Abstract
Key message
Our results confirmed that SlERF.F5 can directly regulate the promoter activity of ACS6 and interact with SlMYC2 to regulate tomato leaf senescence.
The process of plant senescence is complex and highly coordinated, and is regulated by many endogenous and environmental signals. Ethylene and jasmonic acid are well-known senescence inducers, but their molecular mechanisms for inducing leaf senescence have not been fully elucidated. Here, we isolated an ETHYLENE RESPONSE FACTOR F5 (SlERF.F5) from tomato. Silencing of SlERF.F5 causes accelerated senescence induced by age, darkness, ethylene, and jasmonic acid. However, overexpression of SlERF.F5 would not promote senescence. Moreover, SlERF.F5 can regulate the promoter activity of ACS6 in vitro and in vivo. Suppression of SlERF.F5 resulted in increased sensitivity to ethylene and jasmonic acid, decreased accumulation of chlorophyll content, and inhibited the expression of chlorophyll- and light response-related genes. Compared with the wild type, the qRT-PCR analysis showed the expression levels of genes related to the ethylene biosynthesis pathway and the jasmonic acid signaling pathway in SlERF.F5-RNAi lines increased. Yeast two-hybrid experiments showed that SlERF.F5 and SlMYC2 (a transcription factor downstream of the JA receptor) can interact physically, thereby mediating the role of SlERF.F5 in jasmonic acid-induced leaf senescence. Collectively, our research provides new insights into how ethylene and jasmonic acid promote leaf senescence in tomato.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Leaf senescence is a necessary process in the growth and development of plants. During the senescence process, plant leaf cells undergo tremendous changes in the structure, metabolism, and gene expression in a programmed manner (Lim et al. 2007; Zhao et al. 2018). Decomposition of the chloroplast is one of the earliest and most noticeable changes in cell structure. In terms of metabolism, the main differences are the loss of photosynthesis and the hydrolysis of macromolecular substances, such as proteins and nucleic acids (Woo et al. 2013; Lim and Nam 2005). These hydrolyzed molecules are transported to the developing fruits and seeds, which are very important for plant survival and sustainability in annual plants (Woo et al. 2013; Bresson et al. 2018). Leaf senescence is the last step of plant leaf growth and development, and it is affected by growth, hormones, and external environment, such as age, darkness, drought, and pathogen attacks (Lim et al. 2007). Among the many factors that affect leaf senescence, plant hormones are essential, including ethylene, methyl jasmonate (MeJA), abscisic acid (ABA), salicylic acid (SA), and brassinosteroids which can promote senescence. However, cytokinins (CKs), gibberellin acid (GA), and auxin can inhibit senescence (Gan and Amasino 1997; Jibran et al. 2013). But so far, the potential regulatory mechanism of leaf senescence and the role of hormones have not been fully elucidated.
As we all know, ethylene is the most common and very crucial plant hormone. It participates in many growth and development processes, including cell elongation, seed germination, flowering, fruit maturation, organ senescence, and response to stress (Johnson and Ecker 1998). It has a positive regulatory effect on fruit ripening and organ senescence (Abeles et al. 1988). According to reports, many transcription factors related to ethylene play an essential role in plant senescence. For example, in Arabidopsis dark-induced leaf senescence experiments, NO (Nitric Oxide) can regulate EIN2 (ETHYLENE INSENSITIVE 2, a transcription factor for ethylene signaling) to promote senescence (Niu and Guo 2012). EIN3, a key transcription factor for ethylene signaling, is constitutively overexpressed or temporarily activated to accelerate leaf senescence symptoms (Li et al. 2013).
Besides, jasmonic acid (JA) also plays a central role in the senescence of plant leaves induced by darkness. It can influence the expression of various genes to promote senescence (He et al. 2002; Jung et al. 2007). For example, during JA-induced leaf senescence, the level of WRKY57 protein, which can interact with JAZ4/8, is reduced, and the wrky57 mutant produces a typical leaf senescence phenotype in Arabidopsis (Jiang et al. 2014). In JA-induced leaf senescence, Dof2.1 acts as an enhancer, which mainly enhances leaf senescence by promoting SlMYC2 (the helix-loop-helix transcription factor) expression in Arabidopsis (Zhuo et al. 2020). Also, SlMYC2 regulates the root growth and the defense of pathogen infections regulated by JA, and plays a positive regulatory role in JA-induced leaf senescence (Song et al. 2017). According to reports, ethylene and jasmonic acid have been found to coordinate (cooperatively or antagonize) plant growth and development and respond to stress (Li and Guo 2007). In addition, a group of JAZ proteins (JAZ1, JAZ3, and JAZ9) can directly bind to EIN3 and EIL1 involved in ethylene signaling (Zhu et al. 2011). However, the molecular mechanism of their coordination in leaf senescence has been relatively little studied.
The AP2/ERF (APETALA2/Ethylene Response Factor) family is one of the largest plant transcription factors, with approximately 140–280 members in various plants. The functions of many genes in this family have been thoroughly studied, that are mostly related to plant growth and development, biological, and abiotic stresses, and fruit maturation, mainly by controlling the response to various plant hormones (Li et al. 2018; Liu et al. 2016; Nakano et al. 2006). The ethylene response factor (ERF) family belongs to the AP2/ERF superfamily, is characterized by a highly conserved AP2 DNA binding domain consisting of 60–70 amino acid residues (Ohmetakagi and Shinshi 1995; Sakuma et al. 2002). The unique feature of this family is the ability to recognize GCC-box (AGCCGCC) and DRE motif (Ohmetakagi and Shinshi 1990), which confers the ability of ERF transcription factors interacting with other genes to function in many physiological processes. For example, AtERF11 knockout mutants showed increased levels of ACS2/5 expression and ethylene emission (Li et al. 2011b). Silencing of SlERF.A3 (Pit4) inhibited the growth of tomato plants (Ouyang et al. 2016). SlERF52 played a pivotal role in transcriptional regulation in pedicel (Nakano et al. 2014). SlERF6 enhanced the carotenoid and ethylene content and played an important role in fruit ripening (Lee et al. 2012). In recent years, studies have shown that overexpression of SlERF36 can promote flowering and senescence (Upadhyay et al. 2013). However, the studies of AP2/ERF family genes on leaf senescence are still sparse and not deep enough.
Here, a gene from the AP2/ERF family, SlERF.F5, was isolated from tomato (Solanum lycopersicum, Mill. cv. Ailsa Craig, AC++). In this article, a study of SlERF.F5 gene silencing was conducted to clarify the function of SlERF.F5 in tomatoes. Experimental results showed that under normal, dark, and hormone treatment conditions, the SlERF.F5-RNAi lines showed early leaf senescence. The morphological, biochemical, and molecular comparisons of WT and SlERF.F5-RNAi lines further confirmed that silencing of SlERF.F5 can promote senescence. In addition, yeast two-hybrid experiments verified the interaction of SlERF.F5 and SlMYC2, and the dual-luciferase reporter system and yeast one-hybrid verified that ACS6 acts downstream of SlERF.F5. In conclusion, this study provides a basis for studying the role of ERF family genes in leaf senescence, as well as theoretical guidance.
Materials and methods
Plant materials and growth conditions
The WT tomato (Solanum lycopersicon Mill. cv. Ailsa Craig, AC++) and SlERF.F5-RNAi, SlERF.F5-overexpressing transgenic lines were used in this study. Tomato growth conditions: 16 h day (28 °C)/8 h night (18 °C) cycle, greenhouse. To detect the response of SlERF.F5 to plant hormones, 35-day-old WT tomato seedlings were treated with 50 µM abscisic acid (ABA), 50 µM indole 3-acetic acid (IAA), 50 µM jasmonic acid (MeJA), 100 µM gibberellin (GA3), 50 µM 1-aminocyclopropane-1-carboxylate (ACC) and distilled water, respectively. Three biological replicates were performed for each hormone. After 0, 1, 2, 4, 8, 12, and 24 h of treatment, the third leaf was collected from the top of the wild tomato seedlings. To examine the specific expression of SlERF.F5 in tomato, various tissue samples of WT tomato were collected. These tissues include: roots (RT), stems (ST), young leaves (YL), mature leaves (ML), senescent leaves (SL), flowers (FL), immature green (IMG), mature green (MG), break (B), 4 days after break (B + 4), and 7 days after break (B + 7). All these samples were immediately wrapped in foil, frozen with liquid nitrogen and stored in a − 80 °C refrigerator.
Construction of SlERF.F5-RNAi and SlERF.F5-overexpression vectors and plant transformation
To obtain the SlERF.F5-RNAi transgenic lines, the 485 bp fragment of SlERF.F5 was amplified with the primers SlERF.F5-RNAi-F1/R1 (Supplementary Table S1). The amplified products ligated into the pBIN19 vector to form the SlERF.F5-RNAi vector, which can produce hairpin SlERF.F5-specific DNA fragments. The vector construction process was performed according to the previous report (Zhou et al. 2019). To construct the SlERF.F5-overexpressing vector, used primers SlERF.F5-full-F2/R2 (Supplementary Table S1) to amplify the full-length SlERF.F5 cDNA. The amplified products were digested with XbaI/SacI and linked to the plant binary vector pBI121 placed under the control of the CaMV 35S promoter. The constructed vector was transformed into Solanum lycopersicon Mill. cv. Ailsa Craig (WT) by Agrobacterium tumefaciens LBA4404 strain through the freeze–thaw method. Finally, transgenic lines were selected by kanamycin (50 mg/L), and confirmed by PCR using NPTII-F/R primers (Supplementary Table S1).
Total RNA extraction and quantitative reverse-transcription PCR analysis
Total RNA was extracted from stored samples using Trizol reagent (Invitrogen, Shanghai, China). The RNA extraction method was based on previous research (Xie et al. 2014).
Quantitative reverse-transcription PCR (qRT-PCR) was performed by using a CFX96™ RealTime System (Bio-Rad, USA). PCR reaction system: 5 μL enzyme solution (2 × GoTaq®qRT-PCR Master Mix, Promega), 3.5 μL distilled water, 0.5 μL primer pair (10 mM), and 1 μL cDNA. PCR reaction program: 95 °C for 3 min, then at 95 °C for 15 s, and Tm (the most suitable temperature) for 45 s for 40 cycles. SlCAC with relatively stable expression was selected as the internal reference (Nicot et al. 2005), and the expression level of the gene was analyzed using the 2−ΔΔCT method. All samples were repeated three times. The primers used in this experiment were listed in Supplementary Table S1.
Determination of leaf senescence induced by age, darkness, ethylene, and MeJA
For age-dependent leaf senescence, 10-week-age tomato leaves of WT and SlERF.F5-RNAi lines were sampled, and chlorophyll contents were measured, respectively. For dark-induced senescence experiments, 10-week-old plants were selected. Mature leaves of the same location were detached from WT and SlERF.F5-RNAi lines and placed on the filter paper containing 3 mL of distilled water at the bottom of 150 mm Petri dishes, These were placed in a dark environment at 22 °C.
In the experiment of hormone and darkness-induced leaf senescence, mature leaves of WT and RNAi tomato seedlings 10-week-age were collected and placed on a filter paper in a Petri dish. Then 3 mL of distilled water, 100 µM ACC, 50 µM MeJA, or 10 µM AgNO3 was added into the Petri dish, respectively, covered, and placed in a dark environment at 22 °C.
Measurement of total chlorophyll and carotenoids content
To detect the age, darkness, and hormone treatment of the leaf senescence of SlERF.F5-RNAi and WT lines, the contents of chlorophyll and carotenoid were detected. The fresh leaves were weighed, ground thoroughly with liquid nitrogen, and extracted with 80% acetone. The specific experimental process and calculation method were described by Wellburn et al. (Wellburn 1994). Three biological replicates were performed for each experiment.
Measuring malondialdehyde (MDA) and electrolytic leakage
To detect the malondialdehyde (MDA) content, the fresh leaves were thoroughly ground with liquid nitrogen, 0.2 g was weighed into a centrifuge tube, and 4 mL of 10% trichloroacetic acid (TCA) was quickly added, mixed, and centrifuged at 15,000g for 5 min. 1 mL of the supernatant was pipetted into a new centrifuge tube, and then 4 mL of 10% trichloroacetic acid solution containing 0.5% thiobarbituric acid (TBA) was added. After mixing and incubation at 95 °C for half an hour, it was placed in an ice-water mixture to stop the reaction. After 10,000 rpm and 10 min, the absorbance of the supernatant at 532 and 600 nm was measured. Repeat three times for each sample. This method was described by Sanjaya et al. (Sanjaya et al. 2008) and Zhang et al. (Zhang et al. 2009).
To detect the detection of electrical conductivity, tomato leaves of the same size were taken, and after sampling the leaves with a hole punch (avoiding the main vein), 20 round leaves were placed in a tube containing 20 mL ddH2O and soaked at 28 °C for 12 h. The conductivity (R1) was measured. Then the tube was placed in boiling water for 30 min and cooled to 28 °C, and the conductivity (R2) was determined. Relative conductivity = R 1 / R 2*100%.
Superoxide dismutase (SOD)
For the determination of superoxide dismutase (SOD) activity, the WST method was used, and the operation steps refer to the WST method kit instructions.
Yeast two hybrid
The open reading frame of SlERF.F5 was amplified by PCR using primers SlERF.F5-F and SlERF.F5-R (Supplementary Table S1). The PCR product was digested with SmaI and BamHI, and cloned into the pGBKT7 bait vector to obtain the vector SlERF.F5-pGBKT7. At the same time, using the primer pairs SlMYC2-F and SlMYC2-R, the open reading frames of SlMYC2 were amplified by PCR (Supplementary Table S1) and digested with SmaI and BamHI, cloned into the pGADT7 vector to obtain the vectors SlMYC2-pGADT7. Then the constructed vectors were transferred into Y2Hgold, respectively. Yeast two hybrid with bait was plated on SD medium lacking Trp and Leu, and SD medium lacking Trp, His, Ade, and Leu to test the self-activation of SlERF.F5-pGBKT7 and SlMYC2-pGADT7. SlERF.F5-pGBKT7 and SlMYC2-pGADT7 were co-transformed into Y2Hgold. After plating on SD medium lacking Trp and Leu, it was cultured upside down for 3 days. Single colonies on SD medium lacking Trp, His, Ade, and Leu were picked and cultured upside down for 1–2 days. X-a-Gal (QDO/X) was used to judge whether SlERF.F5 can interact with SlMYC2.
Transient expression assay in tobacco leaves
The coding sequence of SlERF.F5 was amplified by PCR using specific primers (Supplementary Table S1) and ligated to the pGreen II 62-SK vector driven by the cauliflower mosaic virus (CaMV) 35S promoter. The promoter fragment of ACS6 was amplified and cloned into pGreen II 0800-LUC. Firefly luciferase and Renilla luciferase were measured using a dual-luciferase assay kit (Promega, USA) according to the manufacturer’s instructions. Three replicate experiments were performed.
Yeast one-hybrid assay
Yeast one-hybrid (Y1H) assays were performed using a Matchmaker Gold Yeast One Hybrid System (TaKaRa). The open reading frame sequence of SlERF.F5 was amplified and transferred into pGADT7 vector to construct the prey vector. The promoter fragment of ACS6 were inserted into pAbAi to construct a bait vector. According to the manufacturer’s instructions, the pAbAi-proACS6 plasmid was linearized and then transformed into the Y1HGold yeast strain. The inhibitory concentration of aureobasidin A (AbA) was screened to avoid self-activation. The prey vector was introduced into the bait yeast strain and screened on the SD/–Leu medium with or without AbA. The pAbAi-p53 and pGADT7-p53 plasmids were used as a positive control. Incubation was done at 30 °C for 2–3 days. All primers are listed in Supplementary Table S1.
Statistical analysis
SPSS 26.0 software was used for statistical analysis. Student’s t test (*P < 0.05, **P < 0.01) was performed to analyze the significant difference. ANOVA statistical analyses were performed using SPSS 26.0. Significant differences (P < 0.05) between treatments, as determined by Tukey’s tests, are indicated with different letters. All measurements were taken from the average of at least three independent biological replicates.
Results
Sequence and phylogenetic tree analyses of SlERF.F5
Based on the Tomato Genome Database (https://solgenomics.net, accession no. Solyc10g009110), sequence analysis showed that SlERF.F5 contained 1466 base pairs (bps) encoding a putative protein of 222 amino acids. SlERF.F5 was named by Liu (Liu et al. 2016). Multi-sequence alignment of proteins was done by DNAMAN. This protein contains a typical AP2 domain, containing three β-sheets and one α-helix (Fig. 1a). Based on previous studies, SlERF.F5 belongs to the class II putative repressor ERFs.
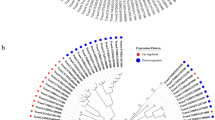
Sequence and expression analysis of SlERF.F5 a Multiple sequence alignment of SlERF.F5 and other ERF proteins.The same amino acids are indicated in black, and its protein sequence has three β-sheets and an α-helix. b Phylogenetic analysis of SlERF.F5 and other ERF proteins was constructed by the neighbor-joining method, bootstrap analysis of 1000 replicates. The accession numbers for the proteins are as follows: AtERF1 (BAA32418.1), AtERF2 (BAA32419.1), Pti4 (NP_001334005.1), LeERF4 (Sl-ERF.B3) (NP_001234313.1), AtERF5 (BAA32422.1), SlERF5 (AS72389.1), SlERF84 (XP_004237817.1), Sl-ERF2 (AAO34704.1), JERF1 (AAK95687.1), SodERF3 (CAM35490.1), AtERF3 (BAA32420.1), AtERF4 (BAA32421.1), SlERF.F5 (NP_001233796.2), NtEREBP5 (AAV54033.1), SlERF52 (BAO18577.1). Stars represent the genes studied in this article
The phylogenetic tree was calculated by MEGA (Molecular Evolutionary Genetics Analysis) version 5.0. Phylogenetic analysis based on full-length sequences of ERF proteins showed that SlERF.F5 was most related to NtEREBP5, followed by AtERF4, AtERF3, and SodERF3 (Fig. 1b). Currently, there is no research on the NtEREBP5 gene. AtERF4 acts as a class II repressor and can be induced by ethylene, jasmonic acid, and abscisic acid (Yang et al. 2005). SodERF3 belongs to class II putative repressor ERF and can bind to GCC-box. Overexpression of SodERF3 improved tolerance to drought and salt in tobacco (Trujillo et al. 2008). In the previous research of SlERF.F5, it was mainly related to stress (Chen et al. 2008), but its role in tomato growth and development has not been reported.
Expression pattern analysis of SlERF.F5
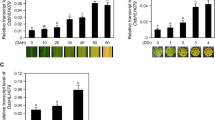
To clarify the potential function of SlERF.F5 in tomato growth and development, the accumulation of its transcripts in various tissues was quantified by qRT-PCR. As shown in Fig. 2a, SlERF.F5 showed the highest transcript accumulation in the B + 4 (4 days after break) fruits, followed by the MG (mature green) fruits and leaves, while relatively low transcript levels were present in the roots and flowers. The specific expression of SlERF.F5 suggested that it may play a role in leaves and fruits. To further study the role of SlERF.F5 in leaf growth and development, the transcription levels of SlERF.F5 in young tomato leaves (Y), mature leaves (M), early senescent leaves (leaf yellowing area > 25%, ES) and late senescent leaves (leaf yellowing area > 50%, LS) were detected (Fig. 2b). The results confirmed that the SlERF.F5 transcripts gradually decreased during leaf development and senescence (Fig. 2c). Besides, photosynthesis genes Cab7 (chlorophyll/binding protein 7), RBCS (ribose bisphosphate carboxylase small chain), and RAV1 (related to ABI3/VP1) were detected, and their expression levels also gradually decreased with leaf senescence (Fig. 2d-f), which was similar to the expression trend of SlERF.F5. In addition, SlSAG12 is an activator for senescence and widely used as a molecular marker for leaf senescenc. In this study, it was specifically expressed in senescent leaves (Fig. 2g), which is consistent with the phenotype. These results indicated that SlERF.F5 may be the repressor of leaf senescence.
Expression pattern of SlERF.F5 a The relative expression patterns of SlERF.F5 in WT. RT root, ST stem, YL young leaf, ML mature leaf, SL senescence leaf, FL flower, IMG immature fruit, MG green fruit, B breaker fruit, B + 4 (4 days after breaker fruit), B + 7 (7 days after breaker fruit). ANOVA statistical analyses were performed using SPSS 26.0. Significant differences (p < 0.05) between treatments, as determined by Tukey’s tests, are indicated with different letters. Data are expressed as the mean ± standard errors for three replicates. b Different development stages of tomato leaves, Y young tomato leaves, M mature leaves, ES (early senescence leaves, yellowing area > 25%) and LS (late senescence leaves, yellowing area > 50%). (c–g) qRT-PCR analysis of the expression levels of SlERF.F5, Cab7, RBCS, Rav1, and SlSAG12 in wild-type tomato leaves at different developmental stages. The experimental results were repeated three times in biology. h qRT-PCR analysis of the expression patterns of SlERF.F5 in response to IAA, ABA, MeJA, GA3, and ACC. The data represent mean from three replicates with three biological repeats. Error bars indicate SE (*P < 0.05, **P < 0.01)
To further investigate the response of SlERF.F5 to hormones, qRT-PCR was performed to examine the expression patterns of SlERF.F5 under different hormone treatments. Fig 2h shows that the accumulation of SlERF.F5 transcripts increased rapidly to the maximum after 1 h of hormone treatment (such as IAA, ABA, MeJA, GA3, and ACC), and with the increase of treatment time, the expression levels of SlERF.F5 gradually decreased, indicating that SlERF.F5 could rapidly respond to IAA, ABA, MeJA, GA3, and ACC hormones.
Silencing of SlERF.F5 accelerates the senescence of tomato leaves
To further clarify the effect of SlERF.F5 on tomato growth and development, the SlERF.F5-RNAi vector was constructed and the transgenic lines were obtained through the Agrobacterium-mediated genetic transformation method. Examination of its silencing efficiency (Fig. 3a) showed that compared with the WT, the expression of SlERF.F5 in the leaves of the five transgenic lines was significantly reduced by 94–98%. Then lines 10, 13, and 16 were selected, and called RNAi10, RNAi13, and RNAi16, respectively, for further research. At the age of 6 weeks of tomato seedlings, the silent lines showed premature senescence (Fig. 3b), and Fig. 3c shows the leaves of the same part of the WT and SlERF.F5-RNAi lines. The leaves of RNAi lines appeared yellow, while the leaves of WT were still greener. To see the color and shape of each leaf more clearly, the leaf was split (Fig. 3d). Compared with WT, the leaves of RNAi plants were yellower, so the chlorophyll content of the leaves was tested. The 6-week-old tomato plants were labeled as the first leaf, the second leaf, and so on, until the sixth leaf. As shown in Fig. 3e, the chlorophyll content of WT leaves was slightly higher than that of RNAi leaves at different leaf ages. Further statistics on the senescence time of tomato leaves revealed that the senescence time of RNAi lines was earlier than that of WT (Fig. 3f). Compared with the WT, leaf senescence time of RNAi lines was advanced by about 1 week. These data demonstrated that silencing of SlERF.F5 can promote the senescence of tomato leaves.
Silencing of SlERF.F5 causes premature senescence of tomato leaves a The expression level of SlERF.F5 in mature leaves of WT and SlERF.F5-RNAi lines. b The senescence phenotype of 10 weeks old WT, RNAi10, RNAi13, and RNAi16 lines. c The senescence phenotype of the fifth leaf of WT, RNAi10, RNAi13, and RNAi16 lines. d Isolation of leaves from 12-week-old WT and RNAi10, RNAi13, RNAi16 lines. e Chlorophyll content of each leaf in WT, RNAi10, RNAi13, and RNAi16 lines. f Leaf senescence time of WT, RNAi10, RNAi13 and RNAi16 lines. The data represent mean from three replicates with three biological repeats. Error bars indicate SE (*P < 0.05, **P < 0.01)
Silencing of SlERF.F5 promotes dark-induced leaf senescence
Dark-induced leaf senescence was a common way to study senescence (Li et al. 2013). To further study the role of SlERF.F5-RNAi in leaf senescence, mature leaves were taken for experiments. As shown in Fig. 4a, the edge of leaves of the SlERF.F5-RNAi lines started to become yellow when treated for 5 days in the dark, and some leaves turned yellow completely after 7 days of treatment, whereas the leaves of the WT were still green at the same time. To further confirm the phenotype of senescence, we measured some physiological indicators related to leaf senescence. First, the chlorophyll contents of the RNAi lines were little higher than that of the WT before treatment. As the dark treatment time increased, the chlorophyll contents of SlERF.F5-RNAi line leaves gradually decreased and were significantly lower than that of WT at 7 d, while the chlorophyll content of WT leaves only decreased slightly during the treatment (Fig. 4b). Second, MDA was an important indicator of membrane damage (Sanjaya et al. 2008). From 0 d to 7 d, MDA contents in the leaves of WT and RNAi lines both increased quickly, but the MDA contents in the leaves of the SlERF.F5-RNAi lines were significantly higher than that of WT at each time point (Fig. 4c). Further, the electrical conductivity was also an indicator of leaf cell membrane damage. At 0 d, the electrical conductivity of the WT leaves was slightly higher than that of the SlERF.F5-RNAi. This may be due to the damage caused by sampling, but at 7 d, the electrical conductivity of RNAi line leaves was higher than that of WT leaves (Fig. 4d). Besides, the activity of superoxide dismutase (SOD) was also an indicator of the degree of damage to the cell membrane. At 0 d, the SOD activity of the RNAi leaves was lower than that of WT, while, at 5 d, the SOD activity of SlERF.F5-RNAi leaves was significantly lower than that of WT (Fig. 4e). These results indicated that silencing of SlERF.F5 gene leads to premature leaf senescence under dark conditions.
Silencing of SlERF.F5 promotes dark-induced leaf senescence and altered the expression of chlorophyll metabolism, ethylene, and jasmonic acid-related genes a The leaves of the WT and SlERF.F5-RNAi lines (RNAi10, RNAi13, RNAi16) were treated in the dark for 0 d, 5 d and 7 d. b Leaf chlorophyll content of WT and RNAi10, RNAi13, RNAi16 lines at 0, 5, and 7 d in the dark. c The MDA (malonaldehyde) content of the leaves of the WT and RNAi10, RNAi13, RNAi6 lines at 0, 5, and 7 d of dark treatment. d Relative conductivity of leaves of WT and RNAi10, RNAi13, and RNAi16 lines in dark treatment for 0 d, 5 d and 7 d. e SOD activity in leaves of WT and RNAi10, RNAi13, and RNAi16 lines at 0 and 5 d after dark treatment. f qRT-PCR analysis of CHLH, CHLM, POR, CAO1, GUN4, PPH, SGR1, AUREA, RBCS, LHCA1 and SlSAG12 expression levels in WT and RNAi10, RNAi13, RNAi16 lines. g qRT-PCR analysis of the expression levels of ethylene biosynthetic pathway genes ACO1, ACS2, ACS4, ACS6, JAZ1, JAZ2, JAZ4, JAZ7, JAZ11, SlMYC2 and COI1 in WT and RNAi10, RNAi13, and RANi16 lines. All data are means (± SE) of three independent biological replicates (*P < 0.05, **P < 0.01)
Silencing of SlERF.F5 affects the expression of chlorophyll, ethylene, and jasmonic acid-related genes
To reveal the possible molecular mechanism of dark-induced leaf senescence in WT and SlERF.F5-RNAi, the transcription levels of chlorophyll-related genes, including magnesium chelatase H subunit (CHLH), Mg protoporphyrin IX methyltransferase (CHLM), protochlorophyllide reductase (POR), and chlorophyllide an oxygenase (CAO1), pheophytin pheophorbide hydrolase (PPH), STAY-GREEN 1 (SGR1), AUREA and the genomes uncoupled 4 (GUN4), were detected in 5 d treatment leaves and they were significantly down-regulated in SlERF.F5-RNAi transgenic lines (Fig. 4f). Besides, the expression level of ribulose bisphosphate carboxylase small chain (RBCS) and light-harvesting protein complex 1 (LHCA1) were also down-regulated in the SlERF.F5-RNAi lines (Fig. 4f). On the contrary, SlSAG12 was significantly up-regulated in the SlERF.F5-RNAi lines compared to WT (Fig. 4f). According to these results, it is speculated that silencing of SlERF.F5 gene affects the expression of some genes in the pathway of chlorophyll, thereby reducing the chlorophyll content, which was one of the reasons for dark-induced leaf senescence.
To investigate ethylene’s role in dark-induced leaf senescence of the WT and SlERF.F5-RNAi lines, some of the ethylene signal synthesis pathway genes were examined. Expression of 1-aminocyclopropane-1-carboxylate oxidase 1 (ACO1), 1-aminocyclopropane-1-carboxylate synthase 2 (ACS2), 1-aminocyclopropane-1-carboxylate synthase 4 (ACS4) and 1-aminocyclopropane-1-carboxylate synthase 6 (ACS6) were up-regulated in the SlERF.F5-RNAi lines compared to WT (Fig. 4g).
Since MeJA plays a positive regulatory role in leaf senescence, in recent years studies have pointed out that SlMYC2 can regulate plant growth and development through physical interaction with EIN3. Thus, the expression levels of MeJA signaling pathway transcription factors were detected in this study. JASMONATE ZIM-domain (JAZ) genes (in. JAZ1, JAZ2, JAZ4, JAZ7, JAZ11) were up-regulated in the SlERF.F5-RNAi lines (Fig. 4g). Besides, SlMYC2 is a transcription factor downstream of the JA receptor, and its expression in the leaves of RNAi lines was higher than that of WT (Fig. 4g). COI1 was an essential regulator of JA-induced leaf senescence, and its expression in the leaves of SlERF.F5-RNAi lines was also higher. These results indicated that silencing of SlERF.F5 gene could increase the gene expression of ethylene biosynthesis, jasmonic acid signal transduction, and receptor downstream transcription factors, which may increase the content of ethylene and jasmonic acid and cause leaf senescence.
Silencing of SlERF.F5 affects the sensitivity of tomato seedlings to ethylene and jasmonic acid
To further clarify the role of ethylene and jasmonic acid in promoting leaf senescence, the triple reaction induced by ACC and a sensitivity test of MeJA were carried out. 0, 5, and 10 μM ACC (1-aminocyclopropane-1-carboxylic acid) were used to treat the germinated seeds of WT and SlERF.F5-RNAi lines. After 5 d of cultivation in the dark, SlERF.F5-RNAi lines showed a slightly lower length of hypocotyls and root and a lighter weight of seedling than that of the WT (Fig. 5a-c), suggesting that silenced-SlERF.F5 seedling was more sensitive to ACC. Besides, compared with WT, the seedling root length of the RNAi lines was shorter at 0 μM (Fig. 5b), and the seedling weight was heavier (Fig. 5c), indicating that under normal circumstances, the seedling growth of the RNAi lines was better than that of WT.
Silencing of SlERF.F5 shows increased sensitivity to ethylene and jasmonic acid a The seeds after 7 days of germination were treated with 0 μM, 5 μM, and 10 μM ACC WT, and RNAi10, RNAi13, RNAi16 lines. (b, c) The root length, hypocotyl, and fresh weight of (a) treated tomato seedlings were measured. d The seeds after 7 days of germination were treated with 0, 10, 20, and 50 μM MeJA WT, and RNAi10, RNAi13, and RNAi16 lines. (e, f) The root length, hypocotyl and fresh weight of (a) treated tomato seedlings were measured. All data are means (± SE) of three independent biological replicates (*P < 0.05, **P < 0.01)
In the MeJA sensitivity experiment, after 7 d of treatment, the length of the hypocotyl, root, and seedling weight of the RNAi lines were lower than those of the WT (Fig. 5d-f). These results indicated that SlERF.F5 silence lines were more sensitive to MeJA.
Ethylene and jasmonic acid accelerate the senescence of SlERF.F5-RNAi leaves in dark conditions
To further verify the role of ethylene and jasmonic acid in leaf senescence, a hormone-induced senescence experiment was carried out. Hormones (ACC and MeJA) were added to the dark-induced leaf senescence experiment. AgNO3 was an inhibitor of ethylene action in plants. The purpose of treatment with AgNO3 and MeJA was to evaluate the effect of jasmonic acid on leaf senescence in the absence of ethylene. The results showed that after 7 d of hormone treatment in dark conditions, the leaves of the SlERF.F5-RNAi lines were yellower than WT (Fig. 6a). Compared with WT, the chlorophyll content of SlERF.F5-RNAi was significantly lower (Fig. 6b), and the MDA content was higher (Fig. 6c). The results demonstrated that both ethylene and jasmonic acid treatments can promote leaf senescence of SlERF.F5-RNAi lines under dark conditions. Also, jasmonic acid alone can also promote leaf senescence of SlERF.F5-RNAi lines, indicating that ethylene and jasmonic acid synergistically promote leaf senescence.
Silencing of SlERF.F5 promotes dark, ethylene and jasmonic acid-induced leaf senescence a Senescence phenotypes of leaves of WT, RNAi10, RNAi13, and RNAi16 lines treated with ACC, MeJA, and AgNO3 + MeJA in the dark. Under dark conditions, the leaves were treated with water (control), 100 μM ACC, or 50 μM MeJA for 7 days. For AgNO3 + MeJA treatment, the leaves were pretreated with 10 μM AgNO3 for 1 h, washed with water, and then treated with 50 μM MeJA in the dark for 7 d. b The chlorophyll content was measured from hormone-treated leaves. c The MDA content was measured from hormone-treated leaves. All data are means (± SE) of three independent biological replicates (*P < 0.05, **P < 0.01)
Overexpression of SlERF.F5 may delay dark-induced leaf senescence
To further verify the function of SlERF.F5 on the leaf senescence, an overexpression vector of SlERF.F5 was constructed and transformed into tomato cotyledons to obtain transgenic lines overexpressing SlERF.F5. qRT-PCR was used to detect the expression level of SlERF.F5. As shown in Fig. 7a, we selected lines 1, 5, and 6 (OE1, OE5, and OE6) with higher expression efficiency for the next experiment. Similarly, a dark-induced leaf senescence experiment was conducted. The leaves of 10-week-old seedlings of WT, SlERF.F5-RNAi, and SlERF.F5-OE lines were harvested, respectively, and the dark-induced senescence experiment was carried out in the same way as above. After 5 d, the leaves of SlERF.F5-RNAi plants began to turn yellow, while no noticeable color change occurred in the leaves of WT and SlERF.F5-OE lines (Fig. 7b). Compared with WT, the chlorophyll and carotenoid content of SlERF.F5-OE lines were slightly higher (Fig. 7c, d). The chlorophyll content of RNAi and overexpressing lines and WT for 5 d after dark treatment were sorted together for comparison (Fig. 7e). The results showed that the chlorophyll content of the leaves of the SlERF.F5-OE lines was slightly higher than that of the WT after dark treatment for 5 d, but at 5 d the greenness of WT leaves remained more than that of SlERF.F5-RNAi lines, and the total chlorophyll content was higher. Also, after 5 d the SOD activity of RNAi leaves was lower than that of WT, whereas the SlERF.F5-OE line leaves showed higher SOD activity than that of WT (Fig. 7f). The above results suggested that suppression of SlERF.F5 can promote leaf senescence, and overexpression of SlERF.F5 might inhibit leaf senescence.
Overexpression of SlERF.F5 does not promote leaf senescence a The expression level of SlERF.F5 in mature leaves of WT and SlERF.F5-OE lines. b Dark-induced leaf senescence for 5 d in WT and RNAi10, RNAi13, RNAi16 lines and OE1, OE5, OE6 lines. c Analysis of chlorophyll content in WT and OE1, OE5, OE6 lines. d Analysis of carotenoid content in WT and OE1, OE5, OE6 lines. e Analysis of chlorophyll content in WT and RNAi10, RNAi13, RNAi16 lines and OE1, OE5, OE6 lines. f Analysis of SOD activity in WT and RNAi10, RNAi13, RNAi16 lines and OE1, OE5, OE6 lines. All data are means (± SE) of three independent biological replicates (*P < 0.05, **P < 0.01)
SlERF.F5 directly inhibits the transcription of ACS6 in tobacco and interacts with SlMYC2
Based on the above research results, it was found that SlERF.F5-RNAi lines promoted leaf senescence under normal growth conditions and under treatment with darkness, ethylene, and jasmonic acid. Given that SlMYC2 was a vital transcription factor downstream of the jasmonic acid receptor and it has a direct relationship with aging, its interaction with SlERF.F5 was the first choice for studying the regulation of aging by ethylene and jasmonic acid. Therefore, the SlMYC2 protein was selected for the yeast two-hybrid experiment. Results showed that the yeast cells co-expressing SlERF.F5-BD and SlMYC2-AD could grow on the quadruple dropout medium (SD/-Leu-Trp-His-Ade), the same as yeast cells carrying pGADT7-T and pGBKT7-53 (positive control) (Fig. 8a), indicating that SlERF.F5 physically interacts with SlMYC2, thereby participating in leaf senescence induced by ethylene and jasmonic acid.
Yeast two-hybrid assay of SlERF.F5 and SlMYC2 protein and tobacco transient expression system assay of SlERF.F5 and ACS6 a Yeast two-hybrid experiment indicated that SlERF.F5 interacted with SlMYC2. Co-transformation of pGADT7-T and pGBKT7-53 as a positive control; co-transformation of pGADT7-T and pGBKT7-Lam as a negative control; single transformation of BD-SlERF.F5 and co-transformation with AD to verify self-activation; co-transformation of BD-SlERF.F5 and AD-SlMYC2 as an experimental group. b The double-reporter plasmid contained with the ACS6 promoter fused to LUC and REN was used as the reporter; the SlERF.F5 driven by CaMV 35S was used as the effector; the empty vector was used as the control. The pACS6 reporter and control constructs for a transactivation assay in tobacco leaves co-transfected. The pACS6 reporter and the p35S::SlERF.F5 effector constructs for a transactivation assay in tobacco leaves co-transfected. LUC, Firefly luciferase; REN, Renilla luciferase; Nos, NOS terminator. c The results of the transactivation activity of tomato SlERF.F5 protein in the transient expression system in N. benthamiana leaves, using a double reporter plasmid containing the ACS6 promoter, the promoter fused to LUC and REN driven by CaMV35S. Control experiments were performed with empty vectors as effector constructs. All data are means (± SE) of three independent biological replicates (*P < 0.05, **P < 0.01). d The interactions between SlERF.F5 and the promoters of ACS6 wer confirmed by the yeast one-hybrid assay. The vector of pAbAi-p53 plus AD-empty acts as the negative control, and pAbAi-p53 plus AD-p53 acts as the positive control
1-Aminocyclopropane-1-carboxylic acid (ACC) synthase (ACS) was the rate-limiting enzyme in the biosynthetic pathway of ethylene, which controls ethylene biosynthesis (Li et al. 2011a). The upstream promoter of ACS6 (− 612–428 bp) contains the DRE motif sequence (Fig. 8b). In this study, the expression level of the ACS6 gene in the leaves of RNAi lines was higher than that of WT. To study whether SlERF.F5 can regulate the activity of the ACS6 promoter, a transient transactivation assay was performed in tobacco (N. benthamiana) leaves. The double-reporter plasmid contained the promoter of ACS6 fused to LUC luciferase and REN luciferase. The effector was SlERF.F5 driven by CaMV 35S; the control construct lacked SlERF.F5 (Fig. 8b). As shown in Fig. 8c, compared with the control, the LUC/REN ratio decreased to approximately 82% in the presence of SlERF.F5. Further, yeast one-hybrid experiments were performed and the results confirmed that SlERF.F5 could bind to the ACS6 promoter (Fig. 8d). These results indicated that the activity of the ACS6 promoter was negatively regulated by SlERF.F5 in vivo and in vitro. Silencing of SlERF.F5 might weaken the negative regulation of SlERF.F5 to ACS6 promoter, subsequently increasing the expression level of ACS6 in RNAi lines, which may increase the ethylene content and promote the early senescence of tomato plants.
Discussion
The functions of AP2/ERF family genes are mainly related to plant growth, biotic and abiotic stress, and fruit ripening. For example, overexpression of SlERF.B3-SRDX leads to a significant delay in fruit ripening time, increased fruit softening, and reduced pigment accumulation (Liu et al. 2014). Overexpression of SlERF5, as a class III ERFs protein, can increase tolerance to drought and salt (Pan et al. 2012). Under iron deficiency conditions, AtERF72 can affect the expression of chlorophyll-degrading genes pheophorbide a oxygenase (PAO) and chlorophyllase (CLH1), and ERF72 can directly bind to the promoter regions of IRT1, HA2, and CLH1, thereby regulating the plant’s lack of response to iron stress (Liu et al. 2017). TERF2/LeERF2 plays a positive regulatory role in ethylene biosynthesis, and it can enhance the freezing resistance of plants (Zhang and Huang 2010). In this study, we found that SlERF.F5 plays a role in tomato leaves. During the growth and development of leaves, the expression levels of SlERF.F5 gene gradually decreased (Fig. 2c). The expression pattern of SlERF.F5 gene in leaf development is similar to that of chloroplast-related genes Cab7, RBCS, and RAV1, but is opposite to that of SlSAG12 (Fig. 2). This experimental results indicated that SlERF.F5 plays a negative role in leaf senescence.
Under conditions induced by age and darkness (Fig. 3, 4), we found that silencing of SlERF.F5 promotes the senescence of tomato leaves (Fig. 3, 4). The chlorophyll content reflects the senescence of the leaves. Compared with WT plants, we found that the chlorophyll content of the leaves of the SlERF.F5-RNAi lines was significantly reduced (Fig. 4b). By detecting chlorophyll-related genes in WT and RNAi lines, it was found that the expression levels of CHLH, CHLM, POR, CAO1, GUN4, PPH, SGR1, RBCS PPH, SGR1, RBCS, ACREA and LHCA1 were reduced in RNAi lines (Fig. 4). In researches related to chlorophyll biosynthesis and degradation genes, compared with WT plants, PPH, PAO, RCCR, and SGR1 in the SlOFP20-OE line was significantly increased. Overexpression of SlOFP20 can regulate chlorophyll accumulation and leaf senescence (Zhou et al. 2019). The expression levels of RBCS1A and CAB1 were examined to investigate the relationship between EIN3 and leaf senescence (Li et al. 2013). During leaf senescence, the expression of SlNAP2 increased. SlNAP2 can activate the expression of SlSGR1 and SlPAO to regulate senescence (Ma et al. 2018). According to previous research, the expression changes of chlorophyll synthesis and metabolism genes were mostly related to leaf senescence. In this article, these genes were down-regulated in senescent leaves of the SlERF.F5-RNAi lines, which clarified the mechanism of leaf senescence from a physiological and molecular level. Besides, we also created the SlERF.F5-OE tomato lines. In the dark-induced senescence experiment, compared with WT plants, the chlorophyll content, SOD activity, and carotenoid content of the SlERF.F5-overexpression lines were slightly higher, and no apparent yellowing phenomenon was observed (Fig. 7). This indicated that overexpression of SlERF.F5 would not promote senescence. On the contrary, leaf senescence may be delayed.
Among the main hormones that affect leaf senescence, ethylene, as a promoter of leaf senescence, plays a significant role in age and darkness-induced senescence. During darkness-induced leaf senescence, ethylene-insensitive mutants (ein2/ore3) act as senescence promoting factors through transcriptional regulation of stress-related responses (Kim et al. 2018). In the ein3 eil1 double mutant, ethylene inhibits the expression of NYE1, NYC1, and PAO containing GCC-box, and EMSA results indicate that EIN3 can directly bind NYE1, NYC1, and PAO promoters and play a central role in ethylene-mediated leaf senescence (Qiu et al. 2015). Therefore, we supposed that ethylene might be responsible for the leaf senescence of SlERF.F5-RNAi lines (Fig. 9). In this study, compared with WT, the expression levels of ACO1, ACS2, ACS4, and ACS6 genes in the ethylene biosynthetic pathway were significantly increased in the SlERF.F5-RNAi lines. Through the tobacco transient expression system, it was found that SlERF.F5 can directly inhibit the promoter activity of ACS6 (Fig. 8, 9), indicating that SlERF.F5 may be a negative regulatory gene in the process of ethylene biosynthesis. These results indicate that the silencing of SlERF.F5 may induce ethylene biosynthesis, thereby promoting senescence.
The proposed model illustrates the regulatory role of SlERF.F5 in leaf senescence. Under age condition, silencing of SlERF.F5 reduces the expression level of chloroplast-related genes, changes the content of chlorophyll, and promotes leaf senescence; under the action of ethylene, silencing of SlERF.F5 increases the expression level of ethylene biosynthesis-related genes, and directly regulating the activity of ACS6 gene may increase the content of ethylene and promote leaf senescence; under the action of jasmonic acid, silencing of SlERF.F5 increases the expression level of jasmonic acid-related genes. On the one hand, silencing of SlERF.F5 will induce the expression of SlMYC2. Both the interaction promotes the senescence of leaves. On the other hand, SlMYC2 can directly activate SGR1 and other chlorophyll catabolic enzyme genes. The dotted line represents the results that have not been confirmed in this experiment, and is only speculation based on other articles. The solid line represents the results that have been confirmed in this experiment
Jasmonic acid is also a promoting factor of leaf senescence, and changes in the expression of jasmonic acid-related genes are critically related to leaf senescence. In research on leaf senescence, the expression of JAZ7 was up-regulated in darkness-induced senescence. The jaz7 mutant showed a large area of yellowing of the leaves. In addition, the double mutants of jaz7 SlMYC2 and jaz7 coi1 showed delayed leaf senescence. In conclusion, JAZ7 protein is a positive regulator of dark-induced leaf senescence (Yu et al. 2016). JAZ4 and JAZ8 can physically interact with WRKY57 and play a negative regulatory role in MeJA-induced leaf senescence (Jiang et al. 2014). In MeJA-induced senescence experiments, MYC5-overexpressing transgenic plants showed early leaf senescence phenotypes, including reduced chlorophyll content. It enhanced JA-regulated senescence-related gene expression (SAG13, SEN4, SAG113, and SAG29) and photosynthesis genes (RBCS and CAB1) (Song et al. 2017). In the mechanism of MeJA-induced leaf senescence, COI1-dependent JA inhibition was considered to be very important (Shan et al. 2011). The experimental results showed that the expression levels of JAZ1, JAZ2, JAZ4, JAZ7, JAZ11 in the jasmonic acid signaling pathway in the SlERF.F5-RNAi lines were significantly increased compared with WT (Fig. 4). Besides, the expression of downstream transcription factors of JA receptor (SlMYC2) and COI1 gene was also increased compared with WT (Fig. 4). Given the role of ERF and jasmonic acid-related genes, we choose SlMYC2 and SlERF.F5 for yeast two-hybrid experiments. The results showed that SlERF.F5 could interact with SlMYC2. The above results indicated that silencing of SlERF.F5 gene might relieve its inhibition to SlMYC2, thereby promoting leaf senescence (Fig. 9). On the other hand, SlMYC2 could directly activate SGR1 and other chlorophyll catabolic enzyme genes during the leaf senescence induced by JA (Zhu et al. 2015). Besides, SlERF.F5 can induce the expression of some JAZ genes in the jasmonic acid signal transduction pathway (JAZ1, JAZ2, JAZ4, JAZ7, JAZ11), among which JAZ7 can interact with COI1 and, or SlMYC2 to regulate leaf senescence induced by darkness (Yu et al. 2016). According to previous reports, increased expression of these genes could promote leaf senescence and may also improve the jasmonic acid content, which may also be one of the reasons for the early senescence of the leaves of the SlERF.F5-RNAi lines.
In the hormone-induced senescence experiment, exogenous addition of ACC and MeJA can induce senescence of SlERF.F5-RNAi leaves. Moreover, the exogenous addition of AgNO3 (an ethylene inhibitor) and MeJA can also induce senescence of SlERF.F5-RNAi leaves. This result indicated that MeJA could also induce senescence in the absence of ethylene (Fig. 6). According to the above results, we can speculate that ethylene and jasmonic acid play a synergistic role in the process of leaf senescence. Overall, the physiological and molecular mechanism analysis showed that SlERF.F5 plays a vital role in regulating the leaf senescence induced by age and darkness. Ethylene and jasmonic acid play a synergistic role in regulating leaf senescence.
References
Abeles FB, Dunn LJ, Morgens P, Callahan A, Dinterman RE, Schmidt J (1988) Induction of 33-Kd and 60-Kd peroxidases during ethylene-induced senescence of cucumber cotyledons. Plant Physiol 87(3):609–615
Bresson J, Bieker S, Riester L, Doll J, Zentgraf U (2018) A guideline for leaf senescence analyses: from quantification to physiological and molecular investigations. J Exp Bot 69(4):769–786
Chen G, Hu Z, Grierson D (2008) Differential regulation of tomato ethylene responsive factor LeERF3b, a putative repressor, and the activator Pti4 in ripening mutants and in response to environmental stresses. J Plant Physiol 165(6):662–670
Gan SS, Amasino RM (1997) Making sense of senescence - molecular genetic regulation and manipulation of leaf senescence. Plant Physiol 113(2):313–319
He YH, Fukushige H, Hildebrand DF, Gan SS (2002) Evidence supporting a role of jasmonic acid in Arabidopsis leaf senescence. Plant Physiol 128(3):876–884
Jiang YJ, Liang G, Yang SZ, Yu DQ (2014) Arabidopsis WRKY57 functions as a node of convergence for jasmonic acid- and auxin-mediated signaling in jasmonic acid-induced leaf senescence. Plant Cell 26(1):230–245
Jibran R, Hunter DA, Dijkwel PP (2013) Hormonal regulation of leaf senescence through integration of developmental and stress signals. Plant Mol Biol 82(6):547–561
Johnson PR, Ecker JR (1998) The ethylene gas signal transduction pathway: a molecular perspective. Annu Rev Genet 32:227–254
Jung C, Lyou SH, Yeu S, Kim MA, Rhee S, Kim M, Lee JS, Do Choi Y, Cheong JJ (2007) Microarray-based screening of jasmonate-responsive genes in Arabidopsis thaliana. Plant Cell Rep 26(7):1053–1063
Kim J, Park SJ, Lee IH, Chu H, Penfold CA, Kim JH, Buchanan-Wollaston V, Nam HG, Woo HR, Lim PO (2018) Comparative transcriptome analysis in Arabidopsis ein2/ore3 and ahk3/ore12 mutants during dark-induced leaf senescence. J Exp Bot 69(12):3023–3036
Lee JM, Joung JG, McQuinn R, Chung MY, Fei Z, Tieman D, Klee H, Giovannoni J (2012) Combined transcriptome, genetic diversity and metabolite profiling in tomato fruit reveals that the ethylene response factor SlERF6 plays an important role in ripening and carotenoid accumulation. Plant J 70(2):191–204
Li HJ, Guo HW (2007) Molecular basis of the ethylene signaling and response pathway in Arabidopsis. J Plant Growth Regul 26(2):106–117
Li L, Zhu B, Yang P, Fu D, Zhu Y, Luo Y (2011a) The regulation mode of RIN transcription factor involved in ethylene biosynthesis in tomato fruit. J Sci Food Agric 91(10):1822–1828
Li Z, Zhang L, Yu Y, Quan R, Zhang Z, Zhang H, Huang R (2011b) The ethylene response factor AtERF11 that is transcriptionally modulated by the bZIP transcription factor HY5 is a crucial repressor for ethylene biosynthesis in Arabidopsis. Plant J 68(1):88–99
Li ZH, Peng JY, Wen X, Guo HW (2013) ETHYLENE-INSENSITIVE3 Is a senescence-associated gene that accelerates age-dependent leaf senescence by directly repressing miR164 transcription in Arabidopsis. Plant Cell 25(9):3311–3328
Li ZJ, Tian YS, Xu J, Fu XY, Gao JJ, Wang B, Han HJ, Wang LJ, Peng RH, Yao QH (2018) A tomato ERF transcription factor, SlERF84, confers enhanced tolerance to drought and salt stress but negatively regulates immunity against pseudomonas syringae pv. tomato DC3000. Plant Physiol Bioch 132:683–695
Lim PO, Nam HG (2005) The molecular and genetic control of leaf senescence and longevity in Arabidopsis. Curr Top Dev Biol 67:49–83
Lim PO, Kim HJ, Nam HG (2007) Leaf senescence. Annu Rev Plant Biol 58:115–136
Liu M, Diretto G, Pirrello J, Roustan JP, Li Z, Giuliano G, Regad F, Bouzayen M (2014) The chimeric repressor version of an ethylene response factor (ERF) family member, Sl-ERF.B3, shows contrasting effects on tomato fruit ripening. New Phytol 203(1):206–218
Liu MC, Gomes BL, Mila I, Purgatto E, Peres LEP, Frasse P, Maza E, Zouine M, Roustan JP, Bouzayen M, Pirrello J (2016) Comprehensive profiling of ethylene response factor expression identifies ripening-associated ERF genes and their link to key regulators of fruit ripening in tomato. Plant Physiol 170(3):1732–1744
Liu W, Li QW, Wang Y, Wu T, Yang YF, Zhang XZ, Han ZH, Xu XF (2017) Ethylene response factor AtERF72 negatively regulates Arabidopsis thaliana response to iron deficiency. Biochem Bioph Res Co 491(3):862–868
Ma X, Zhang Y, Tureckova V, Xue GP, Fernie AR, Mueller-Roeber B, Balazadeh S (2018) The NAC transcription factor SlNAP2 regulates leaf senescence and fruit yield in tomato. Plant Physiol 177(3):1286–1302
Nakano T, Suzuki K, Fujimura T, Shinshi H (2006) Genome-wide analysis of the ERF gene family in Arabidopsis and rice. Plant Physiol 140(2):411–432
Nakano T, Fujisawa M, Shima Y, Ito Y (2014) The AP2/ERF transcription factor SlERF52 functions in flower pedicel abscission in tomato. J Exp Bot 65(12):3111–3119
Nicot N, Hausman JF, Hoffmann L, Evers D (2005) Housekeeping gene selection for real-time RT-PCR normalization in potato during biotic and abiotic stress. J Exp Bot 56(421):2907–2914
Niu YH, Guo FQ (2012) Nitric oxide regulates dark-induced leaf senescence through EIN2 in arabidopsis. J Integr Plant Biol 54(8):516–525
Ohmetakagi M, Shinshi H (1990) Structure and expression of a tobacco beta-1,3-glucanase gene. Plant Mol Biol 15(6):941–946
Ohmetakagi M, Shinshi H (1995) Ethylene-inducible DNA-binding proteins that interact with an ethylene-responsive element. Plant Cell 7(2):173–182
Ouyang Z, Liu S, Huang L, Hong Y, Li X, Huang L, Zhang Y, Zhang H, Li D, Song F (2016) Tomato SlERF.A1, SlERF.B4, SlERF.C3 and SlERF.A3, members of B3 group of ERF Family, are required for resistance to botrytis cinerea. Front Plant Sci 7:1964
Pan Y, Seymour GB, Lu C, Hu Z, Chen X, Chen G (2012) An ethylene response factor (ERF5) promoting adaptation to drought and salt tolerance in tomato. Plant Cell Rep 31(2):349–360
Qiu K, Li Z, Yang Z, Chen J, Wu S, Zhu X, Gao S, Gao J, Ren G, Kuai B, Zhou X (2015) EIN3 and ORE1 accelerate degreening during ethylene-mediated leaf senescence by directly activating chlorophyll catabolic genes in Arabidopsis. PLoS Genet 11(7):e1005399
Sakuma Y, Liu Q, Dubouzet JG, Abe H, Shinozaki K, Yamaguchi-Shinozaki K (2002) DNA-binding specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration- and cold-inducible gene expression. Biochem Bioph Res Co 290(3):998–1009
Sanjaya HPY, Su RC, Ko SS, Tong CG, Yang RY, Chan MT (2008) Overexpression of Arabidopsis thaliana tryptophan synthase beta 1 (AtTSB1) in Arabidopsis and tomato confers tolerance to cadmium stress. Plant Cell Environ 31(8):1074–1085
Shan X, Wang J, Chua L, Jiang D, Peng W, Xie D (2011) The role of Arabidopsis Rubisco activase in jasmonate-induced leaf senescence. Plant Physiol 155(2):751–764
Song SS, Huang H, Wang JJ, Liu B, Qi TC, Xie DX (2017) MYC5 is Involved in jasmonate-regulated plant growth, leaf senescence and defense responses. Plant and Cell Physiol 58(10):1752–1763
Trujillo LE, Sotolongo M, Menendez C, Ochogavia ME, Coll Y, Hernandez I, Borras-Hidalgo O, Thomma BPHJ, Vera P, Hernandez L (2008) SodERF3, a novel sugarcane ethylene responsive factor (ERF), enhances salt and drought tolerance when overexpressed in tobacco plants. Plant Cell Physiol 49(4):512–525
Upadhyay RK, Soni DK, Singh R, Dwivedi UN, Pathre UV, Nath P, Sane AP (2013) SlERF36, an EAR-motif-containing ERF gene from tomato, alters stomatal density and modulates photosynthesis and growth. J Exp Bot 64(11):3237–3247
Wellburn AR (1994) The spectral determination of chlorophyll-a and chlorophhyll-B, as well as total carotenoids, using various solvents with spectrophotometers of different resolution. J Plant Physiol 144(3):307–313
Woo HR, Kim HJ, Nam HG, Lim PO (2013) Plant leaf senescence and death - regulation by multiple layers of control and implications for aging in general. J Cell Sci 126(21):4823–4833
Xie Q, Hu Z, Zhu Z, Dong T, Zhao Z, Cui B, Chen G (2014) Overexpression of a novel MADS-box gene SlFYFL delays senescence, fruit ripening and abscission in tomato. Sci Rep 4:4367
Yang Z, Tian LN, Latoszek-Green M, Brown D, Wu KQ (2005) Arabidopsis ERF4 is a transcriptional repressor capable of modulating ethylene and abscisic acid responses. Plant Mol Biol 58(4):585–596
Yu J, Zhang Y, Di C, Zhang Q, Zhang K, Wang C, You Q, Yan H, Dai SY, Yuan JS, Xu W, Su Z (2016) JAZ7 negatively regulates dark-induced leaf senescence in Arabidopsis. J Exp Bot 67(3):751–762
Zhang ZJ, Huang RF (2010) Enhanced tolerance to freezing in tobacco and tomato overexpressing transcription factor TERF2/LeERF2 is modulated by ethylene biosynthesis. Plant Mol Biol 73(3):241–249
Zhang L, Tian LH, Zhao JF, Song Y, Zhang CJ, Guo Y (2009) Identification of an Apoplastic protein involved in the initial phase of salt stress response in rice root by two-dimensional electrophoresis. Plant Physiol 149(2):916–928
Zhao L, Xia Y, Wu XY, Schippers JHM, Jing HC (2018) Phenotypic analysis and molecular markers of leaf senescence. Methods Mol Biol 1744:35–48
Zhou S, Cheng X, Li F, Feng P, Hu G, Chen G, Xie Q, Hu Z (2019) Overexpression of SlOFP20 in tomato affects plant growth, chlorophyll accumulation, and leaf senescence. Front Plant Sci 10:1510
Zhu ZQ, An FY, Feng Y, Li PP, Xue L, Mu A, Jiang ZQ, Kim JM, To TK, Li W, Zhang XY, Yu Q, Dong Z, Chen WQ, Seki M, Zhou JM, Guo HW (2011) Derepression of ethylene-stabilized transcription factors (EIN3/EIL1) mediates jasmonate and ethylene signaling synergy in Arabidopsis. Proc Natl Acad Sci U S A 108(30):12539–12544
Zhu X, Chen J, Xie Z, Gao J, Ren G, Gao S, Zhou X, Kuai B (2015) Jasmonic acid promotes degreening via MYC2/3/4- and ANAC019/055/072-mediated regulation of major chlorophyll catabolic genes. Plant J 84(3):597–610
Zhuo M, Sakuraba Y, Yanagisawa S (2020) A jasmonate-activated MYC2-Dof2.1-MYC2 transcriptional loop promotes leaf senescence in Arabidopsis. Plant Cell 32(1):242–262
Acknowledgements
This work was supported by the Natural Science Foundation of Chongqing of China (csts2019jcyj-msxmX0094), and the National Natural Science Foundation of China (31872121), and the Innovation project of people returned from studying abroad of Chongqing (cx2019158).
Funding
The authors have not disclosed any funding.
Author information
Authors and Affiliations
Contributions
GC, SZ, and ZH: designed and managed the research work and improved the manuscript. YC, PF, BT, QX: performed the experiments. YC: wrote the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
All authors have read and approved this version of the article and due care has been taken to ensure the integrity of this work. The authors declare that they have no conflict of interest.
Additional information
Communicated by Sukhpreet Sandhu.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, Y., Feng, P., Tang, B. et al. The AP2/ERF transcription factor SlERF.F5 functions in leaf senescence in tomato. Plant Cell Rep 41, 1181–1195 (2022). https://doi.org/10.1007/s00299-022-02846-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-022-02846-1