Abstract
Objectives
To test the hypothesis that a fourfold CAIPIRINHA accelerated, 10-min, high-resolution, isotropic 3D TSE MRI prototype protocol of the ankle derives equal or better quality than a 20-min 2D TSE standard protocol.
Methods
Following internal review board approval and informed consent, 3-Tesla MRI of the ankle was obtained in 24 asymptomatic subjects including 10-min 3D CAIPIRINHA SPACE TSE prototype and 20-min 2D TSE standard protocols. Outcome variables included image quality and visibility of anatomical structures using 5-point Likert scales. Non-parametric statistical testing was used. P values ≤0.001 were considered significant.
Results
Edge sharpness, contrast resolution, uniformity, noise, fat suppression and magic angle effects were without statistical difference on 2D and 3D TSE images (p > 0.035). Fluid was mildly brighter on intermediate-weighted 2D images (p < 0.001), whereas 3D images had substantially less partial volume, chemical shift and no pulsatile-flow artifacts (p < 0.001). Oblique and curved planar 3D images resulted in mildly-to-substantially improved visualization of joints, spring, bifurcate, syndesmotic, collateral and sinus tarsi ligaments, and tendons (p < 0.001, respectively).
Conclusions
3D TSE MRI with CAIPIRINHA acceleration enables high-spatial resolution oblique and curved planar MRI of the ankle and visualization of ligaments, tendons and joints equally well or better than a more time-consuming anisotropic 2D TSE MRI.
Key Points
• High-resolution 3D TSE MRI improves visualization of ankle structures.
• Limitations of current 3D TSE MRI include long scan times.
• 3D CAIPIRINHA SPACE allows now a fourfold-accelerated data acquisition.
• 3D CAIPIRINHA SPACE enables high-spatial-resolution ankle MRI within 10 min.
• 10-min 3D CAIPIRINHA SPACE produces equal-or-better quality than 20-min 2D TSE.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Magnetic resonance imaging (MRI) of the ankle depends heavily on intermediate-weighted and fluid-sensitive fat-suppressed two-dimensional (2D) turbo spin echo (TSE) pulse sequences, which are typically acquired sequentially in orthogonal planes. Limitations of 2D MRI, however, include voxel anisotropy and partial volume effects. 2D MR images are therefore not suitable for curved and oblique planar image reformats, which are desirable for the visualization of the ligaments and tendons about the ankle.
Three-dimensional (3D) isotropic volumetric MRI acquisition techniques, such as Sampling Perfection with Application optimized Contrast using different flip angle Evolutions (SPACE, Siemens Healthcare, Erlangen, Germany), in contrast, offer a small isotropic voxel size through substantially thinner slice partitions, which enable reformation of virtually any imaging plane from a single parent volume. However, drawbacks of 3D TSE have thus far included image blur, low contrast-to-noise ratios, non-uniformity of reformated images and often substantially longer scan times.
One-dimensional (1D) parallel imaging acceleration techniques have been utilized for 3D TSE [1–5], but have been largely limited to acceleration factors of 2 and long acquisition times of high resolution data sets. The CAIPIRINHA (Controlled Aliasing In Parallel Imaging Results IN Higher Acceleration) sampling pattern reduces aliasing artifacts and image noise through optimized use of coil sensitivities and enables parallel imaging acceleration in two dimensions [6–8], which in combination offers the potential for higher acceleration factors and improved image quality of 3D volume acquisitions. To the best of our knowledge, CAIPIRINHA sampling has not been investigated for 3D MRI of the ankle.
Therefore, we prospectively tested the hypothesis that a comprehensive, fourfold accelerated, 10-min, high-resolution, isotropic 3D CAIPIRINHA SPACE MRI ankle prototype protocol derives equal or better quality than a 20-min 2D TSE standard protocol.
Materials and methods
Study design
This prospective, single-center, evidence level 1 [9] study was approved by our institutional review board and complied both with the Declaration of Helsinki and the Health Insurance Portability and Accountability Act of the USA. Written informed consent was obtained from all subjects. Subjects of this study have not been previously reported.
MR imaging technique
MRI exams were performed using a commercially available, clinical wide-bore 3 Tesla MR imaging system (MAGNETOM Skyra, NUMARIS/4 Syngo MR D13A, Siemens Healthcare) and a commercially available foot and ankle surface coil with 16 receiver channels (Siemens Healthcare).
All subjects underwent a single MRI exam of the ankle with a total acquisition time of 30 min, which consisted of a 3D protocol of 10-min acquisition time and 2D protocol of 20-min acquisition time (Table 1). Pulse sequences were acquired in random order. For the 3D protocol, a prototype CAIPIRINHA SPACE sequence was employed to acquire non-fat-suppressed intermediate-weighted and fat-suppressed T2-weighted with fourfold acceleration and isotropic voxel resolution. Our institutional 2D TSE protocol served as the standard of reference.
Subjects
The study population consisted of a consecutive, single group of 24 asymptomatic subjects (12 men, 12 women; mean age, 35 years, range: 24–62 years; mean body mass index, 26 kg/m2, range: 17–33 kg/m2). Inclusion criteria comprised adult age, capacity to give informed consent and asymptomatic ankle joint. Exclusion criteria comprised contraindications for MRI and prior ankle surgery. In all subjects, one randomly chosen ankle joint was imaged. All MRI exams were acquired for research purposes only.
Image evaluation
Two observers with 5 (B.F.) and 10 (J.F.) years of experience in musculoskeletal MRI evaluated the data independently. Intermediate-weighted and fat-saturated T2-weighted 2D and 3D data sets were separated and presented in random order. A 2D data set consisted of non-isotropic axial, sagittal and coronal images. A 3D data set consisted of a single sagittal stack of images. All image annotations were removed and each data set was assigned a unique random number. Readings were performed in a standardized fashion at approximately 1 lux using commercially available picture archiving and communications system software (Vue version 12.1.0.2041, Carestream Health, Rochester, NY, USA) with a 30-in. diagnostic-quality colour liquid crystal display monitor (Barco MDCC-6330, Duluth, GA, USA) calibrated to Digital Imaging and Communications in Medicine standards [10]. 2D data sets were displayed using three viewports in a 2x2 pattern using a predefined hanging protocol. Similarly, 3D data sets were displayed with a predefined hanging protocol using an interactive multiplanar reformation mode and three viewports in a 2x2 pattern. In addition, 3D data sets were viewable in curved planar reformation modes. Observers were free to use their preferred window and level settings, magnification and scrolling mode. The two observers reviewed the datasets independently in eight sessions, which were performed 1 week apart. Following a period of 1 month, both observers repeated their independent assessments.
Outcome variables
Outcome variables were grouped into image quality, visibility of anatomical structures and overall reader satisfaction with regard to the suitability of the respective data set for diagnostic interpretation. Ratings were performed with symmetrical, equidistant 5-point Likert scales, where a rating of 1 denoted ‘very bad’ with resultant complete obscuration of anatomical details, 2 denoted ‘bad’ with partial obscuration of anatomical details, 3 denoted ‘adequate’ with impaired depiction of only fine anatomical detail, 4 denoted ‘good’ with minimal impairment, but preservation of all anatomical detail, and 5 denoted ‘very good’ with unimpaired depiction of all anatomical details.
The image quality assessment included edge sharpness, contrast resolution, fluid brightness, uniformity of image quality between axial, coronal and sagittal planes reformatted from on single 3D TSE data volume and separately acquired corresponding 2D TSE images, fat suppression, image noise, parallel image artifact, pulsatile flow artifact, chemical shift effect, partial volume effect and magic angle effect.
The assessment of anatomical structures included articular cartilage, ligaments and tendons. Articular cartilage of the tibiotalar, posterior subtalar, talocalcaneonavicular, calcaneocuboid, naviculocuneiform and midfoot joints was assessed. The assessment of ligaments included the syndesmotic ligament complex (anterior inferior tibiofibular ligament, interosseous ligament, posterior inferior tibiofibular ligament) [11], lateral collateral ligament complex (anterior talofibular ligament, calcaneofibular ligament, posterior talofibular ligament) [12], medial collateral ligament complex (anterior and posterior tibiotalar ligaments, tibiocalcaneal ligament, tibionavicular ligament, tibiospring ligament) [13], sinus tarsi (cervical ligament, interosseous talocalcaneal ligament and the medial, intermediate and lateral roots of the inferior extensor retinaculum) [14, 15], spring ligament complex (superomedial calcaneonavicular ligament, medioplantar oblique calcaneonavicular ligament, inferoplantar longitudinal calcaneonavicular ligament) [16] and bifurcate ligament complex (calcaneocuboid and calcaneonavicular ligament) [17]. Assessment of tendons included the extensor tendons (anterior tibial tendon, extensor hallucis longus tendon, extensor digitorum longus tendon), peroneal tendons (peroneus brevis and longus tendon), long flexor tendons (posterior tibial tendon, flexor digitorum longus, flexor hallucis longus), Achilles tendon and plantar fascia.
Statistical and quantitative assessments
Statistical analyses were performed using JMP Pro 12.1.0 software (SAS Institute, Cary, NC, USA). Visual ratings are given as median with minimum and maximum in parentheses. In order to detect a difference of 1 of visual ratings, an a priori power calculation based on the Wilcoxon signed-rank test with normal parent distribution was performed and determined a minimum sample size of 24 subjects was needed to achieve a test power of 0.80 with a Bonferroni-corrected alpha error of 0.001. Wilcoxon Signed-Rank Test was used to test for differences of paired ordinal variables. Inter-reader agreement and intra-reader reliability were calculated using Cohen’s weighted kappa test. The grading of interobserver agreement was performed according to the recommendations of Landis and Koch [18]. Wilcoxon Signed-Rank test was used to assess differences of inter- and intraobserver gradings between 2D and 3D data sets. The second readings of both readers were used for data presentation. Due to multiple comparisons, Bonferroni-corrected p-values of 0.001 and less were considered statistically significant.
Results
Image quality
The image quality assessments are given in Table 2. There was good inter-reader agreement (κ = 0.60–0.78) without significant differences between 2D and 3D data sets (p = 0.519). Intrareader reliability was good to very good (κ = 0.69 - 0.85) without significant differences between 2D and 3D data sets (p = 0.764). Edge sharpness, uniformity, contrast resolution and fat suppression were similarly good to very good for 2D TSE and 3D SPACE MR image reformats. There were chemical shift, partial volume effects and pulsatile flow artifacts on 2D MR images, which were minimal or absent on 3D MR images (Figs. 1, 2 and 3). Mild magic angle effects were similarly present on 2D and 3D MR images, specifically in segments of the peroneus longus tendon (Fig. 4). There were no parallel imaging artifacts on 2D and 3D data sets.
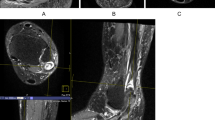
Corresponding 2D TSE and 3D CAIPIRINHA SPACE MR images of the joints and cartilage of the ankle. First and second left column: Sagittal intermediate-weighted and fat-saturated T2-weighted 2D TSE (upper images) and 3D CAIPIRINHA SPACE (lower images) MR images demonstrate the cartilage of tibiotalar (black arrows), posterior subtalar joint (white arrows), and talocalcaneonavicular (gray arrows) joints. The small black arrows show the chemical shift effect on the 2D TSE image (upper image), which is absent on the 3D SPACE image (lower image). Third and fourth column: Axial intermediate-weighted and fat-saturated T2-weighted 2D TSE (upper images) and 3D CAIPIRINHA SPACE (lower images) MR images demonstrate the cartilage of the naviculocuneiform joint (white arrows). The small black arrows show partial volume effects of 2D TSE images, which render the talonavicular joint less distinct.
Corresponding 2D TSE and 3D CAIPIRINHA SPACE MR images of the ligaments of the ankle. First left column: Axial intermediate-weighted 2D TSE (upper image) and 3D CAIPIRINHA SPACE (lower image) MR images demonstrate the anterior talofibular ligament (white arrows). Second column: Coronal fat-saturated T2-weighted 2D TSE (upper image) and 3D CAIPIRINHA SPACE (lower image) MR images demonstrate the posterior talofibular ligament (white arrow). Third column: Sagittal intermediate-weighted 2D TSE (upper image) and 3D CAIPIRINHA SPACE (lower image) MR images demonstrate the calcaneonavicular ligament of the bifurcate ligament complex (white arrows) and lateral root of the inferior extensor retinaculum of the sinus tarsi (black arrows). Fourth column: Sagittal fat-saturated T2-weighted 2D TSE (upper image) and 3D CAIPIRINHA SPACE (lower image) MR images demonstrate the tibiospring ligament of the medial collateral ligament complex (white arrows). The small white arrows show pulsatile flow artifacts on the 2D TSE image (upper image), which are absent on the 3D image (lower image). Fifth column: Axial intermediate-weighted 2D TSE (upper image) and 3D CAIPIRINHA SPACE (lower image) MR images demonstrate the inferoplantar longitudinal calcaneonavicular ligament of the spring ligament complex (black arrows)
Corresponding 2D TSE and 3D CAIPIRINHA SPACE MR images of the tendons of the ankle. First left column: Sagittal fat-saturated T2-weighted 2D TSE (upper image) and 3D CAIPIRINHA SPACE (lower image) MR images demonstrate the Achilles tendon (white arrows) and plantar fascia (gray arrows). Second and third columns: Axial intermediate-weighted and fat-saturated T2-weighted 2D TSE (upper images) and 3D CAIPIRINHA SPACE (lower images) MR images demonstrate the peroneal longus and brevis tendons (white arrows), extensor tendons (black arrows) and the long flexor tendons with tibial neurovascular bundle (gray arrows). Fourth and fifth columns: Sagittal intermediate-weighted and fat-saturated T2-weighted 2D TSE (upper images) and 3D CAIPIRINHA SPACE (lower images) MR images demonstrate the posterior tibial tendon (white arrows), which is less conspicuous on the 2D MR image due to partial volume effects. The small arrows show pulsatile flow artifacts on the 2D TSE images (upper images), which are absent on the 3D images (lower images)
Corresponding sagittal intermediate-weighted and fat-saturated T2-weighted curved planar 3D CAIPIRINHA SPACE MR images of the tendons of the ankle demonstrate the anterior tibial tendon (first left column, white arrows), extensor digitorum longus tendon (second column, white arrows), peroneus longus tendon (third column, white arrows) and posterior tibial tendon (fourth column, white arrows). The small arrows in column 3 show mild segmental signal hyperintensity of the peroneus longus tendon due to presumed magic angle effect
Visibility of anatomical structures
The assessments of joints, ligaments and tendons are given in Tables 3, 4 and 5. There was good inter-reader agreement (κ = 0.62–0.74) without a significant difference between 2D and 3D data sets for joints (p = 0.723), ligaments (p = 0.569) and tendons (p = 0.436). Intra-reader reliability was good (κ = 0.68 - 0.78) without significant differences between 2D and 3D data sets for joints (p = 0.611), ligaments (p = 0.278) and tendons (p = 0.345). Articular cartilage of the midfoot joints was best seen on 3D MR images (Table 3, Fig. 1). The syndesmotic ligaments, lateral collateral ligaments as well as anterior tibiotalar, tibiocalcaneal and tibiospring ligaments, most parts of the sinus tarsi ligaments, bifurcate ligament complex and the two plantar spring ligaments were significantly better (p < 0.001, respectively) seen on 3D MR images (Fig. 2). The peroneal tendons, long flexor tendons as well as the anterior tibial and extensor digitorum longus tendons were best seen on oblique and curved planar 3D MR images (p < 0.001, respectively) (Figs. 2, 3 and 4).
Reader satisfaction
There was no significant difference between the satisfaction ratings of the two readers (p = 0.207). Reader satisfaction ratings were significantly better (p < 0.001) for intermediate-weighted 3D CAIPIRINHA SPACE MR images (average rating, ‘very good’; range, ‘good’–‘very good’) than for intermediate-weighted 2D TSE MR images (‘good’, ‘adequate’–‘very good’), whereas there was no statistically significant (p = 0.420) difference between the reader satisfaction ratings of fat-saturated T2-weighted 2D (‘good’, ‘good’–‘very good’) images and 3D (‘good’, ‘good’–‘very good’) images.
Discussion
The isotropic voxel size and high-spatial resolution of 3D TSE MRI has previously been shown to improve visualization of cartilage [5, 19, 20], ligaments [13, 20, 21] and tendons [22] when compared to 2D TSE MR images. Our study found specifically that the higher volumetric spatial resolution of 3D MRI most benefited the delineation of the midfoot joints, oblique and thin ligaments, and markedly curved tendons. This result is mostly due to the ability to retrospectively create dedicated oblique planar image reformation of ligaments, such as the AFTL [12], and high quality curved planar reformations of markedly-oblique tendons such as the peroneus longus, which is afforded by the small isotropic voxel size.
Compared with previous 3D TSE MRI techniques [20, 22], including FSE-CUBE (GE Healthcare, Little Chalfont, UK), Volume Isotropic Turbo Spin Echo acquisition (VISTA, Philips, Andover, MA, USA), and isoFSE (Hitachi Medical Systems, Twinsburg, OH, USA), our 3D TSE SPACE prototype utilized an integrated CAIPIRINHA sampling pattern, which has previously been used successfully for efficient, isotropic, high-spatial resolution 3D gradient echo MRI. The CAIPIRINHA sampling pattern facilitates optimized use of coil sensitivities and thereby mitigates aliasing artifacts normally seen with high acceleration factors [23]. The CAIPIRINHA sampling pattern allowed for twofold parallel imaging acceleration in two dimensions with an effective acceleration factor of 4 which is double the acceleration as previously possible [20, 22]. This approach afforded efficient data acquisition, absent parallel imaging artifacts and degrading image noise, and true isotropic voxel size of high-spatial resolution with image quality similar or better to our institutional 2D TSE standard, which requires double the scan time.
Based on our clinical efficiency requirements, the goal for our prototype 3D CAIPIRINHA SPACE ankle protocol was a total acquisition time of 10 min, true isotropic voxel size to retain fine anatomical detail, and full Fourier sampling in order to preserve image quality and uniformity of reformations. In addition to the CAIPIRINHA sampling pattern, the naturally-occurring loss of MRI signal through the applied SPAIR fat suppression was compensated for with an increase in voxel lengths of 20% from 0.5 mm to 0.63 mm, which achieved double the voxel volume from 0.125 mm3 to 0.250 mm3 due to the cubic relationship.
An early study utilizing 3D MRI of the ankle showed an increase of scan efficiency through the use of 2D-accelerated parallel imaging with generalized autocalibrating partially parallel acquisitions (GRAPPA) [22]. Together with the use of partial Fourier sampling and long echo trains a scan time of 5 min was achieved for 0.6-mm isotropic 3D-FSE-Cube acquisition of the ankle. However, the authors reported significant degradation of image quality through increased blurring and loss of inter-planar uniformity of the 3D images compared to their 2D FSE institutional standard images. These effects were likely the results of a long echo train of 78, degrading effect of partial Fourier sampling and inherent limitations of GRAPPA, which include aliasing artifacts and increased image noise due to suboptimal use of coil sensitivities. Similarly, the 3D TSE images of another study utilizing an echo train length of 100 [24] showed noticeable blurring. The fourfold acceleration of 3D SPACE through 2D CAIPIRINHA in our study realized an acquisition time of 5 min with low echo train length of 42–54, full Fourier sampling, low image noise, absent parallel imaging artifacts, minimized blurring and high uniformity of image quality among oblique and curved planar reformatted images.
Additionally, the 3D CAIPIRINHA SPACE images in our study had markedly less artifacts than the 2D TSE images. The slice thickness of 3D sequences was 5.4–6.0 times thinner than the conventional 2D TSE sequences, which led to markedly reduced partial volume effects. The lack of partial Fourier sampling contributed to similar image quality across the entire image volume and highly uniform image quality between different reformation planes and types. The use of a high readout bandwidth markedly decreased chemical shift effect. In addition, no pulsatile flow artifacts were present on the 3D SPACE images. The sequence parameters of the 3D parameters were chosen empirically to best match the tissue contrasts of 2D TSE and permit acquisition times of less than 5 min. In a prior study, approximately three times longer repetition times were utilized [22], which does not appear to be required for 3D CAIPIRINHA SPACE.
Despite these advantages, images from the 3D CAIPIRINHA SPACE protocol (0.63-mm slice thickness) did show mildly higher image noise compared to images from the 2D TSE protocol (2.7- and 3.0-mm slice thickness). However, the additional image noise did not compromise the depiction of anatomical structures. This is in keeping with prior investigations [22].
Limitations of this initial study include the study population of asymptomatic subjects, rather than patients with structural abnormalities. As healthy subjects may be more compliant than patients with painful conditions, there is the possibility of a lower rate of degradation of image quality through patient motion in our study. In a sample of 24 test persons and a statistical power of 80%, a p-value larger than 0.05 may not necessarily mean there is no difference between groups. We did not record the length of time needed for image evaluation of 2D and 3D data sets. As standardized hanging protocols were used, the time needed for the evaluation of 2D data sets and 3D data sets in the interactive mode were similar. The creation of curved planar reformations was unique to the 3D data sets and took approximately 20 s for a long structure such as the peroneus longus tendon. In this study, we used our institutional 3T ankle MRI protocol for comparison with the investigational 3D CAIPIRINHA SPACE protocol. We acknowledge that 3T MRI protocols of the ankle vary between institutions and may produce different quality. This first evaluation demonstrated the comparative utility of 3D CAIPIRINHA SPACE in regard to depiction of anatomy and image quality; however, injuries were not evaluated and the diagnostic performance of 3D CAIPIRINHA SPACE is currently not known. The results of this study warrant future clinical trials with the goal of determining the diagnostic performance of 3D CAIPIRINHA SPACE for acute and chronic ankle injuries.
In conclusion, 3D CAIPIRINHA SPACE TSE enables efficient, isotropic MRI of the ankle that facilitates high-spatial resolution oblique and curved planar image reformation of ligaments, tendons and joints, and visualization of critical ankle structures equally well or better than a more time-consuming anisotropic 2D TSE MRI.
References
Bauer JS, Banerjee S, Henning TD, Krug R, Majumdar S, Link TM (2007) Fast high-spatial-resolution MRI of the ankle with parallel imaging using GRAPPA at 3 T. AJR Am J Roentgenol 189:240–245
Kijowski R, Davis KW, Woods MA et al (2009) Knee joint: comprehensive assessment with 3D isotropic resolution fast spin-echo MR imaging--diagnostic performance compared with that of conventional MR imaging at 3.0 T. Radiology 252:486–495
Notohamiprodjo M, Horng A, Kuschel B et al (2012) 3D-imaging of the knee with an optimized 3D-FSE-sequence and a 15-channel knee-coil. Eur J Radiol 81:3441–3449
Notohamiprodjo M, Horng A, Pietschmann MF et al (2009) MRI of the knee at 3T: first clinical results with an isotropic PDfs-weighted 3D-TSE-sequence. Invest Radiol 44:585–597
Ristow O, Steinbach L, Sabo G et al (2009) Isotropic 3D fast spin-echo imaging versus standard 2D imaging at 3.0 T of the knee--image quality and diagnostic performance. Eur Radiol 19:1263–1272
Breuer FA, Blaimer M, Mueller MF et al (2006) Controlled aliasing in volumetric parallel imaging (2D CAIPIRINHA). Magn Reson Med 55:549–556
Wright KL, Harrell MW, Jesberger JA et al (2014) Clinical evaluation of CAIPIRINHA: comparison against a GRAPPA standard. J Magn Reson Imaging 39:189–194
Yutzy SR, Seiberlich N, Duerk JL, Griswold MA (2011) Improvements in multislice parallel imaging using radial CAIPIRINHA. Magn Reson Med 65:1630–1637
Schweitzer ME (2016) Evidence level. J Magn Reson Imaging 43:543
Bidgood WD Jr, Horii SC (1992) Introduction to the ACR-NEMA DICOM standard. Radiographics 12:345–355
Clanton TO, Ho CP, Williams BT et al (2014) Magnetic resonance imaging characterization of individual ankle syndesmosis structures in asymptomatic and surgically treated cohorts. Knee Surg Sports Traumatol Arthrosc. doi:10.1007/s00167-014-3399-1
Boonthathip M, Chen L, Trudell D, Resnick D (2011) Lateral ankle ligaments: MR arthrography with anatomic correlation in cadavers. Clin Imaging 35:42–48
Mengiardi B, Pfirrmann CW, Vienne P, Hodler J, Zanetti M (2007) Medial collateral ligament complex of the ankle: MR appearance in asymptomatic subjects. Radiology 242:817–824
Lektrakul N, Chung CB, Lai Y et al (2001) Tarsal sinus: arthrographic, MR imaging, MR arthrographic, and pathologic findings in cadavers and retrospective study data in patients with sinus tarsi syndrome. Radiology 219:802–810
Thacker P, Mardis N (2013) Ligaments of the tarsal sinus: improved detection, characterisation and significance in the paediatric ankle with 3-D proton density MR imaging. Pediatr Radiol 43:196–201
Mengiardi B, Zanetti M, Schottle PB et al (2005) Spring ligament complex: MR imaging-anatomic correlation and findings in asymptomatic subjects. Radiology 237:242–249
Melao L, Canella C, Weber M, Negrao P, Trudell D, Resnick D (2009) Ligaments of the transverse tarsal joint complex: MRI-anatomic correlation in cadavers. AJR Am J Roentgenol 193:662–671
Landis JR, Koch GG (1977) The measurement of observer agreement for categorical data. Biometrics 33:159–174
Jung JY, Yoon YC, Kwon JW, Ahn JH, Choe BK (2009) Diagnosis of internal derangement of the knee at 3.0-T MR imaging: 3D isotropic intermediate-weighted versus 2D sequences. Radiology 253:780–787
Notohamiprodjo M, Kuschel B, Horng A et al (2012) 3D-MRI of the ankle with optimized 3D-SPACE. Invest Radiol 47:231–239
Rosenberg ZS, Beltran J, Bencardino JT (2000) From the RSNA Refresher Courses. Radiological Society of North America. MR imaging of the ankle and foot. Radiographics 20 Spec No:S153–179
Stevens KJ, Busse RF, Han E et al (2008) Ankle: isotropic MR imaging with 3D-FSE-cube--initial experience in healthy volunteers. Radiology 249:1026–1033
Fritz J, Fritz B, Thawait GG, Meyer H, Gilson WD, Raithel E (2016) Three-Dimensional CAIPIRINHA SPACE TSE for 5-minute high-resolution MRI of the knee. Invest Radiol 51(10):609–617
Yi J, Cha JG, Lee YK, Lee BR, Jeon CH (2016) MRI of the anterior talofibular ligament, talar cartilage and os subfibulare: comparison of isotropic resolution 3D and conventional 2D T2-weighted fast spin-echo sequences at 3.0 T. Skeletal Radiol. doi:10.1007/s00256-016-2367-x
Acknowledgements
The authors would like to thank Zhang Qiong for his work on the pulse sequence prototype.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Guarantor
The scientific guarantor of this publication is Jan Fritz.
Conflict of interest
The authors of this manuscript declare relationships with the following companies: Siemens AG and and Alexion Pharmaceuticals, Inc.
Funding
This study has received funding by Siemens AG.
Statistics and biometry
One of the authors has significant statistical expertise.
Informed consent
Written informed consent was obtained from all subjects in this study.
Ethical approval
Institutional Review Board approval was obtained.
Study subjects or cohorts overlap
No study subjects or cohorts have been previously reported.
Methodology
Prospective, experimental, performed at one institution.
Rights and permissions
About this article
Cite this article
Kalia, V., Fritz, B., Johnson, R. et al. CAIPIRINHA accelerated SPACE enables 10-min isotropic 3D TSE MRI of the ankle for optimized visualization of curved and oblique ligaments and tendons. Eur Radiol 27, 3652–3661 (2017). https://doi.org/10.1007/s00330-017-4734-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-017-4734-y