Abstract
MicroRNAs (miRNAs) are 18–22-nt noncoding RNAs that are involved in post-transcriptional regulation of genes. Oncomirs, a subclass of miRNAs, include genes whose expression, or lack thereof, are associated with cancers. Until the last decade, the domestic dog was an underused model for the study of various human diseases that have genetic components. The dog exhibits marked genetic and physiologic similarity to the human, thereby making it an excellent model for study and treatment of various hereditary diseases. Furthermore, because the dog presents with distinct, spontaneously occurring mammary tumors, it may serve as a model for genetic analysis and treatments of humans with malignant breast tumors. Because miRNAs have been found to act as both tumor suppressors and oncogenes in several different cancers, expression patterns of ten miRNAs (miR-15a, miR-16, miR-17-5p, miR-21, miR-29b, miR-125b, miR-145, miR-155, miR-181b, let-7f) known to be associated with human breast cancers were compared to malignant canine mammary tumors (n = 6) and normal canine mammary tissue (n = 10). Resulting data revealed miR-29b and miR-21 to have a statistically significant (p < 0.05 by MANOVA analysis) upregulation in cancerous samples. The ten canine miRNAs follow the same pattern of expression as in the human, except for miR-145 which does not show a difference in expression between the normal and cancerous canine samples. In addition, when analyzed according to specific cancer phenotypes, miR-15a and miR-16 show a significant downregulation in canine ductal carcinomas while miRsR-181b, -21, -29b, and let-7f show a significant upregulation in canine tubular papillary carcinomas.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
MicroRNAs (miRNAs) are a rapidly expanding class of important, noncoding RNA that have proven regulatory roles in both plants and animals (Ambros 2004; Bartel 2004). Known to work in a post-transcriptional fashion, miRNAs are active in development, apoptosis, differentiation, and tumorigenesis (Ambros 2004; Bartel 2004; Hwang and Mendell 2006). Bioinformatic analysis suggests that there are approximately 1000 miRNA genes in a mammalian genome, and miRNAs are estimated to regulate approximately 30% of all genes (Berezikov et al. 2005; Lewis et al. 2005). In particular, some miRNAs have been labeled “oncomirs” because of their associations with various cancers. This is highlighted by the fact that 50% of the computationally predicted and experimentally proven miRNAs are located within human genomic fragile regions with already proven links between breaks and oncogenesis (Calin et al. 2004b). More specifically, 15 miRNAs are at fragile sites at which alterations and breaks occur frequently, and these aberrations are linked with human breast cancers (Calin et al. 2004b). Other studies have successfully used microarrays and quantitative real-time PCR (qRT-PCR) to “profile” the expression patterns of several different cancers (Blenkiron et al. 2007; Iorio et al. 2005; Jiang et al. 2005; Mattie et al. 2006; Volinia et al. 2006). Breast cancer is currently one of the main foci of these investigations because it is the most often diagnosed cancer in American women (American Cancer Society 2007). The American Cancer Society estimates that 240,510 women would be diagnosed with some form of breast cancer in 2007 and another 40,460 women would die (American Cancer Society 2007).
Breast cancer is not limited to humans. Mammary gland tumors represent almost half (41.7%) of all tumor types of the intact female dogs (Dorn et al. 1968). In fact, a population study found that canine mammary tumors represent 13.4% of all tumors found in the dog (Dorn et al. 1968). Approximately 50% of all canine mammary tumors are malignant. For these malignant variants, surgery is the common treatment and appears to be the only successful treatment option (Lana et al. 2007). Depending on various prognostic factors and tumor-free surgical margins, surgical excision can be curative in many dogs. Key prognostic factors for overall survival time include but are not limited to multiple masses, tumor grade, lymphatic invasion, and distant metastasis (Lana et al. 2007). Perhaps the best characterized prognostic factor is tumor size. For tumors smaller than 3 cm, the median survival time in one study was 22 months. Dogs with a tumor larger than 3 cm had a median survival time of only 14 months (Philibert et al. 2003). Experience with radiation therapy for canine mammary tumors is limited and no definitive data exist. Chemotherapy with agents such as doxorubicin and docetaxel has not radically increased overall survival time (Simon et al. 2006). Hormonal therapy with agents such as tamoxifen does not improve survival and is currently not recommended for use in dogs (Lana et al. 2007). Thus, at this time there are no effective systemic treatment options for dogs with mammary tumors.
There is considerable debate as to whether canine mammary tumors represent a strong model for human breast cancers. Both tumor types are primarily epithelial in nature and both occur spontaneously in the normal population (Dickson et al. 2005; Lana et al. 2007). The biological behavior of both tumors includes local invasion with regional lymph node metastasis and distant metastasis to the lungs (Dickson et al. 2005; Lana et al. 2007). Furthermore, the highly aggressive inflammatory carcinoma is reported in both species. Aberrant expression patterns of proliferating cell nuclear antigen, Bcl-2, cytokeratins, calponin, and the genes p53, Cox-2, and BRCA1 have been found in malignant canine mammary tumors (De Las Mulas et al. 2004; Dore et al. 2003; Kumaraguruparan et al. 2006; Nieto et al. 2003). Many of these cellular markers are used in diagnosis and prognosis of human breast cancers (Dickson et al. 2005). Furthermore, canine mammary tumors and human breast tumors have similar antigenic phenotypes (Mottolese et al. 1994). These similarities support the use of canine mammary tumors as a genetic model for human breast cancers.
In particular, the dog may prove an ideal model for the study of oncomirs. Because mammary cancer is a spontaneously occurring disease in the dog (as opposed to induced rodent models or manipulated in vitro models), the functional genetics can be explored more accurately. The genetic similarities found between the two species suggests conservation of function in that miRNAs may be targeting genes conserved between the human and dog to initiate tumorigenesis or metastasis. Presented here are the expression levels of oncomirs in canine mammary cancer using the highly sensitive qRT-PCR approach.
Materials and methods
Ten human precursor miRNA sequences (hsa-miR-15a, hsa-miR-16, hsa-miR-17-5p, hsa-miR-21, hsa-miR-29b, hsa-miR-125b, hsa-miR-145, hsa-miR-155, hsa-miR-181b, and hsa-let-7f) were obtained from the Sanger miRNA Registry (Griffiths-Jones et al. 2006). These precursor sequences were used to search for the homologous canine sequences using NCBI’s BLAST search program (http://www.ncbi.nlm.nih.gov/genome/seq/BlastGen/BlastGen.cgi?taxid=9615). Identity percentage was calculated by dividing the number of matched bases by the total number of bases in the human precursor. Mature percentages were calculated by dividing the number of matches at the mature sequence by the total number of bases in the mature sequence.
Six primary canine mammary tumors were collected and used in this study (Table 1). Surgical excision of all tumors followed routine surgical guidelines for neoplastic excision. Tumor samples were then stored in RNAlater ® (Ambion Inc., Austin, TX) immediately after surgical excision. Biopsy reports were available for five of the six tumors. Ten normal mammary tissue samples (Table 2) were collected from healthy adult female dogs that underwent an elective ovariohysterectomy (OHE) through Texas A&M University’s Veterinary Medical Teaching Hospital General Surgery Department. Harvesting of normal mammary tissue was approved by the university’s Clinical Research Review Committee (#07-02) and via a signed owner consent form. At the conclusion of the OHE, a 1-cm incision was made over the most prominent mammary gland. A sample of mammary tissue approximately 3 mm × 10 mm was collected and stored in RNAlater. Total RNA was isolated from all samples using the mirVana™ miRNA Isolation Kit (Ambion) following the manufacturer’s recommended protocol. Samples were evaluated for quality and quantity using the NanoDrop® ND-1000 Spectrophotometer (NanoDrop Technologies, Wilmington, DE).
TaqMan® MicroRNA Assays (hsa-miR-15a, hsa-miR-16, hsa-miR-17-5p, hsa-miR-21, hsa-miR-29b, hsa-miR-125b, hsa-miR-145, hsa-miR-155, hsa-miR-181b, and hsa-let-7f) were used to reverse transcribe ten mature miRNA sequences to cDNA (Applied Biosystems, Foster City, CA). Each 15-μl reverse transcription reaction contained 5 μl total RNA diluted to a 1.4-ng/μl concentration, 1 × TaqMan MicroRNA RT primer, 0.25 U RNase inhibitor, 1 × RT buffer, 3.33 U MultiScribe™ reverse transcriptase, and 1 mM dNTPs (ABI). The reverse transcription reactions were run in a BioRad thermocycler (Bio-Rad Laboratories, Hercules, CA) for 30 min at 16°C, 30 min at 42°C, 5 min at 85°C, and then held at 4°C. Controls included reactions with no template, no primer, and the RNU6B TaqMan assay (ABI), which codes for a small RNA that has been validated by ABI as an appropriate control.
The reverse transcription products were diluted 1:15. The 20-μl real-time reaction contained 1.33 μl diluted RT product, 1× TaqMan Universal PCR Master Mix (No AmpErase® UNG), and 1× TaqMan assay, which contained the forward and reverse primers as well as the TaqMan probe. Each reaction was run in triplicate on a 96-well plate. Reactions containing no reverse transcription products served as negative controls. Reactions were incubated in an ABI 7500 Prism Sequence Detection System for 10 min at 95°C, followed by 40 cycles of 95°C for 15 s and 60°C for 1 min.
After completion of the qRT-PCR, the threshold value was automatically configured above the baseline displayed in the amplification plot. Relative quantification of gene expression was evaluated by utilizing the comparative critical threshold (CT). The CT values for each mature miRNA reaction were subtracted from the respective CT value of the RNU6B control, resulting in the ∆CT value. The largest ∆CT value was arbitrarily used as a constant that was subtracted from all other ∆CT values to determine ∆∆CT value. Fold changes were then generated for each mature miRNA by calculating 2-∆∆CT. Expression levels of normal samples were compared to cancerous samples using a standard MANOVA.
Results
Bioinformatics data revealed the ten oncomirs of interest (miR-15a, miR-16, miR-17-5p, miR-21, miR-29b, miR-125b, miR-145, miR-155, miR-181b, let-7f) to have substantial sequence identity between human and dog at the precursor level and complete conservation at the mature level. Sequence identity is presented in Table 3. Based on previous research, the identity found at the mature level permits the use of the ABI TaqMan MicroRNA assays with canine input despite being designed for the corresponding miRNA genes of the human (Boggs et al. 2007). qRT-PCR was chosen as the method of choice to quantify miRNA expression due to its need for very little RNA input as well as the specificity and sensitivity of TaqMan.
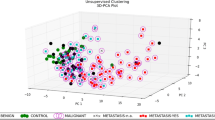
A brief comparison (Fig. 1) of the overall expression of the ten miRNAs studied with qRT-PCR and analyzed with the traditional ΔΔCT method reveals five miRNAs with tumor suppressor activities (miRs-15a, -16, -17-5p, -125b, and -155) and four with oncogenic activity (miRs-21, -29b, -181b, and let-7f). MiR-145 does not seem to differ in expression between normal and malignant mammary tissues. The statistical analysis to compute p values (listed in Table 4) was performed using the multivariate analysis of variance (MANOVA) calculation, a more robust test than the standard t test. MANOVA takes into account that all ten miRNAs were evaluated on the same six samples, and it revealed miR-21 and miR-29b to be significantly different (p ≤ 0.05). However, the biological trends cannot be ignored and are further addressed in the Discussion.
Mammary oncomir expression. Expression levels of ten oncomirs were studied from normal and cancerous canine breast tissue and compared using a MANOVA analysis. (a) Comparison of all six tumor samples versus ten normal samples. (b) Comparison of two tubular papillary carcinoma samples to the ten normals. (c) Comparison of three ductal carcinoma samples to the ten normal samples
Comparison of the samples based on two types of histology (ductal carcinomas and tubular papillary carcinomas) showed that miR-15b and miR-16 have a significantly lower expression in canine ductal carcinoma diagnoses versus normal samples. The ductal carcinomas were identified by neoplastic cells invading the mammary gland. Alternatively, tubular papillary carcinomas invaded the mammary glands, producing abnormal tubular projections into the duct. The tubular papillary tumors had a higher expression of let-7f and miR-181. This is in addition to miR-21 and miR-29b and indicates that these four genes may have oncogenic roles in this particular type of malignancy.
Discussion
It was not surprising to find that the oncomirs of interest are highly conserved between the dog and the human. Previous analysis of the two species revealed that the canine sequence, as a whole, shows great sequence identity to the human, even more than the popular mouse model (Kirkness et al. 2003). The physiology of the dog enables larger and easier-to-obtain specimen collections compared to rodent models. The companion animals also have extraordinary health care, second only to that of the human medicine. The vast similarities in both genetics and physiology and considerable overlap in treatment regimens make the dog an ideal model for many human diseases, including malignant breast cancer.
Like the human, the dog is known to have several risk factors for the development of mammary cancer. All canine mammary tumors appear to be promoted by sex hormones (Lana et al. 2007). Studies have shown that there is a direct correlation between the total amount of estrogen exposure and the risk for mammary tumor development. Dogs that are spayed before their first, second, or third estrus cycle have a risk of 0.5, 8, and 26%, respectively, for developing a mammary tumor (Schneider et al. 1969). This is intriguing because early-aged menopausal women have a significant decrease in risk for development of breast cancer compared to those women who reach menopause at biologically normal times (Dickson et al. 2005).
It must be noted that what is known about the role of estrogen receptors in canine mammary tumors indicates that they may not be good biological models of human mammary cancer. That is, while estrogen clearly promotes the growth of breast cancer in both species (Dickson et al. 2005; Schneider et al. 1969), multiple studies have shown that in true canine mammary malignancies, estrogen receptors are downregulated compared to normal mammary tissue (Illera et al. 2006; Millanta et al. 2005; Nieto et al. 2000). This finding is markedly different when compared to the majority of human postmenopausal breast cancer (Dickson et al. 2005). Consequently, estrogen receptor blockers such as tamoxifen have little effect in the dog. Interestingly, dysplasia and benign canine tumors are characterized by elevations in the estrogen receptors similar to women. Thus, the potential of dogs with mammary tumors to serve as models for study of human breast cancers is not without complication. Even so, the conflicting data pertaining to estrogen receptors is outweighed by the aforementioned shared characteristics between canine and human mammary cancers. These similarities suggest that while treatment response may vary due to the lack of estrogen receptors, the biological development and progression of mammary cancers in these two species is similar enough to enhance the understanding of human breast cancer through the use of the canine model. To this end, evaluation of miRNAs in canine mammary tissue may be a highly productive avenue of investigation.
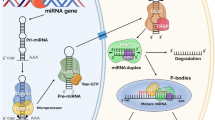
Previous work grouped oncomirs into three categories: tumor suppressors, oncogenes, and those that fulfill both functions (Calin and Croce 2006; Esquela-Kerscher and Slack 2006; Wijnhoven et al. 2007; Zhang et al. 2007). Because miRNAs can bind to more than one mRNA (Fig. 2), it is not only possible but more than likely that they will bind to several mRNAs. Likewise, the miRNAs studied in this article can be placed into one of the three categories.
MicroRNA biogenesis. The primary miRNA (pri-miRA) is transcribed from the genomic DNA. Drosha processes the pri-miRNA into a precursor miRNA (pre-miRNA) which is transported to the cytoplasm by Exportin-5. Dicer further cleaves the hairpin and leaves a short 18–22-nucleotide double-stranded RNA duplex. The RNA-induced silencing complex (RISC) then incorporates the mature strand of the miRNA and binds to the target mRNA in one of two fashions displayed above. The miRNA can then act as a tumor suppressor or oncogene dependent upon the mRNA it targets
Tumor suppressors
The first oncomirs, miRs-15a and 16, were initially discovered to play a role in chronic B-cell lymphocytic leukemia (Calin et al. 2004a). They are clustered together at a known cancer fragile site and act to suppress tumors by repressing BCL2, an anti-apoptotic oncogene (Cimmino et al. 2005). The loss of these two miRNAs results in increased expression of BCL2 and subsequent prosurvival states, a documented event in the majority of chronic cases of B-cell lymphocytic leukemia cases (Cimmino et al. 2005). Decreased expression and a loss in copy number of miR-15a and miR-16 were found in other hematopoetic and solid cancers, including human mammary cancers (Bottoni et al. 2005; Iorio et al. 2005; Mattie et al. 2006; Volinia et al. 2006). The work reported here provides evidence that both miR-15a and miR-16 are generally suppressed in canine mammary carcinomas. However, when the ductal carcinoma phenotype is singled out, the different patterns of expression become significant thereby suggesting that miR-15a and miR-16 may be responsible for the differentiation of the malignancy.
Another known tumor suppressor, miR-125b, also acts to control cell proliferation and regulates the oncogene Lin28 in the human (Lee et al. 2005; Wu and Belasco 2005). Expression of this miRNA in the dog is reduced in malignant samples compared to normal mammary tissues. This is in agreement with microarray, Northern blotting, and comparative genomic hybridization studies done on primary and in vitro samples from the human (Blenkiron et al. 2007; Iorio et al. 2005; Mattie et al. 2006; Volinia et al. 2006; Zhang et al. 2006).
Differential expression of miR-145 was not detected in canine mammary cancer, but it is described as a tumor suppressor and has been found to target FLJ21308 in the human (Kiriakidou et al. 2004). This oncomir is more often associated with colorectal tumors (Bandres et al. 2006), but studies have documented a differential expression in mammary cancer (Iorio et al. 2005; Volinia et al. 2006). It is interesting to note that one group described the expression of both the mature and precursor expression levels of miR-145 in colorectal tumors, but found a difference only at the mature level (Michael et al. 2003). This finding underscores the importance of choosing expression tools that target the “functional element” of miRNAs, i.e., the mature product.
Oncogenes
The oncogenes miR-21 and miR-29b are associated with numerous cancers, including human mammary cancer (Blenkiron et al. 2007; Esquela-Kerscher and Slack 2006; Fabbri et al. 2007; Frankel et al. 2007; Iorio et al. 2005; Mattie et al. 2006; Meng et al. 2007; Mott et al. 2007; Pekarsky et al. 2006; Si et al. 2007; Silveri et al. 2006; Volinia et al. 2006; Zhang et al. 2006; Zhu et al. 2007). Consistently, these two miRNAs are upregulated in analyses of mammary cancers using microarrays, and Zhang et al. (2006) confirmed that these two oncogenes showed an increase in copy number in mammary tumor tissues. In particular, miR-21 targets both PTEN and TPM1, and the loss of this miRNA results in increased caspase activity and subsequent apoptosis (Meng et al. 2007; Zhu et al. 2007). Tcl1 and MCL1 are targets of miR-29b (Mott et al. 2007; Pekarsky et al. 2006). Because these miRNAs have roles in so many cancers, it is reasonable to hypothesize that they have a general role in tumorigenesis rather than a specific role in metastasis or development of mammary cancers. Both miRNAs were significantly upregulated in tumors for this study (p ≤ 0.05). They remained significant even when the tubular papillary carcinoma phenotype was separately analyzed. Because of the increased expression, miR-21 and miR-29b may be ideal targets for in vitro knockdown studies.
Combination tumor suppressors and oncogenes
Some miRNAs serve opposing roles, specifically as both tumor suppressor and oncogene. This is the most interesting and complicated category of miRNAs, and as functional studies continue and more data are generated, it is likely that many more oncomirs will be found to have these dual roles.
Known to suppress the oncogene AIB1, miR-17-5p would be classified in the tumor suppressor category (Hossain et al. 2006). However, it is also known to target E2F1, the transcription factor for c-Myc (Hossain et al. 2006; O’Donnell et al. 2005). Previous work showed increased expression of miR-17-5p in several cancers, and this miRNA is known to specifically accelerate C-MYC-induced tumorigenesis in the mouse (He et al. 2005). Both Iorio et al. (2005) and Volina et al. (2006) used microarray data to suggest that miR-17-5p acts as an oncogene in mammary cancers while many other studies used a variety of techniques to show evidence of miR-17-5p acting as tumor suppressor (Hossain et al. 2006; Lu et al. 2005; O’Donnell et al. 2005; Zhang et al. 2006). In fact, this miRNA is deleted, due to its fragile site position, in 21.9% of breast cancer samples (Zhang et al. 2006). The data from the dog correlate with the majority of the findings when all mammary tumors and tubular papillary carcinomas were analyzed. There does not seem to be a difference in expression in the ductal carcinomas. Because the differences were not statistically significant, additional tumor samples would need to be collected to further investigate the role of miR-17-5p in canine mammary cancer.
In chronic lymphocytic leukemia, miR-181b targets Tcl1, another known oncogene, suggesting that the miRNA is a tumor suppressor (Pekarsky et al. 2006). Other work implicated the gene in differentiation of B cells (Chen and Lodish 2005). In human glioblastoma and pituitary adenomas, miR-181b seems to act in a tumor-suppression fashion (Bottoni et al. 2007; Ciafre et al. 2005). However, Zhang et al. (2006) reported an increased copy number of miR-181b in mammary tumors, and microarray studies showed overexpression of this miRNA in mammary cancer tissue samples. Overall, the canine mammary tumors appear to exhibit increased expression of miR-181b. In the tubular papillary carcinomas, the miRNA has significantly increased expression, indicating that miR-181b may be important in differentiation of mammary cells as well.
Located within the BIC gene, miR-155 is generally thought of as an oncogene because of its increased expression in hemopoietic cancers, thyroid cancers, and lung adenocarcinomas (Costinean et al. 2006; Mattie et al. 2006; Volinia et al. 2006; Zhang et al. 2006). Microarray data showed an increase in expression in multiple tumor samples of different origins, giving further evidence for this oncogenic role (Iorio et al. 2005; Volinia et al. 2006). However, a study of pancreatic cancer showed miR-155 to be downregulated (Roldo et al. 2006). In canine mammary cancers, the miRNA takes on both roles, dependent upon tumor differentiation. The average expression in tubular papillary carcinomas is lower compared to normal mammary tissue, while the ductal carcinomas display a slightly higher expression when compared to the healthy counterparts. It is important to note that only the averages are being compared because the difference of expression was not considered statistically significant. Interestingly, Mattie et al. (2006) found that miR-155 is differentially expressed when comparing estrogen receptor-positive and -negative tumors.
A part of the let-7 family of miRNAs, let-7f is described as a tumor suppressor because it knocks down the oncogene ras (Johnson et al. 2005). The let-7f miRNA is conserved in as distantly related species as the worm, plant, and fruitfly (Pasquinelli et al. 2000). By targeting K-ras and N-ras mRNA for degradation, let-7f’s role in cancer cell proliferation was clear (Johnson et al. 2007). In addition, multiple studies have shown a reduced expression of this miRNA in lung cancer (Jiang et al. 2005; Johnson et al. 2005, 2007). However, one study used the highly sensitive qRT-PCR and showed an upregulation of let-7f in breast cancer cell lines (Jiang et al. 2005). Only the tubular papillary carcinomas of canine mammary cancer showed a statistically significant difference in expression of let-7f, which supports the findings of the qRT-PCR study.
The data reported here were generated using a qRT-PCR approach that has proven specificity and sensitivity. This technique is particularly useful because there is very little RNA obtained from the mammary biopsies and the miRNAs being quantified differ by only one nucleotide. While Northern blots were not performed to confirm the data, previous research has shown the qRT-PCR approach to be more accurate. In addition, RNU6B was chosen as the “housekeeping gene” for two reasons: previous researchers have used the U6 control, making the data comparable, and the expression of RNU6B was found to be relatively consistent among several cancers (Jiang et al. 2005).
Relatively few samples were used in this study because of the difficulty in obtaining tumor and normal tissue from client-owned animals. This means that while most of the miRNAs did not show statistically significant differences, they still may play vital roles in cancer. The analysis of more tumor samples to improve the power of the study is required before any definitive statements on the role of miRNAs in canine mammary cancer are made. This point underscores the need for future collaborations between veterinarians and researchers as well as the establishment of tumor banks.
The fact that the dog appears to follow the trends described in previous literature on human breast cancer research is encouraging as it provides theory for future research that utilizes the canine species as a model. As more samples are collected and in vitro cell lines are established, more canine phenotypes can be explored and compared to the human forms of malignancy beyond just breast cancer. The cell-specific functions should be further investigated and predicted targets can be tested in cell lines derived from canine tumor samples. Elucidating the actions of miRNAs will advance cancer diagnostics, treatment, and survival by providing gene therapy targets of the future. Ideally, the miRNAs found to be involved in tumorigenesis or tumor suppression could serve as targets for gene therapy in both the dog and the human.
References
Ambros V (2004) The function of animal microRNAs. Nature 431:350–355
American Cancer Society (2007) Cancer facts and figures 2007. ACS, Atlanta, GA
Bandres E, Cubedo E, Agirre X, Malumbres R, Zarate R et al (2006) Identification by real-time PCR of 13 mature microRNAs differentially expressed in colorectal cancer and non-tumoral tissues. Mol Cancer 5:29
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Berezikov E, Guryev V, van de Belt J, Wienholds E, Plasterk RHA et al (2005) Phylogenetic shadowing and computational identification of human microRNA genes. Cell 120:21–24
Blenkiron C, Goldstein LD, Thorne NP, Spiteri I, Chin SF et al (2007) MicroRNA expression profiling of human breast cancer identifies new markers of tumour subtype. Genome Biol 8:R214
Boggs RM, Moody JA, Long CR, Tsai KL, Murphy KE (2007) Identification, amplification and characterization of miR-17–92 from canine tissue. Gene 404:25–30
Bottoni A, Piccin D, Tagliati F, Luchin A, Zatelli MC et al (2005) miR-15a and miR-16-1 down-regulation in pituitary adenomas. J Cell Physiol 204:280–285
Bottoni A, Zatelli MC, Ferracin M, Tagliati F, Piccin D et al (2007) Identification of differentially expressed microRNAs by microarray: a possible role for microRNA genes in pituitary adenomas. J Cell Physiol 210:370–377
Calin GA, Croce CM (2006) MicroRNA-cancer connection: the beginning of a new tale. Cancer Res 66:7390–7394
Calin GA, Liu CG, Sevignani C, Ferracin M, Felli N et al (2004a) MicroRNA profiling reveals distinct signatures in B cell chronic lymphocytic leukemias. Proc Natl Acad Sci USA 101:11755–11760
Calin GA, Sevignani C, Dumitru CD, Hyslop T, Noch E et al (2004b) Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc Natl Acad Sci USA 101:2999–3004
Chen CZ, Lodish HF (2005) MicroRNAs as regulators of mammalian hematopoiesis. Semin Immunol 17:155–165
Ciafre SA, Galardi S, Mangiola A, Ferracin M, Liu CG et al (2005) Extensive modulation of a set of microRNAs in primary glioblastoma. Biochem Biophys Res Commun 334:1351–1358
Cimmino A, Calin GA, Fabbri M, Iorio MV, Ferracin M et al (2005) miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc Natl Acad Sci USA 102:13944–13949
Costinean S, Zanesi N, Pekarsky Y, Tili E, Volinia S et al (2006) Pre-B cell proliferation and lymphoblastic leukemia/high-grade lymphoma in E(mu)-miR155 transgenic mice. Proc Natl Acad Sci USA 103:7024–7029
de las Mulas JM, Reymundo C, de los Monteros AE (2004) Calponin expression and myoepithelial cell differentiation in canine, feline and human mammary simple carcinomas. Vet Compar Oncol 2:24–35
Dickson RB, Pestell RG, Lippman ME (2005) Cancer of the breast. In: DeVita VT, Hellman S, Rosenberg S (eds) Cancer: principles and practice of oncology, 7th edn. Lippincott Williams & Wilkins, Philadelphia, pp 1399–1488
Dore M, Lanthier I, Sirois J (2003) Cyclooxygenase-2 expression in canine mammary tumors. Vet Pathol 40:207–212
Dorn CR, Taylor DO, Frye FL, Hibbard HH (1968) Survey of animal neoplasms in Alameda and Contra Costa Counties, California. I. Methodology and description of cases. J Natl Cancer Inst 40:295–305
Esquela-Kerscher A, Slack FJ (2006) Oncomirs—microRNAs with a role in cancer. Nat Rev Cancer 6:259–269
Fabbri M, Garzon R, Cimmino A, Liu Z, Zanesi N et al (2007) MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proc Natl Acad Sci USA 104:15805–15810
Frankel LB, Christoffersen NR, Jacobsen A, Lindow M, Krogh A et al (2007) Programmed cell death 4 (PDCD4) is an important functional target of the microRNA miR-21 in breast cancer cells. J Biol Chem 283:1026–1033
Griffiths-Jones S, Grocock RJ, Dongen Sv, Bateman A, Enright AJ (2006) miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res 34:D140–D144
He L, Thomson JM, Hemann MT, Hernando-Monge E, Mu D et al (2005) A microRNA polycistron as a potential human oncogene. Nature 435:828–833
Hossain A, Kuo MT, Saunders GF (2006) Mir-17–5p regulates breast cancer cell proliferation by inhibiting translation of AIB1 mRNA. Mol Cell Biol 26:8191–8201
Hwang H-W, Mendell JT (2006) MicroRNAs in cell proliferation, cell death, and tumorgenesis. Br J Cancer 94:776–780
Illera JC, Perez-Alenza MD, Nieto A, Jimenez MA, Silvan G et al (2006) Steroids and receptors in canine mammary cancer. Steroids 71:541–548
Iorio MV, Ferracin M, Liu CG, Veronese A, Spizzo R et al (2005) MicroRNA gene expression deregulation in human breast cancer. Cancer Res 65:7065–7070
Jiang J, Lee EJ, Gusev Y, Schmittgen TD (2005) Real-time expression profiling of microRNA precursors in human cancer cell lines. Nucleic Acids Res 33:5394–5403
Johnson CD, Esquela-Kerscher A, Stefani G, Byrom M, Kelnar K et al (2007) The let-7 microRNA represses cell proliferation pathways in human cells. Cancer Res 67:7713–7722
Johnson SM, Grosshans H, Shingara J, Byrom M, Jarvis R et al (2005) RAS is regulated by the let-7 microRNA family. Cell 120:635–647
Kiriakidou M, Nelson PT, Kouranov A, Fitziev P, Bouyioukos C et al (2004) A combined computational-experimental approach predicts human microRNA targets. Genes Dev 18:1165–1178
Kirkness EF, Bafna V, Halpern AL, Levy S, Remington K et al (2003) The dog genome: survey sequencing and comparative analysis. Science 301:1898–1903
Kumaraguruparan R, Prathiba D, Nagini S (2006) Of humans and canines: immunohistochemical analysis of PCNA, Bcl-2, p53, cytokeratin and ER in mammary tumours. Res Vet Sci 81:218–224
Lana SE, Rutteman GR, Withrow SJ (2007) Tumors of the mammary gland. In: Withrow SJ, Vail DM (eds) Withrow & MacEwen’s small animal clinical oncology. Saunders Elsevier, St. Louis, pp 619–636
Lee YS, Kim HK, Chung S, Kim KS, Dutta A (2005) Depletion of human micro-RNA miR-125b reveals that it is critical for the proliferation of differentiated cells but not for the down-regulation of putative targets during differentiation. J Biol Chem 280:16635–16641
Lewis BP, Burge CB, Bartel DP (2005) Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120:15–20
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J et al (2005) MicroRNA expression profiles classify human cancers. Nature 435:834–838
Mattie MD, Benz CC, Bowers J, Sensinger K, Wong L et al (2006) Optimized high-throughput microRNA expression profiling provides novel biomarker assessment of clinical prostate and breast cancer biopsies. Mol Cancer 5:24
Meng F, Henson R, Wehbe-Janek H, Ghoshal K, Jacob ST et al (2007) MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology 133:647–658
Michael MZ, O’Connor SM, van Holst Pellekaan NG, Young GP, James RJ (2003) Reduced accumulation of specific microRNAs in colorectal neoplasia. Mol Cancer Res 1:882–891
Millanta F, Calandrella M, Bari G, Niccolini M, Vannozzi I et al (2005) Comparison of steroid receptor expression in normal, dysplastic, and neoplastic canine and feline mammary tissues. Res Vet Sci 79:225–232
Mott JL, Kobayashi S, Bronk SF, Gores GJ (2007) mir-29 regulates Mcl-1 protein expression and apoptosis. Oncogene 26:6133–6140
Mottolese M, Morelli L, Agrimi U, Benevolo M, Sciarretta F et al (1994) Spontaneous canine mammary tumors. A model for monoclonal antibody diagnosis and treatment of human breast cancer. Lab Invest 71:182–187
Nieto A, Pena L, Perez-Alenza MD, Sanchez MA, Flores JM et al (2000) Immunohistologic detection of estrogen receptor alpha in canine mammary tumors: clinical and pathologic associations and prognostic significance. Vet Pathol 37:239–247
Nieto A, Perez-Alenza MD, Del Castillo N, Tabanera E, Castano M et al (2003) BRCA1 expression in canine mammary dysplasias and tumours: relationship with prognostic variables. J Comp Pathol 128:260–268
O’Donnell KA, Wentzel EA, Zeller KI, Dang CV, Mendell JT (2005) c-Myc-regulated microRNAs modulate E2F1 expression. Nature 435:839–843
Pasquinelli AE, Reinhart BJ, Slack F, Martindale MQ, Kuroda MI et al (2000) Conservation of the sequence and temporal expression of let-7 heterochronic regulatory RNA. Nature 408:86–89
Pekarsky Y, Santanam U, Cimmino A, Palamarchuk A, Efanov A et al (2006) Tcl1 expression in chronic lymphocytic leukemia is regulated by miR-29 and miR-181. Cancer Res 66:11590–11593
Philibert JC, Snyder PW, Glickman N, Glickman LT, Knapp DW et al (2003) Influence of host factors on survival in dogs with malignant mammary gland tumors. J Vet Intern Med 17:102–106
Roldo C, Missiaglia E, Hagan JP, Falconi M, Capelli P et al (2006) MicroRNA expression abnormalities in pancreatic endocrine and acinar tumors are associated with distinctive pathologic features and clinical behavior. J Clin Oncol 24:4677–4684
Schneider R, Dorn CR, Taylor DO (1969) Factors influencing canine mammary cancer development and postsurgical survival. J Natl Cancer Inst 43:1249–1261
Si ML, Zhu S, Wu H, Lu Z, Wu F et al (2007) miR-21-mediated tumor growth. Oncogene 26:2799–2803
Silveri L, Tilly G, Vilotte JL, Le Provost F (2006) MicroRNA involvement in mammary gland development and breast cancer. Reprod Nutr Dev 46:549–556
Simon D, Schoenrock D, Baumgartner W, Nolte I (2006) Postoperative adjuvant treatment of invasive malignant mammary gland tumors in dogs with doxorubicin and docetaxel. J Vet Intern Med 20:1184–1190
Volinia S, Calin GA, Liu CG, Ambs S, Cimmino A et al (2006) A microRNA expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci USA 103:2257–2261
Wijnhoven BP, Michael MZ, Watson DI (2007) MicroRNAs and cancer. Br J Surg 94:23–30
Wu L, Belasco JG (2005) Micro-RNA regulation of the mammalian lin-28 gene during neuronal differentiation of embryonal carcinoma cells. Mol Cell Biol 25:9198–9208
Zhang B, Pan X, Cobb GP, Anderson TA (2007) microRNAs as oncogenes and tumor suppressors. Dev Biol 302:1–12
Zhang L, Huang J, Yang N, Greshock J, Megraw MS et al (2006) microRNAs exhibit high frequency genomic alterations in human cancer. Proc Natl Acad Sci USA 103:9136–9141
Zhu S, Si ML, Wu H, Mo YY (2007) MicroRNA-21 targets the tumor suppressor gene tropomyosin 1 (TPM1). J Biol Chem 282:14328–14336
Acknowledgments
The authors thank the TAMU Statistics help desk for their aide and direction in the analysis portion of this project. Special thanks to Dr. Roy Pool for his help and interpretation of the biopsy reports used in this study.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Boggs, R.M., Wright, Z.M., Stickney, M.J. et al. MicroRNA expression in canine mammary cancer. Mamm Genome 19, 561–569 (2008). https://doi.org/10.1007/s00335-008-9128-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00335-008-9128-7