Abstract
Halophyte Suaeda salsa is native to the saline soil in the Yellow River Delta. Soil salinity can reduce plant productivity and therefore is the most important factor for the degradation of wetlands in the Yellow River Delta. In this work we characterized the salinity-induced effects in S. salsa in terms of metabolic profiling, antioxidant enzyme activities, and gene expression quantification. Our results showed that salinity inhibited plant growth of S. salsa and upregulated gene expression levels of myo-inositol-1-phosphate synthase (INPS), choline monooxygenase (CMO), betaine aldehyde dehydrogenase (BADH), and catalase (CAT), and elevated the activities of superoxide dismutase (SOD), peroxidase (POD), CAT, and glutathione peroxidase (GPx). The significant metabolic responses included the depleted amino acids malate, fumarate, choline, phosphocholine, and elevated betaine and allantoin in the aboveground part of S. salsa seedlings as well as depleted glucose and fructose and elevated proline, citrate, and sucrose in root tissues. Based on these significant biological markers, salinity treatments induced clear osmotic stress (for example, INPS, CMO, BADH, betaine, proline) and oxidative stress (for example, SOD, POD, CAT, GPx activities), disturbed protein biosynthesis/degradation (amino acids and total protein) and energy metabolism (for example, glucose, sucrose, citrate) in S. salsa.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The Chenopodiaceae C3 halophyte Suaeda salsa is the pioneer halophyte in the Yellow River Delta and is widely consumed as a popular vegetable due to its sufficient nutrient components, including 7 vitamins, 18 essential amino acids, 5 essential elements, and other antioxidant ingredients (Wang and others 2007; Zhang and others 2008; Zhao and others 2010). In recent decades, S. salsa has exhibited its important economic value because its seeds contain approximately 40% oil, rich in unsaturated fatty acids, which can be converted to chemical compounds for industrial and pharmaceutical use (Wang and others 2007). For example, Zhang and others (2008) reported that the seed oil of S. salsa could reduce body fat and improve immunity in rats. S. salsa is native to saline soils and even thrives in the intertidal zone of the Yellow River Delta, where soil salinity is often higher than 3% (Song and others 2008).
Salinity is a major adverse environmental constraint to plant productivity, limiting the utilization of about 800 million ha of agricultural land globally (Munns 2005; Owojori and others 2008; Sonjak and others 2009; Dendooven and others 2010; Li and others 2011; Yang and others 2011; Zhang and others 2011a). As estimated, 80,000,000 ha of cultivated land is affected by soil salinity, which corresponds to 5% of all cultivated land (Munns 2002; Askaril and others 2006). High levels of soil salinity can cause water deficit, ion toxicity, and nutrient deficiency leading to molecular damage and even plant death (Tsonev and others 1998; Shao and others 2008; Maggio and others 2010). In addition, salt stress induces oxidative damage to plants by producing reactive oxygen species (ROS) such as the superoxide radical hydrogen peroxide and the hydroxyl radical in plant cells (Borsani and others 2001; Alscher and others 2002; Parida and Jha 2010). Therefore, increasing salinity has become the main adverse factor for the degradation of wetlands in the Yellow River Delta in China (Cui and others 2008). It is necessary to assess the physiological and molecular responses induced by salinity in the native plants in the Yellow River Delta, which could help us better understand the regulative mechanisms of plants to salinity. However, little is known about the metabolic responses of S. salsa to salinity.
Metabolomics, a system biology approach, has been defined as the global profiling of the low-molecular-weight (<1,000 Da) metabolites contained in cells, tissues, biofluids, and even whole organisms (Lindon and others 1999; Davis 2005). The components (metabolites) of the metabolome, as the end-products of metabolisms, can represent the functional responses of a cell. The measurements of large numbers of endogenous metabolites thus provide a broad view of the biochemical status of an organism that can be used to monitor and assess the metabolic responses induced by exogenous factors such as environmental stressors (Fiehn 2002; Pedras and Zheng 2010; Sun and others 2010). Practically, metabolomics can measure metabolic changes with modern analytical techniques such as nuclear magnetic resonance (NMR) spectroscopy and mass spectrometry. Because NMR spectroscopy can analyze all metabolites containing proton(s) simultaneously, NMR-based metabolomics has been widely applied in plant sciences (Xiao and others 2008; Dai and others 2010). Metabolomics of the plant usually involves studies of plant–environment interactions and assessments of plant function and health at the molecular level, and it focuses on the metabolic responses to environmental changes (drought, salinity), abiotic stresses (heavy-metal contaminants), and transgenic events (Aliferis and others 2009; Bundy and others 2009).
In the present study, 1H-NMR-based metabolomics was applied to halophyte S. salsa exposed to two environmentally relevant salinities (170 and 500 mM) to detect the metabolic changes (biomarkers) and then to characterize the salinity-induced effects. In addition, the transcriptional status of some key antioxidant- and osmotic-related genes was measured based on the gene expression levels in plant tissues using the reverse transcription polymerase chain reaction (RT-PCR) technique. The antioxidant status was characterized on the basis of enzymatic activities in S. salsa. Using these combined biochemical parameters, including the metabolic biomarkers, gene expression levels, and antioxidant enzymatic activities; we expected to elucidate the effects of salinity in the halophyte S. salsa.
Materials and Methods
Chemicals
Sodium chloride (NaCl), sodium dihydrogen phosphate (Na2HPO4), disodium hydrogen phosphate (NaH2PO4), and mercury chloride (all analytical grade) were purchased from Guoyao Chemical Co. Ltd. (Shanghai, China). Deuterium oxide (D2O, 99.9% in D) and sodium 3-trimethylsilyl [2,2,3,3-D4] propionate (TSP) were purchased from Cambridge Isotope Laboratories (Miami, FL, USA). The antioxidant enzyme kits were purchased from Nanjing Jiancheng Bioengineering Institute (Nanjing, China).
Cultivation of S. salsa Under Salt Stresses
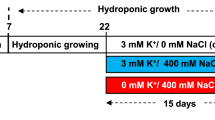
The seeds of S. salsa were collected from the Yellow River Delta in November 2009 and stored in a refrigerator at 4°C for 7 months. The seeds were surface sterilized using 0.5% HgCl2 for 10 min and then washed in sterilized double-distilled water three times. Thirty seeds of similar size were sown in sand in three replicate plastic jugs with a diameter of 20 cm (n = 10, one control and two salinity-exposed groups). The sand was collected from the intertidal zones of the Yellow River Delta and rinsed in diluted nitric acid (1%) to eliminate impurities such as organic matter and metal irons.
The sown S. salsa seeds were irrigated with Hoagland’s nutrient solution. After being sown in plastic jugs for 4 weeks, all the seedlings of the two exposed groups were irrigated with the Hoagland’s nutrient solution containing 170 and 500 mM NaCl, respectively. The two salinities (170 and 500 mM, prepared from NaCl) in the Hoagland’s nutrient solution were similar to the soil salinities of the Yellow River Delta (Wang and others 2007). The culture condition was 28 ± 4°C, photoperiod of 12 h light/12 h darkness, relative humidity of 70%, and photosynthetically active radiation of 600 μmol m−2 s−1. After exposure for 15 days, seedlings (n = 6) from both control and exposed groups were randomly harvested. After a quick measure of the length and weight of the aboveground part of the seedlings, all the plant samples, including the tissues of roots and the aboveground part of seedlings, were flash-frozen in liquid nitrogen and stored at −80°C prior to metabolite extraction, mRNA extraction, and enzymatic assay.
Metabolite Extraction
Polar metabolites were extracted from plant tissues (root and aboveground part) using the solvent system of methanol/water (1/1) as described previously (Kim and Verpoorte 2010; Liu and others 2011a, b). Briefly, the tissue sample was ground in a liquid N2-cooled mortar and pestle. The tissue powder (weighing from 250 to 300 mg per sample) was transferred to a tube containing ~50 ceramic beads of 1 mm diameter, and then thoroughly homogenized in 3.33 ml g−1 methanol/water (1/1) using a high-throughput homogenizer (Precellys 24, Bertin, France). After homogenization, the sample was transferred to an Eppendorf tube and vortexed for 15 s three times. Following centrifugation (3,000×g, 10 min, 4°C), the supernatant was removed and then lyophilized. It was subsequently resuspended in 600 μl of 100 mM of phosphate buffer (Na2HPO4 and NaH2PO4, including 0.5 mM TSP, pH 7.0) in D2O. The mixture was vortexed and then centrifuged at 3,000 g for 5 min at 4°C. The supernatant substance (550 μl) was pipetted into a 5-mm NMR tube prior to NMR analysis.
NMR Analysis
Extracts of S. salsa tissue were analyzed on a Bruker AV 500 NMR spectrometer at 500.18 MHz (at 298 K), as described previously (Zhang and others 2011b, c; Liu and others 2011c). One-dimensional (1D) 1H-NMR spectra were obtained using a 11.9-μs pulse, 6009.6-Hz spectral width, 0.1-s mixing time, and 3.0-s relaxation delay with standard 1D NOESY pulse sequence and 128 transients collected into 16,384 data points. Data sets were zero-filled to 32,768 points, and exponential line broadenings of 0.3 Hz were applied before Fourier transformation. All 1H-NMR spectra were phased, baseline-corrected, and calibrated (TSP at 0.0 ppm) manually using TopSpin (ver. 2.1, Bruker). NMR spectral peaks were assigned following tabulated chemical shifts (Fan 1996) using Chenomx software (evaluation version, Chenomx Inc., Canada).
Spectral Preprocessing and Multivariate Data Analysis
One-dimensional H1-NMR spectra were converted to a format for multivariate analysis using custom-written ProMetab software in Matlab (ver. 7.0; MathsWorks, Natick, MA, USA). Each spectrum was segmented into 0.01-ppm bins between 0.2 and 10.0 ppm, with bins from 4.70 to 5.20 ppm (water) excluded from all the NMR spectra. Bins between 8.32 and 8.35 ppm, between 8.26 and 8.28 ppm, between 8.24 and 8.26 ppm, between 8.18 and 8.20 ppm, between 7.97 and 8.01 ppm, between 7.70 and 7.85 ppm, between 7.52 and 7.56 ppm, and between 6.53 and 6.58 ppm containing pH-sensitive NMR peaks were compressed into single bins. The total spectral area of the remaining bins was normalized to unity to facilitate the comparison between the spectra. All the NMR spectra were generalized log-transformed (glog) with a transformation parameter λ = 1.0 × 10−8, which was optimized using ProMetab software to stabilize the variance across the spectral bins and to increase the weightings of the less intense peaks (Parsons and others 2007).
Principal component analysis (PCA) was used in this work for the separation of control and various salinity-exposed groups. PCA is an exploratory unsupervised pattern recognition (PR) method that calculates inherent variation within the data sets without use of the class membership. The algorithm of PCA calculates the highest amount of correlated variation along PC1, with subsequent PCs containing correspondingly smaller amounts of variance. For each model built, the loading vectors for the PCs can be used for the identification of the contributive metabolites (metabolic biomarkers) for the clusters (Xu 2004). One-way analysis of variance (ANOVA) with Tukey’s test was performed on the PC scores of various groups of samples to test the significance of the separations between the control and salinity-exposed groups. For each model built, the loading vector for the PC could be examined to identify the metabolites that contributed to the clusters. SAM software (Tusher and others 2001) was then used to find significant metabolic differences among salinity-exposed groups with appropriate false discovery rate (FDR) cutoffs. Bins that changed significantly (at FDR < 0.01) were subsequently identified using Chenomx software and isolated. For the identification of significant metabolites, one-way ANOVA was conducted on the ratios of significantly changed (at FDR < 0.01) bin areas (peak intensity) of metabolites to the total spectral area (Katsiadaki and others 2009). These significant metabolites contributed to the separation between control and salinity-treated samples and hence were considered metabolic biomarkers induced by salinity exposures. A P value of 0.05 was considered significant for the ANOVA on the metabolites between control and exposed samples.
Total mRNA Extraction and Gene Quantification
Total RNA from the aboveground part of the S. salsa seedlings was isolated using the TRIzol reagent (Invitrogen). Gene-specific primers for myo-inositol 1-phosphate synthase (INPS), choline monooxygenase (CMO), betaine aldehyde dehydrogenase (BADH), catalase (CAT), glutathione peroxidase (GPx), and the internal control actin were used to amplify amplicons specific for S. salsa. The sequences of primers and the length of amplicons are given in Table 1. The fluorescent real-time quantitative PCR amplifications were carried out in triplicate in a total volume of 20 μl containing 10 μl of 2× SYBR® Premix Ex Taq™ (TaKaRa, Japan), 0.4 μl of 50 × ROX Reference DYE II, 4.8 μl DEPC-treated H2O, 0.4 μl of each primer, 4.0 μl of 1:20 diluted cDNA. The fluorescent real-time quantitative PCR program was as follows: 50°C for 2 min and 95°C for 10 min, followed by 40 cycles of 94°C for 15 s, 58°C for 45 s, 72°C for 30 s. Dissociation curve analysis of amplification products was performed at the end of each PCR to confirm that only one PCR product was amplified and detected. After the PCR program, data were analyzed with the ABI 7500 SDS software (Applied Biosystems). To maintain consistency, the baseline was set automatically by the software. The comparative CT method (2−∆∆CT method) was used to analyze the expression level of the genes (Livak and Schmittgen 2001).
Measurement of Antioxidant Enzyme Activities
The aboveground part of the S. salsa seedlings (n = 6) were ground in liquid nitrogen, and the antioxidant enzyme activities were assayed by a multiscan spectrum microplate spectrophotometer (Infinite M200, TECAN) according to the manufacturer’s protocols using enzyme kits (Jiancheng, Nanjing, China). In this work, the antioxidant enzymes for measuring activity included superoxide dismutase (SOD, EC 1.15.1.1), peroxidase (POD, EC 1.11.1.7), glutathione S-transferases (GST, EC 2.5.1.18), catalase (CAT, EC 1.11.1.6), and glutathione peroxidase (GPx, EC 1.11.1.9). Protein concentration was determined with the Coomassie Brilliant Blue G-250 dye-binding method, using bovine serum albumin as the standard (Bradford 1976). All the enzyme activities were expressed as U mg−1 protein.
Statistical Analysis
Data of growth parameters, enzyme activities, and gene expressions were expressed as the mean ± standard deviation (n = 6). The data were statistically analyzed using the Statistics toolbox in Matlab (ver. 7.0; MathsWorks, Natick, MA). One-way ANOVA (analysis of variance) with Tukey’s test was conducted on the data, and a significant difference was defined as P < 0.05.
Results
Plant Growth
Table 2 gives the weight and length of S. salsa seedlings exposed to various salinities (170 and 500 mM) for 15 days. Plant growth was significantly (ANOVA, P < 0.01) inhibited by exposure to high salinity (500 mM NaCl) in terms of the decreased weight and length. Although the weight and length values from low-salinity-exposed (170 mM NaCl) seedlings were not statistically different (P < 0.05) from those of control samples, the inhibition of plant growth approached statistical significance, with P values of 0.080 and 0.079 for the seedling weight and length, respectively.
Antioxidant Enzyme Activities
After exposure for 15 days, the average antioxidant enzyme activities were elevated in the aboveground parts of seedlings from both groups of salinity-exposed S. salsa samples (Table 3). Except POD, all other antioxidant enzymes, including SOD, GST, CAT, and GPx, indicated clear salinity-dependent trends, which meant that oxidative stresses were induced by salinities in S. salsa. Among these antioxidant enzymes, GST activities showed no statistical significance from either low (170 mM NaCl) or high (500 mM NaCl) salinity-exposed samples.
Metabolic Responses
Figure 1 presents the representative 1H-NMR spectra of tissue extracts from both root and homogeneous aboveground parts from the control group. Several metabolite classes were observed in both root and aboveground tissues, including amino acids (branched chain amino acids [valine, leucine, and isoleucine], alanine, glutamate, aspartate, and tyrosine), energy storage compounds (sucrose, fructose, and glucose), organic osmolytes (betaine), and intermediates in the tricarboxylic acid cycle (succinate, citrate, and malate). Overall, each NMR spectrum was dominated by an organic osmolyte, betaine (3.25 and 3.91 ppm), that is a secondary metabolite that maintains the osmotic balance in S. salsa and was approximately 10–100 times higher than other metabolites in the NMR spectral intensities.
Representative one-dimensional 500-MHz 1H-NMR spectra of tissue extracts from the aboveground part of seedlings (a) and roots (b) of S. salsa using the extraction solvent system of methanol/water (1/1). Metabolite assignments were based on the certain chemical shift(s) of corresponding proton(s). (1) Branched chain amino acids: leucine, isoleucine, and valine, (2) ethanol, (3) lactate, (4) alanine, (5) arginine, (6) glutamate, (7) glutamine, (8) succinate, (9) proline, (10) malate, (11) unknown (2.97 ppm), (12) malonate, (13) choline, (14) phosphocholine, (15) betaine, (16) fructose, (17) sucrose, (18) β-glucose, (19) α-glucose, (20) allantoin, (21) uridine, (22) fumarate, (23) tyrosine, (24) histidine, (25) phenylalanine, (26) tryptophan, (27) hypoxanthine, (28) formate, (29) citrate, (30) γ-aminobutyric acid, and (31) acetate
PCA was conducted on the 1H-NMR spectral data sets from aboveground and root tissue extracts of S. salsa after salinity exposure for 15 days, respectively (Figs. 2, 3). For the aboveground tissue extracts, the control (inverted triangles), low (170 mM NaCl, circles), and high (500 mM NaCl, squares) salinity-exposed samples were significantly (P < 0.05) separated along the PC1 axis from the PC score plots (Fig. 2a). From the corresponding PC1 loading plot, the metabolic profiles of aboveground extracts showed clear increases in betaine and allantoin and decreases in branched chain amino acids (valine, leucine, and isoleucine), lactate, arginine, glutamate, malate, choline, phosphocholine, glucose, fumarate, tyrosine, and phenylalanine in both salinity-exposed samples (Fig. 2b).
PCA scores plot (a) and PC loadings plot (b) from the analysis of the one-dimensional 1H-NMR spectra of tissue extracts from the aboveground part of seedlings of S. salsa from control (triangle), 170 mM NaCl-treated (circle), and 500 mM NaCl-treated (square) groups after exposure for 15 days. Ellipses represented mean ± SD of PC scores along both PC axes for each group (n = 6). Metabolite assignments: (1) branched chain amino acids: leucine, isoleucine, and valine, (2) lactate, (3) arginine, (4) glutamate, (5) glutamine, (6) malate, (7) unknown (2.97 ppm), (8) choline, (9) phosphocholine, (10) betaine, (11) α-glucose, (12) allantoin, (13) fumarate, (14) tyrosine, and (15) phenylalanine
PCA scores plot (A) and PC1 loadings plot (B) from the analysis of the one-dimensional 1H-NMR spectra of tissue extracts from roots of Suaeda salsa from control (triangle), 170 mM NaCl-treated (circle), and 500 mM NaCl-treated (square) groups after exposure for 15 days. Ellipses represented mean ± SD of PC scores along both PC axes for each group (n = 6). Metabolite assignments: (1) branched chain amino acids: leucine, isoleucine, and valine, (2) lactate, (3) proline, (4) glutamine, (5) citrate, (6) unknown (2.97 ppm), (7) choline, (8) phosphocholine, (9) betaine, (10) sucrose, (11) fructose, (12) β-glucose, (13) α-glucose, and (14) phenylalanine
PCA results from the analysis of NMR spectral data of root extracts showed clear separations (P < 0.05) between control and salinity-exposed groups along the PC1 axis from the PC score plots (Fig. 3a). However, the metabolic responses induced by salinities in roots were different from those in aboveground tissues (Fig. 3b). Besides the decreases in branched chain amino acids, lactate, choline, phosphocholine, and phenylalanine, increases in proline, citrate, glucose, and sucrose and a decrease in fructose were uniquely observed in root tissues.
Gene Expression Quantification
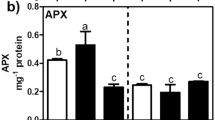
High soil salinity can induce both osmotic and oxidative stresses in plants, including even halophytes (Askaril and others 2006; Song and others 2008; Chen and others 2009). Hereby, the expression levels of three key osmotic regulation-involved genes (INPS, CMO, and BADH) and two antioxidant enzyme genes (CAT and GPx) were quantified using the RT-PCR technique. After salinity exposure for 15 days, the average expression levels of these five genes were upregulated in S. salsa samples (Fig. 4). However, both CMO and BADH gene expressions exhibited significant (P < 0.05) upregulation in the high- (500 mM NaCl) salinity-exposed group. The expression level of the INPS gene was significantly (P < 0.05) increased in low- (170 mM NaCl) salinity-exposed samples. For the antioxidant enzyme genes, only CAT expression levels were significantly (P < 0.01) elevated in both salinity-exposed groups.
Expression levels of osmotic-related enzyme genes INPS, CMO, and BADH (a) and antioxidant enzyme genes CAT and GPx (b) in the aboveground part of seedlings of S. salsa from control, 170 and 500 mM NaCl-treated groups after exposure for 15 days. Data (n = 6) were expressed as mean ± SD, significant difference among groups was tested by one-way analysis of variance with Tukey’s test and indicated by *P < 0.05)
Discussion
As a pioneer halophyte in the saline soil of the Yellow River Delta, S. salsa grows rapidly in moderate-salinity soil and can even survive at extreme salinities (Zhao 1991). Due to its high tolerance to salinity, S. salsa has been widely used as a remediation plant in the degraded wetland to ameliorate the saline soil in the Yellow River Delta. At a density of 15 plants m−2, S. salsa can potentially remove more than 3,000 kg Na+ ha−1 from saline soil, which suggests that it can be used to improve the quality of saline soils (Zhao 1991). Therefore, S. salsa is not only consumed as food, it has been intriguing to researchers in environmental sciences and applied across multiple areas (Zhang and others 2001; Cui and others 2008; Song and others 2008).
High soil salinity can inhibit the growth of various plants, including both nonhalophytes and halophytes (Song and others 2008; Chen and others 2009). This work found salinity-dependent inhibition of the growth of S. salsa (Table 2). In addition, elevation of antioxidant enzyme activities (SOD, POD, CAT, and GPx) indicated obvious oxidative stresses induced in S. salsa by salinity (Table 3). Based on the growth parameters and antioxidant enzyme activities, the two salinities used that reflected environmental conditions could clearly induce physiological and biochemical effects in S. salsa. Therefore, NMR-based metabolomics was used for the detection of metabolic responses of S. salsa induced by salinity.
All the 1H-NMR spectra of both root and aboveground seedling tissues were dominated by betaine, which was approximately 10–100 times higher than other metabolites in the NMR spectral intensities (Fig. 1). As an osmolyte, betaine can be synthesized by cells for protection against osmotic stresses such as drought, high salinity, and high temperature in biological systems, including both plants and animals (marine invertebrates in particular) (Moghaieb and others 2004; Liu and others 2011c; Zhang and others 2011b, c). The pathway of betaine synthesis in plants is short and straightforward: choline monooxygenase (CMO) converts choline (a detectable metabolite in S. salsa, Fig. 1) to betaine aldehyde, and betaine aldehyde dehydrogenase (BADH) converts this product to betaine (Greenway and Osmond 1972; Lee and others 2004; Peel and others 2010). In higher plants, betaine is an important secondary metabolite of alkaloid for protection against osmotic stresses (Greenway and Osmond 1972; Lee and others 2004). Therefore, betaine plays important physiological roles in osmotic regulation and hence was detected at high levels in S. salsa tissues.
From the PCA score plot, clear separations between control and salinity-treated groups were found along the PC1 axis (Figs. 2a, 3a). Usually, there were two kinds of variance in the S. salsa samples: biological variance and technical variance. The biological variance includes both individual variance and salinity-induced variance. Ideally, the average individual variance should be approximately zero because all the S. salsa seeds were randomly collected from thousands of seeds. The technical variance was introduced in the processing procedures of the biological samples by operators and instruments. However, all the biological samples were homogeneously processed by the same operators and instruments, and the technical variance of the samples was similar and not significant. Therefore, the significant variance between the groups of S. salsa samples was induced by salinities, which is seen in the significant separations in Figs. 2a and 3a and the corresponding significant metabolites in Figs. 2b and 3b. The statistical significances between control and salinity-treated groups were all less than 0.05, which demonstrated the significant metabolic changes in salinity-treated S. salsa. For the aboveground parts of S. salsa seedlings, the organic osmolyte betaine was significantly elevated after 170 and 500 mM NaCl treatments for 15 days. Because salinity could induce osmotic stress in plants, it was reasonable that S. salsa accumulated high concentrations of betaine to maintain osmotic balance. Allantoin is the metabolite of hypoxanthine and xanthine, which are catabolic intermediates of pyrimidines and purines (Zhang and others 2011a). Thus, the elevation of allantoin could be related to DNA and RNA biosynthesis/degradation (Zhang and others 2011a). Amino acids, including valine, leucine, isoleucine, glutamate, tyrosine, and phenylalanine, were decreased in the aboveground parts of S. salsa seedlings with salinity exposures. Interestingly, the total protein content in the aboveground parts of S. salsa seedlings was decreased with exposure to increasing salinity (Table 3). Therefore, it could be concluded that both protein biosynthesis and degradation were inhibited by salinity treatments. Malate and fumarate are key intermediates in the tricarboxylic acid (TCA) cycle. The changes in these metabolites indicated salinity-induced disturbances in the TCA cycle that were related to energy metabolism. Choline is the metabolite in betaine synthesis, as mentioned above. In this work, the decrease of choline meant the enhancement of betaine synthesis, which was confirmed by the elevation of betaine in the aboveground part of S. salsa seedlings. Consequently, as the derivative of choline, phosphocholine decreased consistently with the decreased choline. The decrease of glucose in salinity-treated plants after 15 days of exposure indicated the rapid utilization of this compound for the production of other compounds, or a lower production of carbohydrates (Choi and others 2004).
In the S. salsa root tissues that were exposed to salinity treatments, the metabolic responses were different from those in the aboveground part of S. salsa seedlings, in addition to the decreased branched chain amino acids, lactate, choline, and phosphocholine. The increased proline, citrate, and sucrose and decreased glutamine and fructose were uniquely discovered in root tissues. Other metabolites such as malate, fumarate, and allantoin were not significantly altered in root tissues. These salinity-induced metabolic differences between roots and aboveground parts of S. salsa seedlings indicated that there are different regulating mechanisms in various tissues to salinity exposure. The elevation of proline is a common metabolic response of higher plants to water deficit and salinity stress for protecting plant cell membranes and proteins and functioning as a ROS scavenger (Xiao and others 2008; Dai and others 2010). On the other hand, proline is a compatible solute that can be accumulated in plants in response to osmotic stress (Nelson and others 1998). In this work, the increased proline could be considered a metabolic biomarker of salinity exposure in S. salsa root. The increase in citrate, a key intermediate in the TCA cycle, indicated a disturbance in energy metabolism. In plants, sucrose can be commonly converted to glucose and fructose. The decreased glucose and fructose and increased sucrose were discovered in salinity-treated samples herein, indicating the inhibition of conversion of sucrose to glucose and fructose induced by salinity exposures.
Based on the metabolic responses and antioxidant enzyme activities, salinity treatments could induce osmotic and oxidative stresses in S. salsa. To determine whether salinity-induced stress was able to regulate the expression of typical genes responsible for osmotic regulation and antioxidation, the RT-PCR-based assay was performed for the quantification of osmotic regulation-involved genes (INPS, CMO, and BADH) and two antioxidant enzyme genes (CAT and GPx). INPS, CMO, and BADH are involved in the biosynthesis of organic osmolytes, myo-inositol and betaine, which are used to balance osmolarity in S. salsa (Zhang and others 2001). Myo-inositol is biosynthesized by myo-inositol-1-phosphate synthase (INPS, BE644574). As reported by Nelson and others 1998, INPS regulates the myo-inositol biosynthesis pathway (Nelson and others 1998), which is a specific pathway in salt tolerance in the common ice plant (Loewus and Dickinson 1982; Vernon and Bohnert 1992), and its expression can be feedback-inhibited by myo-inositol. In higher plants, betaine is synthesized by oxidation of choline: choline → betaine aldehyde → betaine. The first step is catalyzed by choline monooxygenase (CMO, AW991015) and the second step is catalyzed by betaine aldehyde dehydrogenase (BADH, DQ641924). CAT (AW990998) and GPx (AW991114) are common antioxidant enzymes that protect cells from oxidative stresses. Our analysis showed that expressions of the genes INPS, CMO, BADH, CAT, and GPx were upregulated under both 170 and 500 mM NaCl exposures (Fig. 4), which confirmed the osmotic and oxidative stresses in S. salsa induced by salinities. For CAT and GPx, however, the gene expression levels were not correlated with the corresponding activities in S. salsa samples. The lack of correlation between gene expression and enzyme activity is not surprising, because gene expression is characterized by only one isoenzyme of one gene family, whereas enzyme measurements typically include all expressed members of such a family. Moreover, enzyme activities can be affected by a number of feedback regulations, so that a good correlation is not always found. In addition, because the concentrations of betaine in S. salsa samples were difficult to accurately quantify using NMR-based metabolomics, good correlation between betaine contents and CMO or BADH expression levels was not found.
In conclusion, we analyzed the metabolic profiles, antioxidant enzyme activities, and gene expressions of S. salsa that was exposed to environmentally relevant salinity levels (170 and 500 mM NaCl) for 15 days. The significant metabolic responses included decreased amino acids, choline, and phosphocholine and increased betaine and allantoin in aboveground parts of S. salsa seedlings. The root tissues of S. salsa indicated uniquely elevated glutamate, citrate, and sucrose and depleted glucose and fructose. The upregulated expression levels of INPS, CMO, BADH, and CAT were found in the aboveground part of S. salsa seedlings after salinity exposures, together with increased activities of antioxidant enzymes, including SOD, GST, POD, CAT, and GPx. Overall, these results indicated osmotic and oxidative stresses, disturbed protein biosynthesis/degradation, and energy metabolism in S. salsa exposed to salinities. A further proteomic study should be conducted to better understand the salt-responsive mechanisms in S. salsa.
References
Aliferis KA, Materzok S, Paziotou GN, Chrysayi-Tokousbalides M (2009) Lemna minor L. as a model organism for ecotoxicological studies performing 1H NMR fingerprinting. Chemosphere 76:967–973
Alscher RG, Erturk N, Heath LS (2002) Role of superoxide dismutases (SODs) in controlling oxidative stress in plants. J Exp Bot 53:1331–1341
Askaril H, Edqvist J, Hajheidaril M, Kafi M, Salekdeh GH (2006) Effects of salinity levels on proteome of Suaeda aegyptiaca leaves. Proteomics 6:2542–2554
Borsani O, Valpuesta V, Botella MA (2001) Evidence for a role of salicylic acid in the oxidative damage generated by NaCl and osmotic stress in Arabidopsis Seedlings. Plant Physiol 126:1024–1030
Bradford M (1976) A rapid and sensitive method for the quantification of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Bundy JG, Davey MP, Viant MR (2009) Environmental metabolomics: a critical review and future perspectives. Metabolomics 5:3–21
Chen H, Jiang JG, Wu GH (2009) Effects of salinity changes on the growth of Dunaliella salina and its isozyme activities of glycerol-3-phosphate dehydrogenase. J Agric Food Chem 57:6178–6182
Choi HK, Choi YH, Verberne MC, Lefeber AWM, Erkelens C, Verpoorte R (2004) Metabolic fingerprinting of wild type and transgenic tobacco plants by 1HNMR and multivariate analysis technique. Phytochemistry 65:857–864
Cui BS, He Q, Zhao XS (2008) Ecological thresholds of Suaeda salsa to the environmental gradients of water table depth and soil salinity. Acta Ecol Sin 28:1408–1418
Dai H, Xiao C, Liu H, Hao F, Tang H (2010) Combined NMR and LC-DAD-MS analysis reveals comprehensive metabonomic variations for three phenotypic cultivars of Salvia Miltiorrhiza Bunge. J Proteome Res 9:1565–1578
Davis B (2005) Growing pains for metabolomics. The Scientist 19:25–28
Dendooven L, Alcántara-Hernández RJ, Valenzuela-Encinas C, Luna-Guido M, Perez-Guevara F, Marsch R (2010) Dynamics of carbon and nitrogen in an extreme alkaline saline soil: a review. Soil Biol Biochem 42:865–877
Fan WMT (1996) Metabolite profiling by one- and two-dimensional NMR analysis of complex mixtures. Prog Nucl Magn Reson Spectrosc 28:161–219
Fiehn O (2002) Metabolomics: the link between genotypes and phenotypes. Plant Mol Biol 48:155–171
Greenway H, Osmond CB (1972) Salt responses of enzymes from species differing in salt tolerance. Plant Physiol 49:256–259
Katsiadaki I, Williams TD, Ball JS, Bean TP, Sanders MB, Wu H, Santos EM, Brown MM, Baker P, Ortega F, Falciani F, Craft JA, Tyler CR, Viant MR, Chipman JK (2009) Hepatic transcriptomic and metabolomic responses in the Stickleback (Gasterosteus aculeatus) exposed to ethinyl-estradiol. Aquat Toxicol 97:174–187
Kim HK, Verpoorte R (2010) Sample preparation for plant metabolomics. Phytochem Anal 21:4–13
Lee MB, Blunt JW, Lever M, Georgea PM (2004) A nuclear-magnetic-resonance-based assay for betaine–homocysteine methyltransferase activity. Anal Biochem 330:199–205
Li W, Wang D, Jin T, Chang Q, Yin D, Xu S, Liu B, Liu L (2011) The vacuolar Na+/H+ antiporter gene SsNHX1 from the halophyte Salsola soda confers salt tolerance in transgenic alfalfa (Medicago sativa L.). Plant Mol Biol Rep 29:278–290
Lindon JC, Nicholson JK, Everett JR (1999) NMR spectroscopy of biofluid. Ann Rep NMR Spectrosc 38:1–88
Liu X, Yang C, Zhang L, Li L, Liu S, Yu J, You L, Zhou D, Xia C, Zhao J, Wu H (2011a) Metabolic profiling of cadmium-induced effects in one pioneer intertidal halophyte Suaeda salsa by NMR-based metabolomics. Ecotoxicology 20:1422–1432
Liu X, Zhang L, You L, Wu H, Zhao J, Cong M, Li F, Wang Q, Li L, Li C, Han G, Wang G, Xia C, Yu J (2011b) Metabolomic study on the halophyte Suaeda salsa in the Yellow River Delta. CLEAN Soil Air Water 39:720–727
Liu X, Zhang L, You L, Yu J, Zhao J, Li L, Wang Q, Li F, Liu D, Wu H (2011c) Differential toxicological effects induced by mercury in gills from three pedigrees of Manila clam Ruditapes philippinarum by NMR-based metabolomics. Ecotoxicology 20:177–186
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25:402–408
Loewus FA, Dickinson MW (1982) Cyclitols. In: Loewus FA, Tanner W (eds) Plant carbohydrates. Springer, Berlin, pp 193–216
Maggio A, Barbieri G, Raimondi G, De Pascale S (2010) Contrasting effects of GA3 treatments on tomato plants exposed to increasing salinity. J Plant Growth Regul 29:63–72
Moghaieb REA, Saneoka H, Fujita K (2004) Effect of salinity on osmotic adjustment, glycine betaine accumulation and the betaine aldehyde dehydrogenase gene expression in two halophytic plants, Salicornia europaea and Suaeda maritime. Plant Sci 166:1345–1349
Munns R (2002) Comparative physiology of salt and water stress. Plant Cell Environ 25:239–250
Munns R (2005) Genes and salt tolerance: bringing them together. New Phytol 167:645–663
Nelson DE, Rammesmayer G, Bohnert HJ (1998) Regulation of cell-specific insitol metabolism and transport in plant salt tolerance. Plant Cell 10:753–764
Owojori OJ, Reinecke AJ, Rozanov AB (2008) Effects of salinity on partitioning, uptake and toxicity of zinc in the earthworm Eisenia fetida. Soil Biol Biochem 40:2385–2393
Parida AK, Jha B (2010) Antioxidative defense potential to salinity in the euhalophyte Salicornia brachiata. J Plant Growth Regul 29:137–148
Parsons HM, Ludwig C, Gunther UL, Viant MR (2007) Improved classification accuracy in 1- and 2-dimensional NMR metabolomics data using the variance stabilising generalised logarithm transformation. BMC Bioinformatics 8:234
Pedras MSC, Zheng QA (2010) Metabolic responses of Thellungiella halophila/salsuginea to biotic and abiotic stresses: metabolite profiles and quantitative analyses. Phytochemistry 71:581–589
Peel GJ, Mickelbart MV, Rhodes D (2010) Choline metabolism in glycine betaine accumulating and non-accumulating near-isogenic lines of Zea mays and Sorghum bicolor. Phytochemistry 71:404–414
Shao G, Chen M, Wang W, Zhang G (2008) The effect of salinity pretreatment on Cd accumulation and Cd-induced stress in BADH-transgenic and nontransgenic rice seedlings. J Plant Growth Regul 27:205–210
Song J, Fan H, Zhao YY, Jia YH, Du XH, Wang BS (2008) Effect of salinity on germination, seedling emergence, seedling growth and ion accumulation of a euhalophyte Suaeda salsa in an intertidal zone and on saline inland. Aquat Bot 88:331–337
Sonjak S, Udovic M, Wraber T, Likar M, Marjana Regvar M (2009) Diversity of halophytes and identification of arbuscular mycorrhizal fungi colonising their roots in an abandoned and sustained part of Secovlje salterns. Soil Biol Biochem 41:1847–1856
Sun X, Zhang J, Zhang H, Ni Y, Zhang Q, Chen J, Guan Y (2010) The responses of Arabidopsis thaliana to cadmium exposure explored via metabolite profiling. Chemosphere 78:840–845
Tsonev TD, Lazova GN, Stoinova ZG, Popova LP (1998) A possible role for jasmonic acid in adaptation of barley seedlings to salinity stress. J Plant Growth Regul 17:153–159
Tusher VG, Tibshirani R, Chu G (2001) Significance analysis of microarrays applied to the ionizing radiation response. Proc Nat Acad Sci USA 98:5116–5121
Vernon DM, Bohnert HJ (1992) A novel methyl transferase induced by osmotic stress in the facultative halophyte Mesembryanthemum crystallinum. EMBO J 11:2077–2086
Wang CQ, Chen M, Wang BS (2007) Betacyanin accumulation in the leaves of C3 halophyte Suaeda salsa L. is induced by watering roots with H2O2. Plant Sci 172:1–7
Xiao C, Dai H, Liu H, Wang Y, Tang HR (2008) Revealing the metabonomic variation of rosemary extracts using 1H NMR spectroscopy and multivariate data analysis. J Agric Food Chem 56:10142–10153
Xu L (2004) Methods of chemometrics. Science Press, Beijing, pp 221–227
Yang G, Zhou R, Tang T, Chen X, Ouyang J, He L, Li W, Chen S, Guo M, Li X, Zhong C, Shi S (2011) Gene expression profiles in response to salt stress in Hibiscus tiliaceus. Plant Mol Biol Rep 29:609–617
Zhang L, Ma XL, Zhang Q, Ma CL, Wang PP, Sun YF, Zhao YX, Zhang H (2001) Expressed sequence tags from a NaCl-treated Suaeda salsa cDNA library. Gene 267:193–200
Zhang JY, Li MH, Xu LM, Wang ZJ (2008) Effect of Suaeda seed oil on blood-fat and immunologic function of mouse. Occup Health 24:1529–1530 (in Chinese)
Zhang JT, Zhang Y, Du YY, Chen SY, Tang HR (2011a) Dynamic metabonomic responses of tobacco (Nicotiana tabacum) plants to salt stress. J Proteome Res 10:1904–1914
Zhang L, Liu X, You L, Zhou D, Wang Q, Li F, Cong M, Li L, Zhao J, Liu D, Yu J, Wu H (2011b) Benzo(a)pyrene-induced metabolic responses in Manila clam Ruditapes philippinarum by proton nuclear magnetic resonance (1H NMR) based metabolomics. Environ Toxicol Pharmacol 32:218–225
Zhang L, Liu X, You L, Zhou D, Wu H, Li L, Zhao J, Feng J, Yu J (2011c) Metabolic responses in gills of Manila clam Ruditapes philippinarum exposed to copper using NMR-based metabolomics. Mar Environ Res 72:33–39
Zhao KF (1991) Desalinization of saline soils by Suaeda salsa. Plant Soil 135:303–305
Zhao HL, Ma YZ, Li JX, Piao L (2010) Study on edible value of Suaeda salsa. J Anhui Agric Sci 38:14350–14351 [in Chinese]
Acknowledgments
We thank Dr. Mark Viant (School of Bioscience, The University of Birmingham) for use of the software ProMetab. This research was supported by the Project of National Science & Technology Pillar Program in “12th Five Year” Period (2011BAC02B01), The 100 Talents Program of the Chinese Academy of Sciences, Innovation Programs of the Chinese Academy of Sciences (KZCX2-YW-223 and KZCX2-YW-225), and Technology Development Program Projects of Shandong Province (2008GG20005006 and 2008GG3NS0700), and in part by the CAS/SAFEA International Partnership Program for Creative Research Teams.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wu, H., Liu, X., You, L. et al. Effects of Salinity on Metabolic Profiles, Gene Expressions, and Antioxidant Enzymes in Halophyte Suaeda salsa . J Plant Growth Regul 31, 332–341 (2012). https://doi.org/10.1007/s00344-011-9244-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00344-011-9244-6