Abstract
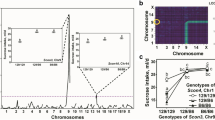
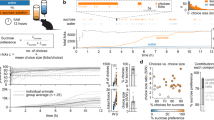
Individual variability in sucrose consumption is prominent in humans and other species. To investigate the genetic contribution to this complex behavior, we conducted behavioral, electrophysiological, and genetic studies, using male progeny of two inbred mouse strains (C57BL/6ByJ [B6] and 129/J [129]) and their F2 hybrids. Two loci on Chromosome (Chr) 4 were responsible for over 50% of the genetic variability in sucrose intake. These loci apparently modulated intake by altering peripheral neural responses to sucrose. One locus affected the response threshold, whereas the other affected the response magnitude. These findings suggest that the majority of difference in sucrose intake between male B6 and 129 mice is due to polymorphisms of two genes that influence receptor or peripheral nervous system activity.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Bachmanov AA, Reed DR, Tordoff MG, Price RA, Beaucharap GK (1996a) Intake of ethanol, sodium chloride, sucrose, critic acid and quinine hydrochloride solutions by mice: A genetic analysis. Behav Genet 26, 563–573
Bachmanov AA, Tordoff MG, Beauchamp GK (1996b) Ethanol consumption and taste preferences in C57BL/6ByJ and 129/J mice. Alcohol Clin Exp Res 20, 201–206
Belknap JK, Crabbe JC, Young ER (1993) Voluntary consumption of ethanol in 15 inbred mouse strains. Psychopharmacol 112, 503–510
Berettini WH, Ferraro TN, Alexander RC, Buchberg AM, Vogel WH (1994) Quantitative trait loci mapping of three loci controlling morphine preference using inbred mouse strains. Nature Genet 7, 54–58
Blass EM, Shah A (1995) Pain-reducing properties of sucrose in human newborns. Chem Senses 20, 29–35
Blizard DA, McClearn GE (1996) Sweet genes and ethanol intake in mice. Alcohol Clin Exp Res 20, 25A
Bolles RC (1991) ed. The Hedonics of Taste. (Hillsdale, NJ, Lawrence Erlbaum.)
Danciger M, Farber DB, Peyser M, Kozak CA (1990) The gene for the beta-subunit of retinal transducin (Gnb-1) maps to distal mouse chromosome 4, and related sequences map to mouse chromosomes 5 and 8. Genomics 6, 428–435
Davis JD, Smith GP (1988) Analysis of lick rate measures: the positive and negative feedback effects of carbohydrates on eating. Appetite 11, 229–238
Dietrich W, Katz H, Lincoln SE, Shin H-S, Friedman J, Dracopoli N, Lander ES (1992) A genetic map of the mouse suitable for typing intraspecific crosses. Genetics 131, 423–447
Dietrich WF, Miller JC, Steen RG, Merchant M, Damron D, Nahf R, Gross A, Joyce DC, Wessel M, Dredge RG, Marquis A, Stein LD, Goodman N, Page DC, Lander ES (1994). A genetic map of the mouse with 4,006 simple sequence length polymorphisms. Nature Genet 7, 220–245
Dobbing J (1987) ed. Sweetness. (London, Springer-Verlag.)
Forgie ML, Beyerstein BL, Alexander BK (1988) Contributions of taste factors and gender to opioid preference in C57BL and DBA mice Psychopharmacol 95, 237–244
Fuller JL (1974) Single-locus control of saccharin preference in mice. J Hered 65, 33–36
George SR, Roldan L, Lui A, Naranjo CA (1991) Endogenous opioids are involved in the genetically determined high preference for ethanol consumption. Alcohol Clin Exp Res 15, 668–672
Gosnell BA, Majchrzak MJ (1989) Centrally administered opioid peptides stimulate saccharin intake in nondeprived rats. Pharmacol Biochem Behav 33, 805–810
Hogan B, Costantini F, Lacy E (1986) Manipulating the Mouse Embryo: A Laboratory Manual. (Cold Spring Harbor, NY, Cold Spring Harbor Press)
Koch JE, Glass MJ, Cooper ML, Bodnar RJ (1995) Alterations in deprivation, glucoprivic and sucrose intake following general, mu and kappa opioid antagonists in the hypothalamic paraventricular nucleus of rats. Neuroscience 66, 951–957
Lander ES, Botstein D (1989) Mapping Mendelian factors underlying quantitative traits using RFLP linkage maps. Genetics 121, 185–199
Lander E, Green P, Abrahamson J, Barlow A, Daley M, Lincoln S, Newburg L (1987) MAPMAKER: An interactive complex package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1, 174–181
Lander ES, Kruglyak L (1995) Genetic dissection of complex traits: guidelines for interpreting and reporting linkage findings. Nature Genetics 11, 241–247
Lush IE (1989) The genetics of tasting in mice. VI. Saccharin, acesulfame, dulcin and sucrose. Genet Res 53, 95–99
Lush IE, Hornigold N, King P, Stoye JP (1995) The genetics of tasting in mice. VII. Glycine revisited, and the chromosomal location of Sac and Soa. Genet Res 66, 167–174
Melchoir JC, Rigaurd D, Colas-Linhart N, Petiet A, Girard A, Apfelbaum M (1991) Immunoreactive beta-endorphin increases after an aspartame chocolate drink in healthy human subjects. Physiol Behav 50, 941–944
Melo JA, Shendure J, Pociask K, Silver LM (1996) Identification of sexspecific quantitative trait loci controlling alcohol preference in C57B176 mice. Nature Genet 13, 147–153
Ninomiya Y, Fukami Y, Yamazaki K, Beauchamp GK (1996) Amiloride inhibition of chorda tympani responses to NaCl and its temperature dependency in mice. Brain Res 708, 153–158
Ninomiya Y, Sako N, Katsukawa H, Funakoshi M (1991) Taste receptor mechanisms influenced by a gene on chromosome 4 in mice. In Chemical Senses, vol. 3: Genetics of Perception and Communication, CJ Wysocki, MR Kare, eds.(New York, NY, Marcel Dekker) pp 267–278
Pfaffmann C (1959) The sense of taste. In Handbook of Physiology, Section 1: Neurophysiology. J Field, HW Magoun, VE Hall, eds. (Washington, DC, American Physiological Society) vol. I, pp 507–533
Phillips TJ, Crabbe JO, Metten P, Belknap JK (1994) Localization of genes affecting alcohol drinking in mice. Alcohol Clin Exp Res 18, 931–941
Wong GT, Gannon KS, Margolskee RF (1996) Transduction of bitter and sweet taste by gustducin. Nature 381, 796–800
Author information
Authors and Affiliations
Additional information
An erratum to this article is available at http://dx.doi.org/10.1007/PL00006942.
Rights and permissions
About this article
Cite this article
Bachmanov, A.A., Reed, D.R., Ninomiya, Y. et al. Sucrose consumption in mice: Major influence of two genetic Loci affecting peripheral sensory responses. Mammalian Genome 8, 545–548 (1997). https://doi.org/10.1007/s003359900500
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s003359900500