Abstract
Microorganisms have a crucial role to play in the cycling of nutrients within glacial environments. These systems are often nutrient-limited, and so biogeochemical reactions, which ensure the availability of nutrients for microbial communities, are critical for the maintenance of these systems. This study uses molecular biology to characterise the supraglacial cryoconite microbial communities that are capable of cycling carbon and nitrogen in a range of glacial environments. Organisms with the potential to photosynthesise were identified, including Cyanobacteria, Actinobacteria, Betaproteobacteria, Stramenopiles and Haptophyceae. Organisms with the potential to perform nitrification and denitrification processes were also identified and featured Betaproteobacteria, Alphaproteobacteria, Thaumarchaeota and Cyanobacteria. While it is unlikely that the chemical and physical parameters of the supraglacial environment will facilitate optimal rates of all of the nitrogen-related biogeochemical processes, the transport of these cryoconite communities to downstream locations, where more favourable conditions may prevail, will perhaps provide a valuable inoculation of microorganisms with the genetic potential to catalyse these reactions elsewhere.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Microbial communities have an essential role to play in the cycling of nutrients in all ecosystems (reviewed in Zehr et al. 2003). In the cryosphere, microorganisms are thought to perform biogeochemical cycles within soils (Nordin et al. 2004), snow (Jones and Deblois 1986), lakes (Canfield and Green 1985) and both in supraglacial (Hodson et al. 2005; Telling et al. 2010, 2011) and subglacial (Tranter et al. 1994; Hodson et al. 2005; Barker et al. 2006; Wynn et al. 2006, 2007; Boyd et al. 2011) environments. Supraglacial microbial communities, for example, within the snowpack or within cryoconite debris (0.1–3 mm dark, granular aggregates: Gerdel and Drouet 1960; Takeuchi et al. 2001b), are known to cycle carbon through photosynthesis and respiration pathways (Fogg 1967; Säwström et al. 2002; Mueller et al. 2005; Foreman et al. 2007; Hodson et al. 2007; Stibal and Tranter 2007; Anesio et al. 2009; Hodson et al. 2010a, b; Telling et al. 2010). They are also thought to contribute towards nitrogen cycling within the glacial environment through the catalysis of nitrification (Hodson et al. 2005; Wynn et al. 2007) and nitrogen fixation (Telling et al. 2011). It has been suggested that the transport of organic carbon and nitrogen, associated with supraglacial microbial communities, to the subglacial environment provides a key substrate for microbial biochemical processes in the subglacial environment (Hodson et al. 2005; Wynn et al. 2007). Cryoconite holes (water filled depressions upon the glacier surface containing a layer of cryoconite debris) have been identified as being important hydrological and biological systems within glacial environments, providing refuge from the extreme conditions of the cryosphere (reviewed in MacDonell and Fitzsimons 2008). Within cryoconite holes, and in particular within cryoconite debris, diverse communities of bacterial, eukaryotic and archaeal microorganisms exist (Säwström et al. 2002; Edwards et al. 2011; Cameron et al. 2012). These communities have been found to be metabolically active during the summer (Säwström et al. 2002; Hodson et al. 2007; Stibal and Tranter 2007; Anesio et al. 2009; Hodson et al. 2010a, b; Telling et al. 2010, 2011).
Organisms found within cryoconite communities can potentially be linked into a multi-trophic web (Säwström et al. 2002). At the base of this food web, bacterial and eukaryotic autotrophic organisms exist, with the potential to fix atmospheric carbon dioxide into biologically available organic carbon, through photosynthesis. The first rate limiting step of photosynthesis is catalysed by the ribulose-1,5-biphosphate carboxylase/oxygenase (RubisCO) enzyme (Ellis 1979), of which the large subunits of the predominant form (form I; Spiridonova et al. 2004; Selesi et al. 2005) are encoded by cbbL genes (Kusian and Bowien 1997: otherwise named as the rbcL gene in older nomenclature or when referring to eukaryotic organisms; Tabita 1988). CbbL genes have been found within green-like autotrophic bacterial groups (including plants, green algae, Cyanobacteria and representatives of some Alpha-, Beta- and Gammaproteobacteria), red-like autotrophic bacterial groups (including non-green algae and representatives of some Alpha- and Betaproteobacteria; Watson and Tabita 1997; Spiridonova et al. 2004; Selesi et al. 2005) and autotrophic eukaryotes.
In the nitrogen cycle, nitrogen gas is fixed into ammonia (NH3) by nitrogen fixation which is catalysed by the nitrogenase enzyme (Howard and Rees 1996; Fig. 1); encoded by the nitrogen fixation (nif) gene (Zehr et al. 2003). Ammonia readily converts to ammonium (NH +4 , ionised ammonia) under acidic pH conditions (Howard and Rees 1996). The collective process of nitrification is an energy-producing reaction involving the aerobic oxidation of ionised ammonia into nitrite (NO −2 ) (by ammonia oxidation) and nitrite into nitrate (NO −3 ) (by nitrite oxidation; reviewed by Bothe et al. 2007). The first step of ammonia oxidation is catalysed by the ammonia monooxygenase protein (Hollocher et al. 1981), the first subunit of which is encoded by the amoA gene (McTavish et al. 1993). The reduction of nitrate to nitrite is catalysed by nitrate reductase proteins that are either membrane-bound (encoded by the nar operon; Warnecke-Eberz and Friedrich 1993) or are located within the periplasm (encoded by the nap gene; Siddiqui et al. 1993). Once nitrite is formed, it can be reduced further by one of the three main anaerobic pathways: (1) the multistep reduction of nitrite to form dinitrogen gas – termed denitrification (Zumft 1997), (2) the formation of ammonium by dissimilatory nitrate reduction to ammonium (DNRA) (Knowles 1982) or (3) the coupling of ammonium oxidation to the reduction of nitrite to form dinitrogen gas by anaerobic ammonium oxidation (anammox) (Mulder et al. 1995; van de Graaf et al. 1995). In denitrification, nitrite is reduced to nitric oxide (NO) using either of two nitrite reductase proteins, NirS and NirK (encoded by the nitrite respiration genes, nirS and nirK); nitric oxide is then reduced to nitrous oxide (N2O) using nitric oxide reductase (encoded by the nitric oxide respiration gene, nor) and nitrous oxide is reduced to dinitrogen gas using nitrous oxide reductase (encoded by the nitrous oxide respiration gene, nosZ; Zumft 1997). DNRA is catalysed by formate dehydrogenases, encoded by the nrfA gene (Darwin et al. 1993), and the anammox pathway is partially catalysed by hydroxylamine oxidoreductase, which is encoded by the hao gene (Schalk et al. 2000; Strous et al. 2006).
Major biochemical processes of the nitrogen cycle. The functional genes that were targeted within this study are shown in lowercase italics (with the exception of the nor gene which is expressed as qnor). The specific gene locus that was targeted is described after the functional gene in uppercase and subscript italics. The dashed arrow depicts the non-microbially mediated transformation of ammonia to ammonium
This study aims to establish whether genes encoding some of the key enzymes, introduced above, that catalyse the biogeochemical reactions of the carbon and nitrogen cycle are detectable in cryoconite debris. The identification of these genes within cryoconite communities establishes the genetic potential for these processes to occur and highlights the possible role of cryoconite communities in the nutrient cycling of the supraglacial environment. Specifically, genes associated with photosynthesis, nitrogen fixation, nitrification, denitrification and DNRA were examined. Furthermore, the diversity of organisms containing these genes was investigated through sequence analysis, allowing for the diversity of organisms that may be important in catalysing these biogeochemical reactions within cryoconite hole environments to be examined.
Materials and methods
Sampling sites and cryoconite hole sampling
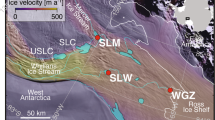
Cryoconite was sampled from individual holes at 9 glacial locations in Northern and Southern Hemisphere Polar regions (Table 1; Fig. 2). At all locations, one hole was sampled, with the exception of the Svalbard Midtre Lovénbreen location, where four holes were sampled. Cryoconite granules were extracted using a sterile plastic pasteur pipette. Material was placed into sterile Eppendorf tubes and any excess liquid was removed. Samples were frozen at −20 °C on return to the field laboratories. Material was transported back to the UK frozen and on arrival, the samples were stored at −80 °C. Table 2 shows the chemical properties of the cryoconite water that bathes the cryoconite debris. Nutrients were generally measured at sub-mg L−1 levels, suggesting that the cryoconite samples tended to be bathed in very dilute glacial meltwaters. The exception was Signy Island, where snowmelt is heavily enriched in atmospherically derived marine aerosol and nutrients from a local penguin colony (most notably inorganic nitrogen and phosphorous; Hodson 2006). Elsewhere, PO 3−4 levels are very often below the detection limit of conventional ion chromatography (ca. 0.01 mg L−1). Levels of DOC are not well known at many of the study sites, but most likely lie in the range 0.1–1.0 mg L−1 according to studies elsewhere (e.g. Priscu and Christner 2004).
Location of cryoconite sampling sites in a Arctic and b Antarctic regions. Sampling locations: Greenland: South West Ice Sheet (G-K) and Kronprins Christian Land (G-Kp), Svalbard: Longyearbreen (S-L), Rieperbreen (S-R), Foxfonna (S-F), Midtre Lovénbreen (S-M) and Vestfonna (S-V), Norway: Jostedalsbreen Ice Cap (N-J), and Antarctic Signy Island (A-S)
Nucleic acid extractions from cryoconite granules
Genomic DNA was extracted from cryoconite using the PowerSoil™ DNA isolation kit (Mo Bio laboratories, Cambridge, UK) in accordance with the manufacturer’s instructions and using approximately 0.8 g (dry weight) of solid cryoconite material. DNA was eluted into a final volume of 30 μL of nuclease-free water (Ambion, Warrington, UK).
Polymerase chain reaction (PCR) amplification
Functional genes were amplified using polymerase chain reaction (PCR). Reactions (50 μL) comprised 0.4 μM of each forward and reverse oligonucleotide primer, 200 μM of each deoxyribonucleotide triphosphate (dNTPs), 1 mM MgCl2, 1× buffer (Bioline, London, UK), 2.5 U of Taq polymerase (Bioline) and 1 μL of DNA. The primers and PCR cycling conditions are detailed in Table 3. PCR products were visualised by gel electrophoresis with ethidium bromide staining to ensure the correct size fragment was amplified.
Clone library construction, sequencing and analysis
PCR products were purified using the QIAquick PCR purification kit (Qiagen), ligated into the pCR2.1®-TOPO® TA cloning vector (Invitrogen, Paisley, UK) and transformed into One Shot® Chemically Competent Escherichia coli TOP10 F’ cells (Invitrogen) in accordance with the manufacturer’s instructions. Transformed cells were plated on Luria–Bertani (LB) agar containing ampicillin (50 μg mL−1) and X-gal (20 μg mL−1) and incubated overnight at 37 °C. White colonies were picked at random and used to inoculate 100 μL of LB broth containing ampicillin. After 2-h incubation at 37 °C, 1 μL of each culture was used in a PCR to amplify insert DNA using vector-specific primers T3 (5′-ATT AAC CCT CAC TAA AGG GA-3′) and T7 (5′-TAA TAC GAC TCA CTA TAG GG-3′) (Invitrogen). The amplified vector inserts were purified using SureClean (Bioline) and were sequenced using a BigDye® Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems) with T3 and T7 primers. Sequencing was performed in an ABI 3730 Genetic Analyser. Sequences were edited manually using Chromas Pro version 1.34 (http://www.technelysium.com.au). Chimeras were identified using Mallard version 1.02 (Ashelford et al. 2006). FASTA formatted sequences were translated into protein sequences using ExPASy (Gasteiger et al. 2003), and protein sequences were compared to the GenBank database using Basic Local Alignment Search Tool for protein (BLASTp; Altschul et al. 1990). Bioedit version 7.0.5.3 (Hall 1999) was used to align the library of sequences to themselves and to the top BLAST hits using ClustalW (Thompson et al. 1994), which also presented an additional opportunity to highlight and alter any errors within the sequence reading. Mega version 4.1 (Tamura et al. 2007) was used to calculate evolutionary distances and create phylogenetic trees, using the Neighbour-Joining method (Saitou and Nei 1987) with bootstrap analysis (1,000 replicates).
Nucleotide sequence accession numbers
Nucleotide sequences were submitted to the EMBL nucleotide sequence database. The following accession numbers were created: cbblR: HE774752–HE774812, cbblG: HE774813–HE774836, rbcL: HE774837–HE774905, napA: HE774906–HE774915, narG: HE774916–HE775005, qnorB: HE775006–HE775016, nosZ: HE775017–HE775086, amoA: HE792983–HE793010, nirS: HE793011–HE793030.
Results
Distribution and diversity of cbbL genes within cryoconite communities
To investigate the presence and diversity of non-green bacterial primary producers, partial cbbLR sequences, encoding the large subunit of form I red-like RubisCO, were amplified from all of the cryoconite communities studied (five communities were investigated from three different glacial locations), using primers designed by Selesi et al. (2005) (Table 3). When gene products, amplified from a Svalbard Midtre Lovénbreen and an Antarctic Signy Island cryoconite community, were sequenced, the phylogenetic diversity of these two communities differed (Fig. 3). The majority of the Midtre Lovénbreen gene sequences (97 %) were most closely related (74–76 % identity) to cbbL gene sequences from an Actinobacteria Mycobacterium organism (accession number: EU026272, Park et al. 2009), while only 6 % of the Signy Island sequences were most closely related to this sequence (with each clone sequence sharing 75 % identity to the database sequence). The majority of the Signy Island sequences (72 %) were most closely related (80–100 % identity) to sequences of the cbbL gene from Betaproteobacteria Burkholderiales, for example, Ralstonia eutropha (accession number: AM260480, Pohlmann et al. 2007). Both the Signy Island library and the Midtre Lovénbreen library each contained a single clone that was most closely related to Alphaproteobacteria Rhizobiales cbbL sequences (82 and 81 % identity, respectively; Fig. 3).
Neighbour-joining phylogeny of CbbLR protein sequences, cloned from Antarctic Signy Island (A-S1) and Svalbard Midtre Lovénbreen (S-M2) locations and from closest-related GenBank database sequences. The number of clones within each collapsed tree branch is indicated. Closest-related database sequences (with accession number) are indicated by a circle. Bootstrap values of >75 % are shown (from 1,000 replicates). Scale bar represents 5 % sequence divergence
Amplifications of the cbbLG gene, encoding the large subunit of form I green-like RubisCO, yielded PCR products of the correct size in all of the communities studied (six communities were investigated from three different glacial locations) when amplified using the RubIg primer set (Spiridonova et al. 2004; Table 3). When products from a Svalbard Midtre Lovénbreen community were sequenced, the majority of the clones (96 %) were found to be most closely related to cyanobacterial cbbL gene sequences (Fig. 4). 84 % of the clones were most closely related to Cyanobacteria Oscillatoriales Leptolyngbya sequences (e.g. accession number: AB075914, Tomitani et al. 2006, 83–86 % identity), while 12 % of the clones were most closely related to a Cyanobacteria Nostoc sequence (accession number: AB075918, Tomitani et al. 2006, 88–90 % identity). One clone showed 90 % identity to a eukaryotic haptophyceae Pavlova lutheri sequence (accession number: AY119785, Yoon et al. 2002; Fig. 4).
Neighbour-joining phylogeny of CbbLG protein sequences, cloned from Svalbard Midtre Lovénbreen (S-M2) using RubIg oligonucleotide primers (Spiridonova et al. 2004) and from closest-related GenBank database sequences. The number of clones within each collapsed branch is indicated. Closest related database sequences (with accession number) are indicated by a circle. Bootstrap values of >75 % are shown (from 1,000 replicates). Scale bar represents 5 % sequence divergence
Primer sets, designed to amplify genes encoding the large subunit of RubisCO within eukaryotes (Wawrik et al. 2002) were used to investigate the presence and diversity of these genes within cryoconite communities. rbcL genes were amplified from all of the communities investigated (six communities were investigated from three different glacial locations; Table 4). Clones from an Arctic (S-M2) and an Antarctic (A-S1) community were sequenced and revealed distinct communities. Clones from the Svalbard Midtre Lovénbreen community were all closely related to Stramenopiles Xanthophyceae organisms, with the majority of the clones (82.5 %) having 98–99 % identity to Botrydiopsis constricta rbcL sequences (accession number: AJ579566, Negrisolo et al. 2004). Clones from the Antarctic Signy Island community contained sequences related to a greater range of taxa, including those from Stramenopiles and Haptophyceae (Fig. 5). Of the Stramenopiles identified within the A-S1 community, clones with similarities to Xanthophyceae, Synurophyceae and Chrysophyceae sequences were identified (Fig. 5).
Neighbour-joining phylogeny of RbcL protein sequences, cloned from Antarctic Signy Island (A-S1) and Svalbard Midtre Lovénbreen (S-M2) locations and from closest-related GenBank database sequences. The number of clones within each collapsed branch is indicated. Closest-related database sequences (with accession number) are indicated by a circle. Bootstrap values of >75 % are shown (from 1,000 replicates). Scale bar represents 2 % sequence divergence
Distribution and diversity of nitrogen cycling functional genes within cryoconite communities
Nitrogen fixation
The presence of a nitrogen fixing community within cryoconite was investigated through the identification of the nifH gene, one of three genes encoding the nitrogen fixating nitrogenase enzyme. Despite several attempts to optimise PCR conditions, amplification of the nifH gene from cryoconite communities was unsuccessful when degenerate oligonucleotide primers, designed by Zehr and McReynolds (1989), were used (six communities were investigated from five different glacial locations; Table 4).
Nitrification
To investigate the presence and diversity of a nitrifying community, bacterial and archaeal amoA genes, encoding ammonia monooxygenase enzymes that catalyse ammonia oxidation, were amplified from cryoconite. Batcerial- and archaeal-related PCR products were amplified from all 10 of the communities investigated (from eight different glacial locations); however, both bacterial- and archaeal-specific amoA gene primers produced amplicons of multiple lengths from each of the communities studied (Table 4). When PCR products from the Signy Island community (A-S1) were excised and sequenced, bacterial- and archaeal-specific amoA genes were confirmed to be present. Bacterial-related amoA genes were similarly isolated and sequenced from the Svalbard Rieperbreen community. Purification and sequencing of archaeal-specific amoA gene products (amoAarc; Treusch et al. 2005) of the correct size [around 532 base pairs (bp)] from the A-S1 community resulted in five clones, all of which were closely related (sharing 93–96 % identity) to amoA genes of uncultured Crenarchaeote (now Thaumarchaeota) clones (e.g. accession number: EU022817, Santoro et al. 2008). Amplification and sequencing of bacterial-specific amoA genes of the correct size (approximately 470 bp), using primers designed by Rotthauwe et al. (1997) (amoAbac), from an Antarctic Signy Island (A-S1) and a Svalbard Rieperbreen (S-R1) community was successfully undertaken. Sequencing of these bacterial amoA products revealed clones that were all closely related (92–97 % identity) to amoA genes from uncultured ammonia-oxidising Betaproteobacteria (e.g. accession number: EF615145, Kim et al. 2008) or from uncultured Betaproteobacteria belonging to the Nitrosomonadaceae family (e.g. accession number: AY189142, Mintie et al. 2003; Fig. 6).
Neighbour-joining phylogeny of AmoAbac protein sequences, cloned from Antarctic Signy Island (A-S1) and Svalbard Reiperbreen (S-R1) locations and closest-related GenBank database sequences. The number of clones within each collapsed branch is indicated. Closest related database sequences (with accession number) are indicated by a circle, and the environmental origin of uncultured clones is detailed. Bootstrap values of >75 % are shown (from 1,000 replicates). Scale bar represents 2 % sequence divergence
Nitrate reduction
Cryoconite communities were analysed for the presence and diversity of genes encoding protein products essential for denitrification and DNRA and for the presence of 16S rRNA genes of anammox bacteria. Genes encoding periplasmic and membrane-bound nitrate reductase (napA and narG, respectively) were amplified from cryoconite to investigate the presence and diversity of communities capable of reducing nitrate to nitrite. napA and/or narG genes were amplified from all ten of the cryoconite communities investigated (from eight different glacial locations; Table 4). A clone library was made of napA genes from a Svalbard Foxfonna community (S-F2), in which napA clone sequences shared 76–84 % identity to the most closely related cultured sequences, which belonged to either Alpha-, Beta- or Gammaproteobacteria classes. Clone libraries of the narG gene were constructed from an Antarctic Signy Island community and from Svalbard communities, including Midtre Lovénbreen, Foxfonna, Rieperbreen and Vestfonna. When clone sequences were compared to the GenBank database, the sequences that were most closely related (89–94 % identity) were those of narG genes belonging to unidentified or uncultured bacterial clones with no classification descriptions (e.g. accession number: EU052949, Strief et al. unpublished; Fig. 7). Lower levels of similarity (80–91 % identity) were seen between clone sequences and sequences related to those from Alphaproteobacteria, for example the Rhodobacterales Paracoccus denitrificans (accession number: CP000490, Copeland et al. unpublished) and the Rhizobiales Bradyrhizobium (accession number: CP000494, Giraud et al. 2007), as well as to Betaproteobacteria Burkholderiales, for example Polaromonas naphthalenivorans (accession number: CP000529, Copeland et al. unpublished; Fig. 7).
Neighbour-joining phylogeny of NarG protein sequences, cloned from Antarctic Signy Island (A-S1), Svalbard Reiperbreen (S-R1), Svalbard Midtre Lovénbreen (S-M5) and Svalbard Vestfonna (S-V1) locations and closest-related GenBank database sequences. The number of clones within each collapsed branch is indicated. Closest-related database sequences (with accession number) are indicated by a circle, and the environmental origin of uncultured clones is detailed. Bootstrap values of >75 % are shown (from 1,000 replicates). Scale bar represents 5 % sequence divergence
Genes associated with the reduction of nitrite were also investigated within cryoconite communities. nirS genes, encoding nitrite reductase proteins, which are essential for denitrification, were amplified from all seven of the communities investigated (from four different glacial locations; Table 4). However, PCR products from several of the communities contained fragments of multiple lengths, one of which was of the expected 858 bp length. PCR products from an Antarctic Signy Island (A-S1) and a Svalbard Foxfonna (S-F2) community were sequenced and were found to be most closely related (79–100 % identity) to nirS sequences from uncultured bacterial clones (e.g. accession number: AJ440485, Nogales et al. 2002). Cryoconite nirS sequences were identified with lower levels of identity (80–93 % identity) to members of four orders; Betaproteobacteria Rhodocyclales, for example Thauera terpenica (accession number: AY078266, Song and Ward 2006), Betaproteobacteria Burkholderiales, for example Acidovorax (accession number: AY078273, Song and Ward 2006), Alphaproteobacteria Rhizobiales, for example Bradyrhizobium japonicum (accession number: BA000040, Kaneko et al. 2002) and Gammaproteobacteria Pseudomonadales, for example Pseudomonas syringae (accession number: AE016853, Buell et al. 2003; Fig. 8).
Neighbour-joining phylogeny of NirS protein sequences, cloned from Antarctic Signy Island (A-S1) and Svalbard Foxfonna (S-F2) locations and closest-related GenBank database sequences. The number of clones within each collapsed branch is indicated. Closest-related database sequences (with accession number) are indicated by a circle, and the environmental origin of uncultured clones is detailed. Bootstrap values of >75 % are shown (from 1,000 replicates). Scale bar represents 5 % sequence divergence
The presence of nrfA genes within cryoconite communities, which encode nitrite reductase enzymes for DNRA, was investigated. PCR products of varying sizes were amplified from several communities when nrfA gene-specific primers were used. However, attempts to sequence these products were unsuccessful (eleven communities from eight glacial locations were attempted; Table 4). Molecular markers of the anammox reaction (identified using the Brod primer sequence set which targets the 16S rRNA genes of anammox-related bacteria (Penton et al. 2006) were not amplified from any of the cryoconite communities investigated within this study (nine communities from eight glacial locations were attempted; Table 4).
The nitric oxide reductase gene (qnorB), whose protein product is responsible for catalysing the reduction of nitric oxide to nitrous oxide, was amplified and sequenced from a Svalbard Foxfonna cryoconite community (S-F2) (Table 4). When sequences were compared to the GenBank database, similarities to the most closely related sequences ranged from 79 to 83 %. Three sequences with over 80 % identity to the cloned sequences were identified, including the Alphaproteobacteria Xanthobacter autotrophicus (accession number: CP000781, Copeland et al. unpublished, 81 % identity), the Deltaproteobacteria Sorangium cellulosum (accession number: AM746676, Schneiker et al. 2007, 81 % identity) and the Gammaproteobacteria Actinobacillus succinogenes (accession number: CP000746, Copeland et al. unpublished, 82 % identity). Further sequences, most closely related to uncultured bacteria clones, were additionally identified.
The nitrous oxide reductase gene (nosZ), whose protein product catalyses the reduction of nitrous oxide to nitrogen gas, was amplified and sequenced from four cryoconite communities (Antarctic Signy Island, Norway Jostedalsbreen, Svalbard Foxfonna and Svalbard Vestfonna). PCR products from a further six communities from four different glacial locations were amplified and were found to contain products of multiple lengths, one of which was a product of the expected length (1,021 bp; Table 4). nosZ genes were not amplified from the Greenland Kronprins Christian Land community (G-Kp2). When cloned sequences were compared to sequences in GenBank, highest similarities (89–98 % identity) were found to nosZ sequences from uncultured organisms of unknown lineage (e.g. accession number: AM419672, Braker et al. unpublished; Fig. 9). Matches to sequences of classified organisms with over 80 % similarity to the nosZ clones included Betaproteobacteria Burkholderiales (e.g. Ralstonia pickettii, accession number: CP001069, Lucas et al. unpublished, 80 % identity), Betaproteobacteria Rhodocyclales (e.g. Azoarcus, accession number: AM406670, Krause et al. 2006, 80–81 % identity) and Alphaproteobacteria Rhizobiales (Rhodopseudomonas palustris, accession number: CP000301, Copeland et al. unpublished, 82–84 % identity; Fig. 9).
Neighbour-joining phylogeny of NosZ protein sequences, cloned from Antarctic Signy Island (A-S1), Svalbard Foxfonna (S-F2), Svalbard Vestfonna (S-V1) and Norway Jostedalsbreen (N-J1) locations and closest-related GenBank database sequences. The number of clones within each collapsed branch is indicated. Closest-related database sequences (with accession number) are indicated by a circle, and the environmental origin of uncultured clones is detailed. Bootstrap values of >75 % are shown (from 1,000 replicates). Scale bar represents 20 % sequence divergence
Discussion
Genes encoding the large subunit of green-like and red-like bacterial form I RubisCO and eukaryotic form I RubisCO were identified through PCR amplification within all of the cryoconite communities investigated from Antarctic, Svalbard and Norway locations. Functional genes, associated with the carbon cycling of glacial microbial communities, have not been reported previously. However, molecular and observational diversity studies of cryoconite communities have revealed the presence of photosynthetic organisms, including Cyanobacteria and Chloroplastida (Gerdel and Drouet 1960; Christner et al. 2003; Mueller and Pollard 2004; Edwards et al. 2011). Additionally, studies measuring the rates of photosynthesis, respiration and protein synthesis (Säwström et al. 2002; Foreman et al. 2007; Hodson et al. 2007; Stibal and Tranter 2007; Anesio et al. 2009; Telling et al. 2010), and the acquisition, storage and loss of carbon from cryoconite holes (Lafrenière and Sharp 2004; Stibal et al. 2008) have revealed that carbon is imported, cycled within and exported out of this ecosystem during summer. Clones related to members of the Cyanobacteria Oscillatoriales and Nostocales orders were amplified from a Svalbard Midtre Lovénbreen community (S-M2) using primer sets designed to amplify the cbbLG gene. Other cryoconite diversity studies have also identified these Cyanobacterial classes (Christner et al. 2003; Porazinska et al. 2004; Mueller and Pollard 2004; Edwards et al. 2011; Cameron et al. 2012). Autotrophic bacteria containing red-like type I RubisCO cbbLR genes were identified within Burkholderiales, Actinobacteria and Rhizobiales taxa, which have been previously identified within the A-S1 and/or S-M2 communities via 16S rRNA gene analysis (Cameron et al. 2012). Additionally, clones related to the Betaproteobacteria Nitrosomonadales order, which have not been previously described with respect to cryoconite microbial communities, were identified within the A-S1 community.
The presence of eukaryotic autotrophs within cryoconite communities was investigated using primers designed to amplify the rbcL gene. Clones related to Stramenopile species, including Synurophyceae, Chrysophyceae and Xanthophyceae classes, were identified within the A-S1 and S-M2 communities, and clones related to Haptophyta Pavlovales were identified within the A-S1 community. Clones related to these organisms have been previously identified through 18S rRNA gene analysis of the same cryoconite communities (Cameron et al. 2012), and similarly, Haptophyta-related clones have been identified within an Antarctic Dry Valley cryoconite community (Christner et al. 2003).
Cyanobacteria have been noted within all cryoconite diversity studies (e.g. Gerdel and Drouet 1960; Wharton et al. 1985; Takeuchi et al. 2001a; Christner et al. 2003), and several studies have described cryoconite communities as being dominated by these organisms (Vincent et al. 2000; Säwström et al. 2002; Mueller and Pollard 2004). In addition to these Cyanobacteria, this current study has identified other organisms, such as photosynthetic Stramenopiles and Proteobacteria, which may also photosynthesize to produce biologically available carbon.
Genes encoding the nitrogenase enzyme for the catalysis of nitrogen fixation were unsuccessfully amplified from any of the cryoconite communities studied, regardless of the primer set having been designed to and having previously targeted Cyanobacterial nifH genes (Zehr and McReynolds 1989). Despite this, 16S rRNA gene diversity studies (Cameron et al. 2012) and functional gene studies of the cbbLG gene (presented here) identified the presence of Cyanobacteria species belonging to the Nostocaceae family within cryoconite originating from Antarctica, Svalbard and Greenland. Nostocaceae-related organisms contain heterocyst cells that are capable of nitrogen fixation (Tomitani et al. 2006). Furthermore, previous 16S rRNA gene diversity studies of cryoconite communities (e.g. Christner et al. 2003; Cameron et al. 2012) and this current study have revealed a diversity of Alphaproteobacteria, Betaproteobacteria, Gammaproteobacteria, Firmicutes and Archaea, a proportion of which may have the genetic potential to fix nitrogen. Further investigations using alternative nifH primer designs may be successful in identifying the range of organisms within these communities that can fix nitrogen. Although the presence of nitrogen fixation genes found within communities associated with supraglacial environments has not previously been published, Arctic subglacial environments (Boyd et al. 2011), Antarctic microbial mats (Jungblut and Neilan 2009) and microorganisms present within the ice cover of an Antarctic frozen lake (Olson et al. 1998) have been found to contain nifH genes. The glacial snowpack has been recognised as a major store and source of nutrients such as ammonium (Tranter et al. 1993; Kuhn 2001; Hodson et al. 2005). If cryoconite communities, containing nifH genes, are able to actively fix nitrogen (as demonstrated upon Midtre Lovénbreen; Telling et al. 2011 and upon Leverette glacier, Greenland; Telling et al. open discussion article), these supraglacial niches will contribute a further source of ammonium to downstream habitats.
Nitrification genes were amplified and sequenced from organisms related to Betaproteobacteria from both Arctic and Antarctic cryoconite communities and to archaeal Thaumarchaeota species from Antarctic cryoconite communities. The genetic potential for nitrification and measurements of nitrification process rates have not been previously studied within the supraglacial environment. However, nutrient budget studies have found that microbially mediated nitrification occurs in the region between the supraglacial snowpack and the ice margin, most likely within sedimentary environments such as cryoconite, subglacial till, lateral moraine and talus slopes (Tranter et al. 1994; Hodson et al. 2005; Wynn et al. 2007; Hodson et al. 2010a, b). Its occurrence in debris-poor habitats such as the snowpack is suspected, but nutrient addition experiments remain unequivocal (Wynn et al. 2007). Interestingly, an Arctic subglacial study (Boyd et al. 2011) identified similar communities of bacterial ammonia-oxidising amoA genes as have been found within this current study. In addition, these subglacial communities have been found to contain archaeal amoA genes (Boyd et al. 2011). The similarity of these ammonia-oxidising communities, between supraglacial and subglacial environments, is suggestive that supraglacial systems, such as cryoconite holes, act as microbial sources to the subglacial ecosystem.
Genes encoding enzymes to catalyse the four dissimilatory steps of denitrification, enabling the reduction of nitrate to dinitrogen gas, were amplified from several cryoconite communities of Arctic and Antarctic origin. The amplification and sequencing of these denitrifying functional genes revealed a diversity of Betaproteobacteria Burkholderiales, capable of catalysing each of the four stages of denitrification. Additionally, clones relating to Betaproteobacteria Rhodocyclales and Alphaproteobacteria Rhizobiales and Rhodobacterales were also identified through the amplification of two or more denitrifying functional genes. Denitrification processes have not been previously noted within supraglacial environments. However, subglacial microbial communities have been thought to reduce or deplete surface originating nitrate stocks through denitrification (Tranter et al. 1994; Hodson et al. 2005). Stable isotope measurements of 15N–NO3 presented by Wynn et al. (2006) also provide compelling evidence for the occurrence of this process at Midtre Lovénbreen. A Canadian Arctic subglacial narG gene study similarly found a high diversity of organisms capable of nitrate reduction by nitrate reductase (Boyd et al. 2011). In this subglacial community analysis by Boyd et al., many clones were featured with a high similarity to Polaromonas naphthalenivorans, as was similarly found within this present study.
Given the typical environmental conditions of cryoconite water (an oxidising environment with Eh values of over 300 mV; Foreman et al. 2007), the reductive process of denitrification seems an unlikely process to occur here. This is especially true when compared to the low redox environment of the subglacial environment (Eh = 90 mV; Wynn et al. 2007; Mikucki et al. 2009). However, the centre of each cryoconite granule is often a region of dark decomposing matter (Takeuchi et al. 2001a; Langford et al. 2011), which may provide an anoxic microzone for denitrification to occur. Similarly, cryoconite communities, carrying the genetic ability for denitrification, may be flushed into subglacial zones, where they may commence the denitrification process and seed the subglacial ecosystem. Interestingly, microbial communities found on the soils of a glacial forefield have been found to contain genes responsible for denitrification (Deiglmayr et al. 2006; Kandeler et al. 2006). Furthermore, within this current study, several Arctic and Antarctic narG sequences were most closely matched to narG sequences originating from a glacial forefield (accession number: DQ233263; Deiglmayr et al. 2006). Thus, it may be possible that the transportation of cryoconite either through the subglacial environment to the glacial forefield, or directly from the glacier surface during ice retreat, may contribute to the colonisation of these denitrifying communities (Porazinska et al. 2004; Foreman et al. 2007; Řehák et al. 2007; Hodson et al. 2008; Schütte et al. 2009). Similarly, the aeolian and aqueous transportation of fragments of local, and perhaps distant, niches onto the glacial surface are likely to be a major seedling factor of cryoconite communities and thus will have an influence over their functional potential (Mueller et al. 2001; Cameron et al. 2012). However, many of the sites in this current study are in close proximity to maritime, soil, lake, river and/or rock environments, and thus, the physical and chemical conditions of the supraglacial environment may be so different that they render alien populations from the surrounding environment inactive.
Genes encoding the periplasmic nitrite reductase enzyme, which catalyses the ammonification of nitrite (DNRA), were not identified by gene sequence analysis within this study. The inability to amplify nrfA genes raises the question of whether these systems do in fact have the genetic potential to process the direct reduction of nitrite to ammonium, or whether these genes were simply not detected due to methodological reasons (von Wintzingerode et al. 1997). DNRA processes may compete with denitrification under anaerobic conditions, influenced by nitrate concentrations and the environmental redox potential (Matheson et al. 2002; Dong et al. 2009). As with denitrification processes, DNRA reactions may be limited or absent within cryoconite holes. However, even the dispersal of small numbers of DNRA-capable organisms to other niches (with lower redox potentials) may enable these communities to thrive. Within the low nutrient abundance that is typical of glacial environments (Table 2), DNRA processes, which prevent the loss of nitrogen from the biosphere, would be advantageous over denitrification (Matheson et al. 2002). Anammox catalysing communities were not identified within cryoconite. However, the Brod primer set (Penton et al. 2006), which was used for analysis, was designed to target 16S rRNA gene sequences of the Planctomycetes phylum, which have not previously been identified within cryoconite communities. Thus, further studies relating to the detection of anammox pathways within these systems are urged in the future.
Conclusion
Genes associated with photosynthesis, nitrification and denitrification were identified within all of the Arctic and Antarctic communities investigated. Additionally, organisms with the ability to perform nitrogen fixation were identified in previous diversity studies. Sequence data from functional gene analyses revealed the potential photosynthetic importance of Cyanobacteria, Actinobacteria, Betaproteobacteria, Stramenopiles and Haptophyceae within cryoconite. Similarly, Betaproteobacteria and Thaumarchaeota organisms, containing genes for nitrification, and Alphaproteobacteria and Betaproteobacteria, carrying genes for nitrate reduction via denitrification, were identified as being potentially important for nitrogen cycling within cryoconite communities. Although the process rates of the biogeochemical reactions under investigation within this current study are unknown, the genetic potential of these communities to undertake microbially mediated carbon and nitrogen cycling is an important one. In addition, the eventual transportation of cryoconite communities to other niches within the biologically sparse glacial environment (Porazinska et al. 2004; Foreman et al. 2007; Stibal et al. 2008; Schütte et al. 2009) may provide a valuable source of organisms with the potential to perform biogeochemical cycling processes elsewhere.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Bio 215:403–410
Anesio AM, Hodson AJ, Fritz A, Psenner R, Sattler B (2009) High microbial activity on glaciers: importance to the global carbon cycle. Glob Change Bio 15:955–960
Ashelford KE, Chuzhanova NA, Fry JC, Jones AJ, Weightman AJ (2006) New screening software shows that most recent large 16S rRNA gene clone libraries contain chimeras. Appl Env Microbiol 72:5734–5741
Barker J, Sharp M, Fitzsimons S, Turner R (2006) Abundance and dynamics of dissolved organic carbon in glacier systems. Arct Antarct Alp Res 38:163–172
Bothe H, Ferguson SJ, Newton WE (2007) Biology of the nitrogen cycle. Elsevier, Oxford
Boyd ES, Lange RK, Mitchell AC, Havig JR, Hamilton TL, Lafrenière MJ, Shock EL, Peters JW, Skidmore M (2011) Diversity, abundance, and potential activity of nitrifying and nitrate-reducing microbial assemblages in a subglacial ecosystem. Appl Environ Microbiol 77:4778–4787
Braker G, Tiedje JM (2003) Nitric oxide reductase (norB) genes from pure cultures and environmental samples. Appl Environ Microbiol 69:3476–3483
Braker G, Fesefeldt A, Witzel KP (1998) Development of PCR primer systems for amplification of nitrite reductase genes (nirK and nirS) to detect denitrifying bacteria in environmental samples. Appl Environ Microbiol 64:3769–3775
Buell CR, Joardar V, Lindeberg M, Selengut J, Paulsen IT, Gwinn ML, Dodson RJ, Deboy RT, Durkin AS, Kolonay JF, Madupu R, Daugherty S, Brinkac L, Beanan MJ, Haft DH, Nelson WC, Davidsen T, Zafar N, Zhou L, Liu J, Yuan Q, Khouri H, Fedorova N, Tran B, Russell D, Berry K, Utterback T, Van Aken SE, Feldblyum TV, D’Ascenzo M, Deng WL, Ramos AR, Alfano JR, Cartinhour S, Chatterjee AK, Delaney TP, Lazarowitz SG, Martin GB, Schneider DJ, Tang X, Bender CL, White O, Fraser CM, Collmer A (2003) The complete genome sequence of the Arabidopsis and tomato pathogen Pseudomonas syringae pv. tomato DC3000. Proc Natl Acad Sci USA 100:10181–10186
Cameron KA, Hodson AJ, Osborn AM (2012) Structure and diversity of bacterial, eukaryotic and archaeal communities in glacial cryoconite holes from the Arctic and the Antarctic. Appl Environ Microbiol. doi:10.1111/j.1574-6941.2011.01277.x
Canfield DE, Green WJ (1985) The cycling of nutrients in a closed-basin Antarctic lake: Lake Vanda. Biogeochem 1:233–256
Christner BC, Kvitko BH, Reeve JN (2003) Molecular identification of bacteria and eukarya inhabiting an Antarctic cryoconite hole. Extremophiles 7:177–183
Clausen HB, Stampe M, Hammer CU, Hvidberg CS, Dahl-Jensen D, Steffensen JP (2001) Glaciological and chemical studies on ice cores from Hans Tausen Iskappe, Greenland. Meddelelser om Grønland Geosci 39:123–149
Darwin A, Hussain H, Griffiths L, Sambongi Y, Busby S, Cole J (1993) Regulation and sequence of the structural gene for cytochrome c552 from Escherichia coli: not a hexahaem but a 50 kDa tetrahaem nitrite reductase. Mol Microbiol 9:1255–1265
Deiglmayr K, Philippot L, Tscherko D, Kandeler E (2006) Microbial succession of nitrate-reducing bacteria in the rhizosphere of Poa alpina across a glacier foreland in the Central Alps. Environ Microbiol 8:1600–1612
Dong LF, Smith CJ, Papaspyrou S, Stott A, Osborn AM, Nedwell DB (2009) Changes in benthic denitrification, nitrate ammonification, and anammox process rates and nitrate and nitrite reductase gene abundances along an estuarine nutrient gradient (the Colne Estuary, United Kingdom). Appl Environ Microbiol 75:3171–3179
Edwards A, Anesio AM, Rassner SM, Sattler B, Hubbard B, Perkins WT, Young M, Griffith GW (2011) Possible interactions between bacterial diversity, microbial activity and supraglacial hydrology of cryoconite holes in Svalbard. ISME J 5:150–160
Ellis RJ (1979) The most abundant protein in the world. Trends Biochem Sci 4:241–244
Flanagana D, Gregorya L, Cartera J, Karakas-Sena A, Richardsona D, Spiroa S (2006) Detection of genes for periplasmic nitrate reductase in nitrate respiring bacteria and in community DNA. FEMS Microbiol Lett 177:263–270
Fogg GE (1967) Observations on the snow algae of the South Orkney Islands. Philos Trans R Soc Lond B Biol Sc Biol Sci 252:279–287
Foreman CM, Sattler B, Mikucki JA, Porazinska DL, Priscu JC (2007) Metabolic activity and diversity of cryoconites in the Taylor Valley, Antarctica. J Geophys Res. doi:10.1029/2006JG000358
Gasteiger E, Gattiker A, Hoogland C, Ivanyi I, Appel RD, Bairoch A (2003) ExPASy: the proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res 31:3784–3788
Gerdel RW, Drouet F (1960) The cryoconite of the Thule Area, Greenland. Trans Am Microsc Soc 79:256–272
Giraud E, Moulin L, Vallenet D, Barbe V, Cytryn E, Avarre JC, Jaubert M, Simon D, Cartieaux F, Prin Y, Bena G, Hannibal L, Fardoux J, Kojadinovic M, Vuillet L, Lajus A, Cruveiller S, Rouy Z, Mangenot S, Segurens B, Dossat C, Franck WL, Chang WS, Saunders E, Bruce D, Richardson P, Normand P, Dreyfus B, Pignol D, Stacey G, Emerich D, Verméglio A, Médigue C, Sadowsky M (2007) Legumes symbioses: absence of nod genes in photosynthetic Bradyrhizobia. Science 316:1307–1312
Hall T (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hodson AJ (2006) Biogeochemistry of snowmelt in an Antarctic glacial ecosystem. Water Resour Res. doi:10.1029/2005WR004311
Hodson AJ, Mumford PN, Kohler J, Wynn PM (2005) The High Arctic glacial ecosystem: new insights from nutrient budgets. Biogeochem 72:233–256
Hodson A, Anesio AM, Ng F, Watson R, Quirk J, Irvine-Fynn T, Dye A, Clark C, McCloy P, Kohler J, Sattler B (2007) A glacier respires: quantifying the distribution and respiration CO2 flux of cryoconite across an entire Arctic supraglacial ecosystem. J Geophys Res. doi:10.1029/2007JG000452
Hodson A, Anesio AM, Tranter M, Fountain A, Osborn M, Priscu J, Laybourn-Parry J, Sattler B (2008) Glacial ecosystems. Ecol Monogr 78:41–67
Hodson A, Cameron K, Bøggild C, Irvine-Fynn T, Langford H, Pearce D, Banwart S (2010a) The structure, biological activity and biogeochemistry of cryoconite aggregates upon an Arctic valley glacier: Longyearbreen, Svalbard. J Glaciol 56:349–362
Hodson A, Bøggild C, Hanna E, Huybrechts P, Langford H, Cameron K, Houldsworth A (2010b) The cryoconite ecosystem on the Greenland ice sheet. Ann Glaciol 51:123–129
Hollocher TC, Tate ME, Nicholas DJ (1981) Oxidation of ammonia by Nitrosomonas europaea. Definite 18O-tracer evidence that hydroxylamine formation involves a monooxygenase. J Biol Chem 256:10834–10836
Howard JB, Rees DC (1996) Structural basis of biological nitrogen fixation. Chem Rev 96:2965–2982
Hunter EM, Mills HJ, Kostka JE (2006) Microbial community diversity associated with carbon and nitrogen cycling in permeable shelf sediments. Appl Environ Microbiol 72:5689–5701
Jones H, Deblois C (1986) Chemical dynamics of N-containing ionic species in a Boreal forest snowcover during the spring melt period. Hydrol Process 1:271–282
Jungblut AD, Neilan BA (2009) nifH gene diversity and expression in a microbial mat community on the McMurdo Ice Shelf, Antarctica. Antarct Sci 22:117–122
Kandeler E, Deiglmayr K, Tscherko D, Bru D, Philippot L (2006) Abundance of narG, nirS, nirK and nosZ genes of denitrifying bacteria during primary successions of a glacier foreland. Appl Environ Microbiol 72:5957–5962
Kaneko T, Nakamura Y, Sato S, Minamisawa K, Uchiumi T, Sasamoto S, Watanabe A, Idesawa K, Iriguchi M, Kawashima K, Kohara M, Matsumoto M, Shimpo S, Tsuruoka H, Wada T, Yamada M, Tabata S (2002) Complete genomic sequence of nitrogen-fixing symbiotic bacterium Bradyrhizobium japonicum USDA 110. DNA Res 9:189–197
Kim O, Junier P, Imhoff J, Witcel K (2008) Comparative analysis of ammonia monooxygenase (amoA) genes in the water column and sediment–water interface of two lakes and the Baltic Sea. FEMS Microbiol Ecol 66:367–378
Knowles R (1982) Denitrification. Microbiol Mol Biol Rev 46:43–70
Krause A, Ramakumar A, Bartels D, Battistoni F, Bekel T, Boch J, Bohm M, Friedrich F, Hurek T, Krause L, Linke B, McHardy AC, Sarkar A, Schneiker S, Syed AA, Thauer R, Vorhölter FJ, Weidner S, Pühler A, Reinhold-Hurek B, Kaiser O, Goesmann A (2006) Complete genome of the mutualistic, N2-fixing grass endophyte Azoarcus sp. strain BH72. Nat Biotechnol 24:1384–1390
Kuhn M (2001) The nutrient cycle through snow and ice, a review. Aquat Sci-Res Across Boundaries 63:150–167
Kusian B, Bowien B (1997) Organization and regulation of cbb CO2 assimilation genes in autotrophic bacteria. FEMS Microbiol Rev 21:135–155
Lafrenière M, Sharp M (2004) The concentration and fluorescence of dissolved organic carbon (DOC) in glacial and nonglacial catchments: interpreting hydrological flow routing and DOC sources. Arct Antarct Alp Res 36:156–165
Langford H, Hodson A, Banwart S (2011) Using FTIR spectroscopy to characterise the soil mineralogy and geochemistry of cryoconite from Aldegondabreen glacier, Svalbard. Appl Geochem 26:S206–S209
MacDonell S, Fitzsimons S (2008) The formation and hydrological significance of cryoconite holes. Prog Phys Geogr 32:595–610
Matheson FE, Nguyen ML, Cooper AB, Burt TP, Bull DC (2002) Fate of 15N-nitrate in unplanted, planted and harvested riparian wetland soil microcosms. Ecol Eng 19:249–264
Matoba S, Narita H, Motoyama H, Kamiyama K, Watanabe O (2002) Ice core chemistry of Vestfonna Ice Cap in Svalbard, Norway. J Geophys Res. doi:10.1029/2002JD002205
McTavish H, Fuchs JA, Hooper AB (1993) Sequence of the gene coding for ammonia monooxygenase in Nitrosomonas europaea. J Bacteriol 175:2436–2444
Mikucki JA, Pearson A, Johnston DT, Turchyn AV, Farquhar J, Schrag DP, Anbar AD, Priscu JC, Lee PA (2009) A contemporary microbially maintained subglacial ferrous “ocean”. Sci 324:397–400
Mintie AT, Heichen RS, Cromack K Jr, Myrold DD, Bottomley PJ (2003) Ammonia-oxidizing bacteria along meadow-to-forest transects in the Oregon Cascade Mountains. Appl Environ Microbiol 69:3129–3136
Mueller DR, Pollard WH (2004) Gradient analysis of cryoconite ecosystems from two polar glaciers. Polar Biol 27:66–74
Mueller DR, Vincent WF, Pollard WH, Fritsen CH (2001) Glacial cryoconite ecosystems: a bipolar comparison of algal communities and habitats. Nova Hedwig Beih 123:173–197
Mueller DR, Vincent WF, Bonilla S, Laurion I (2005) Extremotrophs, extremophiles and broadband pigmentation strategies in a High Arctic ice shelf ecosystem. FEMS Microbiol Ecol 53:73–87
Mulder A, van de Graaf AA, Robertson LA, Kuenen JG (1995) Anaerobic ammonium oxidation discovered in a denitrifying fluidized bed reactor. FEMS Microbiol Ecol 16:177–184
Negrisolo E, Maistro S, Incarbone M, Moro I, la Valle L, Broady PA, Andreoli C (2004) Morphological convergence characterizes the evolution of Xanthophyceae (Heterokontophyta): evidence from nuclear SSU rDNA and plastidial rbcL genes. Mol Phylogenet Evol 33:156–170
Nogales B, Timmis KN, Nedwell DB, Osborn AM (2002) Detection and diversity of expressed denitrification genes in estuarine sediments after reverse transcription-PCR amplification from mRNA. Appl Environ Microbiol 68:5017–5025
Nordin A, Schmidt IK, Shaver GR (2004) Nitrogen uptake by Arctic soil microbes and plants in relation to soil nitrogen supply. Ecology 85:955–962
Olson JB, Steppe TF, Litaker RW, Paerl JW (1998) N2-fixing microbial consortia associated with the ice cover of Lake Bonney, Antarctica. Microb Ecol 36:231–238
Park SW, Hwang EH, Jang HS, Lee JH, Kang BS, Oh JI, Kim YM (2009) Presence of duplicate genes encoding a phylogenetically new subgroup of form I Ribulose 1,5-bisphosphate carboxylase/oxygenase in Mycobacterium sp. strain JC1 DSM 3803. Res Microbiol 160:159–165
Paul JH, Albin A, Boris W (2000) Micro- and macrodiversity in rbcL sequences in ambient phytoplankton populations from the southeastern Gulf of Mexico. Marine Ecol Prog Ser 198:9–18
Penton CR, Devol AH, Tiedje JM (2006) Molecular evidence for the broad distribution of anaerobic ammonium-oxidizing bacteria in freshwater and marine sediments. Appl Environ Microbiol 72:6829–6832
Philippot L, Piutti S, Martin-Laurent F, Hallet S, Germon JC (2002) Molecular analysis of the nitrate-reducing community from unplanted and maize-planted soils. Appl Environ Microbiol 68:6121–6128
Pohlmann A, Fricke WF, Reinecke F, Kusian B, Liesegang H, Cramm R, Eitinger T, Ewering C, Pötter M, Schwartz E, Strittmatter A, Voß I, Gottschalk G, Steinbüchel A, Friedrich B, Bowien B (2007) Genome sequence of the bioplastic-producing “Knallgas” bacterium Ralstonia eutropha H16. Nat Biotechnol 25:478
Porazinska DL, Fountain AG, Nylen TH, Tranter M, Virginia RA, Wall DH (2004) The biodiversity and biogeochemistry of cryoconite holes from McMurdo Dry Valley glaciers, Antarctica. Arct Antarct Alp Res 36:84–91
Priscu JC, Christner BC (2004) Earths icy bioshpere. In: Bull AT (ed) Microbial diversity and bioprospecting. ASM Press, Washington
Řehák J, Řehák S, Stibal M, Řeháková K, Šabacká M, Kostka S (2007) Glacier caves and drainage systems of the northern part of Hornsund area, southwest Spitsbergen, Svalbard. In: 8th GLACKIPR symposium, Sosnowiec, Poland, p 111
Rotthauwe JH, Witzel KP, Liesack W (1997) The ammonia monooxygenase structural gene amoA as a functional marker: molecular fine-scale analysis of natural ammonia-oxidizing populations. Appl Env Microbiol 63:4704–4712
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Santoro A, Francis C, Sieyes N, Boehm A (2008) Shifts in the relative abundance of ammonia-oxidizing bacteria and archaea across physicochemical gradients in a subterranean estuary. Env Microbiol 10:1068–1079
Säwström C, Mumford P, Marshall W, Hodson A, Laybourn-Parry J (2002) The microbial communities and primary productivity of cryoconite holes in an Arctic glacier (Svalbard 79 degrees N). Polar Biol 25:591–596
Scala DJ, Kerkhof LJ (1998) Nitrous oxide reductase (nosZ) gene-specific PCR primers for detection of denitrifiers and three nosZ genes from marine sediments. FEMS Microbiol Lett 162:61–68
Schalk J, de Vries S, Kuenen JG, Jetten MSM (2000) Involvement of a novel hydroxylamine oxidoreductase in anaerobic ammonium oxidation. Biochem 39:5405–5412
Schneiker S, Perlova O, Kaiser O, Gerth K, Alici A, Altmeyer MO, Bartels D, Bekel T, Beyer S, Bode E, Bode HB, Bolten CJ, Choudhuri JV, Doss S, Elnakady YA, Frank B, Gaigalat L, Goesmann A, Groeger C, Gross F, Jelsbak L, Jelsbak L, Kalinowski J, Kegler C, Knauber T, Konietzny S, Kopp M, Krause L, Krug D, Linke B, Mahmud T, Martinez-Arias R, McHardy AC, Merai M, Meyer F, Mormann S, Muñoz-Dorado J, Perez J, Pradella S, Rachid S, Raddatz G, Rosenau F, Rückert C, Sasse F, Scharfe M, Schuster SC, Suen G, Treuner-Lange A, Velicer GJ, Vorhölter FJ, Weissman KJ, Welch RD, Wenzel SC, Whitworth DE, Wilhelm S, Wittmann C, Blöcker H, Pühler A, Müller R (2007) Complete genome sequence of the Myxobacterium Sorangium cellulosum. Nat Biotechnol 25:1281–1289
Schütte UME, Abdo Z, Bent SJ, Williams CJ, Schneider GM, Solheim B, Forney LJ (2009) Bacterial succession in a glacier foreland of the High Arctic. ISME J 3:1258–1268
Selesi D, Schmid M, Hartmann A (2005) Diversity of green-like and red-like Ribulose-1,5-bisphosphate carboxylase/oxygenase large-subunit genes (cbbL) in differently managed agricultural soils. Appl Env Microbiol 71:175–184
Siddiqui RA, Warnecke-Eberz U, Hengsberger A, Schneider B, Kostka S, Friedrich B (1993) Structure and function of a periplasmic nitrate reductase in Alcaligenes eutrophus H16. J Bacteriol 175:5867–5876
Smith CJ, Nedwell DB, Dong LF, Osborn AM (2007) Diversity and abundance of nitrate reductase genes (narG and napA), nitrite reductase genes (nirS and nrfA), and their transcripts in estuarine sediments. Appl Environ Microbiol 73:3612–3622
Song B, Ward B (2006) Nitrite reductase genes in halobenzoate degrading denitrifying bacteria. FEMS Microbiol Ecol 43:349–357
Spiridonova EM, Berg IA, Kolganova TV, Ivanovsky RN, Kuznetsov BB, Tourova TP (2004) An oligonucleotide primer system for amplification of the Ribulose-1,5-bisphosphate carboxylase/oxygenase genes of bacteria of various taxonomic groups. Microbiol 73:316–325
Stibal M, Tranter M (2007) Laboratory investigation of inorganic carbon uptake by cryoconite debris from Werenskioldbreen, Svalbard. J Geophys Res. doi:10.1029/2007JG000429
Stibal M, Tranter M, Benning LG, Rehak J (2008) Microbial primary production on an Arctic glacier is insignificant in comparison with allochthonous organic carbon input. Env Microbiol 10:2172–2178
Strous M, Pelletier E, Mangenot S, Rattei T, Lehner A, Taylor MW, Horn M, Daims H, Bartol-Mavel D, Wincker P, Barbe V, Fonknechten N, Vallenet D, Segurens B, Schenowitz-Truong C, Médigue C, Collingro A, Snel B, Dutilh BE, Op den Camp HJ, van der Drift C, Cirpus I, van de Pas-Schoonen KT, Harhangi HR, van Niftrik L, Schmid M, Keltjens J, van de Vossenberg J, Kartal B, Meier H, Frishman D, Huynen MA, Mewes HW, Weissenbach J, Jetten MS, Wagner M, Le Paslier D (2006) Deciphering the evolution and metabolism of an anammox bacterium from a community genome. Nat 440:790–794
Tabita FR (1988) Molecular and cellular regulation of autotrophic carbon dioxide fixation in microorganisms. Microbiol Rev 52:155–189
Takeuchi N, Kohshima S, Goto-Azuma K, Koerner RM (2001a) Biological characteristics of dark colored material (cryoconite) on Canadian Arctic glaciers (Devon and Penny ice caps) Mem. Natl Inst Polar Res 54:495–505
Takeuchi N, Kohshima S, Seko K (2001b) Structure, formation, and darkening process of albedo-reducing material (cryoconite) on a Himalayan glacier: a granular algal mat growing on the glacier. Arct Antarct Alp Res 33:115–122
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Telling J, Anesio AM, Hawkings J, Tranter M, Wadham JL, Hodson AJ, Irvine-Fynn T, Yallop ML (2010) Measuring rates of gross photosynthesis and net community production in cryoconite holes: a comparison of field methods. Ann Glaciol 51:153–162
Telling J, Anesio AM, Tranter M, Irvine-Fynn T, Hodson AJ, Butler C, Wadham JL (2011) Nitrogen fixation on Arctic glaciers, Svalbard. J Geophys Res. doi:10.1029/2010JG001632
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Tomitani A, Knoll AH, Cavanaugh CM, Ohno T (2006) The evolutionary diversification of Cyanobacteria: molecular-phylogenetic and paleontological perspectives. Proc Natl Acad Sci USA 103:5442–5447
Tranter M, Brown G, Raiswell R, Sharp M, Gurnell A (1993) A conceptual model of solute acquisition by Alpine glacial meltwaters. J Glaciol 39:573–581
Tranter M, Brown G, Hodson A, Gurnell A, Sharp M (1994) Variations in the nitrate concentration of glacial runoff in Alpine and sub-Polar environments. Snow and ice covers: interactions with the atmosphere and ecosystems. In: Proceedings of Yokohama Symposia J2 and J5, July 1993. IAHS Publ. no. 423, vol 223, pp 299–311
Treusch AH, Leininger S, Kletzin A, Schuster SC, Klenk HP, Schleper C (2005) Novel genes for nitrite reductase and Amo-related proteins indicate a role of uncultivated mesophilic Crenarchaeota in nitrogen cycling. Environ Microbiol 7:1985–1995
van de Graaf AA, Mulder A, de Bruijn P, Jetten MS, Robertson LA, Kuenen JG (1995) Anaerobic oxidation of ammonium is a biologically mediated process. Appl Environ Microbiol 61:1246–1251
Vincent WF, Gibson JAE, Pienitz R, Villeneuve V, Broady PA, Hamilton PB, Howard-Williams C (2000) Ice shelf microbial ecosystems in the High Arctic and implications for life on snowball earth. Naturwissenschaften 87:137–141
von Wintzingerode F, Göbel UB, Stackebrandt E (1997) Determination of microbial diversity in environmental samples: pitfalls of PCR-based rRNA analysis. FEMS Microbiol Rev 21:213–229
Warnecke-Eberz U, Friedrich B (1993) Three nitrate reductase activities in Alcaligenes eutrophus. Arch Microbiol 159:405–409
Watson GM, Tabita FR (1997) Microbial Ribulose 1,5-bisphosphate carboxylase/oxygenase: a molecule for phylogenetic and enzymological investigation. FEMS Microbiol Lett 146:13–22
Wawrik B, Paul JH, Tabita FR (2002) Real-time PCR quantification of rbcL (ribulose-1,5-bisphosphate carboxylase/oxygenase) mRNA in diatoms and pelagophytes. Appl Environ Microbiol 68:3771–3779
Wharton RA Jr, McKay CP, Simmons GM Jr, Parker BC (1985) Cryoconite holes on glaciers. BioSci 35:499–503
Wynn P, Hodson A, Heaton T (2006) Chemical and isotopic switching within the subglacial environment of a High Arctic glacier. Biogeochem 78:173–193
Wynn PM, Hodson AJ, Heaton THE, Chenery SR (2007) Nitrate production beneath a High Arctic glacier, Svalbard. Chem Geol 244:88–102
Yoon HS, Hackett JD, Bhattacharya D (2002) A single origin of the peridinin- and fucoxanthin-containing plastids in Dinoflagellates through tertiary endosymbiosis. Proc Natl Acad Sci USA 99:11724–11729
Zehr JP, McReynolds LA (1989) Use of degenerate oligonucleotides for amplification of the nifH gene from the marine Cyanobacterium Trichodesmium thiebautii. Appl Environ Microbiol 55:2522–2526
Zehr JP, Jenkins BD, Short SM, Steward GF (2003) Nitrogenase gene diversity and microbial community structure: a cross-system comparison. Environ Microbiol 5:539–554
Zumft WG (1997) Cell biology and molecular basis of denitrification. Microbiol Mol Biol Rev 61:533–616
Acknowledgments
KC was funded by a Ph.D. studentship awarded by the University of Sheffield. This research was supported by a Leverhulme Research Fellowship (RF/4/RFG/2007/0398) awarded to AJH and by the NERC for providing access to the NERC Arctic Research Station. The authors would like to acknowledge the support of Nick Cox, Steve Marshall and Rob Smith at the NERC Arctic Station, Ny Ålesund, Svalbard and Monica Kristensen and Jacob Yde for support during Svalbard and Greenland fieldwork. KC was supported during manuscript preparation by NSF-OPP grant 0739783 (awarded to Karen Junge) and NSF-OPP grant 1023462 (awarded to Karen Junge and Ronald Sletten).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Cameron, K.A., Hodson, A.J. & Osborn, A.M. Carbon and nitrogen biogeochemical cycling potentials of supraglacial cryoconite communities. Polar Biol 35, 1375–1393 (2012). https://doi.org/10.1007/s00300-012-1178-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00300-012-1178-3