Abstract
Freesia hybrida is an important worldwide cut flower, especially in America and Europe. For efficient regeneration of this flower from young inflorescence and rachillae in tetraploid, we developed a simple in vitro micropropagation protocol. Explants of Freesia hybrida can regenerate plantlets through somatic embryogenesis via two kinds of pathways, that is, directly from the epidermal cells or indirectly from an embryonic callus, depending on the exogenous plant growth regulators (PGRs) used in the culture media. In direct embryogenesis, when the explants were cultured on Murashige and Skoog (MS) medium supplemented with 11.43 μM indole acetic acid (IAA) and 4.44 μM 6-benzylaminopurine (6-BA), the induction rate was 84% for young inflorescence and 100% for rachillae. After the multishoots were subcultured on the rooting MS medium containing 1.08 μM α-naphthalene acetic acid (NAA), the rooting rate was close to 100%. In indirect embryogenesis, embryonic calluses were formed when the culture medium contained 22.22 μM 6-BA and 4.52 μM 2,4-dichlorophenoxy acetic acid (2,4-D), and the induction rate was 92.4% for young inflorescence and 100% for rachillae. After the embryonic calluses were transferred to the medium supplemented with 11.43 μM IAA and 13.33 μM 6-BA, they could develop into plantlets with roots. In assessing the two regeneration pathways in terms of genetic and epigenetic fidelity of the regenerants, two kinds of molecular markers [amplified fragment length polymorphism (AFLP) and methylation-sensitive amplified polymorphism (MSAP)] were employed. The AFLP analysis used 20 primer pairs that yielded 916 scorable bands among the donor plant and 11 regenerants from direct embryogenesis, of which 8 (0.87%) were polymorphic. The regenerants from indirect embryogenesis had 1075 clear bands of which 3 (0.27%) were polymorphic scorable bands from 18 primer pairs. Moreover, the variant band patterns included two types, that is, loss-of-original and gain-of-novel bands. MSAP analysis revealed that tissue culturing of the flower induced DNA cytosine methylation alterations in both CG and CNG levels and patterns compared with the donor plant. The variation rate was 1.1 and 1.3% for the direct and indirect embryogenesis pathways, respectively. The findings show that tissue culture of flowering plants is a form of stress which can induce some heritable epigenetic variations and should be considered in future long-term genotype preservation programs of Freesia hybrida.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Freesia species has been grown widely as a cut flower and its popularity is increasing. Current market research shows that over 110 million Freesia stems are sold in the UK each year (http://www.flowers.org.uk/public/flower_search_by_name.php). Studies show that it originated from the southern region of Africa and was first imported to Europe at the end of the 19th century. Now it has become one of the most popular flowers in the world. Somatic embryogenesis has been induced in the tissue culture of diploid Freesia refracta either directly from the epidermal cells of young leaves and inflorescences or indirectly via intervening callus (Wang and others 1990). However, tissue culture in the tetraploid Freesia hybrida, which accounts for the largest proportion of the Freesia cut flowers in the tissue culture market, has not been reported so far.
Knowledge of the optimal conditions favoring differentiation under in vitro conditions is an important step in the application of in vitro manipulation techniques in higher plants. Successful plant regeneration depends on factors such as the genotype, explant type, age of the donor plants, the number of subcultures (Jain 1998; Veilleux and Johnson 1998), and the composition of medium, especially plant growth regulators (PGRs). Two major PGRs are auxin and cytokinin (Skoog and Miller 1957).
In vitro clonal propagation techniques provide the ability to efficiently multiply and maintain large numbers of elite genotypes. However, tissue culture-induced phenotypic and genotypic variations, collectively termed “somaclonal variation,” are commonly observed in plants and are at least partly due to in vitro-induced stress (Larkin and Scowcroft 1981; Evans and others 1984). The molecular basis of somaclonal variation is not precisely known; however, both genetic and epigenetic mechanisms are thought to play a role. Somaclonal variation may arise as a result of point mutations, rearrangements in nuclear or organellar DNA (Phillips and others 1994), the activation of mobile elements (Kidwell and Lisch 1997; Kazazian 2004; Liu and others 2004), ploidy (Sunderland 1977; Bayliss 1980; Creissen and Karp 1985), or epigenetic changes (Jaligot and others 2000; Kumar and Mathur 2004; Gostimsky and others 2005; Kuznetsova and others 2005) causing deviations from a desired phenotype quality or standard. In principle, somaclonal variation is defined primarily as epigenetic change that alters gene expression patterns without changes in the DNA sequence (Russo and others 1996; Kaeppler and others 2000). One chief contributor implicated in epigenetics is DNA methylation. In higher plants, cytosine is primarily methylated in a CG dinucleotide context and CNG sites (Gruenbaum and others 1981) and, less abundantly, in CNN sequences (Tariq and Paszkowski 2004).
In this study, the amplified fragment length polymorphism (AFLP) technique was used to investigate genetic changes induced by tissue culture. The AFLP method is based on the principle of selectively amplifying a subset of restriction fragments from a complex mixture of DNA fragments obtained after digestion of genomic DNA with restriction endonuclease (Vos and others 1995). To determine the extent of DNA methylation changes and the presence of methylated CCGG sites, the methylation-sensitive amplified polymorphism (MSAP) technique (Xiong and others 1999) was used; it is a modification of the AFLP method that makes use of the differential sensitivity of a pair of isoschizomers, MspI and HpaII, to cytosine methylation. The objectives of this study were (1) to develop a highly efficient and relatively stable tissue culture protocol for routine plant regeneration in the tetraploid Freesia hybrida and (2) to assess the occurrence and extent of genetic and epigenetic instabilities induced by tissue culture using AFLP and MSAP molecular markers so as to be incorporated in future long-term genotype preservation programs of Freesia hybrida.
Materials and Methods
Tissue Culture
Two kinds of Freesia cultivars were used for tissue culture, red Freesia hybrida were for direct embryogenesis and yellow Freesia hybrida were for indirect embryogenesis. Young inflorescences with rachillae were obtained from greenhouse-grown plants and were surface sterilized with 70% ethanol followed by immersion in 0.1% mercuric chloride for about 4 min with frequent agitation, and then rinsed five times successively using sterile distilled water. Before being cultivated onto the solid MS medium, the inflorescences and rachillae were dissected and cut into 2-3-mm-thick segments by removal of surrounding leaves. The medium was supplemented with different concentrations of plant growth regulators (PGRs), 2.5% sucrose, and 0.6% agar, and the pH was adjusted to 5.8. All cultures were incubated in growth chambers at 24 ± 3°C, with a 12-h photoperiod and a photon flux density of 1600 μE m−2 s−1. Subculture was under taken every 5 weeks.
Plantlets regenerated from both direct embryogenesis medium without inducing roots by NAA and indirect embryogenesis medium were used for AFLP and MSAP analysis.
AFLP and MSAP Analysis
Genomic DNA was extracted from expanded leaves of the donor plants and regenerated plants using a modified cetyl trimethyl ammonium bromide (CTAB) method. A standard AFLP protocol with minor modifications was followed (Wang and others 2005). One preselective and 23 selective primer combinations were used (Table 1). MSAP was performed using HpaII and MspI, a pair of isoschizomers that recognize the same restriction site (5′-CCGG) but have different sensitivity to methylation of the cytosines. Specifically, HpaII will not cut if either of the cytosines is fully methylated (double-strand); in contrast, MspI will not cut only if the external cytosine is fully or hemimethylated (McClelland and others 1994). Thus, for a given DNA sample, two major methylation states at the CCGG sites, (1) full methylation of the internal C, which are bands absent from HpaII but present in the corresponding MspI digest, that is, pattern H/M = −/+, and (2) hemimethylation of the external C, which are bands present in HpaII but absent from the corresponding MspI digest, that is, pattern H/M = +/−, will be readily recognized in the MSAP profiles (Reyna-Lopez and others 1997; Cervera and others 2002). Nonetheless, full methylation of the external cytosine or both cytosines indigested by both HpaII and MspI are invisible; thus, the level of cytosine is lower than reality. One pair of preselective and 18 pairs of selective primers were used (Table 1). The restriction enzymes EcoRI, MseI, HpaII, and MspI and T4 ligase were purchased from New England Biolabs Inc. (Ipswich, MA; http://www.neb.com). The amplification products of AFLP or MSAP were resolved by denaturing polyacrylamide gel electrophoresis (5% acrylamide) and visualized by silver staining. Only clear and completely reproducible bands were scored.
Data Analysis
The AFLP or MSAP bands were scored as a binary character for absence (0) or presence (1). To rule out confounding effects of nucleotide sequence change at the CCGG sites in the computed MSAP data, the method described by Cervera and others (2002) was used.
Results
In Vitro Micropropagation of Freesia hybrida
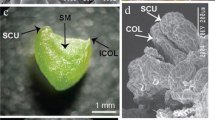
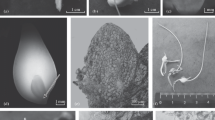
Explants of Freesia hybrida can regenerate plantlets through somatic embryogenesis via two kinds of pathways, that is, directly from the epidermal cells or indirectly from an embryonic callus stage, depending on the exogenous PGRs used in the culture media. We found that segments of young inflorescences and rachillae were responsive for embryoids then developing into multishoots and embryonic calluses. During the direct somatic embryogenesis pathway, globular embryoids (Fig. 1a) emerged at the periphery of explants after 7-10 days in culture on the MS medium containing IAA and 6-BA combinations at different concentrations. After 50 days, multishoots appeared (Fig. 1b). Plantlets with roots (Fig. 1c) were regenerated 30 days after transferring the multishoots onto the MS basal medium containing only 1.08 μM NAA; the rooting rate was close to 100%. When the same explants were cultivated on the medium supplemented with 6-BA and 2,4-D combinations, big masses of pale yellow embryonic calli (Fig. 1d) were induced after 25-30 days in culture; they developed into single or multiple shoots (Fig. 1e) about 30 days after subculturing on the differentiation medium containing 11.43 μM IAA and 13.33 μM 6-BA. Multiple plantlets (Fig. 1f) bearing roots were obtained after one more month in culture.
Table 2 shows that different types and concentrations of PRG combinations may induce different regeneration pathways and the induction rate was also dissimilar. By testing a series of embryoids and callogenic media, we found that three IAA and 6-BA combinations resulted in direct embryogenesis. Indeed, when the IAA:6-BA value was 11.43:4.44 μM, the induction rate was the highest: 84% for young inflorescences and 100% for young rachillae. As for indirect embryogenesis, MS medium supplemented with 6-BA and 2,4-D was found to be more suitable, with the highest induction rate of 92.4% for young inflorescences and 100% for young rachillae. Obviously, the use of young rachis as explants was more efficient than inflorescences in causing induction.
Assessment of Genetic and Epigenetic Instabilities in Regenerants of Freesia hybrida by Two Different Types of Molecular Markers
AFLP Analysis
Twenty-three primer pairs for selective amplification were selected among 45 combinations (9 MseI + 3 primers combined with 5 EcoRI + 3 primers, see Table 1) for AFLP analysis.
Regenerants from Direct Embryogenesis
Twenty primer pairs yielded 916 scorable bands when the donor plant and 11 regenerants from direct embryogenesis were analyzed (Table 3), of which 8 (0.87%) were polymorphic. The average number of bands generated by each primer pair was 45.08 (range = 22-70), with an average of 0.4 polymorphic bands (range = 0–2). The polymorphic bands included only gain-of-novel bands (Fig. 2) in the regenerants, and three of the eight novel bands were singletons, that is, variant band(s) present or absent in a single regenerant for a given primer pair.
Regenerants from Indirect Embryogenesis
When regenerants from indirect embryogenesis were analyzed, 1075 with 3 (0.27%) polymorphic scorable bands were obtained using 18 primer pairs (Table 4), and the average number of bands was 59.72 (range = 38-83). Contrasting with the results from direct embryogenesis, the variant band patterns included two types: (1) loss-of-original bands seen in Fig. 3 and (2) gain-of-novel bands, of which two were singletons. Furthermore, we noticed that the variant bands existed only in a few of the regenerants.
MSAP Analysis
Based on the same criteria for AFLP primer selections, 18 selective amplification primer pairs for MSAP were chosen among 70 combinations (10 HpaII/MspI + 3 primers combined with 7 EcoRI + 3 primers; Table 1).
Regenerants from Direct Embryogenesis
By using 12 pairs of selected EcoRI + HpaII/MspI primer combinations, 541-544 clear and reproducible bands were amplified in DNA from the donor plant and 15 regenerants, of which 11 plant samples were used as previously described in the AFLP Analysis section. By tabulating the number of bands representing the various types of MSAP pattern, the levels of CG, CNG, and simultaneous CG/CNG methylation were calculated (Table 5). The levels of all three kinds of methylation in the 15 regenerated plantlets tended to be quite similar to the methylation level of the donor plant, though with a little reduction. We also examined the MSAP patterns in a locus-specific manner. Table 6 shows that (i) both hypo- and hypermethylation occurred in regenerated plants, and the total DNA methylation variation rate was 1.1%. (ii) The level of hypomethylation was higher than hypermethylation (0.73 vs. 0.37%). (iii) As for hypomethylation, the number of variant bands for both CNG and CG hypomethylation (3) was greater than for only CG hypomethylation alone (1).
Regenerants from Indirect Embryogenesis
Eighteen pairs of selected EcoRI + HpaII/MspI primer combinations were used for MSAP analysis. A total of 689 scorable bands were amplified from the same set of plant samples used for AFLP analysis. The number of nonmethylated, CG, CNG, and simultaneous CG/CNG methylation sites was calculated based on MSAP profiles (Table 7). The donor plant had a total methylation level of 9.8%, comprising 6.1, 2.9, and 0.8% at CG, CNG, and simultaneous CG/CNG methylation sites, respectively. Compared with the donor plant, 11 regenerated plantlets showed a similar total methylation level although it was lower (except Reg 10); the lowest was 9.2%.
MSAP profiling also allowed comparison of the cytosine methylation patterns in a locus-specific manner. By using the donor plant as a control, all scorable MSAP loci in the studied samples were classified into four major groups: CG, CNG, CG/CNG hypomethylation, and CG/CNG hypermethylation (Fig. 4). As seen in Table 8, the number of CG, CNG, CG/CNG hypomethylation, and CG/CNG hypermethylation variant bands was 2, 2, 4, and 1, respectively. Obviously, the hypomethylation level was higher than hypermethylation level, and the global methylation variation rate was 1.3%.
An example of MSAP profiles of the donor plant and 11 selected regenerants (Reg 1-11) in Freesia hybrida by primer combination EcoRI + ACG/HpaII(MspI) + TTA. E + H, the combination of EcoRI and HpaII; E + M, the combination of EcoRI and MspI. Some typical methylation and changing methylation patterns, as detailed in the text, are marked by arrowheads
Discussion
Culture systems aimed at inducing embryogenesis have been developed for many angiosperm and gymnosperm species (Brown and others 1995; Dunstan and others 1995; Krishnaraj and Vasil 1995; Thorpe and Stasolla 2001). To date, the only conclusive evidence on this subject is that a cell culture has to be grown in a medium containing a high concentration of auxin to become embryogenic. Due to its characteristics as an important cut flower in the world, especially in Europe and America, and the apparent potential held by modern biotechnology for further improvement, establishing an efficient, stable, and simple in vitro micropropagation protocol in Freesia hybrida is of interest. This study demonstrates the feasibility of regenerating plantlets from isolated young inflorescences and rachillae via both direct and indirect somatic embryogenesis. Conceivable advantages of using young inflorescences and rachillae as explants may include high totipotency of its constituent primordial cells, easier decontamination, and lower incidence of virus infection in the regenerants.
Plant growth regulators can affect the rate of somaclonal variation both directly (Stover 1987) and indirectly by increasing the multiplication rate and inducing adventitious shoots (Damasco and others 1998; Bairu and others 2006). We observed that the concentrations of 2,4-D played a critical role in inducing embryonic calli and in maintaining regeneration potential during subcultures in Freesia hybrida. In addition, IAA and 6-BA were both suitable for direct embryogenesis and transferring indirect embryogenesis to the direct pathway. Similar studies were done in E. pulcherrima (Osternack and others 1999), Dendrathema grandiflora (May and Trigiano 1991), and in several other cereal crops, including barley and maize (Bregitzer and others 1995; Bohorova and others 1995; Carvalho and others 1997).
We detected both genetic and epigenetic changes in randomly chosen regenerants of Freesia hybrida, although the possibility that such changes were due to preexisting heterozygosity or natural mutations in the explants was minimized by using explants taken from a single donor plant (Meins 1983). Therefore, it can be concluded that all scored changes were more likely to have resulted from genetic and epigenetic instabilities induced by the tissue culture stress process. Similar research has been done in callus and regenerants of maize (Brown 1989; Brown and others 1991; Kaeppler and Phillips 1993b), in progeny of regenerated rice plants (Brown and others 1990; Muller and others 1990), and in a continuously proliferating, dedifferentiated cell suspension culture of Arabidopsis (Tanurdzic and others 2008). Usually AFLP is more likely to find band losses, but minor losses were detected in our research; thus, there is the very likely possibility that this Freesia is tetraploid. In mammals, DNA methylation is normally confined to one of the alleles related to imprinting and X-chromosome inactivation (Feil 2006). When four alleles are present in Freesia, most losses of bands may not be visible as other copies of the alleles are still present.
Somaclonal variation has been documented to occur frequently in plant tissue culture but the underlying mechanism remains largely obscure (Kaeppler and others 2000; Xu and others 2004; Peredo and others 2006). The heritability of epigenetic changes can be thought of as a precursor for genetic change, especially when stress conditions are maintained. Furthermore, the epigenetic mechanisms play a fundamental role in somaclonal variation in that they may induce a broad range of genomic mutations, from single-point mutations to chromosome breakage and polyploidy (Kaeppler and others 2000). Hence, epigenetic variation is deemed to be more frequent than genetic variation. Indeed, this was the observed scenario in this study of the in vitro micropropagation protocol of Freesia hybrida in which we found methylation polymorphism to be greater than DNA polymorphism. For example, during indirect embryogenesis, the level of methylation variation was 1.3% versus 0.27% in DNA polymorphism. The same results were obtained in rice (Ashikawa 2001), Arabidopsis thaliana (Cervera and others 2002), cotton (Keyte and others 2006), and maize (Kaeppler and Phillips 1993b). On the other hand, there are several studies indicating that there are intimate correlations between frequencies of genetic changes and DNA methylation alterations (Guo and others 2006, 2007; Li and others 2007). This suggests that the genetic and epigenetic instabilities occurring under tissue culture conditions may share a common mutagenic mechanism.
The induction of somatic embryogenesis consists of the termination of a current gene expression pattern in the explant’s tissue and its replacement with an embryogenic gene expression program. It has been proposed that PGRs play a central role in mediating the signal transduction cascade leading to the reprogramming of gene expression (Arnold and others 2002). One possible mechanism for regulation of current gene expression is DNA methylation, which is influenced by auxins (LoSchiavo and others 1989; Chakrabarty and others 2003). Similar methylation alterations have also been observed in rose somatic embryos (Xu and others 2004), potato microplants (Joyce and Cassells 2002), germinating pepper seeds (Portis and others 2004), micropropagated banana (Peraza-Echeverria and others 2001; Baurens and others 2003), and oil palm (Jaligot and others 2000). Some alterations were reported to have occurred upon exposure to in vitro conditions (Phillips and others 1994) and some by wounding (Kaeppler and Phillips 1993a). The alteration in cytosine methylation level in Freesia hybrida occurred in regenerants, with a general trend of decreased methylation in the second plantlet regenerating pathway. Similar findings of a general decrease in methylation levels associated with tissue culture have been reported previously in several plants, including maize (Kaeppler and Philipps 1993b), soybean (Quemada and others 1987), oil palm (Jaligot and others 2000; Matthes and others 2001), and barley (Li and others 2007), although increased methylation was also reported in some cases (Cecchini and others 1992; Smulders and others 1995; Parra and others 2001; Xu and others 2004). Recent studies by Tanurdzic and others (2008) have shown that stress due to cell culture causes epigenetic changes in which the euchromatin becomes hypermethylated while some hypomethylation takes place in the heterochromatin indicating a search for a new genomic stability after the induced stress.
In general, the levels of cytosine methylation polymorphism and DNA polymorphism were very low in this study: 1.1 and 0.87% and 1.3 and 0.27% for direct and indirect embryogenesis, respectively. Hence, it can be concluded that the mode of regeneration does not have a significant effect on the balance between sequence and methylation state changes induced by a tissue culture process. This finding contradicts the commonly held belief that plants regenerated without a passage through a callus phase are less prone to somaclonal variation than those emerging after a period in callus (Thomas and others 1982; Young and others 1999).
In conclusion, to the best of our knowledge this is the first report on tetraploid Freesia hybrida in vitro micropropagation that comprises two efficient and simple protocols via direct and indirect somatic embryogenesis pathways. Furthermore, AFLP and MSAP analysis showed that regenerants were true-to-type of their donor plants with respect to genetic and epigenetic fidelity (Li and others 2006).
References
Arnold S, Sabala I, Bozhkov P, Dyachok J, Filonova L (2002) Developmental pathways of somatic embryogenesis. Plant Cell Tissue Organ Cult 69:233–249
Ashikawa I (2001) Surveying CpG methylation at 5′-CCGG in the genomes of rice cultivars. Plant Mol Biol 45:31–39
Bairu MW, Fennell CW, van Staden J (2006) The effect of plant growth regulators on somaclonal variation in Cavendish banana (Musa AAA cv’.Zelig’). Sci Hortic 108:347–351
Baurens FC, Bonnot F, Bienvenu D, Causse S, Legavre T (2003) Using SD-AFLP and MSAP to assess CCGG methylation in the banana genome. Plant Mol Biol Rep 21:339–348
Bayliss MW (1980) Chromosomal variation in tissue culture. Int Rev Cytol Suppl 11A:113–144
Bohorova NE, Luna B, Briton RM, Huerta LD, Hoisington DA (1995) Regeneration potential of tropical, and subtropical, midaltitude, and highland maize inbreds. Maydica 40:275–281
Bregitzer P, Campbell RD, Wu Y (1995) Plant regeneration from barley callus: effects of 2,4-dichlorphenoxy acetic acid and phenylacetic acid. Plant Cell Tissue Organ Cult 43:229–235
Brown PTH (1989) DNA methylation in plants and its role in tissue culture. Genome 31(2):717–729
Brown PTH, Kyozuka J, Sukekiyo Y, Kimura Y, Shimamoto K, Lörz H (1990) Molecular changes in protoplast-derived rice plants. Mol Gen Genet 223:324–328
Brown PTH, Gobel E, Lorz H (1991) RFLP analysis of Zea mays callus cultures and their regenerated plants. Theor Appl Genet 81:227–232
Brown DCW, Finstad KI, Watson EM (1995) Somatic embryogenesis in herbaceous species. In: Thorpe TA (ed) In vitro embryogenesis in plants. Kluwer, Dordrecht, The Netherlands, pp 345–415
Carvalho CHS, Bohorova N, Bordallo PN, Abreu LL, Valicente FH, Bressan W, Paiva E (1997) Type II callus production and plant regeneration in tropical maize genotypes. Plant Cell Rep 17:73–76
Cecchini E, Natali L, Cavallini A, Durante M (1992) DNA variations in regenerated plants of pea (Pisum sativum L.). Theor Appl Genet 84:874–879
Cervera MT, Ruiz-Garcia L, Martinez-Zapater JM (2002) Analysis of DNA methylation in Arabidopsis thaliana based on methylation sensitive AFLP markers. Mol Genet Genomics 268:543–552
Chakrabarty D, Yu KW, Paek KY (2003) Detection of DNA methylation changes during somatic embryogenesis of Siberian ginseng (Eleuterococcus senticosus). Plant Sci 165:61–68
Creissen SS, Karp A (1985) Karyotypic changes in potato plants regenerated from protoplasts. Plant Cell Tissue Organ Cult 4:171–182
Damasco OP, Smith MK, Adkins SW, Godwin ID (1998) Use of SCAR based marker for early detection of dwarf off-types in micropropagated ‘Cavendish’ bananas. Acta Hort 461:157–164
Dunstan DI, Tautorus TE, Thorpe TA (1995) Somatic embryogenesis in woody plants. In: Thorpe TA (ed) In vitro embryogenesis in plants. Kluwer, Dordrecht, The Netherlands, pp 471–541
Evans DA, Sharp WR, Medina-Filho HP (1984) Somaclonal and gametoclonal variation. Am J Bot 77:759–774
Feil R (2006) Environmental and nutritional effects on the epigenetic regulation of genes. Mutat Res 600:46–57
Gostimsky SA, Kokaeva ZG, Konovalov FA (2005) Studying plant genome variation using molecular markers. Russ J Genet 41:378–388
Gruenbaum Y, Naveh-Many T, Cedar A, Razin A (1981) Sequence specificity of methylation in higher plant DNA. Nature 292:860–862
Guo WL, Gong L, Ding ZF, Li YD, Li FX, Zhao SP, Liu B (2006) Genomic instability in phenotypically normal regenerants of medicinal plant Codonopsis lanceolata Benth. et Hook. f., as revealed by ISSR and RAPD markers. Plant Cell Rep 25:896–906
Guo WL, Wu R, Zhang YF, Liu XM, Wang HY, Gong L, Zhang ZH, Liu B (2007) Tissue culture-induced locus-specific alteration in DNA methylation and its correlation with genetic variation in Codonopsis lanceolata Benth. et Hook. f. Plant Cell Rep 26:1297–1307
Jain SM (1998) Plant biotechnology and mutagenesis for sustainable crop improvement In: Behl RK, Singh DK, Lodhi GP (eds) Crop improvement for stress tolerance. New Delhi, India: CCSHAU, Hissar & MMB, pp 218–232
Jaligot E, Rival A, Beule T, Dussert S, Verdeil JL (2000) Somaclonal variation in oil palm (Elaeis guineensis Jacq.): the DNA methylation hypothesis. Plant Cell Rep 19:684–690
Joyce SM, Cassells AC (2002) Variation in potato microplant morphology in vitro and DNA methylation. Plant Cell Tissue Organ Cult 70:125–137
Kaeppler SM, Phillips RL (1993a) DNA methylation and tissue culture-induced variation in plants. In Vitro Cell Dev Biol Plant 29:125–130
Kaeppler SM, Phillips RL (1993b) Tissue culture-induced DNA methylation variation in maize. Agric Sci 90:8773–8776
Kaeppler SM, Kaeppler HF, Rhee Y (2000) Epigenetic aspects of somaclonal variation in plants. Plant Mol Biol 43:179–188
Kazazian HH (2004) Mobile elements: drivers of genome evolution. Science 303:1626–1632
Keyte AL, Percifield R, Liu B, Wendel JF (2006) Intraspecific DNA methylation polymorphism in cotton (Gossypium hirsutum). J Hered 97:444–450
Kidwell AG, Lisch D (1997) Transposable elements as source of variation in animals and plants. Proc Natl Acad Sci USA 94:7704–7711
Krishnaraj S, Vasil IK (1995) Somatic embryogenesis in herbaceous monocots. In: Thorpe TA (ed) In vitro embryogenesis in plants. Kluwer, Dordrecht, The Netherlands, pp 417–471
Kumar PS, Mathur VL (2004) Chromosomal instability in callus culture of Pisum sativum. Plant Cell Tissue Organ Cult 78:267–271
Kuznetsova OI, Ash OA, Hartina GA, Gostimskij SA (2005) RAPD and ISSR analyses of regenerated pea Pisum sativum L. plants. Russ J Genet 41:60–65
Larkin PJ, Scowcroft WR (1981) Somaclonal variation—a novel source of variability from cell cultures for plant improvement. Theor Appl Genet 60:443–455
Li Y, Guo W, Liu X, Shan X, Li F, Zhang Z, Liu B (2006) Efficient micropropagation of Japanese Photinia [Photinia glabra (Thunb.) Maxim.] retaining genetic and epigenetic stability. Propag Ornam Plants 6:149–155
Li X, Yu X, Wang N, Feng Q, Dong Z, Liu L, Shen J, Liu B (2007) Genetic and epigenetic instabilities induced by tissue culture in wild barley (Hordeum brevisubulatum (Trin.) Link). Plant Cell Tissue Organ Cult 90:153–168
Liu ZL, Han FP, Tan M, Shan XH, Dong YZ, Wang XZ, Fedak G, Hao S, Liu B (2004) Activation of a rice endogenous retrotransposon Tos17 in tissue culture is accompanied by cytosine demethylation and causes heritable alteration in methylation pattern of flanking genomic regions. Theor Appl Genet 109:200–209
LoSchiavo F, Pitto L, Giuliano G, Torti G, Nuti-Ronchi V, Marazziti D, Vergara R, Orselli S, Terzi M (1989) DNA methylation of embryogenic carrot cell cultures and its variations as caused by mutation, differentiation, hormones and hypomethylating drugs. Theor Appl Genet 77:325–331
Matthes M, Singh R, Cheah SC, Karp A (2001) Variation in oil palm (Elais guineensis Jacq.) tissue culture-derived regenerants revealed by AFLPs with methylation sensitive enzymes. Theor Appl Genet 102:971–979
May RA, Trigiano RN (1991) Somatic embryogenesis and plant regeneration from leaves of Dendrathema grandiflora. J Am Soc Hortic Sci 116:366–371
McClelland M, Nelson M, Raschke E (1994) Effect of site-specific modification on restriction endonucleases and DNA modification methyltransferases. Nucleic Acids Res 22:3640–3659
Meins F Jr (1983) Heritable variation in plant cell culture. Annu Rev Plant Physiol 34:327–346
Muller E, Brown PTH, Hartke S, Lorz H (1990) DNA variation in tissue-culture-derived rice plants. Theor Appl Genet 80:673–679
Osternack N, Saare-Surminski K, Preil W, Lieberei R (1999) Induction of somatic embryos, adventitious shoots and roots in hypocotyls tissue of Euphorbia pulcherrima Willd. Ex Klotzsch: comparative studies on embryogenic and organogenic competence. J Appl Bot 73:197–201
Parra R, Pastor MT, Pérez-Payá E, Amo-Marco JB (2001) Effect of in vitro shoot multiplication and somatic embryogenesis on 5-methylcytosine content in DNA of Myrtus communis L. Plant Growth Regul 33:131–136
Peraza-Echeverria S, Herrera-Valencia VA, James-Kay A (2001) Detection of DNA methylation changes in micropropagated banana plants using methylation-sensitive amplification polymorphism (MSAP). Plant Sci 161:359–367
Peredo EL, Ángeles Revilla M, Arroyo-García R (2006) Assessment of genetic and epigenetic variation in hop plants regenerated from sequential subcultures of organogenic call. J Plant Physiol 163:1071–1079
Phillips MH, Keppler SM, Olhoft P (1994) Genetic variability of plant tissue cultures: breakdown of normal control. Proc Natl Acad Sci USA 91:5222–5226
Portis E, Acquadro A, Comino C, Lanteri S (2004) Analysis of DNA methylation during germination of pepper (Capsicum annuum L.) seeds using methylation-sensitive amplification polymorphism (MSAP). Plant Sci 166:169–178
Quemada H, Roth EK, Lark KG (1987) Changes in methylation status of tissue cultured soybean cells detected by digestion with the restriction enzymes Hpa II and Msp I. Plant Cell Rep 6:63–66
Reyna-Lopez GE, Simpson J, Ruiz-Herrera J (1997) Differences in DNA methylation patterns are detectable during the dimorphic transition of fungi by amplification of restriction polymorphisms. Mol Gen Genet 253:703–710
Russo VEA, Martienssen RA, Riggs AD (1996) Epigenetic mechanisms of gene regulation. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Skoog F, Miller CO (1957) Chemical regulation of growth and organ formation in plant tissues cultured in vitro. Symp Soc Exp Biol 54:118–130
Smulders MJM, Rus-Kortekaas W, Vosman B (1995) Tissue culture-induced DNA methylation polymorphisms in repetitive DNA of tomato calli and regenerated plants. Theor Appl Genet 91:1257–1264
Stover RH (1987) Somaclonal variation in Grand Naine and Saba bananas in the nursery and in the field. In: Persley GJ, De Langhe EA (eds), Bananas and plantain breeding strategies. Proceedings of an international workshop, No. 21, Cairns, Australia, 13–17 October 1986. Australian Centre for International Agricultural Research (ACIAR), Canberra, pp 136–139
Sunderland N (1977) Nuclear cytology. In: Street HE (ed) Plant tissue and cell culture. Blackwell, Oxford, pp 177–205
Tanurdzic M, Vaughn MW, Jiang H, Lee TJ, Slotkin RK, Sosinski B, Thompson WF, Doerge RW, Martienssen RA (2008) Epigenomic consequences of immortalized plant cell suspension culture. PLoS Biol 6(12):2880–2895
Tariq M, Paszkowski J (2004) DNA and histone methylation in plants. Trends Genet 20(6):244–251
Thomas E, Bright SWJ, Franklin J, Lancaster V, Miflin BJ (1982) Variation amongst protoplast-derived potato plants (Solanum tuberosum cv, “Maris Bar”). Theor Appl Genet 62:65–68
Thorpe TA, Stasolla C (2001) Somatic embryogenesis. In: Bhojwani SS, Soh WH (eds) Current trends in the embryology of angiosperms. Kluwer, Dordrecht, The Netherlands, pp 279–336
Veilleux RE, Johnson ATT (1998) Somaclonal variation: molecular analysis, transformation, interaction, and utilization. Plant Breed Rev 16:229–268
Vos P, Hogers R, Bleeker M, Reijans M, Vandelee T, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP—a new technique for DNA-fingerprinting. Nucleic Acids Res 23:4407–4414
Wang L, Huang B, He M, Hao S (1990) Somatic embryogenesis and its hormonal regulation in tissue cultures of Freesia refracta. Ann Bot Lond 65:271–276
Wang YM, Dong ZY, Zhang ZJ, Lin XY, Shen Y, Zhou D, Liu B (2005) Extensive de novo genomic variation in rice induced by introgression from wild rice (Zizania latifolia Griseb.). Genetics 170:1945–1956
Xiong LZ, Xu CG, Maroff MAS, Zhang QF (1999) Patterns of cytosine methylation in an elite rice hybrid and its parental lines, detected by a methylation-sensitive amplification polymorphism technique. Mol Gen Genet 261:439–446
Xu ML, Li XQ, Korban SS (2004) DNA-methylation alterations and exchanges during in vitro cellular differentiation in rose (Rosa hybrida L.). Theor Appl Genet 109:899–910
Young WP, Schupp JM, Keim P (1999) DNA methylation and AFLP marker distribution in the soybean genome. Theor Appl Genet 99:785–792
Acknowledgments
This study was supported by the Program for Changjiang Scholars and Innovative Research Team (PCSIRT) in University (#IRT0519), the National Natural Science Foundation of China (30970280), and a grant of the Jilin provincial government of China (20085030).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gao, X., Yang, D., Cao, D. et al. In Vitro Micropropagation of Freesia hybrida and the Assessment of Genetic and Epigenetic Stability in Regenerated Plantlets. J Plant Growth Regul 29, 257–267 (2010). https://doi.org/10.1007/s00344-009-9133-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00344-009-9133-4