccinellia tenuiflora showed that, under salt stress, expression of PtSOS1, PtHKT1;5 and PtNHX1 were significantly increased within 6 h, and then decreased after 24 h (Zhang and others 2017). These results indicated that such genes may act as instantaneous factors in salt response.
At present, some RLKs have been reported to regulate the salt stress response, such as OsSIK1 and OsGIRL1 in rice, as well as GsSRK and GsRLCK in Glycine soja (Ouyang and others 2010; Sun and others 2013a, b; Park and others 2014). Transgenic plants with overexpression of OsSIK1, GsSRK, or GsRLCK showed higher tolerance to salt and drought stresses than control plants, whereas Arabidopsis plants with overexpressed OsGIRL1 displayed a hypersensitive response to salt stress. In our data, expression of the OsSIK1 ortholog gene Sobic.001G085700 showed no difference in any of the comparisons. However, Sobic.004G091300, an ortholog gene of OsGIRL1, was significantly down-regulated in HN-48h/HN-CK and HN-72h/HN-CK (|log2FC| = − 1.85 and − 3.03). This result was in line with the manifestation of OsGIRL1 and indicated that Sobic.004G091300 could be a marker gene in salt resistance.
In plants, salt stress signaling always transduces via abscisic acid-dependent and acid-independent pathways, as well as several other factors. HAB1 and RD29B have been reported to function in response to abiotic stress (Dugas and others 2011; Gelli and others 2014). In our results, expressions of their ortholog genes (Sobic.003G242200 and Sobic.001G200700) were dramatically up-regulated after salt treatment, and expression change of the RD29B-like gene (Sobic.001G200700) even reached more than 24 fold. Moreover, transcription factors including MYB, NAC, and ERF were highly abundant in our results. These transcription factors are relatively widespread, and have significant functions in the abiotic stress response in Arabidopsis and rice (Hu and others 2006; Dai and others 2007; Abogadallah and others 2011; Hao and others 2011; Yang and others 2012; Licausi and others 2013; Rong and others 2014). Thus, these transcription factors discovered in this study may be ideal targets for salt-resistant breeding in plants. In addition, RNA-seq data and qRT-PCR results showed that expression of some genes in 623B contrasted with those in HN, demonstrating the different performances between the two cultivars.
In summary, we conducted a comprehensive analysis of sorghum under salt stress using RNA-seq. Hundreds of SNPs and DEGs were identified, and dozens of DEGs related to salt tolerance were discovered. The common DEGs, GO terms, COG classifications, and KEGG pathways demonstrated that both 623B and HN shared similar strategies under salt treatment, whereas the distinct expression pattern and classification of genes indicated the difference between the two cultivars in response to salt stress, which may result in salt sensitivity of 623B and salt tolerance of HN. This transcriptome study develops the molecular basis for understanding the specific salt-tolerance mechanisms in sorghum, and will also provide valuable genetic resources for salt-tolerance breeding in other species.
Availability of Supporting Data
The raw data files for this RNA-seq during our experiment are deposited in NCBI (BioProject: PRJNA395348).
References
Abogadallah GM, Nada RM, Malinowski R, Quick P (2011) Overexpression of HARDY, an AP2/ERF gene from Arabidopsis, improves drought and salt tolerance by reducing transpiration and sodium uptake in transgenic Trifolium alexandrinum L. Planta 233(6):1265–1276
Anders S, Huber W (2010) Differential expression analysis for sequence count data. Genome Biol 11(10):R106
Apse MP, Aharon GS, Snedden WA, Blumwald E (1999) Salt tolerance conferred by overexpression of a vacuolar Na+/H+ antiport in Arabidopsis. Science 285(5431):1256–1258
Apweiler R, Bairoch A, Wu CH, Barker WC, Boeckmann B, Ferro S, Gasteiger E, Huang H, Lopez R, Magrane M, Martin MJ, Natale DA, O’Donovan C, Redaschi N, Yeh LS (2004) UniProt: the universal protein knowledge base. Nucleic Acids Res 32(Database issue):D115–D119
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25(1):25–29
Calvino M, Messing J (2012) Sweet sorghum as a model system for bioenergy crops. Curr Opin Biotechnol 23(3):323–329
Carpita NC, McCann MC (2008) Maize and sorghum: genetic resources for bioenergy grasses. Trends Plant Sci 13(8):415–420
Chopra R, Burow G, Hayes C, Emendack Y, Xin Z, Burke J (2015) Transcriptome profiling and validation of gene based single nucleotide polymorphisms (SNPs) in sorghum genotypes with contrasting responses to cold stress. BMC Genom 16(1):1040
Dai X, Xu Y, Ma Q, Xu W, Wang T, Xue Y, Chong K (2007) Overexpression of an R1R2R3 MYB gene, OsMYB3R-2, increases tolerance to freezing, drought, and salt stress in transgenic Arabidopsis. Plant Physiol 143(4):1739–1751
Deinlein U, Stephan AB, Horie T, Luo W, Xu G, Schroeder JI (2014) Plant salt-tolerance mechanisms. Trends Plant Sci 19(6):371–379
Dugas DV, Monaco MK, Olsen A, Klein RR, Kumari S, Ware D, Klein PE (2011) Functional annotation of the transcriptome of Sorghum bicolor in response to osmotic stress and abscisic acid. BMC Genom 12:514
Gelli M, Duo Y, Konda AR, Zhang C, Holding D, Dweikat I (2014) Identification of differentially expressed genes between sorghum genotypes with contrasting nitrogen stress tolerance by genome-wide transcriptional profiling. BMC Genom 15:179
Golldack D, Lüking I, Yang O (2011) Plant tolerance to drought and salinity: stress regulating transcription factors and their functional significance in the cellular transcriptional network. Plant Cell Rep 30(8):1383–1391
Hao YJ, Wei W, Song QX, Chen HW, Zhang YQ, Wang F, Zou HF, Lei G, Tian AG, Zhang WK, Ma B, Zhang JS, Chen SY (2011) Soybean NAC transcription factors promote abiotic stress tolerance and lateral root formation in transgenic plants. Plant J 68(2):302–313
Hasegawa PM, Bressan RA, Zhu JK, Bohnert HJ (2000) Plant cellular and molecular responses to high salinity. Annu Rev Plant Physiol Plant Mol Biol 51:463–499
Hu H, Dai M, Yao J, Xiao B, Li X, Zhang Q, Xiong L (2006) Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice. Proc Natl Acad Sci USA 103(35):12987–12992
Johnson SM, Lim FL, Finkler A, Fromm H, Slabas AR, Knight MR (2014) Transcriptomic analysis of Sorghum bicolor responding to combined heat and drought stress. BMC Genom 15:456
Kanehisa M, Goto S, Kawashima S, Okuno Y, Hattori M (2004) The KEGG resource for deciphering the genome. Nucleic Acids Res 32(Database issue):D277–D280
Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL (2013) TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14(4):R36
Lease K, Ingham E, Walker JC (1998) Challenges in understanding RLK function. Curr Opin Plant Biol 1(5):388–392
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25(16):2078–2079
Licausi F, Ohme-Takagi M, Perata P (2013) APETALA2/ethylene responsive factor (AP2/ERF) transcription factors: mediators of stress responses and developmental programs. New Phytol 199(3):639–649
Lin HX, Zhu MZ, Yano M, Gao JP, Liang ZW, Su WA, Hu XH, Ren ZH, Chao DY (2004) QTLs for Na+ and K+ uptake of the shoots and roots controlling rice salt tolerance. Theor Appl Genet 108(2):253–260
Mace ES, Tai S, Gilding EK, Li Y, Prentis PJ, Bian L, Campbell BC, Hu W, Innes DJ, Han X, Cruickshank A, Dai C, Frere C, Zhang H, Hunt CH, Wang X, Shatte T, Wang M, Su Z, Li J, Lin X, Godwin ID, Jordan DR, Wang J (2013) Whole-genome sequencing reveals untapped genetic potential in Africa’s indigenous cereal crop sorghum. Nat Commun 4:2320
Monreal JA, Arias-Baldrich C, Pérez-Montaño F, Gandullo J, Echevarría C, García-Mauriño S (2013) Factors involved in the rise of phosphoenolpyruvate carboxylase-kinase activity caused by salinity in sorghum leaves. Planta 237(5):1401–1413
Munns R, Gilliham M (2015) Salinity tolerance of crops—what is the cost? New Phytol 208(3):668–673
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681
Osakabe Y, Yamaguchi-Shinozaki K, Shinozaki K, Tran LS (2013) Sensing the environment: key roles of membrane-localized kinases in plant perception and response to abiotic stress. J Exp Bot 64(2):445–458
Ouyang SQ, Liu YF, Liu P, Lei G, He SJ, Ma B, Zhang WK, Zhang JS, Chen SY (2010) Receptor-like kinase OsSIK1 improves drought and salt stress tolerance in rice (Oryza sativa) plants. Plant J 62(2):316–329
Park S, Moon JC, Park YC, Kim JH, Kim DS, Jang CS (2014) Molecular dissection of the response of a rice leucine-rich repeat receptor-like kinase (LRR-RLK) gene to abiotic stresses. J Plant Physiol 171(17):1645–1653
Paterson AH, Bowers JE, Bruggmann R, Dubchak I, Grimwood J, Gundlach H, Haberer G, Hellsten U, Mitros T, Poliakov A, Schmutz J, Spannagl M, Tang H, Wang X, Wicker T, Bharti AK, Chapman J, Feltus FA, Gowik U, Grigoriev IV, Lyons E, Maher CA, Martis M, Narechania A, Otillar RP, Penning BW, Salamov AA, Wang Y, Zhang L, Carpita NC, Freeling M, Gingle AR, Hash CT, Keller B, Klein P, Kresovich S, McCann MC, Ming R, Peterson DG, Mehboob ur R, Ware D, Westhoff P, Mayer KF, Messing J, Rokhsar DS (2009) The Sorghum bicolor genome and the diversification of grasses. Nature 457(7229):551–556
Platten JD, Cotsaftis O, Berthomieu P, Bohnert H, Davenport RJ, Fairbairn DJ, Horie T, Leigh RA, Lin HX, Luan S, Mäser P, Pantoja O, Rodríguez-Navarro A, Schachtman DP, Schroeder JI, Sentenac H, Uozumi N, Véry AA, Zhu JK, Dennis ES, Tester M (2006) Nomenclature for HKT transporters, key determinants of plant salinity tolerance. Trends Plant Sci 11:372–374
Pruitt KD, Tatusova T, Maglott DR (2005) Ncbi reference sequence (refseq): a curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Res 33(Database issue):D501–D504
Rong W, Qi L, Wang A, Ye X, Du L, Liang H, Xin Z, Zhang Z (2014) The ERF transcription factor TaERF3 promotes tolerance to salt and drought stresses in wheat. Plant Biotechnol J 12(4):468–479
Sayyad-Amin P, Jahansooz MR, Borzouei A, Ajili F (2016) Changes in photosynthetic pigments and chlorophyll-a fluorescence attributes of sweet-forage and grain sorghum cultivars under salt stress. J Biol Phys 42(4):601–620
Shi H, Ishitani M, Kim C, Zhu JK (2000) The Arabidopsis thaliana salt tolerance gene SOS1 encodes a putative Na+/H+ antiporter. Proc Natl Acad Sci USA 97(12):6896–6901
Shi H, Lee BH, Wu SJ, Zhu JK (2003) Overexpression of a plasma membrane Na+/H+ antiporter gene improves salt tolerance in Arabidopsis thaliana. Nat Biotechnol 21(1):81–85
Sui N, Yang Z, Liu M, Wang B (2015) Identification and transcriptomic profiling of genes involved in increasing sugar content during salt stress in sweet sorghum leaves. BMC Genom 16:534
Sun X, Sun M, Luo X, Ding X, Cai H, Bai X, Liu X, Zhu Y (2013a) A Glycine soja ABA-responsive receptor-like cytoplasmic kinase, GsRLCK, positively controls plant tolerance to salt and drought stresses. Planta 237(6):1527–1545
Sun XL, Yu QY, Tang LL, Ji W, Bai X, Cai H, Liu XF, Ding XD, Zhu YM (2013b) GsSRK, a G-type lectin S-receptor-like serine/threonine protein kinase, is a positive regulator of plant tolerance to salt stress. J Plant Physiol 170(5):505–515
Surender Reddy P, Jogeswar G, Rasineni GK, Maheswari M, Reddy AR, Varshney RK, Kavi Kishor PB (2015) Proline over-accumulation alleviates salt stress and protects photosynthetic and antioxidant enzyme activities in transgenic sorghum [Sorghum bicolor (L.) Moench]. Plant Physiol Biochem 94:104–113
Tatusov RL, Galperin MY, Natale DA, Koonin EV (2000) The COG database: a tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res 28(1):33–36
Xiong L, Schumaker KS, Zhu JK (2002) Cell signaling during cold, drought, and salt stress. Plant Cell 14(Suppl):S165–S183
Yang A, Dai X, Zhang WH (2012) A R2R3-type MYB gene, OsMYB2, is involved in salt, cold, and dehydration tolerance in rice. J Exp Bot 63(7):2541–2556
Yang QS, Gao J, He WD, Dou TX, Ding LJ, Wu JH, Li CY, Peng XX, Zhang S, Yi GJ (2015) Comparative transcriptomics analysis reveals difference of key gene expression between banana and plantain in response to cold stress. BMC Genom 16:446
Yokoi S, Quintero FJ, Cubero B, Ruiz MT, Bressan RA, Hasegawa PM, Pardo JM (2002) Differential expression and function of Arabidopsis thaliana NHX Na+/H+ antiporters in the salt stress response. Plant J 30(5):529–539
Zahaf O, Blanchet S, de Zelicourt A, Alunni B, Plet J, Laffont C, de Lorenzo L, Imbeaud S, Ichante JL, Diet A, Badri M, Zabalza A, Gonzalez EM, Delacroix H, Gruber V, Frugier F, Crespi M (2012) Comparative transcriptomic analysis of salt adaptation in roots of contrasting Medicago truncatula genotypes. Mol Plant 5(5):1068–1081
Zhang G, Guo G, Hu X, Zhang Y, Li Q, Li R, Zhuang R, Lu Z, He Z, Fang X, Chen L, Tian W, Tao Y, Kristiansen K, Zhang X, Li S, Yang H, Wang J, Wang J (2010) Deep RNA sequencing at single base-pair resolution reveals high complexity of the rice transcriptome. Genome Res 20(5):646–654
Zhang WD, Wang P, Bao Z, Ma Q, Duan LJ, Bao AK, Zhang JL, Wang SM (2017) SOS1, HKT1;5, and NHX1 synergistically modulate Na+ homeostasis in the halophytic grass Puccinellia tenuiflora. Front Plant Sci 8:576
Zhu JK (2001) Plant salt tolerance. Trends Plant Sci 6(2):66–71
Acknowledgements
This study was supported by grants from Hebei Province Key Basic Research Foundation (15962905D).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: JC, JC. Performed the experiments: JC, GR, HQ, XX, JC. Analyzed the data: JC, GR, LH, and JC. Contributed reagents/materials/analysis tools: GR, HQ, and XX.
Corresponding author
Ethics declarations
Conflict of interest
The authors declared that they had no conflicting interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Cui, J., Ren, G., Qiao, H. et al. Comparative Transcriptome Analysis of Seedling Stage of Two Sorghum Cultivars Under Salt Stress. J Plant Growth Regul 37, 986–998 (2018). https://doi.org/10.1007/s00344-018-9796-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00344-018-9796-9