Abstract
Using lines of mice having undergone long-term selection for high and low growth, a large-sample (n = ~1,000 F2) experiment was conducted to gain further understanding of the genetic architecture of complex polygenic traits. Composite interval mapping on data from male F2 mice (n = 552) detected 50 QTL on 15 chromosomes impacting weights of various organ and adipose subcomponents of growth, including heart, liver, kidney, spleen, testis, and subcutaneous and epididymal fat depots. Nearly all aggregate growth QTL could be interpreted in terms of the organ and fat subcomponents measured. More than 25% of QTL detected map to MMU2, accentuating the relevance of this chromosome to growth and fatness in the context of this cross. Regions of MMU7, 15, and 17 also emerged as important obesity “hot-spots.” Average degrees of directional dominance are close to additivity, matching expectations for body composition traits. A strong QTL congruency is evident among heart, liver, kidney, and spleen weights. Liver and testis are organs whose genetic architectures are, respectively, most and least aligned with that for aggregate body weight. In this study, growth and body weight are interpreted in terms of organ subcomponents underlying the macro aggregate traits, and anchored on the corresponding genomic locations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
An estimated 65% of U.S. adults are overweight, and 31% are obese (Flegal et al. 2002). Overweight and obesity substantially increase the risk of hypertension, dyslipidemia, type 2 diabetes, coronary heart disease, stroke, gallbladder disease, osteoarthritis, sleep apnea and respiratory problems, and endometrial, breast, prostate, and colon cancers, combining to form the single largest cause of death in developed countries (NHLBI, 1998). While molecular biology has yielded significant advancements in understanding weight regulation at the metabolic and physiological levels, an alarmingly low number of cases of obesity in humans can be attributed to mutations within genes exerting effects in well-characterized, energy-balance pathways (Chagnon et al. 2003).
Carcass and body composition traits also constitute extremely important considerations of modern livestock production systems where consumer health concerns and marketing perspectives play increasingly prominent roles (Lasater and Kingsbery 1987; Smith 1991). The problem of excess fat in livestock and poultry carcasses is ubiquitous and has serious consequences for the animal industry at four levels: health perceptions of consumers; wasteful production of an undesired biological component; labor costs of trimming waste fat; and lower biological efficiencies of fatter animals (Eisen 1989).
Understanding the complex genetic architecture underlying quantitative trait variation for growth and body weight in mice has recently been advanced by use of crosses of large sample sizes, with an emphasis on localizing and defining the nature of QTL for these traits (e.g., Vaughn et al. 1999; Rocha et al. 2004a). While many QTL studies have focused on body fat as a biomedically and agriculturally relevant trait (e.g., reviewed by Pomp 1997; Brockmann and Bevova 2002), fewer investigations have evaluated components of body weight with a more global approach encompassing body fatness as well as organ weights and estimates of lean mass.
We created and evaluated a large sample from an F2 cross between lines of mice that had undergone long-term selection for rapid growth rate and low body weight to identify QTL for a variety of complex traits, including body weight and growth (Rocha et al. 2004a). In the present study we report on components of growth including weights of many organs and indicators of body fatness and leanness in males of the F2.
Materials and methods
Relevant information pertaining to the parental high growth (M16i) and low body weight (L6) selection lines, development of the F2 intercross population, and marker genotyping have been presented in a companion paper (Rocha et al. 2004a). Only methods relevant to the specific organ and body composition trait phenotypes evaluated and their statistical data analysis will be described here.
Phenotypes
Ten-week-old males (n = 552) from the F2 cross described in Rocha et al. (2004a) were weighed (WT10wk) and euthanized by cervical dislocation. Heart (HRT), liver (LIV), spleen (SPL), right kidney (KID), right testis (TES), right hindlimb subcutaneous fat pad (SCF), and right epididymal fat pad (EPF) were immediately excised and wet weights recorded. All organs were returned to the body cavity except for the spleen, which was used for subsequent DNA extraction. Gastrointestinal contents were flushed with tap water, blotted, and wet body weight (w) was recorded. The carcass was then lyophilized to a constant weight (d), and percentage water (PWAT), a proxy for leanness of the organism, was calculated as 100 × (w − d)/w. Two additional traits analyzed were FAT, the sum of SCF and EPF, and SCF/EPF, their ratio. Least squares means for related traits for the grandparental M16i and L6 lines were, respectively, 57.9 vs. 18.4 g for WT10wk, 7.92 vs. 6.00% for LIV, 0.23 vs. 0.39% for TES, and 0.80 vs. 0.67% for EPF, with weights of organs expressed as percentages of WT10wk.
Data analyses
Descriptive statistics and phenotypic correlations for organ and adipose traits are presented in Tables 1 and 2. Appropriate statistical models were identified for each trait by fitting generalized linear models (PROC GLM; SAS Institute Inc. 1985, 1996), including fixed effects of replicate/parity, full-sib family/litter, and respective interactions, if significant. Residuals were analyzed with QTL Cartographer (Version 1.15; Basten et al. 2001) to perform composite interval mapping (CIM; Zeng 1993, 1994). With the exception of SCF/EPF, logarithmic transformation was employed prior to composite interval mapping to correct patterns of heteroscedasticity. Day of tissue harvest was a significant source of phenotypic variation and was included in the statistical models as a nested term within replicate. Interestingly, animals born in litters from the second parity of F1 dams tended to exhibit heavier organs but were leaner, an effect that was particularly noticeable for the subcutaneous fat pad (data not shown).
QTL analyses were also performed with three sets of covariate adjustments: 1) WT10wk; 2) FAT; and 3) both WT10wk and FAT. The purpose of these adjustments was to attempt to identify alternate sets of QTL involved in different pathways responsible for the fluctuating patterns of phenotypic and genetic correlations observed among the various traits. In the body weight-adjusted models, relationships of organ and fat weights with WT10wk were always linear and positive (as also evident from phenotypic correlations). Estimates of regression coefficients for WT10wk were negative for PWAT and SCF/EPF. The former was expected, since heavier animals are fatter and PWAT is a proxy for leanness. The latter result may indicate that increased body weight leads to higher rate of deposition of epididymal versus subcutaneous fat. For the FAT-adjusted models, quadratic terms were used in some cases owing to minor deviations from linearity. The sign of the regression coefficient for FAT changed from positive to negative when WT10wk was also used as a covariate, indicating that at a constant body weight, fatter animals tended to have smaller organ weights.
The threshold adopted in the forward-backward stepwise regression factor-selection procedures for CIM was 0.001 for statistical models not including adjustment for WT10wk, and 0.01 when such adjustments were included. A 10-cM window size was used. Permutations were performed for organ and fat traits, and when similar results were observed for all traits, their permutations were pooled, leading to adoption of LOD = 3.3 for the 5% genomewise significance threshold (Churchill and Doerge 1994; see Rocha et al. 2003a).
Results
A summary of all identified QTL and related QTL detected by other studies in the same genomic regions are presented in Table 3. To avoid duplication, results from models including adjustment for WT10wk (designated as HRT%, KID%, etc.; Table 3) are presented only when they identified QTL not detected from the unadjusted models. Models including adjustment for FAT identified QTL results similar to those found when not including such adjustments, so they will not be considered further. Results for the composite trait FAT (sum of SCF and EPF) simply reproduced QTL that were detected for the sub-components (subcutaneous and epididymal fat pads), so they will not be reported. Furthermore, no QTL were detected for SCF/EPF, or for WT10wk once this trait was adjusted for all organ weights.
A total of 50 significant QTL were detected, and each was assigned a locus symbol following MGD guidelines (Table 3). QTL (n) were detected for HRT (3), TES (3), LIV (10), KID (7), SPL (9), SCF (4), EPF (5), and PWAT (1). Additional QTL (n) were detected when traits were adjusted for WT10wk, including HRT% (1), KID% (2), SCF% (1),and EPF% (4). For unadjusted traits, the vast majority of QTL exhibited additive gene action with the M16i allele1 increasing organ and fat pad weights. For traits adjusted for WT10wk, M16i alleles had both positive and negative additive effects depending upon the locus (see Table 3).
Tables 4 and 5 summarize QTL information across traits, while Table 2 presents the QTL-congruencies [QTL common to two traits (overlapping confidence intervals) as a proportion of the total number of QTL detected for the two traits] in relation to the respective phenotypic correlations. Large QTL-congruencies among heart, liver, kidney, and spleen weights are particularly evident, in agreement with phenotypic correlations. Liver weight appears to be controlled by a genetic architecture that is most aligned with body weight, while the reverse is true for testis weight, where no QTL congruencies were found with any other trait.
QTL for most organ and all fat pad weights display a prevalence for additive gene action. Testis and spleen weights, as might be expected given their respective roles in reproduction and disease resistance, both exhibited relatively strong directional dominance. The limited explorations of epistasis that were conducted (see Rocha et al. 2004a for methods) revealed little evidence for interactions among QTL affecting individual organ and fat pad weights.
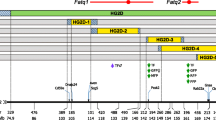
Nearly 25% (12) of detected QTL are located on Chromosome (Chr) 2, reflecting the large effects detected for aggregate body weight (Rocha et al. 2004a) and reproduction (Rocha et al. 2004c) traits. Likelihood plots for Chr 2 QTL for fat traits are provided in Fig. 1. QTL peaks for fat traits appear to match the peak representing the more distal QTL for WT10wk (Rocha et al. 2004a). This potentially pleiotropic locus likely reflects patterns of fat deposition with increasing maturity and the strong correlation between weight and fat in mice.
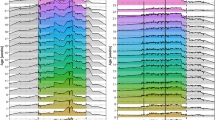
Fitting of alternative statistical models, with or without covariate adjustments, provides in some cases indirect evidence for existence of multiple linked QTL for the same trait. One such example for the case of potentially two QTL for SPL detected on MMU1 is illustrated in Fig. 2. This plot could represent a statistical artifact from the fitting of simple QTL models. However, the fact that adjustment for body weight differentially impacts the two putative and linked QTL lends some support to the possibility that these are in fact two distinct yet linked QTL, one impacting SPL independent of body weight. Formal tests comparing two QTL models relative to single QTL models will be employed in the future across this entire data set.
Figures 3 and 4 schematically summarize the genomic bases of components of 10-week body weight. Only rarely were QTL detected for organ and fat traits that did not overlap with the confidence intervals of QTL that had previously been detected for aggregate growth traits (Rocha et al. 2004a). The unique relevance of Chr 2 in the context of this cross is further demonstrated.
Chromosomes responsible for 10-week body weight, as assessed from the results of Rocha et al. (2004a), and corresponding assignments in terms of organ and fat subcomponents based on overlapping confidence intervals. In some cases, unequal relative areas represent different numbers of QTL detected for the different traits. On MMU1, two QTL for spleen were found, while other organs are represented by a single QTL. On MMU2 all organs are represented by two QTL except for SCF (subcutaneous fat), for which only one QTL was localized.
A) relative numbers of QTL associated with each of the subcomponent organ and fat traits (under the broad general assumption that body composition evaluated at 10 weeks harbors relevant information regarding body composition changes in each growth period); and B), relative contributions of each chromosome to the total growth process in terms of QTL numbers for all organs.
Discussion
Obesity exacts an immense economic and social toll, accounting for ~7% of US health expenditures and leading to potential bias and discrimination in employment, education, and health care (NHLBI 1998). It is an exceptionally complex disorder, with relatively equal contributions of multifactorial genetic susceptibility and interacting environmental factors (Pomp 1997). Predisposition to obesity results from combinations of relatively small effects of genetic variations within a large number of polygenes (QTL; Brockmann and Bevova 2002) of yet unidentified function.
The present results, in combination with a concurrent analysis of growth and body weight (Rocha et al. 2004a), have added new information to the predisposition map for mouse body weight regulation, and have further partitioned body weight QTL in terms of adipose tissue and weights of various organs. The vast majority of growth QTL, impacting different age periods, can be interpreted in terms of fat and organ subcomponents measured. Muscle, bone, gut, skin, and various small organs were not measured in this experiment. Many studies (Belknap et al. 1992; Brockmann et al. 2000; Williams 2000; Airey et al. 2001; Williams et al. 2001; Leamy et al. 2002) have identified QTL for several of these unmeasured subcomponents which match some of the growth QTL identified in the present study that have unknown underlying subcomponents. Additionally, only a few QTL (and primarily those with small effects) detected for organ subcomponents in this study did not overlap with confidence intervals of QTL that were detected for the aggregate growth traits (Rocha et al. 2004a).
Our results strongly re-emphasize the observation of Mehrabian et al. (1998) regarding commonality of genomic regions impacting obesity traits across many different genetic backgrounds. The QTL for fatness we report on Chr 2, 7, 15, and 17 appear to be obesity genomic hot-spots, although caution is required when assuming that the underlying genes in each region are identical in each genetic background.
Large effects of QTL impacting aggregate growth of the organism, growth of most organs, and especially fatness are present on mouse Chr 2. In regard to obesity predisposition, distal Chr 2 appears to be one of the most relevant regions of the mouse genome. Not only is this region well populated with multiple body weight and fatness QTL as determined from crosses employing different approaches and genetic backgrounds (Lembertas et al. 1997; Pomp 1997; Mehrabian et al. 1998; Schadt et al. 2003a), the QTL harbored in this region in the M16 selection line appear to have among the largest effects of any body weight regulating polygenes ever localized. Since distal MMU2 and HSA20 (from p13 through q13) form continuous stretches of synteny between the two species, identification of the genes underlying obesity QTL in this region of the mouse represents an excellent opportunity for comparative mapping and application of murine models to studies of human health. The importance of this is amplified by the findings from several groups that the homologous region of HSA20 harbors genes influencing predisposition to obesity and diabetes in humans (see Chagnon et al. 2003).
Efforts to identify and characterize the M16-based distal MMU2 weight and fat QTL are under way (Jerez-Timaure et al. 2002; Pomp et al. 2002). Our detailed studies in this region using M16i and an M16i.B6-(D2Mit306–D2Mit52) congenic line (referred to as MB2) strongly indicate the presence of multiple linked QTL with effects on weight and fatness (Jerez-Timaure and Pomp, unpublished). Efforts at positional cloning are further complicated by the fact that this region of the mouse genome is known to contain many genes involved in regulation of energy balance. For these reasons, success in this endeavor will likely require experimental approaches that combine functional analyses with the resolving power of recombination.
A new paradigm for bridging the gap between our knowledge of the physiology and predisposition of obesity is to combine QTL mapping with large-scale gene expression analysis. Transcriptome mapping, also called “genetical genomics” (Jansen and Nap 2001; Jansen 2003), treats levels of gene expression of any particular gene measured across different individuals as an expression-level polymorphism that in principle reflects the underlying genetic variation (Dumas et al. 2000; Jansen and Nap 2001; Doerge 2002). This type of analysis was pioneered by de Vienne et al. (1994), using proteomic evaluation [later extended by the same group to the transcriptome (Consoli et al. 2002)] in an F2 population of maize. Transcriptome mapping is a powerful mechanism to dissect complex traits and make more efficient the selection of candidate genes underlying predisposition loci, with recent successful implementation in yeast (Brem et al. 2002), Drosophila (Wayne and McIntyre 2002), and mice (Schadt et al. 2003a, 2003b).
By identifying expression QTL that coincided with a QTL peak for, and had correlations with, subcutaneous fat mass, Schadt et al. (2003a) proposed the NM_025575 and NM_15731 genes as primary candidates for the distal MMU2 QTL for dietary-induced obesity described by Lembertas et al. (1977). These candidate genes also possibly overlap with the obesity QTL we identified in this region. Important distinctions exist between the results from Schadt et al. (2003a) and those from the present experiment. Distinctly different genetic models were used. Obesity in the M16 line does not require high levels of dietary fat. Indeed, no interactions between dietary fat consumption and effects of MMU2 QTL on body fat levels were found when comparing the M16i selection line and MB2. This lack of interaction is noteworthy given that the donor line for MB2 is C57, which is one of the strains used in the Schadt et al. (2003a) study, and which is among the strains most responsive to dietary-induced obesity (West et al. 1992). Nevertheless, localization of candidate genes identified by Schadt et al. (2003a) at ~165 MB on distal MMU2 aligns with the QTL peak for the Epfq2 locus, representing the more distal of the two obesity QTL found in our study and the one with significantly smaller effects. However, preliminary data from our subsequent fine-mapping efforts with MB2 indicate that the best supported position for Epfq2 may be at ~159 MB (Jerez-Timaure and Pomp, unpublished).
QTL for specific organ weights that support findings from independent studies are those for testis weight on the proximal region of MMU13, spleen weight on MMU1 (more proximal of the two detected), and kidney weight on MMU11. In this respect, MMU11 is second in importance to MMU2 in regard to effects on components of body weight in the context of this cross. Surprisingly, the only QTL detected for percentage water in the carcass did not match any of the QTL detected for fat traits, which may reflect the different nature of the two phenotypes. PWAT exhibits much less variability than SCF or EPF and is impacted not only by fat content but also by protein and mineral content. Nevertheless, results from other studies provide some support for the existence of a QTL for PWAT on MMU1, through both an impact on fat content (Purcell-Huynh et al. 1995; Machleder et al. 1997; Moody et al. 1999; Suto et al. 1999; Taylor et al. 2001) and/or mineral content (Leamy et al. 2002).
Liver and testis are those organs whose weights have the largest and the smallest numbers of QTL, respectively. Liver weight is the most aligned with growth of the overall organism, while testis weight is the least aligned with overall growth. Strong QTL congruencies exist among liver, heart, kidney, and spleen weights, in agreement with phenotypic correlations and paralleling results reported by other studies that have undertaken detailed organ subcomponent analyses (Kramer et al. 1998; Brockmann et al. 2000; Leamy et al. 2002). The latter two studies, which included comprehensive QTL analyses for organ and fat subcomponents, corroborate many of our present findings. Such agreement is not surprising, especially in regard to Leamy et al. (2002) and Pomp (1997), who analyzed data from a cross that also utilized the M16i selection line.
An interesting finding from this study concerns how different covariate adjustments may in some cases provide strong evidence supporting the existence of multiple linked QTL. Average additive QTL effects and average variance QTL effects for body composition traits are relatively larger than those reported for the aggregate growth traits (Rocha et al. 2004a). This may represent differences in genetic architecture of the traits, but more likely reflects the smaller sample size in the body composition analysis, which will tend to lead to estimates having an upward bias (Beavis 1998; Melchinger et al. 1998; Utz et al. 2000).
Directional dominance is not expected to play much of a role for the organ and fat traits studied at 10 weeks of age, as heterosis observed for corresponding traits in livestock species is usually minimal. Estimates of dominance effects match that expectation, with small average values reflecting predominantly additive gene action. Testis and spleen weights provide exceptions. This would perhaps be expected for testis weight, given the classical relevance of dominance and heterosis for reproductive (fitness) traits. Regarding spleen weight, the high average dominance values may be related to the involvement of this organ in disease-resistance mechanisms, which are traits for which dominance and heterosis are generally important.
Comparisons of the total amount of variance accounted for by all QTL, when fitted together, and the sum of the individual fractions of variance explained by individual QTL provide limited evidence about the relevance of epistatic interactions on phenotypic expression of these traits (i.e., with no epistasis, the two values should coincide). On the basis of this approach, organ and fat subcomponents of body weight appear to be traits for which epistasis is especially not important. However, validation of this finding requires more thorough statistical approaches (i.e., fitting of multiple QTL models).
While molecular biology has yielded advancements in understanding weight regulation at the metabolic and physiological levels [see reviews by Bray and Tartaglia (2000) and Barsh and Schwartz (2002)], the genetic architecture of obesity remains essentially undefined. And while QTL maps for obesity-related traits are well represented in both mice (Brockmann and Bevova 2002) and humans (Chagnon et al 2003), only a few isolated cases of obesity in humans can be attributed to mutations within genes exerting effects in well-characterized, energy-balance pathways (Chagnon et al. 2003). This gap between our knowledge of physiological mechanisms underlying obesity and the nature of genetic predisposition to obesity greatly impairs the ability to identify relevant QTL, which in turn limits development of gene-based tools for diagnosing, treating, and eventually preventing obesity and related diseases. Our future objective is to use the M16 obesity model to integrate genetics and physiology and resolve the macro level of phenotypic expression in terms of successive subcomponent layers (e.g., de Vienne et al. 1994; Flint and Mott 2001; Schadt et al. 2003a, 2003b) to the level of integrated and interacting pathways of biochemical networks. When this approach is anchored with a sound understanding of the corresponding genomic organization responsible for the operation of physiological systems, resolution at the level of single genes or integrated sets of interacting genes may become more readily attainable.
References
DC Airey L Lu RW Williams (2001) ArticleTitleGenetic control of the mouse cerebellum: identification of quantitative trait loci modulating size and architecture. J Neurosc 21 5099–5109 Occurrence Handle1:CAS:528:DC%2BD3MXkvFSrsrw%3D
GS Barsh MW Schwartz (2002) ArticleTitleGenetic approaches to studying energy balance: perception and integration. Nat Rev Genet 3 589–600 Occurrence Handle1:CAS:528:DC%2BD38XlslamsLo%3D Occurrence Handle12154382
CJ Basten BS Weir Z-B Zeng (2001) QTL Cartograher, version 1.15 North Carolina State University Raleigh, NC
WD Beavis (1998) Power, precision, and accuracy. AH Paterson (Eds) Molecular Dissection of Complex Traits. CRC Press Boca Raton, Fla 145–162
JK Belknap TJ Phillips LA O’Toole (1992) ArticleTitleQuantitative trait loci associated with brain weight in the BXD/Ty recombinant inbred mouse strains. Brain Res Bull 29 337–344 Occurrence Handle10.1016/0361-9230(92)90065-6 Occurrence Handle1:STN:280:ByyD3cjhs1w%3D Occurrence Handle1393606
GA Bray LA Tartaglia (2000) ArticleTitleMedicinal strategies in the treatment of obesity. Nature 404 672–677 Occurrence Handle1:CAS:528:DC%2BD3cXis1Gruro%3D Occurrence Handle10766254
RB Brem G Yvert R Clinton L Kruglyak (2002) ArticleTitleGenetic dissection of transcriptional regulation in budding yeast. Science 296 752–755 Occurrence Handle10.1126/science.1069516 Occurrence Handle1:CAS:528:DC%2BD38XjtlOnt70%3D Occurrence Handle11923494
GA Brockmann MR Bevova (2002) ArticleTitleUsing mouse models to dissect the genetics of obesity. Trends Genet 18 367–376 Occurrence Handle10.1016/S0168-9525(02)02703-8 Occurrence Handle1:CAS:528:DC%2BD38XlsVOlsrY%3D Occurrence Handle12127777
GA Brockmann U Renne K Kopplow P Das (1998a) ArticleTitleGenetic markers for the detection of quantitative trait loci with special consideration of body weight and fat. Acta Theriol 43 IssueIDSuppl 5 53–62
GA Brockmann CS Haley U Renne SA Knott M Schwerin (1998b) ArticleTitleQuantitative trait loci affecting body weight and fatness from a mouse line selected for extreme high growth. Genetics 150 369–381 Occurrence Handle1:CAS:528:DyaK1cXmtFKgsLc%3D
GA Brockmann J Kratzsch CS Haley U Renne M Schwerin et al. (2000) ArticleTitleSingle QTL effects, epistasis, and pleiotropy account for two-thirds of the phenotypic F2 variance of growth and obesity in DU6i × DBA/2 mice. Genome Res 10 1941–1957 Occurrence Handle11116089
YC Chagnon T Rankinen EE Snyder SJ Weisnagel L Perusse et al. (2003) ArticleTitleThe human obesity gene map: the 2002 update. Obes Res 11 313–367 Occurrence Handle1:CAS:528:DC%2BD3sXisVKgtLY%3D Occurrence Handle12634430
GA Churchill RW Doerge (1994) ArticleTitleEmpirical threshold values for quantitative trait mapping. Genetics 138 963–971 Occurrence Handle1:STN:280:ByqC2MvntFw%3D Occurrence Handle7851788
L Consoli A Lefevre M Zivy D de Vienne C Damerval (2002) ArticleTitleQTL analysis of proteome and transcriptome variations for dissecting the genetic architecture of complex traits in maize. Plant Mol Biol 48 575–581 Occurrence Handle10.1023/A:1014840810203 Occurrence Handle1:CAS:528:DC%2BD38XjsFWrtro%3D Occurrence Handle11999835
D de Vienne A Maurice JM Josse A Leonardi C Damerval (1994) ArticleTitleMapping factors controlling genetic expression. Cell Mol Biol (Noisy-le-Grand) 40 29–39 Occurrence Handle1:CAS:528:DyaK2cXmsleisLw%3D
RW Doerge (2002) ArticleTitleMapping and analysis of quantitative trait loci in experimental populations. Nat Rev Genet 3 43–52 Occurrence Handle1:CAS:528:DC%2BD38XhsV2gsbY%3D Occurrence Handle11823790
P Dumas Y Sun G Corbell S Trenblay Z Pausova et al. (2000) ArticleTitleMapping of quantitative trait loci (QTL) of differential stress gene expression in rat recombinant inbred strains. Hypertens 18 545–551 Occurrence Handle10.1097/00004872-200018050-00006 Occurrence Handle1:CAS:528:DC%2BD3cXktVOiuro%3D Occurrence Handle10826556
EJ Eisen (1989) ArticleTitleSelection experiments for body composition in mice and rats: a review. Livest Prod Sci 23 17–32 Occurrence Handle10.1016/0301-6226(89)90003-1
DS Falconer TF Mackay (1996) Introduction to Quantitative Genetics. Longman Group Ltd. Harlow, UK
JS Fisler CH Warden (1997) ArticleTitleMapping of mouse obesity genes: a generic approach to a complex trait. J Nutr 127 IssueIDSuppl 1909S–1916S Occurrence Handle1:CAS:528:DyaK2sXlvFGltLo%3D Occurrence Handle9278581
KM Flegal MD Carroll CL Ogden CL Johnson (2002) ArticleTitlePrevalence and trends in obesity among US adults, 1999–2000. JAMA (J am Med Assoc) 288 1723–1727 Occurrence Handle10.1001/jama.288.14.1723
J Flint R Mott (2001) ArticleTitleFinding the molecular basis of quantitative traits: successes and pitfalls. Nat Rev Genet 2 437–445 Occurrence Handle10.1038/35076585 Occurrence Handle1:CAS:528:DC%2BD3MXktlKqurY%3D Occurrence Handle11389460
L Gu MW Johnson AJ Lusis (1999) ArticleTitleQuantitative trait locus analysis of plasma lipoprotein levels in an autoimmune mouse model. Arterioscler Thromb Vase Biol 19 442–453 Occurrence Handle1:CAS:528:DyaK1MXhsFSjur8%3D
S Horvat L Bunger VM Falconer P Mackay A Law et al. (2000) ArticleTitleMapping of obesity QTLs in a cross between mouse lines divergently selected on fat content. Mamm Genome 11 2–7 Occurrence Handle1:CAS:528:DC%2BD3cXos12iug%3D%3D Occurrence Handle10602985
OA Lakoubova H Dushkin DR Beier (1995) ArticleTitleLocalization of a murine recessive polycystic kidney disease mutation and modifying loci that affect disease severity. Genomics 26 107–114 Occurrence Handle10.1016/0888-7543(95)80088-4 Occurrence Handle7782068
RC Jansen (2003) ArticleTitleStudying complex biological systems using multifactorial perturbation. Nat Rev Genet 4 145–151 Occurrence Handle10.1038/nrg996 Occurrence Handle1:CAS:528:DC%2BD3sXnsFaruw%3D%3D Occurrence Handle12560811
RC Jansen JP Nap (2001) ArticleTitleGenetical genomics: the added value from segregation. Trends Genet 17 388–391 Occurrence Handle10.1016/S0168-9525(01)02310-1 Occurrence Handle1:CAS:528:DC%2BD3MXksVyntLo%3D Occurrence Handle11418218
N Jerez-Timaure D Pomp EJ Eisen (2002) ArticleTitleConfirmation of quantitative trait loci (QTL) affecting body weight and fatness in a congenic line of mice. J Anim Sci 80 IssueIDSuppl 2 45 Occurrence Handle11833536
PD Keightley KH Morris A Ishikawa VM Falconer F Oliver (1998) ArticleTitleTest of candidate gene-quantitative trait locus association applied to fatness in mice. Heredity 81 630–637 Occurrence Handle10.1038/sj.hdy.6884500 Occurrence Handle9885188
MG Kramer TT Vaughn LS Pletscher KK Ellison E Adams et al. (1998) ArticleTitleGenetic variation in body weight gain and composition in the intercross of Large (LG/J) and Small (SM/J) inbred strains of mice. Genet Mol Biol 21 211–218
L Lasater B Kingsbery (1987) The New Cattle Industry, 1st ed. Kingsbery Communications Woodinville, Washington
LJ Leamy D Pomp EJ Eisen JM Cheverud (2002) ArticleTitlePleiotropy of quantitative trait loci for organ weights and limb bone lengths in mice. Physiol Genomics 10 21–29 Occurrence Handle1:CAS:528:DC%2BD38XmsFaitrY%3D Occurrence Handle12118102
AV Lembertas L Perusse YC Chagnon JS Fisler CH Warden et al. (1997) ArticleTitleIdentification of an obesity quantitative trait locus on mouse chromosome 2 and evidence of linkage to body fat and insulin on the human homologous region 20q. J Clin Invest 100 1240–1247 Occurrence Handle1:CAS:528:DyaK2sXlvFOgsbY%3D Occurrence Handle9276742
I Le Roy S Tordjman DM Samour H Degrelle PL Roubertoux (2001) ArticleTitleGenetic architecture of testis and seminal vesicle weights in mice. Genetics 158 333–340 Occurrence Handle1:CAS:528:DC%2BD3MXkt1Ojsbo%3D Occurrence Handle11333241
D Machleder B Ivandic C Welch L Castellani K Reue et al. (1997) ArticleTitleComplex genetic control of HDL levels in mice in response to an atherogenic diet. J Clin Invest 99 1406–1419 Occurrence Handle1:CAS:528:DyaK2sXitVWktLo%3D Occurrence Handle9077551
M Mehrabian P-Z Wen J Fisler RC Davis AJ Lusis (1998) ArticleTitleGenetic loci controlling body fat, lipoprotein metabolism, and insulin levels in a multifactorial mouse model. J Clin Invest 101 2485–2496 Occurrence Handle1:CAS:528:DyaK1cXjvVajsbs%3D Occurrence Handle9616220
M Mehrabian LW Castellani P-Z Wen J Wong T Rithaporn et al. (2000) ArticleTitleGenetic control of HDL levels and composition in an interspecific mouse cross (CAST/Ei × C57BL/6J). J Lipid Res 41 1936–1946 Occurrence Handle1:CAS:528:DC%2BD3MXhsVOktw%3D%3D Occurrence Handle11108726
AE Melchinger HF Utz CC Schon (1998) ArticleTitleQuantitative trait locus (QTL) mapping using different testers and independent population samples in maize reveals low power of QTL detection and large bias in estimates of QTL effects. Genetics 149 383–403 Occurrence Handle1:CAS:528:DyaK1cXks1ektb8%3D Occurrence Handle9584111
DE Moody D Pomp MK Nielsen LD Van Vleck (1999) ArticleTitleIdentification of quantitative trait loci influencing traits related to energy balance in selection and inbred lines of mice. Genetics 152 699–711
NHLBI (1998) Clinical Guidelines on the Identification, Evaluation, and Treatment of Overweight and Obesity in Adults: The Evidence Report. NIH Publication No. 98-4083
D Pomp (1997) ArticleTitleGenetic dissection of obesity in polygenic animal models. Behav Genet 27 285–306 Occurrence Handle10.1023/A:1025631813018 Occurrence Handle1:STN:280:DyaK1c7otVSrtQ%3D%3D Occurrence Handle9519558
D Pomp N Jerez-Timaure MF Allan EJ Eisen (2002) ArticleTitleIntegrated genomic, proteomic and metabolomic dissection of polygenic selection response for murine growth and fatness. Proc 7th World Congr Genet Appl Livest Prod 32 447–450
DA Purcell-Huynh A Weinreb LW Castellani M Mehrabian MH Doolittle et al. (1995) ArticleTitleAnalysis of a genetic cross between inbred mouse strains NZB/BINJ and SM/J using a complete linkage map approach. J Clin Invest 96 1845–1858 Occurrence Handle1:CAS:528:DyaK2MXosFyhtro%3D Occurrence Handle7560076
AW Roberts M Hasegawa D Metcalf SJ Foote (2000) ArticleTitleIdentification of a genetic locus modulating splenomegaly induced by granulocyte colony-stimulating factor in mice. Leukemia 14 657–661 Occurrence Handle10.1038/sj.leu.2401735 Occurrence Handle1:CAS:528:DC%2BD3cXivFOhu74%3D Occurrence Handle10764151
JL Rocha EJ Eisen LD Van Vleck D Pomp (2004a) ArticleTitleA large-sample QTL study in mice: I. Growth. Mamm Genome 15 83–99
JL Rocha F Siewerdt EJ Eisen LD Van Vleck D Pomp (2004c) ArticleTitleA large-sample QTL study in mice: III. Reproduction. Mamm Genome . .
InstitutionalAuthorNameSAS Institute Inc. (1985) SAS User’s Guide: Basics. SAS Cary, N.C.
InstitutionalAuthorNameSAS Institute Inc. (1996) SAS System for Mixed Models SAS Cary, N.C.
EE Schadt SA Monks TA Drake AJ Lusis N Che et al. (2003a) ArticleTitleGenetics of gene expression surveyed in maize, mouse and man. Nature 422 297–302 Occurrence Handle1:CAS:528:DC%2BD3sXitFKmuro%3D
EE Schadt SA Monks SH Friend (2003b) ArticleTitleA new paradigm for drug discovery: integrating clinical, genetic, genomic and molecular phenotype data to identify drug targets. Biochem Soc Trans 31 437–443 Occurrence Handle1:CAS:528:DC%2BD3sXisFGjt7k%3D
GC Smith (1991) Responding to a changing consumer: repositioning beef in the diet—safety, nutrient content and palatability. The King Ranch and the Texas A&M University System: Partners in Progress since 1891 Texas A&M University College Station, TX M1–M11
JL Spearow PA Nutson WS Mailliard M Porter M Barkley (1999) ArticleTitleMapping genes that control hormone-induced ovulation rate in mice. Biol Reprod 61 857–872 Occurrence Handle1:CAS:528:DyaK1MXmtlajsLk%3D Occurrence Handle10491617
J Suto S Matsuura H Yamanaka K Sekikawa (1999) ArticleTitleQuantitative trait loci that regulate plasma lipid concentration in hereditary obese KK and KK-Ay mice. Biochim Biophys Acta 1453 385–395 Occurrence Handle10.1016/S0925-4439(99)00013-7 Occurrence Handle1:CAS:528:DyaK1MXitFWrsLY%3D Occurrence Handle10101257
BA Taylor SJ Phillips (1996) ArticleTitleDetection of obesity QTLs on mouse chromosomes 1 and 7 by selective DNA pooling. Genomics 34 389–398
BA Taylor SJ Phillips (1997) ArticleTitleObesity QTLs on mouse chromosomes 2 and 17. Genomics 43 249–257
BA Taylor LM Tarantino SJ Phillips (1999) ArticleTitleGender-influenced obesity QTLs identified in a cross involving the KK type II diabetes-prone mouse strain. Mamm Genome 10 963–968
BA Taylor C Wnek D Schroeder SJ Phillips (2001) ArticleTitleMultiple obesity QTLs identified in an intercross between the NZO (New Zealand obese) and the SM (small) mouse strains. Mamm Genome 12 95–103
HF Utz AE Melchinger CC Schon (2000) ArticleTitleBias and sampling error of the estimated proportion of genotypic variance explained by quantitative trait loci determined from experimental data in maize using cross validation and validation with independent samples. Genetics 154 1839–1849 Occurrence Handle10866652
TT Vaughn LS Pletscher A Peripato KK Ellison E Adams et al. (1999) ArticleTitleMapping quantitative trait loci for murine growth: a closer look at genetic architecture. Genet Res Camb 74 313–322 Occurrence Handle10.1017/S0016672399004103 Occurrence Handle1:CAS:528:DC%2BD3cXhs1Grtbc%3D
CH Warden JS Fisler SM Shoemaker P-Z Wen KL Svenson et al. (1995) ArticleTitleIdentification of four chromosomal loci determining obesity in a multifactorial mouse model. J Clin Invest 95 1545–1552 Occurrence Handle1:CAS:528:DyaK2MXkslyktrw%3D Occurrence Handle7706460
ML Wayne LM McIntyre (2002) ArticleTitleCombining mapping and arraying: an approach to candidate gene identification. Proc Natl Acad Sci USA 99 14903–14906 Occurrence Handle10.1073/pnas.222549199 Occurrence Handle1:CAS:528:DC%2BD38Xpt1yrtLw%3D Occurrence Handle12415114
DB West CN Boozer DL Moody RL Atkinson (1992) ArticleTitleDietary obesity in nine inbred mouse strains. Am J Physiol 262 R1025–R1032 Occurrence Handle1:STN:280:By2A387jtlE%3D Occurrence Handle1621856
DB West JG Lefevre B York GE Truett (1994) ArticleTitleDietary obesity linked to genetic loci on chromosomes 9 and 15 in a polygenic mouse model. J Clin Invest 94 1410–1416 Occurrence Handle1:CAS:528:DyaK2cXmvFCiu70%3D Occurrence Handle7929816
DB West GE Truett JG Lefevre B York (1995) ArticleTitleLoci on chromosomes 4, 7, 9,12, & 15 control a significant proportion of the dietary obesity phenotype in the mouse. FASEB J 9 A722
RW Williams (2000) Mapping genes that modulate mouse brain development: a quantitative genetic approach. AM Goffinet P Rakic (Eds) Mouse Brain Development, Results and Problems in Cell Differentiation, vol 30 Springer-Verlag Berlin, Germany 21–49
RW Williams DC Airey A Kulkarni G Zhou L Lu (2001) ArticleTitleGenetic dissection of the olfactory bulbs of mice: QTLs on four chromosomes modulate bulb size. Behav Genet 31 61–77 Occurrence Handle10.1023/A:1010209925783 Occurrence Handle1:STN:280:DC%2BD3MvoslOhuw%3D%3D Occurrence Handle11529276
B York K Lei DB West (1996) ArticleTitleSensitivity to dietary obesity linked to a locus on chromosome 15 in a CAST/Ei × C57BL/6J F2 intercross. Mamm Genome 7 677–681
Z-B Zeng (1993) ArticleTitleTheoretical basis for separation of multiple linked gene effects in mapping quantitative trait loci. Proc Natl Acad Sci USA 90 10972–10976 Occurrence Handle1:CAS:528:DyaK2cXkslWlsQ%3D%3D Occurrence Handle8248199
Z-B Zeng (1994) ArticleTitlePrecision mapping of quantitative trait loci. Genetics 136 1457–1468 Occurrence Handle1:CAS:528:DyaK2MXhtFantw%3D%3D Occurrence Handle8013918
V Zidek A Musilova J Pintir M Simakova M Pravenec (1998) ArticleTitleGenetic dissection of testicular weight in the mouse with the BXD recombinant inbred strains. Mamm Genome 9 503–505 Occurrence Handle10.1007/s003359900808 Occurrence Handle1:CAS:528:DyaK1cXks1Cnu7c%3D Occurrence Handle9657844
Acknowledgements
We gratefully acknowledge Mary Ann Cushman, Stephenie Foster, and Grady Beck for collection of genotypic data. J.L. Rocha acknowledges the support of the Portuguese Foundation for Science and Technology. This research is a contribution of the University of Nebraska Agricultural Research Division (Lincoln, neb.; Journal Series No. 14110) and the North Carolina Agricultural Research Service, and was supported in part by funds provided through the Hatch Act. This research was also partially based upon work supported by the National Science Foundation under Grant No. 0091900 (Nebraska EPSCOR infrastructure improvement grant).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rocha, J.L., Eisen, E.J., Dale Van Vleck, L. et al. A large-sample QTL study in mice: II. Body composition . Mamm Genome 15, 100–113 (2004). https://doi.org/10.1007/s00335-003-2308-6
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00335-003-2308-6