Abstract
Objective
To investigate the efficacy of contrast-enhanced computed tomography (CECT)–based radiomics signatures for preoperative prediction of pathological grades of hepatocellular carcinoma (HCC) via machine learning.
Methods
In this single-center retrospective study, data collected from 297 consecutive subjects with HCC were allocated to training dataset (n = 237) and test dataset (n = 60). Manual segmentation of lesion sites was performed with ITK-SNAP, the radiomics features were extracted by the Pyradiomics, and radiomics signatures were synthesized using recursive feature elimination (RFE) method. The prediction models for pathological grading of HCC were established by using eXtreme Gradient Boosting (XGBoost). The performance of the models was evaluated using the AUC along with 95% confidence intervals (CIs) and standard deviation, sensitivity, specificity, and accuracy.
Results
The radiomics signatures were found highly efficient for machine learning to differentiate high-grade HCC from low-grade HCC. For the clinical factors, when they were merely applied to train a machine learning model, the model achieved an AUC of 0.6698, along with 95% CI and standard deviation of 0.5307–0.8089 and 0.0710, respectively (sensitivity, 0.6522; specificity, 0.4595; accuracy, 0.5333). Meanwhile, when the radiomics signatures were applied in association with clinical factors to train a machine learning model, the performance of the model remarkably increased with AUC of 0.8014, along with 95% CI and standard deviation of 0.6899–0.9129 and 0.0569, respectively (sensitivity, 0.6522; specificity, 0.7297; accuracy, 0.7000).
Conclusions
The radiomics signatures could non-invasively explore the underlying association between CECT images and pathological grades of HCC.
Key Points
• The radiomics signatures may non-invasively explore the underlying association between CECT images and pathological grades of HCC via machine learning.
• The radiomics signatures of CECT images may enhance the prediction performance of pathological grading of HCC, and further validation is required.
• The features extracted from arterial phase CECT images may be more reliable than venous phase CECT images for predicting pathological grades of HCC.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Hepatocellular carcinoma (HCC) is the commonest type of primary liver cancer [1], and the second leading cause of cancer-related death worldwide [2]. The global incidence of HCC shows an increasing trend, especially in China where over 50% of cases can be found [3]. Surgery is the mainstay of HCC treatment among all the therapeutic strategies, including liver transplantation (LT), radiofrequency ablation (RFA), and microwave ablation (MWA) [4, 5]. Clinically, it has been demonstrated that HCC is prone to recurrence and metastasis with poor prognosis [6]. Intrahepatic recurrence of the tumor is correlated with pathological grade. To be more specific, the intrahepatic recurrence rate increases along with the tumor grade [7,8,9]. For a higher intrahepatic recurrence rate, special treatments are required during the operation and for follow-up [10]. Therefore, accurate prediction of pathological grade of HCC is of supreme importance for making clinical decisions and prescribing the most appropriate treatment.
The histological grade, which typically represents the biological behavior of tumors, is the gold standard for diagnosing the majority of solid organ neoplasms; thus, it can provide valuable prognostic information [9]. According to the clinical guidelines, needle biopsy, a routine method for preoperative evaluation of malignant tumors, is not recommended for diagnosing HCC because of the risk of possible complications and sample errors [1, 11, 12]. How can we predict the pathological grades of HCC with medical imaging using non-invasive diagnostic tools? The solution to this question is of great clinical significance for the management of patients with HCC, as well as precision medicine.
Radiomics is a high-throughput quantitative feature extraction method that converts images into mineable data [13]. Radiomics has been shown to provide new perspectives on precision medicine in oncologic practice related to disease-free survival [14], metastases prediction [15], and therapeutic response assessment [16]. Medical imaging plays a substantial role in the characterization of HCC [17]. It has been reported that medical imaging is closely associated with pathological grades of HCC. Oh et al analyzed preoperative arterial phase CT images to find out the relationship between the texture features and histologic grade of HCC; however, their results lacked validation because of the limited number of cases [18]. Wu et al predicted the grade of HCC via non-contrast-enhanced magnetic resonance imaging (MRI)–based radiomics signatures [19].
To our knowledge, there is no strong evidence to demonstrate that CECT-based radiomics signatures could be employed to predict pathological grades of HCC patients. Therefore, the purpose of the present study was to investigate whether CECT-based radiomics signatures could be applied to preoperatively predict pathological grades of HCC.
Materials and methods
Study population
A consecutive series of patients who were pathologically diagnosed with HCC at Henan Provincial People’s Hospital (Zhengzhou, China) from February 2012 to August 2016 were enrolled in the single-center retrospective study. The inclusion criteria were as follows: (1) patients who underwent partial hepatectomy and were pathologically confirmed with HCC, (2) patients who underwent multiphasic dynamic CECT of liver within 1 week before surgery, (3) patients who received no previous treatment, including LT, MWA, or RFA, (4) the quality of medical imaging met the requirement of analysis, and (5) patients with no other types of cancer. The study was approved by the Ethics Committee of Henan Provincial People’s Hospital (Zhengzhou, China), and the subjects signed the written informed consent. Figure 1 shows the inclusion and exclusion processes.
Image acquisition
All patients underwent multiphasic dynamic CECT of liver using one of the multidetector row CT units (Brilliance 16, Philips or LightSpeed VCT, GE Healthcare, or Discovery CT750 HD, GE Healthcare, or SOMATOM Definition Flash, Siemens) in two modalities, which were called arterial phase (AP) and venous phase (VP). 53, 72, 91, and 81 patients were scanned on Brilliance 16, LightSpeed VCT, Discovery CT750 HD, and SOMATOM Definition Flash, respectively. AP- and VP-CECT were performed after 25 and 60 s of delay after intravenous injection of contrast agent (1.5 ml/kg; Ultravist 370, Bayer HealthCare Pharmaceuticals Inc.) at a rate of 3.0 ml/s using a pump injector (Ulrich CT Plus 150, Ulrich Medical). Sensitive information was removed, all images were anonymous and stored in digital imaging and communications in medicine (DICOM) format [20]. The parameters of the CECT sequences are summarized in Table 1.
Clinicopathological characteristics of patients
The clinical data of each HCC patient were collected from electronic health record (EHR), which included demographic characteristics (age, gender), image characteristics (hepatocirrhosis, lymph node enlargement, pseudocapsule, and portal vein tumor thrombosis), and laboratory tests (hepatitis B surface antigen (HBsAg) or hepatitis C antibody (HCVab) status, alpha-fetoprotein (AFP), alanine aminotransferase (ALT), aspartate aminotransferase (AST), and gamma-glutamyl transpeptidase (GGT)).
The pathological data were retrieved from the Pathology Information Management System, in which the pathological grades were noted. The pathological grading system for HCC was Edmondson-Steiner (ES), which typically allocated grades as low (grade 1 or 2) and high grades (grade 3 or 4) [9, 21].
Tumor segmentation
Manual segmentation of lesion sites on CECT images was performed with ITK-SNAP v.3.6.0 software, which provided semi-automatic segmentation using active contour methods, as well as manual delineation and image navigation [22].
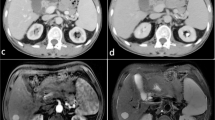
First, the AP- and VP-CECT images were loaded into the ITK-SNAP software. Second, with the supervision of a radiologist with 20 years of experience, the region of interest (ROI) that covered the whole lesion was manually delineated on each slice by a radiologist with over 8 years of experience in abdominal imaging. Both radiologists have 3 years of experience in the software. The representative results of segmentation are illustrated in Fig. 2.
Image preprocessing, feature extraction, and selection
To obtain knowledge from high-dimensional data, we adopted a radiomics approach. Radiomics features of each patient were extracted from both AP- and VP-CECT images using Pyradiomics v.2.1.2 package [23]. To eliminate the variance caused by different scanner acquisitions, avoid anisotropic resolution, and improve the reproducibility, image preprocessing was implemented for each patient by a researcher with 10 years of experience: normalized by centering to the mean standard deviation, resampled to voxel size of 1 × 1 × 1 mm3 using B-Spline interpolation, and gray-level discretized by a fixed bin width of 25 in the histogram [24].
The extracted features were classified into seven categories as follows: I, first-order statistics; II, shape features (3D); III, gray-level co-occurrence matrix (GLCM); IV, gray-level size zone matrix (GLSZM); V, gray-level run length matrix (GLRLM); VI, neighboring gray tone difference matrix (NGTDM); VII, gray-level dependence matrix (GLDM). Besides, 17 filters were applied to the original images, and yielded derived images for each patient. All the classes of features, with the exception of shape features, were calculated on both the original images and the derived images. The details of the various features and filters are present in Supplementary Appendix 1.
The values of extracted features were not on the same scale because of the differences in units and range among radiomics features. Feature scaling method was employed before dimensionality reduction to ensure that the range of radiomics features was relatively uniform. The scaling process was described below. First, the mean value of the features was subtracted. Second, the standard deviation of the features was divided by the results. Median and mode of the continuous and discrete variables were calculated, respectively, and then they were used to fill the corresponding missing data.
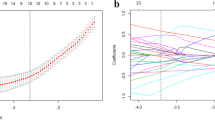
It could be inefficient for computation with high-dimensional features, and in the worst case, it could lead to overfitting [25]. Therefore, recursive feature elimination (RFE), which is capable of reducing dimension of feature, was utilized to select the most informative and valuable features [26]. The feature selection process is described as follows: First, calculate the importance of each feature. Then, remove the least important features from the current set of features. These steps should be recursively repeated until the number of remaining features reaches the required number of features.
Reproducibility of image features
To evaluate the reproducibility of the extracted features, the radiologist with 20 years of experience delineated ROIs in 30 randomly selected HCC patients. Intraclass correlation coefficient (ICC) was calculated to indicate the reliability of feature reproducibility [27]. The ICC is a statistical measure between 0 and 1, where 0 indicates no and 1 indicates perfect reliability [28]. Only the features with an ICC value equal to or higher than 0.80 were included in the further feature selection process [29].
Statistical analysis and establishment of a model based on machine learning
Characteristics of the study population were compared between training dataset and test dataset. Continuous variables were analyzed with the Shapiro–Wilk test to determine their distributions. Non-normally distributed data were expressed as median (25th to 75th percentile). Categorical variables were presented as frequencies and percentages. Differences in clinical characteristics were analyzed by t test for continuous variables and χ2 test for categorical variables. P < 0.05 was considered statistically significant. Statistical analyses were performed using SAS 9.4.
In machine learning, the classification is considered as a supervised learning task of inferring a function from labeled training data [30]. In the present study, we used XGBoost, which is a machine learning technique that assembles weak prediction models to establish a robust prediction model [31]. Python scikit-learn 0.18 package was employed to establish and evaluate the model. The dataset was randomly divided into a training dataset and test dataset using stratified sampling based on the pathological grade of patients. 5-fold cross-validation and grid search methods were applied to select the optimal model parameters [32]. This process can be generally described as follows. In the beginning, the training dataset was randomly divided into five groups with an equal number of images, and the parameters of XGBoost were randomly initialized. During training, for each step, one group among the five groups was selected as the validation dataset, while the other four groups were training dataset.
Firstly, a clinical model was established using patients’ clinicopathological characteristics, such as age, gender, hepatocirrhosis, lymph node enlargement, pseudocapsule, portal vein tumor thrombosis, HBsAg or HCVab status, AFP, ALT, AST, and GGT. Secondly, radiomics signature models based on features of AP- and VP-CECT images were built. Then, a combined model integrating radiomics signature and all clinicopathological characteristics was established.
Evaluating performance of the model
The test dataset was applied to evaluate performance of the model. The performance of both training and test datasets was evaluated using AUC along with 95% confidence intervals (CIs) and standard errors, sensitivity, specificity, and accuracy. p value was computed to measure the fitting degree of the model. The Checklist for AI in Medical Imaging (CLAIM) is available in Supplementary Appendix 2 [33]. The entire process is outlined in Fig. 3.
Results
Clinicopathological characteristics of the patients
Two hundred ninety-seven patients were enrolled and divided into training dataset (n = 237) and test dataset (n = 60). The clinicopathological characteristics of HCC patients are listed in Table 2. There were 212 (89.5%) male patients in the training dataset and 54 (90.0%) male patients in the test dataset. The median age of the patients in the training dataset and the test dataset was 53 (range, 25–71 years old) and 57 (range, 28–74 years old) years old, respectively. The number of patients with low-grade HCC was 148 (62.4%) in the training dataset and 37 (61.7%) in the test dataset. There was no significant difference in the clinicopathological characteristics between the training dataset and the test dataset (all p > 0.05).
Feature extraction and selection
A total of 3376 features were extracted from CECT images. For both AP- and VP-CECT images, 1688 features were extracted from original and derived images, respectively. Among those features extracted from original images, the number of seven aforementioned categories was 18, 14, 24, 14, 16, 16, and 5 for first-order statistics, shape-based, GLCM, GLDM, GLRLM, GLSZM, and NGTDM, respectively. The extracted features are listed in Supplementary Appendix 3.
Among all the extracted features, 232 features were excluded because the ICC values were less than 0.80. Based on AP, VP, and combined AP and VP (AP-VP), 12, 34, and 50 features were selected, respectively. The results of feature selection and feature importance are listed in Supplementary Appendix 4.
Predictive ability of radiomics signatures and the clinical factors
The radiomics signatures based on AP-CECT images and AP-VP-CECT images showed high-performance in differentiating high-grade HCC from low-grade HCC, with an average AUC of 0.7186 (95% CI, 0.5900–0.8472) and 0.7579 (95% CI, 0.6296–0.8863), respectively. The AUC of the VP-CECT images was 0.6128 (95% CI, 0.4522–0.7734, p > 0.0125) in the test dataset. The AUC of the clinical model was 0.6689 (95% CI, 0.5307–0.8089, p > 0.0125) in the test dataset.
Predictive performance of the clinical model combined with the radiomics signatures
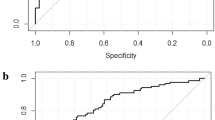
As mentioned previously, for the radiomics signatures, all the three cases are taken into account. When clinical factors were combined with AP only, the model reached the AUC of 0.7286 (95% CI, 0.5989–0.8582), when clinical factors were combined with VP, the model attained an AUC of 0.6769 (95% CI, 0.5352–0.8185). On the other hand, when clinical factors are combined with AP-VP simultaneously, the model outperformed compared with others with an improved AUC of 0.8014 (95% CI, 0.6899–0.9129). The ROC curves with confidence intervals are shown in Fig. 4 and Supplementary Appendix 5. The detailed prediction performance of the radiomics signatures, the clinical model, and the combined model is presented in Table 3.
Discussion
In the present study, we employed CECT-based radiomics signatures and XGBoost classifier for preoperative prediction of pathological grades of HCC. The radiomics signatures based on AP- and VP-CECT images were respectively extracted, and RFE method was used to reduce dimension of features. Based on AP, VP, and AP-VP, 12, 34, and 50 features were selected, respectively. Additionally, XGBoost classifier with 5-fold cross-validation strategy was utilized to train the classification model. The results showed that the radiomics signatures based on AP-CECT images could successfully classify pathological grades of HCC in both the training dataset (p < 0.0125) and the test dataset (p < 0.0125). No significant differences were observed in VP-CECT images and clinical factors (p > 0.0125) for the discrimination of low-grade HCC and high-grade HCC in the test dataset. Because the arterial blood supply of HCC was sufficient, AP could deliver signals with higher intensity in the tumor parenchyma compared with VP. Consequently, the features extracted from AP-CECT images were more reliable for predicting pathological grades of HCC, which is consistent with the results of previous research [34]. Hence, it can be concluded that the AP-CECT images are more significant to explore the heterogeneity of HCC. Accordingly, the radiomics signatures could be used to capture insignificant differences in signal intensities in tumors with different pathological grades, which is in accordance with Oh et al’s outcomes [18]. In addition, we investigated the prediction performance of the established model (based on AP- and VP-CECT images), the clinical model, and the clinical model combined with the radiomics signatures. The results indicated that the model which combined clinical factors with AP- and VP-CECT images outperformed compared with other models. The abovementioned results are in agreement with Wu et al’s study in which the association between MRI-based radiomics signatures and HCC grade was explored [19].
Radiomics is an emerging field, aiming to explore the underlying relationship between medical images and phenotypic characteristics of tumor cells in a non-invasive manner [35]. Previous studies demonstrated that radiomics is crucial in classifying tumor grade. Bektas et al reported that the machine learning–based CT texture analysis could be a promising non-invasive method for the prediction of Fuhrman nuclear grade for clear cell renal carcinoma (cc-RCC). They employed five classifiers to train the models. Especially, four-fifth of the patients could be correctly classified using the SVM classifier, and the model obtained an AUC of 0.860 [29]. Sun et al compared the prediction performance of frequently utilized radiomics feature selection and classification methods in glioma grading; radiomics features of 210 high-grade and 75 low-grade glioma subjects were extracted in the four types of MRI modality images. The performance of sixteen feature selection methods and fifteen classifiers were evaluated. The results showed that the tumor region was more relevant to the tumor grade than other regions, and multiparametric MRI modalities could boost the prediction performance [36]. Wu et al investigated the clinical significance of MRI-based radiomics signatures for the preoperative prediction of HCC grade, and the AUC of model using radiomics signatures and clinical factors was 0.800. The study showed that the combination of the radiomics signatures with clinical factors could be useful for the prediction of HCC grade, and both the AFP level and radiomics signatures were independent predictors [19]. To our knowledge, only one study concentrated on the CT texture to predict histological grade of HCC, and texture analysis was performed on the original and derived MRI images of eighty-one patients. The results showed that the textures of AP-CECT images were significantly correlated with histological grade of HCC [18].
Despite the novelties of the present research, there are several limitations. First, the experiment lacks external validation since it is a single-center cohort study; thus, the reproducibility and generalizability of our findings remain to be verified. Therefore, further multicenter cohorts should be conducted. Second, our study included only HCC, because it is the most common type of primary liver cancer. Third, multimodal radiomics models were not applied in this study, because we could not harvest other imaging modals such as ultrasound and MRI. Forth, different CT scanners were used to collect imaging data, and different technical parameters were used. Finally, although ICC test was carried out, bias is still possible in selection process for training and test datasets.
In summary, the machine learning–based CECT radiomics analysis could improve accuracy of a prediction model, and may non-invasively explore the underlying association between images and pathological grades of HCC.
Abbreviations
- 3D:
-
Three-dimensional
- AFP:
-
Alpha-fetoprotein
- ALT:
-
Alanine aminotransferase
- AP:
-
Arterial phase
- AST:
-
Aspartate aminotransferase
- cc-RCC:
-
Clear cell renal carcinoma
- CECT:
-
Contrast-enhanced computed tomography
- CI:
-
Confidence interval
- CLAIM:
-
Checklist for AI in Medical Imaging
- DICOM:
-
Digital imaging and communications in medicine
- EHR:
-
Electronic health records
- ES:
-
Edmondson-Steiner
- GGT:
-
Gamma-glutamyl transpeptidase
- GLCM:
-
Gray-level co-occurrence matrix
- GLDM:
-
Gray-level dependence matrix
- GLRLM:
-
Gray-level run length matrix
- GLSZM:
-
Gray-level size zone matrix
- HBsAg:
-
Hepatitis B surface antigen
- HCVab:
-
Hepatitis C antibody
- ICC:
-
Intraclass correlation coefficient
- LT:
-
Liver transplantation
- MRI:
-
Magnetic resonance imaging
- MWA:
-
Microwave ablation
- NGTDM:
-
Neighboring gray tone difference matrix
- RFA:
-
Radiofrequency ablation
- RFE:
-
Recursive feature elimination
- ROI:
-
Region of interest
- SVM:
-
Support vector machine
- VP:
-
Venous phase
- XGBoost:
-
eXtreme Gradient Boosting
References
European Association for the Study of the Liver (2018) EASL clinical practice guidelines: management of hepatocellular carcinoma. J. Hepatol. 69(1):182–236
Njei B, Rotman Y, Ditah I, Lim JK (2015) Emerging trends in hepatocellular carcinoma incidence and mortality. Hepatology 61(1):191–199
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68(6):394–424
Lencioni R, Crocetti L (2012) Local-regional treatment of hepatocellular carcinoma. Radiology 262(1):43–58
Forner A, Reig M, Bruix J (2018) Hepatocellular carcinoma. Lancet 391(10127):1301–1314
Wang Y-Y, Zhong J-H, Su Z-Y et al (2016) Albumin-bilirubin versus Child-Pugh score as a predictor of outcome after liver resection for hepatocellular carcinoma. Br J Surg 103(6):725–734
Sasaki A, Kai S, Iwashita Y, Hirano S, Ohta M, Kitano S (2005) Microsatellite distribution and indication for locoregional therapy in small hepatocellular carcinoma. Cancer 103(2):299–306
Weiss LM, Medeiros LJ, Vickery AL (1989) Pathologic features of prognostic significance in adrenocortical carcinoma. Am J Surg Pathol 13(3):202–206
Martins-Filho SN, Paiva C, Azevedo RS, Alves VAF (2017) Histological grading of hepatocellular carcinoma-a systematic review of literature. Front Med (Lausanne) 4:193
Bruix J, Sherman M (2005) Management of hepatocellular carcinoma. Hepatology 42(5):1208–1236
Llovet JM, Zucman-Rossi J, Pikarsky E et al (2016) Hepatocellular carcinoma. Nat Rev Dis Primers 2:16018
Robert M, Sofair AN, Thomas A et al (2009) A comparison of hepatopathologists’ and community pathologists' review of liver biopsy specimens from patients with hepatitis C. Clin Gastroenterol Hepatol 7(3):335–338
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: images are more than pictures, they are data. Radiology 278(2):563–577
Huang Y, Liu Z, He L et al (2016) Radiomics signature: a potential biomarker for the prediction of disease-free survival in early-stage (I or II) non-small cell lung cancer. Radiology 281(3):947–957
Ortiz-Ramón R, Larroza A, Ruiz-España S, Arana E, Moratal D (2018) Classifying brain metastases by their primary site of origin using a radiomics approach based on texture analysis: a feasibility study. Eur Radiol 28(11):4514–4523
Jin X, Zheng X, Chen D et al (2019) Prediction of response after chemoradiation for esophageal cancer using a combination of dosimetry and CT radiomics. Eur Radiol 29(11):6080–6088
Hennedige T, Venkatesh SK (2013) Imaging of hepatocellular carcinoma: diagnosis, staging and treatment monitoring. Cancer Imaging 12:530–547
Oh J, Lee JM, Park J et al (2019) Hepatocellular carcinoma: texture analysis of preoperative computed tomography images can provide markers of tumor grade and disease-free survival. Korean J Radiol 20(4):569–579
Wu M, Tan H, Gao F et al (2019) Predicting the grade of hepatocellular carcinoma based on non-contrast-enhanced MRI radiomics signature. Eur Radiol 29(6):2802–2811
Willemink MJ, Koszek WA, Hardell C et al (2020) Preparing medical imaging data for machine learning. Radiology 295(1):4–15
Edmondson HA, Steiner PE (1954) Primary carcinoma of the liver: a study of 100 cases among 48,900 necropsies. Cancer 7(3):462–503
Yushkevich PA, Piven J, Hazlett HC et al (2006) User-guided 3D active contour segmentation of anatomical structures: significantly improved efficiency and reliability. Neuroimage 31(3):1116–1128
van Griethuysen JJM, Fedorov A, Parmar C et al (2017) Computational radiomics system to decode the radiographic phenotype. Cancer Res. 77(21):e104–e107
Leijenaar RTH, Nalbantov G, Carvalho S et al (2015) The effect of SUV discretization in quantitative FDG-PET radiomics: the need for standardized methodology in tumor texture analysis. Sci Rep 5:11075
E L, Lu L, Li L, Yang H, Schwartz LH, Zhao B (2019) Radiomics for classification of lung cancer histological subtypes based on nonenhanced computed tomography. Acad Radiol 26(9):1245–1252
Guyon I, Weston J, Barnhill S, Vapnik V (2002) Gene Selection for Cancer Classification using Support Vector Machines. Mach Learn 46(1/3):389–422
Koo TK, Li MY (2016) A guideline of selecting and reporting intraclass correlation coefficients for reliability research. J Chiropr Med 15(2):155–163
Leijenaar RTH, Carvalho S, Velazquez ER et al (2013) Stability of FDG-PET Radiomics features: an integrated analysis of test-retest and inter-observer variability. Acta Oncol. 52(7):1391–1397
Bektas CT, Kocak B, Yardimci AH et al (2019) Clear cell renal cell carcinoma: machine learning-based quantitative computed tomography texture analysis for prediction of Fuhrman nuclear grade. Eur Radiol 29(3):1153–1163
Singh A (2019) Foundations of machine learning. SSRN J. https://doi.org/10.2139/ssrn.3399990
Chen T, Guestrin C (2016) XGBoost: a scalable tree boosting system. In: Krishnapuram B, Shah M, Smola A, Aggarwal C, Shen D, Rastogi R (eds) Proceedings of the 22nd ACM SIGKDD international conference on knowledge discovery and data mining. ACM, New York, pp 785–794
Akinkunmi M (2019) Introduction to statistics using R. Morgan & Claypool, San Rafael California
Mongan J, Moy L, Kahn CE (2020) Checklist for artificial intelligence in medical imaging (CLAIM): a guide for authors and reviewers. Radiol Artif Intell 2(2):e200029
Zhou W, Zhang L, Wang K et al (2017) Malignancy characterization of hepatocellular carcinomas based on texture analysis of contrast-enhanced MR images. J Magn Reson Imaging 45(5):1476–1484
Parmar C, Grossmann P, Bussink J, Lambin P, Aerts HJWL (2015) Machine learning methods for quantitative radiomic biomarkers. Sci Rep 5:13087
Sun P, Wang D, Mok VC, Shi L (2019) Comparison of feature selection methods and machine learning classifiers for radiomics analysis in glioma grading. IEEE Access 7:102010–102020
Funding
The study was funded by the National Natural Science Foundation of China (NO. 71974065) and Key R & D and promotion projects in Henan Province (NO. 182400410172).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Guarantor
The scientific guarantor of this publication is Jingdong Ma.
Conflict of interest
The authors of this manuscript declare no relationships with any companies, whose products or services may be related to the subject matter of the article.
Statistics and biometry
No complex statistical methods were necessary for this paper.
Informed consent
Written informed consent was obtained from all subjects (patients) in this study.
Ethical approval
Institutional Review Board approval was obtained.
Methodology
• Retrospective
• Diagnostic or prognostic study
• Performed at one institution
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PNG 493 kb)
ESM 2
(PNG 1033 kb)
ESM 3
(PNG 794 kb)
ESM 4
(PNG 1026 kb)
ESM 5
(PNG 718 kb)
ESM 6
(PNG 1026 kb)
ESM 7
(PNG 1537 kb)
ESM 8
Details of the various features and filters (DOCX 13 kb)
ESM 9
The Checklist for AI in Medical Imaging (CLAIM) (DOCX 24 kb)
ESM 10
The list of the extracted features in our study (CSV 127 kb)
ESM 11
The results of feature selection and feature importance. The results of feature selection of AP images, VP images, clinical factors, AP-VP images, combined clinical factors and AP images, combined clinical factors and VP images, combined clinical factors and AP-VP images are shown in an order from A to G, respectively.5.The ROC curves with confidence intervalof AP images, VP images, clinical factors, AP-VP images, combined clinical factors and AP images, combined clinical factors and VP images are shown in an order from A to F, respectively. (PNG 6385 kb)
Rights and permissions
About this article
Cite this article
Mao, B., Zhang, L., Ning, P. et al. Preoperative prediction for pathological grade of hepatocellular carcinoma via machine learning–based radiomics. Eur Radiol 30, 6924–6932 (2020). https://doi.org/10.1007/s00330-020-07056-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-020-07056-5