Abstract
Actinobacillus pleuropneumoniae is among the most important pathogens worldwide in pig production. The agent can cause severe economic losses due to decreased performance, acute or chronic pleuropneumonia and an increased incidence of death. Therapeutics cannot be used in a sustainable manner, and vaccination is not always available, but discovering more about host defence and disease mechanisms might lead to new methods of prophylaxis. The aim of the present study was to detect quantitative trait loci (QTL) associated with resistance/susceptibility to A. pleuropneumoniae. Under controlled conditions, 170 F2 animals of a Hampshire/Landrace family, with known differences in founder populations regarding A. pleuropneumoniae resistance, were challenged with an A. pleuropneumoniae serotype 7 aerosol followed by a detailed clinical, radiographic, ultrasonographic, pathological and bacteriological examination. F2 pigs were genotyped with 159 microsatellite markers. Significant QTL were identified on Sus scrofa chromosomes (SSC) 2, 6, 12, 13, 16, 17 and 18. They explained 6–22 % of phenotypic variance. One QTL on SSC2 reached significance on a genome-wide level for five associated phenotypic traits. A multiple regression analysis revealed a combinatory effect of markers SWR345 (SSC2) and S0143 (SSC12) on Respiratory Health Score, Clinical Score and the occurrence of death. The results indicate the genetic background of A. pleuropneumoniae resistance in swine and provide new insights into the genetic architecture of resistance/susceptibility to porcine pleuropneumonia. The results will be helpful in identifying the underlying genes and mechanisms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Actinobacillus pleuropneumoniae is the causative agent of porcine pleuropneumonia. It is characterised by a hemorrhagic and necrotizing pneumonia and fibrinous pleuritis (Haesebrouck et al. 1997). Peracute deaths, acute disease with severe clinical and pathologic-anatomical outcomes, chronic pleuritis in slaughter pigs and subclinical cases with declined performance (Gottschalk and Taylor 2006) all cause extensive losses in pig production worldwide (Noyes et al. 1990).
Several different strategies for combatting porcine pleuropneumonia are currently pursued, which include age-segregated rearing (Cleveland-Nielsen et al. 2002), vaccination and application of antibiotics. However, efficacy of vaccination strategies is often hampered by limited cross-serovar protection (Higgins et al. 1985; Fenwick and Henry 1994).
New subunit vaccines appear to convey better cross-protection (Haesebrouck et al. 1997), and several reports described the advantage of using a commercially available subunit vaccine containing Apx toxins and an outer membrane protein (Tumamao et al. 2004; Shao et al. 2010). However, despite vaccination, pigs were not protected against the disease, and therefore, other disease prevention measures and treatments such as antibiotics still remain essential in controlling A. pleuropneumoniae infections (Jirawattanapong et al. 2008; Sjölund and Wallgreen Sjölund and Wallgren 2010).
Increasing use of antibiotics may provoke the emergence of antimicrobial resistant strains (White et al. 2002), and first antimicrobial resistance of A. pleuropneumoniae isolates has been reported (Gutiérrez-Martín et al. 2006). Thus, for future prophylaxis and control of the disease, it is essential to develop antibiotic independent measures. In addition, there is a strongly increasing demand for pork from healthy animals not subjected to treatment with antibiotic drugs (Van Oirschot 1994). Reduction of pain, suffering and damage in farmed pigs is a further serious issue that needs to be addressed with regard to swine health and infection (Reiner 2009).
In this context, the use of natural resistance and breeding selection towards more disease resistant pigs might be advantageous. Detected gene variants could be used in marker-assisted or genomic selection. Additionally, they would reveal new insights into pathogenesis and host defence, potentially inspiring new prophylactic strategies.
Mapping of quantitative trait loci (QTL) might be one approach towards the identification of relevant genes and causal polymorphisms for important traits (Haley and Andersson 1997).
Previous research has shown that differences in the immune capacity between breeds can be mapped (Edfors-Lilja et al. 1998; Reiner 2009). However, studies of the resistance or susceptibility of pigs to infectious agents are still rare, although investigations of resistance to pseudorabies virus, Sarcocystis miescheriana (Reiner et al. 2002, 2007) and the porcine reproductive and respiratory syndrome virus (e.g. Lunney et al. 2011; Boddicker et al. 2012) have been conducted. Concerning A. pleuropneumoniae, differences in susceptibility among pigs of different populations have been shown in a detailed and comprehensive clinical study by Hoeltig et al. (2009). Gregersen et al. (2010) and Sorensen et al. (2011) mapped QTL for chronic pleuropneumonia and ventro-cranial chronic pleuritis, respectively, providing further evidence for genetic involvement in lung lesions with at least partial involvement of A. pleuropneumoniae.
Based on the populations characterised as distinct regarding A. pleuropneumoniae resistance by Hoeltig et al. (2009), the aim of the present study was to map QTL for the acute onset of pleuropneumonia in precisely characterised F2 animals under environmentally standardised conditions and to gain new insights into the genetics of disease and defence against A. pleuropneumoniae infection.
Material and methods
Experimental pigs
The experimental pigs used in this study emerged from a three generation family. A Hampshire boar was mated to 2 Landrace sows to produce 21 F1 animals, from which 2 F1 boars and 9 F1 sows were crossed, avoiding full-sib matings. The boar was derived from a Hampshire population that had been identified as relatively resistant to infection with the same A. pleuropneumoniae strain used in the present study (Hoeltig et al. 2009). The sows were taken from a population of the German Landrace that was shown to be most susceptible to A. pleuropneumoniae by the same recent study (Hoeltig et al. 2009). F1 animals were born at a German breeding company and then transferred to the Clinic for Small Ruminants and Swine, University of Veterinary Medicine, Foundation, Hannover, Germany. Here, 170 F2 pigs from 17 litters were born, raised and studied. Before entering the challenge study, all pigs were tested for absence of A. pleuropneumonie-specific antibodies (by Apx IIA and Apx IVA ELISA, Leiner et al. 1999; Dreyfus et al. 2004). In addition, all pigs were tested negative for Mycoplasma hyopneumoniae and PRRSV (HerdChek, IDEXX laboratories, Wetbrook, ME, USA) by antibody ELISA and for Influenza A by means of a haemagglutination inhibition test (Hoeltig et al. 2009). All F2 pigs were clinically healthy before entering the challenge study.
The piglets were weaned at the age of 4 weeks, fed with a standardised diet and housed under standardised containment level 2 conditions with exclusive access only through a gated hygiene system.
All animals were cared for in accordance with the principles outlined in the European Convention for the Protection of Vertebrate Animals (approval number: 33.9-42502-04-05/919).
Infection and clinical examination
Actinobacillus pleuropneumonie serotype 7 (Jacobsen et al. 1996; Maas et al. 2006) was obtained from the Institute for Microbiology at the University of Veterinary Medicine, Foundation, Hannover, Germany. Infection and clinical examination were established and improved in a recent project (Hoeltig et al. 2009). In short, pigs were challenged at 7 weeks with a well-introduced A. pleuropneumoniae-containing aerosol (Baltes et al. 2001, 2003a, b; Tonpitak et al. 2002; Jacobsen et al. 2005; Hoeltig et al. 2009). The clinical status was monitored from the day before infection (day 0) to day 7 post infectionem (p.i.) and consisted of: (i) an evaluation of the pigs’ general appearance (posture, behaviour, feed intake, temperature, vomiting), (ii) a detailed examination of the pigs’ respiratory tract (breathing noise, dyspnoea, respiratory frequency, coughing, cyanosis), and (iii) the measurement of oxygen saturation by pulse oximetry (for details, see Hoeltig et al. 2009). Parameters with two manifestations (vomiting) were scored with 0 or 4 points, parameters with three manifestations (dyspnoea) with 0, 2 or 4 points, parameters with four manifestations (breathing noise, coughing and feed intake) with 0, 1.33, 2.67 or 4 points, and the remaining parameters with five manifestations with 0, 1, 2, 3 or 4 points. The points were added and divided by 11 to consider the 11 clinical parameters shown under (i), (ii) and (iii) equally. These points produced the daily clinical score, with a score of 0 for a healthy animal and a maximum score of 4 for a severely diseased animal. Deceased animals were scored with 5 points. The clinical score for each day was calculated as the sum of the daily clinical scores from day 1 to the current day.
Additionally, ultrasonographic and radiographic examination was applied on days 0 (before infection) and 7 p.i. to evaluate lung status (for further details see Hoeltig et al. 2009). For ultrasonography, both lungs were divided into 32 areas and 0 (normal lung tissue) to 8 (no unaltered lung tissue) points were assigned per area. Radiography was carried out in two views (latero-lateral and dorso-ventral; Convix Generator 360, Fa. Picker Int., Munich, Germany, 400 mAs, 110 kV, 1,000 ms) using a shutter priority (Precimat, Fa. Picker Int.). For the radiographic score, each lung was divided into eight sections. Bronchial, alveolar and interstitial patterns, cardiac and diaphragm silhouette and sequesters were analysed with a point range from 0 to 3. The radiographic score was the total of the scores for all observed alterations, with a score of 50 being lethal.
If a pig died between infection and necropsy, then this event was recorded
Pathological examination
On day 7 p.i., pigs were euthanized, and a pathological examination was performed. From all lungs, lesions were scored using the Lung Lesion Score (LLS) proposed by Hannan et al. (1982) (Fig. 1). In brief, areas of non-physiological consistence were recorded in a schematic porcine lung map. All seven lobes were divided into triangles. The number of damaged triangles per lobe were counted to determine the lobe’s value, where 5 was the maximum score. The values of all seven lobes were added to give the LLS, with a maximum of 35.
Scoring of lung lesions. a Picture (A.1) and Scoring Scheme (A.2) of a lung without lung lesions 7 days after infection. In the scoring scheme, the lung is divided into seven lung lobs indicated by the white numbers. Each lung lobe is schematized by different numbers of triangles. b Picture (B.1) and Scoring Scheme (B.2) of a lung with severe lesions 7 days after infection. The Scoring scheme shows marked triangles in accordance to the appearance of lung lesions upon the picture [Lobus accessorius (number 7) is not visible]. c Calculation formula of the LLS. At first a LLS is calculated for each lung lobe by the number of marked triangles and the number of triangles of the lung lobe; then, the LLS of all lung lobes are summated. The factor five in the formula determines the maximum score points of each lung lobe, and so the maximum LLS of the lung is 35
Blood, liver and spleen samples were taken for DNA preparation
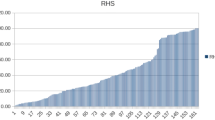
Findings from clinical, radiographic and ultrasonographic examinations and the LLS were combined to give the Respiratory Health Score (RHS) (Hoeltig et al. 2009). Based on the RHS, two groups of 50 pigs each were formed, representing highest and lowest scores, respectively (RHS*: 0/1).
Bacteriological examination
Bacterial colonisation of the lungs was semi-quantified using the Re-isolation Score (RIS) (Maas et al. 2006). Each of the seven lung lobes, the tonsils and one lung lymph node were given score points: 0 (no growth of A. pleuropneumoniae), 1 (colonies growing in the swabbed area only), 2 (colonies growing in the swabbed area and in the first fractionated streak) to 3 (A. pleuropneumoniae colonies growing in the swabbed area and in both fractionated streaks), producing a quantitative cultural score (RIS) with values from 0–27 (summed up score of nine tissues). Tissue was taken from precisely defined positions and cultured on modified Columbia agar (Jacobsen and Nielsen 1995).
Markers and linkage map
Genomic DNA was extracted from blood, liver or spleen tissue of experimental pigs using a commercial DNA isolation kit (Analytik Jena, Germany), which proved its suitability for DNA isolation in former studies. DNA quality and quantity was measured with an Ultrospec 1100 pro photometer (GE Healthcare Europe GmbH, Freiburg, Germany).
Informative marker loci were selected to cover all autosomes at average intervals of 15 cM. Although genome wide 60 SNP chips have been available for the pig since 2009 (Ramos et al. 2009), we decided to work with microsatellites because they were well established in our laboratory. From 491 microsatellite markers selected from public sources (www.ncbi.nlm.nih.gov; www.usda.gov), only 155 markers were informative and used to genotype the F2 families (see Supplementary Table 1). Another four markers (CD14, TNF, IFNAR1 and CD40) developed for this study were found to be associated with some candidate genes. Genetic positions of these markers were calculated with the Kosambi function of CRI-MAP, Version 2.4 (Green et al. 1990). Positions of the other markers were obtained from NCBI, UniSTS (http://www.ncbi.nlm.nih.gov/genome/sts), providing a much higher confidence than those obtained from single studies. Most of the terminal markers from public sources were non-informative in this study. Therefore, the initial position of markers on SSC1, 5, 6, 7, 8, 9, 12, 14, 15, and 16 is different from 0 cM.
Statistical analysis
Phenotypic data were analysed using IBM-SPSS, Release 20.0 (IBM, Herrenberg, Germany). Means and standard deviations were calculated using the Descriptives procedure. Occurrence of death after infection was specified as a percentage of animals within groups. Significances for this binomial trait were tested with Kruskal–Wallis test for independent samples. The 50 pigs with the highest and the lowest RHS were classified as high and low responders, respectively. The remaining pigs were classified as intermediate. Responsiveness was included into the new variable RHS*, including 1 (low responders), 2 (intermediate) and 3 (high responders).
The study was conducted consecutively with one or two litters per run. The age of piglets within the single study group was identical. Thus, the effect of age was contained in the effect of the study group. Means, standard errors and significances of clinical, pathological and microbiological traits were calculated with univariate variance analysis with RHS*, sex and study group as fixed effects, following the model:
Effects of QTL-linked markers on phenotypic traits were tested using multiple regression analysis; differences in mean values were calculated by means of the general linear model described before, with inclusion of the fixed effects of the genotypes of markers Swr345 (SSC2) and S0143 (SSC12).
QTL analysis
QTL analysis was performed using the Web-based application ‘Grid QTL’ (Seaton et al. 2006). Analyses were based on a least-square method developed for crosses between inbred and outbred lines (Haley et al. 1994). In the general linear model, the fixed effects of sex and study group were included. Chromosome-wide and genome-wide significance levels were estimated by means of permutation tests (Churchill and Doerge 1994) with 1,000 permutations. F values were calculated as average from all 18 autosomes. F values were 5.4 (p ≤ 0.05) and 7.7 (p ≤ 0.01) for chromosome-wide significance and 9.8 (p ≤ 0.05) for the genome-wide significance threshold. To avoid violating the assumption that marker genotypes associated with multiple QTL are uncorrelated, the QTL analysis was performed sequentially (Holl et al. 2004). The highest QTL effect of the first run was included into the model as a background effect in the second run, and both QTL were included as background effects in the third run and so on. QTL analysis for one trait was stopped when the last run did not identify a new QTL with an F value ≥5.4. Positive values with additive effects indicate that Hampshire alleles result in higher values than Landrace alleles and vice versa.
Effects of QTL-linked markers on phenotypic traits were tested by multiple regression analysis in IBM-SPSS, Release 20.0 (IBM, Herrenberg, Germany). Instead of using the original allele length of the microsatellite markers, marker information was prepared for calculation as homozygous Hampshire derived (1), homozygous German Landrace derived (3) or heterozygous (2). Significant were only markers Swr345 (SSC2) and S0143 (SSC12). Thus, these two markers were combined to generate nine genotypes which were included in the general linear model as described before.
Results
Phenotypes
All pigs included into the study were clinically healthy before infection (day 0) with clinical scores of 0 to 0.6 (mean 0.06), sonographic scores between 0 and 2 (mean 0.12) and a radiographic score of 0. All pigs were tested free from antibodies against A. pleuropneumoniae, M. hyopneumoniae, Influenza A and PRRSV. There was neither elevated temperature, nor dyspnoea, coughing nor apathy detectable.
Upon challenge with A. pleuropnemoniae, clinical scores increased steadily from day 1 to day 7 p. i. Besides overall scores, the coefficients of variation increased too. Day 7 was characterised by comet tail artefacts, interruption of the pleural line, liverlike parenchyma texture, abscesses, bronchography and increasing lung opacity in sonographic and radiographic examinations, which are typical for porcine pleuropneumonia.
Due to severe clinical symptoms, five pigs died or were euthanized before day 7. Nine from 35 points and 9 from 27 points were reached in LLS, and RIS obtained during necropsy of the pigs on day 7 p.i. Although infected, some pigs did not develop scores higher than 0 in any of the parameters examined. The overall RHS varied between 0 and 66.7. Ranking scores between RHS and the other clinical parameters were between 0.345 (occurrence of death) and 0.714 (LLS) (Table 3), indicating different aspects of the disease represented by the different scores. Correlations between phenotypic traits obtained after challenge were mainly significant (p < 0.001). Traits obtained before infection were not correlated with traits after infection. There was a weak correlation between death because of infection (0/1) and the other scores. Contribution rates of the correlations between the RHS and the major clinical, pathological and microbiological traits varied between 41.5 and 91 % (Table 2).
Clinical, pathological and microbiological traits of the F2 pigs showed a high degree of variation (Table 1), with coefficients of variation between 0.8 and 3.6. None of the scores obtained were normally distributed within the F2 pigs.
Score variation was significantly affected by response (low/intermediate/high) and run (14 groups of one or two litters consecutively housed and challenged), but not by sex of the piglets. A comparison of trait values in low, intermediate and high responders is given in Table 3. There were no differences between low, intermediate and high responders for all traits obtained before infection. However, the 50 low and high responders were significantly different at p ≤ 0.001 for each of the traits obtained after infection. Scores within high responders showed a normal distribution and scores within intermediate responders showed widely normal distribution. However, scores within low responders were still highly variable.
QTL
QTL analysis mapped a total of 7 QTL, significant at least on a chromosome-wide level (p < 0.05), linked with the clinical score (increasing in significance from day 1 p.i. to day 7 p.i.), with the radiographic score on day 7, with the occurence of death (0/1), and with the RHS (Table 4). The confidence interval of 14 cM was lowest in association with the clinical score 7 d.p.i. Interestingly, Hampshire alleles at this position were linked with a higher degree of susceptibility of the F2 pigs.
Clinical Score and RHS were further influenced by a QTL on SSC17; this effect appeared late after infection and again, unfavourable scores were linked with Hampshire alleles. The sonographic scores and RHS were further linked with a QTL on SSC12. Favourable alleles came from Hampshire. Interestingly, this QTL showed linkage to the sonographic score prior to infection. Sonographic score was further linked to QTL on SSC6 and 12. The RIS was linked with two QTL on SSC16 and 18, and the LLS correlated with a QTL on SSC18. Grouping of individuals (RHS*) into low and high responders was linked with two QTL on SSC13 and 18. QTL explained between 6 and 22 % of total F2 variance. Except for the QTL on SSC2, confidence intervals were generally wide.
QTL on SSC2, 6, 12 and 17 peaked at the positions of markers Swr345 (SSC2), KVL3285 (SSC6), S0143 (SSC12) and Swn335 (SSC17). Multiple regression analyses of markers showed significant effects for Swr345 and S0143 on RHS, Clinical Score on day 7 and the occurrence of death. Both markers had a significant combinatory effect on all three phenotypic traits (Fig. 2a–c). The occurrence of death was linked with the homozygous presence of the unfavourable allele in one marker with at least one unfavourable allele in the other marker (heterozygous or homozygous), while in all other cases no death occurred (Fig. 2a). The same principle was found for Clinical Score (day 7) (Fig. 2b) and RHS (Fig. 2c). Effects of the single markers and combined effects of both markers were statistically significant (p < 0.001) in all three phenotypic traits. The contribution ratio was 0.351. Favourable alleles of Swr345 (SSC2) came from the German Landrace, those of S0143 (SSC12) from Hampshire.
Combinatory effects of markers Swr345 and S0143 on phenotypic traits: occurrence of death (a), Clinical Score (b), and RHS (c). H alleles of the Hampshire breed, L allele of the German Landrace. RHS Respiratory Health Score. Results are based on 170 pigs. Values of the y axis refer to the percentage of dead pigs, the clinical score and the RHS score, respectively, in each of the nine genotypes combined from markers Swr345 (SSC2) and S0143 (SSC12)
Discussion
Several studies confirmed the low susceptibility of a Hampshire pig population to a challenge with A. pleuropneumoniae. A population of German Landrace pigs has to be considered as most susceptible, as only pigs of this breeding line died or had to be euthanized due to severe clinical symptoms in preceding experiments (Hoeltig et al. 2009). We used serotype 7, but we expect similar results upon infection with other A. pleuropneumoniae serotypes, because the general mechanisms of pathogenicity are species-specific. However, the genetic mechanisms responsible for the breed differences are still unknown.
The intention of the present study was to map chromosomal areas linked to the observed differences in relative susceptibility/resistance to A. pleuropneumoniae. The QTL have been mapped in a model of maximal accuracy of phenotypes and minimal animal numbers to address concerns of animal welfare. Founder pigs have been chosen based on the previous comparison of different pig breeds under the same experimental conditions (Hoeltig et al. 2009). Although certain populations of German Landrace proved to be most and a population of Hampshire to be least susceptible to A. pleuropneumoniae, segregation of resistance/susceptibility is generally given in each of the populations. Gregersen et al. (2010) investigated 7,300 slaughter pigs in the field for symptoms of the chronic outcome of A. pleuropneumoniae infection, i.e. dorso-caudal chronic pleuritis. Their study revealed six QTL significant on a chromosome-wide level. The QTL on SSC2, 12, 13 and 18 were confirmed by the present study. Confidence intervals were similar in both studies. A comparison of both studies demonstrates the power of extensive individual phenotyping in QTL analysis, especially regarding traits of disease resistance. Differences in maternal immunity, immunity of the infected slaughter pigs, infection pressure in herds, infection time point and individual stressors are the environmental factors that most often interfere and lead to false classifications of an individual’s resistance/susceptibility. The challenge model of the present study used the same environment, time point and dosage for infection, along with a very detailed and precisely conducted phenotyping.
Following previous study, acute lung damage appears to be decisive for the course of disease, suggesting a major role of the non-adaptive immune system in the defence of pleuropneumonia. Thus, the present experiment could be terminated on day 4 p.i. without loss of information (Hoeltig et al. 2009). We have, however, prolonged the experiment until day 7 p.i., and the data showed a steady increase in the power of the resulting QTL for the clinical score. This gradient of QTL power argues for the chance of improved mapping results, if sampling had been performed later than day 7 p.i., at least with this characteristic.
Our study shows for the first time QTL that are linked to the acute development of pleuropneumonia after standardised infection. The study detected 7 QTL located on SSC2, 6, 12, 13, 17 and 18.
The QTL on SSC2 covers the region of recently described QTL for the resistance to Sarcocystis miescherina (Reiner et al. 2007), pseudorabies (Reiner et al. 2002) and ventro-cranial chronic pleuritis (Sorensen et al. 2011). Candidate genes on SSC2 are interleukin 9 (IL9; Sorensen et al. 2011) and CD14 (cluster of differentiation 14; Tsai et al. 2011). IL 9 variants have been described together with human lung function variability (Aschard et al. 2009). CD14 cooperates with MD-2 and TLR4 to mediate the innate immune response to bacterial lipopolysaccharides (LPS), one of the major virulence factors of A. pleuropneumoniae and a key response of the host defence (Cho et al. 2005). Candidate genes for the SSC12 QTL are STAT3, STAT5A, STAT5B (signal transducer and activator of transcription 3, 5A and 5B). Members of the STAT gene family are of specific importance in immunesystem-related gene regulation (Casanova et al. 2012). The QTL on SSC13 is located in the area of the transferrin gene (TF). TF is an important factor in pleuropneumonia, depending on the disposability of iron for the bacteria (Chiers et al. 2010). The existence of TF variants has been described (Danilowicz et al. 2009). Different QTL involved in RHS/RHS*, mapped on the basis of 170 and 50/50 pigs, might serve as further evidence for alternative pathways in A. pleuropneumoniae defence in different animals. Together with the known pathophysiological effects of TF, these results argue, however, for the consideration of this region in further studies.
QTL on SSC8 and 14 were only detected by the group of Gregersen et al. (2010), QTL on SSC6, 16 and 17 only by the present study. This difference and the occurrence of QTL almost exclusively in singular families in the study by Gregersen et al. (2010) point towards distinct genes that might be involved in A. pleuropneumoniae defence.
A batch of factors involved in innate immunity has been mapped to SSC18, including TLR9, IL10 and C3C (Wimmers et al. 2008a, b, Uddin et al. 2011). These candidate genes have been identified as significant factors modulating the innate immune system after contact with tetanus toxoid and Porcine Reproductive and Respiratory Syndrome Virus (PRRSV) (Wimmers et al. 2008a, b). Additionally, the QTL on SSC18 overlaps with a QTL for lymphocyte numbers in a chronical disease model (Reiner et al. 2008). SNPs have been identified on SSC18 with association to CD4 + CD8−, CD4+ and CD4 + CD8+ lymphocyte subpopulation fractions (Lu et al. 2012), implicating a possible role of the adaptive arm of immunity.
The most prominent effect in the present study was that any pig which died from A. pleuropneumoniae infection was homozygous for one founder breed in one of the markers Swr345 (SSC2) or S0143 (SSC12) and carried at least one unfavourable allele in the corresponding marker. Interestingly, favourable effects regarding the outcome of A. pleuropneumonia infection were linked with alleles from Hampshire in S0143 and alleles from German Landrace in Swr345. This aspect adds further impact to the involvement of different genes and pathways in A. pleuropneumoniae defence, as discussed above. Although the founder pigs were derived from subpopulations with marked differences in susceptibility to pleuropneumonia, the actual founder pigs could not be tested individually. Thus, there might be individuals within both populations, allowing for more marked inter-population differences than those selected for the present study.
The effects of these markers are significant in the F2 population of the present study. Thus, both markers are linked with highly interesting causal gene variants that need to be defined by further studies. Because of the genetic nature of F2 crossbreds, linkage between markers and causal variants might, however, be too loose to be applied in commercial populations, without further testing.
Although the present study was based on a relatively low number of animals (n = 170) and markers (n = 159), the high accuracy of phenotypes led to a high conformity with results obtained with 7,300 pigs and a 7k porcine SNP chip (Gregersen et al. 2010) with respect to location, power and confidence intervals of the QTL. Nevertheless, some QTL might have been underestimated or even escaped from mapping because of high distances between some markers in the present study. Thus, future studies should involve available genome wide SNP chips (e.g. 60k; Ramos et al. 2009) or genotyping by sequencing (Spindel et al. 2013). Because of the relatively (SSC2) and absolutely (the other chromosomes) wide confidence intervals, fine mapping of candidate genes cannot be based solely on the results of the present study. However, combining these positional data with functional data should help to restrict functional candidate genes and lead to the discovery and use of those genes, involved in A. pleuropneumoniae resistance/susceptibility. While absolute resistance or susceptibility can hardly be assumed from the complexity of disease and defence mechanisms, even relative resistance would be extremely constructive for worldwide swine production. The RISs of the most resistant pigs of the present study argue for an overall reduction in the herds’ infectious pressure, a major factor for the spread of infectious diseases within and between herds (Reiner 2009). Clinical scores, RHS and LLSs of the most resistant 50 pigs proof their successful defence of A. pleuropneumoniae. Although selecting for a unique disease resistance might also lead to susceptibility to other diseases, the knowledge of the causal gene variants and their application in future genomic selection strategies could reduce disease incidence, improve performance and product quality and, by reducing the necessity of antibiotic treatment, would improve consumer protection and animal welfare. A lot of information will be needed, before a clear picture can be drawn for a distinct resistance in the future.
References
Aschard H, Bouzigon E, Corda E, Ulgen A, Dizier MH, Gormand F, Lathrop M, Kauffmann F, Demenais F, EGEA Cooperative Group (2009) Sex-specific effect of IL9 polymorphisms on lung function and polysensitization. Genes Immun 10:559–565
Baltes N, Topitak W, Gerlach GF, Hennig-Pauka I, Hoffmann-Moujahid M, Ganter M, Rothkotter HJ (2001) Actinobacillus pleuropneumoniae iron transport and urease activity: effects on bacterial virulence and host immune response. Infect Immun 69:472–478
Baltes N, Hennig-Pauka I, Jacobsen I, Gruber AD, Gerlach GF (2003a) Identification of dimethyl sulfoxide reductase in Actinobacillus pleuropneumoniae and its role in infection. Infect Immun 71:6784–6792
Baltes N, Tonpitak W, Hennig-Pauka I, Gruber AD, Gerlach GF (2003b) Actinobacillus pleuropneumoniae serotype 7 siderophore receptor FhuA is not required for virulence. FEMS Microbiol Lett 220:41–48
Boddicker N, Rowland R, Lunney JK, Garrick DJ, Reecy J, Dekkers JCM (2012) A major QTL associated with host response to porcine reproductive and respiratory syndrome virus challenge. J Anim Sci 90:1733–1746
Casanova JL, Holland SM, Notarangelo LD (2012) Inborn errors of human JAKs and STATs. Immunity 36:515–528
Chiers K, De Waele T, Pasmans F, Ducatelle R, Hasebrouck F (2010) Virulence factors of Actinobacillus pleuropneumoniae involved in colonization, persistence and induction of lesions in its porcine host. Vet Res 41:65
Cho WS, Jung K, Kim J, Ha Y, Chae C (2005) Expression of mRNA encoding interleukin (IL)-10, IL-12p35 and IL-12p40 in lungs from pigs experimentally infected with Actinobacillus pleuropneumoniae. Vet Res Commun 29:111–122
Churchill GA, Doerge RW (1994) Empirical threshold values for quantitative trait mapping. Genetics 138:963–971
Cleveland-Nielsen A, Nielsen EO, Ersbol AK (2002) Chronic pleuritis in Danish slaughter pig herds. Prev Vet Med 55:121–135
Danilowicz E, Martinez-Arias R, Dolf G, Singh M, Probst I, Tümmler B, Höltig D, Waldmann KH, Gerlach GF, Stanke F, Leeb T, The IRAS Consortium (2009) Characterization of the porcine transferrin gene (TF) and its association with disease severity following an experimental Actinobacillus pleuropneumoniae infection. Anim Genet 41:424–427
Dreyfus A, Schaller A, Nivollet S, Segers RPAM, Kobisch M, Mieli L, Soerensen V, Hussy D, Miserez R, Zimmermann W, Inderbitzin F, Frey J (2004) Use of recombinant ApxIV in serodiagnosis of Actinobacillus pleuropneumoniae infections, development and prevalidation of the Apx IV Elisa. Vet Microbiol 99:227–238
Edfors-Lilja I, Wattrang E, Marklund L, Moller M, Andersson-Eklund L, Andersson L, Fossum C (1998) Mapping quantitative trait loci for immune capacity in the pig. J Immunol 161:829–835
Fenwick B, Henry S (1994) Porcine pleuropneumonia. J Am Vet Med Assoc 204:1334–1340
Gottschalk M, Taylor DJ (2006) Actinobacillus pleuropneumoniae. In: Taylor DJ, Straw BE, Zimmermann JJ, D′Allaire S (eds) Diseases of swine. Blackwell Publishing, Ames, pp 563–576
Green P, Falls K, Crooks S (1990) Documentation for CRI-MAP, version 2.4. Washington University School of Medicine, St. Louis
Gregersen VR, Sørensen KK, Christensen OF, Busch ME, Vingborg RKK, Velander IH, Lund MS, Bendixen C (2010) Identification of QTL for dorso-caudal chronic pleuritis in 12 crossbred porcine families. Anim Genet 41:509–514
Gutiérrez-Martín CB, García Del Blanco N, Blanco M, Navas J, Rodríguez-Ferri EF (2006) Changes in antimicrobial susceptibility of Actinobacillus pleuropneumoniae isolated from pigs in Spain during the last decade. Vet Microbiol 115:218–222
Haesebrouck F, Chiers K, Van Overbeke I, Ducatelle R (1997) Actinobacillus pleuropneumoniae infections in pigs: the role of virulence factors in pathogenesis and protection. Vet Microbiol 58:239–249
Haley CS, Andersson L (1997) Linkage mapping of quantitative trait loci in plants and animals. In: Dear PH (ed) Genome mapping. IRL Press, Oxford, pp 49–71
Haley CS, Knott SA, Elsen J-M (1994) Mapping quantitative trait loci in crosses between outbred lines using least squares. Genetics 136:1195–1207
Hannan PC, Bhogal BS, Fish JP (1982) Tylosin tartrate and tiamutilin effects on experimental piglet pneumonia induced with pig lung homogenate containing mycoplasmas, bacteria and viruses. Res Vet Sci 33:76–88
Higgins R, Lariviere S, Mittal KR, Martineau GP, Rousseau P, Cameron J (1985) Evaluation of a killed vaccine against porcine pleuropneumoniae due to Haemophilus pleuropneumoniae. Can Vet J 26:86–89
Hoeltig D, Hennig-Pauka I, Thies K, Rehm T, Beyerbach M, Strutzberg-Minder K, Gerlach GF, Waldmann KH, FUGATO-consortium IRAS (2009) A novel Respiratory Health Score (RHS) supports a role of acute lung damage and pig breed in the course of an Actinobacillus pleuropneumoniae infection. BMC Vet Res 5:14
Holl JW, Cassady JP, Johnson RK (2004) A genome scan for quantitative trait loci and imprinted regions affecting reproduction in pigs. J Anim Sci 82:3421–3429
Jacobsen MJ, Nielsen JP (1995) Development and evaluation of a selective and indicative medium for isolation of Actinobacillus pleuropneumoniae. Vet Microbiol 47:191–197
Jacobsen MJ, Nielsen JP, Nielsen R (1996) Comparison of virulence of different Actinobacillus pleuropneumoniae serotypes and biotypes using an aerosol infection model. Vet Microbiol 49:159–168
Jacobsen I, Hennig-Pauka I, Baltes N, Trost M, Gerlach GF (2005) Enzymes involved in anaerobic respiration appear to play a role in Actinobacillus pleuropneumoniae virulence. Infect Immun 73:226–234
Jirawattanapong P, Stockhofe-Zurwieden N, van Leengoed L, Wisselink H, Raymakers R, Cruijsen T, van der Peet-Schwering C, Nielen M, van Nes A (2008) Pleuritis in slaughter pigs: relations between lung lesions and bacteriology in 10 herds with high pleuritis. Res Vet Sci 88:11–15
Leiner G, Franz B, Strutzberg K, Gerlach GF (1999) A novel enzymelinked immunosorbent assay using the recombinant Actinobacillus pleuropneumoniae ApxII antigen for diagnosis of pleuropneumonia in pig herds. Clin Diagn Lab Immunol 6:630–632
Lu X, Fu WX, Luo YR, Ding XD, Zhou JP, Liu Y, Liu JF, Zhang Q (2012) Genome-wide association study for T lymphocyte subpopulations in swine. Genomics 13:488
Lunney JK, Steibel JP, Reecy JM, Fritz E, Rothschild MF, Kerrigan M, Trible B, Rowland RRR (2011) Probing genetic control of swine responses to PRRSV infection: current progress of the PRRS host genetics consortium. BMC Proc 5(Suppl 4):30
Maas A, Meens J, Baltes N, Hennig-Pauka I, Gerlach GF (2006) Development of a DIVA subjunit vaccine against Actinobacillus pleuropneumoniae infection. Vaccine 24:7226–7237
Noyes EP, Feeney D, Pijoan C (1990) Comparison of the effect of pneumonia detected during lifetime with pneumonia detected at slaughter on growth in swine. J Am Vet Med Assoc 197:1025–1029
Ramos AM, Crooijmans RPMA, Affara NA, Amaral AJ, Archibald AL et al (2009) Design of a high density SNP genotyping assay in the pig using SNPs identified and characterized by next generation sequencing technology. PLoS One 4:e6524. doi:10.1371/journal.pone.0006524
Reiner G (2009) Investigations on genetic disease resistance in swine—a contribution to the reduction of pain, suffering and damage in farm animals. Appl Anim Behav Sci 118:217–221
Reiner G, Melchinger E, Kramarova M, Pfaff E, Büttner M, Saalmüller A, Geldermann H (2002) Detection of quantitative trait loci for resistance/susceptibility to the pseudorabies virus in swine. J Gen Virol 83:167–172
Reiner G, Kliemt D, Willems H, Berge T, Fischer R, Köhler F, Hepp S, Hertrampf B, Daugschies A, Geldermann H, Mackenstedt U, Zahner H (2007) Mapping of quantitative trait loci affecting resistance/susceptibility to Sarcocystis miescheriana in swine. Genomics 89:638–646
Reiner G, Fischer R, Hepp S, Berge T, Köhler F, Willems H (2008) Quantitative trait loci for white blood cell numbers in swine. Anim Genet 39:163–168
Seaton G, Hernandez J, Grunchec JA, White I, Allen J, De Koning DJ, Wei W, Berry D, Haley C, Knott S (2006) GridQTL: a grid portal for QTL mapping of compute intensive datasets. Proceedings of the 8th world congress on genetics applied to livestock production, Belo Horizonte, Brazil, 13–18 Aug 2006
Shao M, Wang Y, Wang C, Guo Y, Peng Y, Liu J, Li G, Liu H, Liu S (2010) Evaluation of multicomponent recombinant vaccines against Actinobacillus pleuropneumoniae in mice. Acta Veterinaria Scand 52:52
Sjölund S, Wallgren P (2010) Field experience with two different vaccination strategies aiming to control infections with Actinobacillus pleuropneumoniae in a fattening pig herd. Acta Vet Scand 52:23
Sorensen KK, Gregersen VR, Christensen OF, Velander IH, Bendixen C (2011) Genomic regions associated with ventro-cranial chronic pleuritis in pig. Anim Breed Genet 128:314–318
Spindel J, Wright M, Chen C, Cobb J, Gage J, Harrington S, Lorieux M, Ahmadi N, McCouch S (2013) Bridging the genotyping gap: using genotyping by sequencing (GBS) to add high-density SNP markers and new value to traditional bi-parental mapping and breeding populations. Theor Appl Genet 126:2699–2716
Tonpitak W, Baltes N, Hennig-Pauka I, Gerlach GF (2002) Construction of an Actinobacillus pleuropneumoniae serotype 2 prototype live negative-marker vaccine. Infect Immun 70:7120–7125
Tsai TH, Chen SF, Huang TY, Tzeng CF, Chiang AS, Kou YR, Lee TS, Shyue SK (2011) Impaired CD14 and CD36 expression, bacterial clearance, and Toll-like receptor 4YMYD88 signaling in Caveolin-1Y deleted macrophages and mice. Shock 35:92–99
Tumamao JQ, Bowles RE, van den Bosch H, Klaasen HL, Fenwick BW, Storie GJ, Blackall PJ (2004) Comparison of the efficacy of a subunit and a live streptomycin-dependent porcine pleuropneumonia vaccine. Aust Vet J 82:370–374
Uddin MJ, Cinar MU, Große-Brinkhaus C, Tesfaye D, Tholen E, Juengst H, Looft C, Wimmers K, Phatsara C, Schellander K (2011) Mapping quantitative trait loci for innate immune response in the pig. Int J Immunogenet 36:121–131
Van Oirschot JT (1994) Vaccination in food animal populations. Vaccine 12:415–418
White DG, Zhao S, Simjee S, Wagner DD, McDermott PF (2002) Antimicrobial resistance of food born pathogens. Microbes Infect 4:405–412
Wimmers K, Jonas E, Schreinemachers HJ, Tesfaye D, Ponsuksili S, Tholen E, Juengst H, Schellander K, Phatsara C (2008a) Verification of chromosomal regions affecting the innate immunity in pigs using linkage mapping. Dev Biol (Basel) 132:279–283
Wimmers K, Murani E, Schellander K, Ponsuksili S (2008b) QTL for traits related to humoral immune response estimated from data of a porcine F2 resource population. Int J Immunogenet 36:141–151
Acknowledgments
The authors thank the German Ministry of Education and Research (BMBF) and the Development Association for Biotechnology (FBF) for financial support.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Reiner, G., Bertsch, N., Hoeltig, D. et al. Identification of QTL affecting resistance/susceptibility to acute Actinobacillus pleuropneumoniae infection in swine. Mamm Genome 25, 180–191 (2014). https://doi.org/10.1007/s00335-013-9497-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00335-013-9497-4