Abstract
Key message
DgD27 was cloned from D. grandiflorum for the first time and played an important role in shoot branching of chrysanthemum.
Abstract
Shoot branching plays an important role in determining plant architecture. D27 was previously proven to be involved in the strigolactone biosynthetic pathway in rice, Arabidopsis, and Medicago. To investigate the role of D27 in shoot branching of chrysanthemum, we isolated the D27 homolog DgD27. Functional analysis showed that DgD27 was a plastid-localized protein that restored the phenotype of Arabidopsis d27-1. Gene expression analysis revealed that DgD27 was expressed at the highest levels in stem, and was up-regulated by exogenous auxin. Decapitation could down-regulate DgD27 expression, but this effect could be restored by exogenous auxin. DgD27 expression was significantly down-regulated by dark treatment in axillary buds. In addition, DgD27 transcripts produced rapid responses in shoots and roots under conditions of phosphate absence, but only mild variation in responses in buds, stems, and roots with low nitrogen treatment. DgBRC1 transcripts also showed the same response in buds under low nitrogen conditions. Under phosphate deficiency, indole-3-acetic acid (IAA) levels increased, zeatin riboside levels decreased, and abscisic acid (ABA) levels increased in the shoot, while both IAA and ABA levels increased in the shoot under low nitrogen treatments. Gibberellin acid levels were unaffected by phosphate deficiency and low nitrogen treatments. Taken together, these results demonstrated the diverse roles of DgD27 in response to physiological controls in chrysanthemum shoot branching.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Shoot branching plays an important role in determining plant architecture, and is a highly plastic determinant of plant shape, allowing plants to respond to environmental stresses (Evers et al. 2011). Shoot branching is controlled by complex interactions between genetic, hormonal (e.g., auxins, cytokinins, strigolactones), developmental (bud formation, bud outgrowth), and environmental (e.g., light, phosphorus, nitrogen) factors (Schmitz and Theres 1999; Beveridge et al. 2003; McSteen and Leyser 2005; Ongaro et al. 2008; Kebrom et al. 2013). Recent studies on plant hormones have focused on how auxins, strigolactones (SLs), and cytokinins (CKs) interact to regulate bud outgrowth and shoot branching (Leyser 2009; Ruyter-Spira et al. 2011; Kebrom et al. 2013; Ward et al. 2013).
A series of increasingly branching mutants in Arabidopsis thaliana, Oryza sativa, Petunia hybrida, and Pisum sativum have been identified in which SLs act as endogenous signals, regulating plant development by mediating the inhibition of axillary bud outgrowth (for review, see Al-Babili and Bouwmeester 2015). Through analysis of these mutants, seven genes of either the SL biosynthesis or signaling pathway have been identified. Four components of SL biosynthesis pathway include D27, CCD7, CCD8, and MAX1. D27 encodes a novel chloroplast-located, iron-containing protein, which acts upstream of MAX1 in the SL biosynthesis pathway, and the d27 mutant has a relatively highly tillered and dwarf phenotype (Lin et al. 2009; Liu et al. 2011; Waters et al. 2012a); CCD7 and CCD8 encode divergent plastidic carotenoid cleavage dioxygenases that function in the chloroplast, while CCD7s cleave different 9-cis-carotenes, and CCD8 introduce a series of oxygen molecules (Sorefan et al. 2003; Booker et al. 2004; Snowden et al. 2005; Auldridge et al. 2006; Ledger et al. 2010; Guan et al. 2012; Kohlen et al. 2012; Pasare et al. 2013; Bruno et al. 2014; Lauressergues et al. 2015); MAX1 encodes a cytochrome P450 monooxygenase, and works downstream of CCD7 and CCD8 to catalyze the oxidation of the SL precursor carlactone to the first real SL (Booker et al. 2005; Lazar and Goodman 2006; Challis et al. 2013; Abe et al. 2014; Zhang et al. 2014). Another three components of the SL signaling pathway include MAX2, D14, and D53. MAX2 encodes an F-box protein, that functions in signaling pathways downstream of SLs and responds to SLs (Stirnberg et al. 2002; Johnson et al. 2006). D14 encodes an α/β-fold hydrolase, which is proposed to be specific to signaling via MAX2 that mediates both SL and karrikin signaling (Arite et al. 2009; Hamiaux et al. 2012; Waters et al. 2012b). Finally, D53 encodes a protein that shares predicted features with the class I Clp ATPase proteins, and acts as a repressor of SL signaling (Jiang et al. 2013; Zhou et al. 2013). In addition, BRC1 acts downstream of the SLs pathway, locally regulating shoot branching (Minakuchi et al. 2010; Braun et al. 2012; Chen et al. 2013).
Auxins inhibit bud outgrowth and are synthesized in the shoot apex and transported basipetally in the polar auxin transport stream (PATS), while CKs and SLs are transported acropetally in the xylem. Auxins act indirectly on entering the bud through two mechanisms: the auxin transport canalization model and the second messenger model. The auxin transport canalization model works as follows: if it is assumed that during active growth, buds establish their own PATS into the main stem, then high auxin concentrations in the main stem can prevent bud activation by reducing the sink strength of the main stem for auxins, thereby preventing the canalization of auxin transport out of the bud (Bennett et al. 2006; Prusinkiewicz et al. 2009; Balla et al. 2011). In the second messenger model, auxins in the main stem can regulate the synthesis of CKs or SLs, which act as auxin second messengers within the bud and regulate branching (Tanaka et al. 2006; Brewer et al. 2009, 2015). Auxins can increase the expression of SL biosynthetic genes (Hayward et al. 2009), and can also negatively regulate CK content (Tanaka et al. 2006). CKs are directly transported into axillary buds to regulate outgrowth (Li et al. 1995). SLs regulate xylem sap CK (X-CK) levels through a feedback signal (Beveridge et al. 2000; Morris et al. 2001; Foo et al. 2005). SLs and CKs can also act antagonistically on pea bud growth (Dun et al. 2012). Auxins, CKs, and SLs interact in multiple feedback loops, and provide a robust balance in the regulation of shoot branching (Ferguson and Beveridge 2009; Domagalska and Leyser 2011; Ward et al. 2013).
The biosynthesis of SLs is responsive to environmental conditions, such as light, phosphate (P) and/or nitrogen (N), for regulating plant architecture. Light can affect SL levels or signaling and is particularly relevant to shading responses (Koltai et al. 2011). SLs may act downstream of the phytochrome B (phyB)-dependent response to both low red: far-red (R:FR) ratios and high R:FR ratios (Finlayson et al. 2010). P or N limitations in plants cause increased SL levels in the roots (Yoneyama et al. 2007a, b; Umehara et al. 2010; Kohlen et al. 2011; Mayzlish-Gati et al. 2012; Yoneyama et al. 2012). The different responses of SL expression to P or N deficiency are related to the nutrient acquisition strategies of plants (Yoneyama et al. 2012). Auxin signaling increases under low P conditions (López-Bucio et al. 2002; Pérez-Torres et al. 2008), while N deficiency decreases cytokinin levels, and the addition of cytokinin can counteract the root response to low phosphate (Martín et al. 2000). In rice, N deficiency increased indole-3-acetic acid (IAA) levels both in shoot and root in a split-root system, while P deficiency had no effect on shoot and root (Yoneyama et al. 2015).
D27 encodes an iron-containing protein in rice, which has homologs in Arabidopsis (AtD27 gene) and in Medicago truncatula (MtD27 gene) (Lin et al. 2009; Liu et al. 2011; Waters et al. 2012a). In rice, D27 is expressed mainly in the vascular cells of shoots and roots (Lin et al. 2009). In M. truncatula and rice, D27 transcripts are directly regulated by NSP1 and NSP2 (Liu et al. 2011). In-vitro evidence suggests that D27 is a β-carotene isomerase that converts all-trans-β-carotene into 9-cis-β-carotene (Alder et al. 2012). In Arabidopsis, AtD27 is localized to the plastid, and its transcripts are subject to both local feedback and auxin-dependent signals (Waters et al. 2012a).
Chrysanthemum (Dendranthema grandiflorum) is one of the most important commercial cut-flowers, however, most cultivars of cut-flower chrysanthemums produce a number of branches, that must be removed to maintain single cut-flowers, and this practice increases production costs. Therefore, regulating the pattern of shoot branching is a prerequisite for the molecular breeding of chrysanthemum. However, the molecular mechanism of how SLs regulate shoot branching in chrysanthemum is still unclear. In previous studies, SLs directly inhibited bud activity, and the expression of SLs biosynthetic genes were down-regulated by exogenous SL, but overridden by apical auxin application. SLs also down-regulate the biosynthesis of CKs. In addition, SLs contributed to chrysanthemum shoot branching control in response to P starvation conditions in a systemic way. The related genes DgCCD7, DgCCD8, DgMAX2, DgD14, and DgBRC1 have been identified (Liang et al. 2010; Chen et al. 2013; Dong et al. 2013; Wen et al. 2015; Xi et al. 2015).
In this study, we identified DgD27, a D27 orthologous gene of the DUF 4033 superfamily, in chrysanthemum (D. grandiflorum ‘Jinba’), and found that the expression of DgD27 was inhibited by decapitation and induced by auxins. Furthermore, DgD27 could produce a rapid response to light/dark, phosphate free, and low N (LN) treatments. These findings provide new insights into the dynamics of the putative SL biosynthesis component DgD27 in chrysanthemum.
Materials and methods
Plant materials and growth conditions
Chrysanthemum plantlets were propagated under sterile conditions in jars containing MS agar medium (Murashige and Skoog 1962), and then grown in a tissue culture room at 22–25 °C with a photoperiod of 16/8 h light/dark and a light intensity of 100–120 μmol·m−2·s−1.
Hormone stocks
Naphthaleneacetic acid (NAA; N0640, Sigma-Aldrich, Shanghai, China) was dissolved in 70 % ethanol, and GR24 (LeadGen Labs, Orange CT, USA) was dissolved in acetone. 6-Benzylaminopurine (6-BA; B3408, Sigma-Aldrich, Shanghai, China) was dissolved in 1 M NaOH, while indole-3-acetic acid (IAA; I5148, Sigma-Aldrich, Shanghai, China) and gibberellin acid (GA3; G7645, Sigma-Aldrich, Shanghai, China) were dissolved in ethanol.
Isolation of the full-length coding sequence for DgD27
Total RNA was extracted from stems with TRIzol Reagent (15596-026; Life Technologies/Invitrogen) and cDNA synthesis was performed using RevertAid First Strand cDNA Synthesis Kit (#K1621; Thermo Scientific). Primers were designed for DgD27 cloning based on the sequence regions of D27 genes that are conserved among Arabidopsis AtD27, Medicago MtD27, and rice D27 genes. After obtaining a conserved domain fragment using forward primer P_for and reverse primer P_rev, the 3′ fragment of DgD27 was amplified by the rapid amplification of cDNA Ends (RACE) method, using the 3′ RACE primers 3′-race1 and 3′-race2, and the 5′ fragment of DgD27 amplified in the same way using the 5′ RACE primers 5′-race1 and 5′-race2. The products, amplified using PrimeSTAR HS DNA Polymerase (R010A; TaKaRa, Dalian, China), were cloned into pMD18-T vector (D101A; TaKaRa, Dalian, China) and verified by sequencing (Zhongke Xilin Biotechnology, Beijing, China). Sequence alignment and phylogenic analysis were performed using the ClustalW and ESPript programs (http://www.genome.jp/tools/clustalw/) and the MEGA5.0 program (http://www.megasoftware.net/) respectively. Genomic DNA was extracted from the shoot apex and young leaves using the CTAB method. The primers used were gD27_for and gD27_rev. Primer sequences are listed in Table S1.
Subcellular localization
For construction of the 35S::DgD27-GFP reporter plasmids, the open reading frame (ORF) of DgD27 was amplified with primers SalI_for and SmaI_rev and was cloned into the binary vector pEZS-NL. Plastids are positively identified by co-expression of a protein fusion between the cRecA gene from Arabidopsis and RFP (Auldridge et al. 2006). Transformation into onion (Allium cepa) was performed as described previously (Varagona et al. 1992). After over night incubation in the dark, the GFP signal and RFP signal were examined under a confocal microscope at excitation wavelengths of 488 and 543 nm, respectively. Onion peels were unfolded in water and then viewed under a confocal laser scanning microscope (Nikon); images were acquired using EZ-C1 FreeViewer software (Nikon). Primer sequences are listed in Table S1.
Vector construction and plant transformation
For complementation experiments, the ORF of DgD27 was amplified with primers XbaI_for and SpeI_rev and was cloned into vector p-Super1300+. The resulting constructs were transformed into A. thaliana mutant Atd27-1 plants via Agrobacterium using the floral-dip method (Clough and Bent 1998). Independent transformants were screened on MS medium containing 70 mg/L Hygromycin B. Independent homozygous T3 lines with single insertion sites were used for the branching phenotype analysis. Amplification of Ubiquitin gene was performed using 30 cycles as a normalization control, and the expression of DgD27 was performed using 32 cycles. Primer sequences used for semi-quantitative reverse transcription PCR (RT-PCR) are listed in Table S1. Primers used for analysis of DgD27 or Ubiquitin were DgD27_for and DgD27_rev; UBQ_for and UBQ_rev. Primer sequences are listed in Table S1.
Split-plate and two-bud section system
The split-plate system was modified from that described previously for Arabidopsis and chrysanthemum (Chatfield et al. 2000; Chen et al. 2013). We removed a 10-mm wide strip of medium from the centre of a plate containing 30 mL solidified MS medium. Using a micro-pipette, we then injected 5 μM NAA or an equal volume of ethanol into the upper side of the media block, and 5 μM GR24 or an equal volume of acetone into the basal side. The plates were then left at 4 °C for the last 72 h to allow the hormone to diffuse evenly throughout the media. Chrysanthemum seedlings grown to 10–12 cm high in sterile condition had two nodes (node n and node n + 1) and a stem after decapitation or an intact shoot apex. After hormones had diffused evenly throughout the media, two-bud sections were cut from the chrysanthemum seedlings, and then inserted into media. The petri dishes were then held vertically in the culture room. The “control” chrysanthemum seedlings had two nodes and a stem after decapitation; those with an intact shoot apex were named “intact.” Topical buds (are equal to node n), basal buds (are equal to node n + 1), and stems were harvested separately 6 h after treatment for analysis of DgD27 transcripts. For each sample, 10–12 plants were collected. All experiments were repeated for 3 biological replicates. All samples were snap-frozen in liquid nitrogen, and stored at −80 °C.
Hormone treatments
The chrysanthemum cuttings (7 cm in height) were initially grown in vermiculite for 10 days, then plants were transferred into pots in a greenhouse, and hormone treatments began after 2 weeks (when plants were about 10 cm in height). Once every 2 days, 20 mL solution per 12 plants was sprayed over whole plants including leaves, buds, and stems. The solution contained 50 mg/L IAA, GA, or 6-BA with 0.5 % Tween-20, respectively, and water containing 0.5 % Tween-20 was used as control. The upper one-third section of plant axillary buds and stems was harvested 0, 1, 6, 24, and 48 h after treatments began, each containing tissue originating from 10 to 12 plants. All experiments were repeated for 3 biological replicates. Tissues were snap-frozen in liquid nitrogen, and stored at −80 °C.
Plant decapitation and exogenous auxin expression assay
The chrysanthemum cuttings (7 cm in height) were grown initially in vermiculite for 10 days, then plants were transferred into pots in a greenhouse, and the plant decapitation assay and exogenous auxin assay were performed once the cuttings reached 15 cm in height (about 3.5 weeks). The plant shoot apex was either decapitated or left intact. For the decapitated stems, Eppendorf tubes containing MS agar medium with 5 μM NAA or an equal volume of ethanol were placed over the residual stem after decapitation. Bud 1 was adjacent to the cut site with 2, 3, 4 progressively further away, the stem 1, 2, 3 and 4 mean for internodes, below the numbered bud. Buds and stems were harvested at 0, 1, 6, 24, 48 h after treatments start, snap-frozen in liquid nitrogen, and stored at −80 °C. Three replicates were used for each sample, with 12–15 plants used per replicate.
Light/dark treatments
The chrysanthemum seedlings were grown to 2–3 new leaves in sterile conditions. Two weeks after subculture, the light/dark treatments were initiated. For the light treatment, the seedlings were subjected to a normal 16/8 h light/dark photoperiod, and to a 16/8 h dark/dark photoperiod for the dark treatment. Axillary buds and stems were harvested after 5 days, snap-frozen in liquid nitrogen, and stored at −80 °C. Three replicates were done for each sample, with 10–12 plants used per replicate. Phenotype observation was performed after 5 days, and measurements were recorded for each plant’s height, internode length, stem diameter, length of leaf, and width of leaf. The data presented show average values of 10–12 plants.
RNA extraction and quantitative real-time PCR (qRT-PCR) Analysis
Plant samples of 100–150 mg, each containing tissue originating from 10 to 15 plants, were harvested and total RNA was isolated using TRIzol Reagent (Invitrogen, USA). RNA integrity was examined at 260 and 280 nm by NanoDrop 2000 (UV–Vis; Thermo Fisher Scientific Inc., USA) and checked on a gel electrophoresis in 1.5 % agarose gels with 1.5 μL RNA. The RNA (1 μg) of each tissue was reverse transcribed using FastQuant RT Kit (with gDNase; KR106; Tiangen, China). The resultant cDNA was diluted tenfold and kept at −20 °C for qRT-PCR analysis. qRT-PCR was performed on a StepOnePlus™ Real-time PCR detection system (Applied Biosystems, Foster City, CA, USA) using KAPA™ SYBR Fast qPCR Master Mix (Microread, Beijing, China). Each reaction consisted of 2 μL of cDNA template, 0.4 μL of each primer, 0.4 μL of ROX High, and 10 μL master mix in a final volume of 20 μL. The reaction procedure was as follows: denaturation at 95 °C for 20 s, followed by 40 cycles of 5 s at 95 °C and 20 s at 58 °C. The chrysanthemum β-Actin gene was used as an internal control for normalization, and the data were analyzed by OneStepPlus™ software (Applied Biosystems). The relative expression of the detected genes was calculated using the relative 2−ΔΔCT method. The experiments were repeated twice with three replicates. Primers used for were DgD27_for and DgD27_rev; DgBRC1_for and DgBRC1_rev; β-Actin_for and β-Actin_rev. Primer sequences are listed in Table S1.
Hydroponic culture
Chrysanthemum seedlings were grown to 8–10 cm high in sterile conditions in 3 weeks. Their roots were then washed free of MS agar medium after opening the caps 1 day, and the plants were transferred to a hydroponic solution consisting of an improved Hoagland’s solution. P was supplied as KH2PO4 and N was supplied as NH4NO3. The pH of all solutions was adjusted to 6.0 with 3 M KOH. The final concentrations in the different solutions were 1.0 mM P and 17.0 mM N (normal solution; that is, the P/N recovery solution), 0 mM P and 17.0 mM N (P absence), and 1.0 mM P and 0.17 mM N (LN). The plants were grown in a growth chamber with a 16/8 h photoperiod at a light intensity of 100–120 μmol m−2 s−1 at 25/20 °C.The absent P or LN treatments were initiated 12 days after the transfer, then plants were removed to the P/N recovery solution after 2 weeks. The upper one-third section of the plants was regarded as “bud up” compared with the lower one-third of plants, which were regarded as “bud base.” Stem has the same status as bud. Leaf, stem up/base, bud up/base, and root tissue were harvested at 0, 1, 10, 24, 48 h after treatments started, snap-frozen in liquid nitrogen, and stored at −80 °C. Three replicates were done for each sample, with 10–12 plants used per replicate.
Hormone quantification
The snap-frozen samples (200 mg) from the light/dark treatments and P absence/LN treatments were ground into fine powder with quartz sand, and dissolved in 3.0 mL of phosphate-buffered saline (PBS) containing 0.1 % (v/v) Tween-20 and 0.1 % (w/v) gelatin (pH 7.5) to quantify free IAA, zeatin riboside (ZR), abscisic acid (ABA), and GA3 by the enzyme-linked immunosorbent assay (ELISA), following the protocol described previously (Zhao et al. 2006). The mouse monoclonal antigen and corresponding antibodies were produced at the Center of Crop Chemical Control, China Agricultural University, China (Weiler et al. 1981; Wang et al. 2012). Calculations of the ELISA data were performed as described by Weiler et al. (1981). Three replicates were done for each sample.
Statistical analysis
All experiments were replicated three times to confirm the results. The data presented are mean ± standard error from a typical single experiment. ANOVA was conducted, followed by a Duncan’s test. Different letters indicate significant differences (P < 0.05) between different treatments.
Results
Identification of the D27 gene homologue from chrysanthemum
To address the regulatory role of the D27 gene during shoot branching of chrysanthemum, its putative ortholog, DgD27, was isolated, and full-length cDNA of DgD27 was isolated by 5′ and 3′ RACE PCR. The full transcript of DgD27 (accession number KR029722) is 1123 bp in length and contains an 822 bp ORF encoding a predicted protein of 273 amino acids, a 70 bp 5′ untranslated region (UTR), and a 231 bp 3′ UTR. The predicted amino acid sequence of the conserved domain belongs to the DUF 4033 superfamily, a domain of unknown function. The genomic fragment corresponding to the DgD27 gene isolated by PCR comprised 2653 bp, and the DgD27 gene was shown to have a seven-exon structure, comprising 288, 140, 89, 62, 109, 93, and 41 bp in size respectively (Fig. 1a).
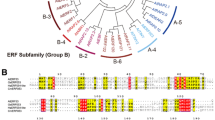
Sequence analysis of the full-length cDNA named DgD27. a Structure of the DgD27 gene. b Alignment of the predicted amino acid sequences of DgD27 compared with homologs in Arabidopsis (AtD27), corn (ZmD27), rice (OsD27), and Medicago (MtD27). The alignment was generated using ClustalW and ESPript (http://espript.ibcp.fr/ESPript/ESPript/). c Phylogenetic analysis of response regulators from a range of plant species. The aligned sequences were used to construct a phylogenetic tree using MEGA5.0. Accession numbers are listed in Table S2
Amino acid sequence comparisons between DgD27 and its orthologs from Medicago, Arabidopsis, rice, and maize showed that the predicted DgD27 has a 46.35 % sequence identity to MtD27, 37.91 % identity to AtD27, 40 % identity to OsD27, and 41.82 % identity to ZmD27 (Fig. 1b). To explore the evolutionary relationship among DgD27 genes from various plant species, we performed phylogenetic analysis using MEGA5.0 software. Our analysis showed that DgD27 is more closely related to MtD27 and AtD27, which belong to clade 1 of the DWARF27 clade (Waters et al. 2012a) (Fig. 1c).
Tissue specificity of DgD27 expression
We used qRT-PCR to determine the expression pattern of DgD27 in axillary bud, leaf (petiole removed), petiole, stem, shoot apex and root during the vegetative phase of chrysanthemum. As shown in Fig. 2, the highest expression of DgD27 was found in stem, followed by in shoot apex, axillary bud, root, and it was only weakly expressed in leaf and petiole. The expression pattern was similar to that reported for another study of D27 in rice (Lin et al. 2009), indicating that DgD27 expression is strongly correlated with shoot branching development.
DgD27 is localized to the plastid
To further characterize the function of DgD27, we investigated the subcellular localization of DgD27. The ORF of DgD27 was fused to green fluorescent protein (GFP) in the pEZS-NL vector, and the construct 35S::DgD27-GFP was bombarded into onion epidermal cells. The co-expressed marker was used to mark the plastids as negative controls. We found that the control displayed red fluorescence throughout the plastids (Fig. 3b), while the GFP fluorescence fully overlapped with the red fluorescence from the co-expressed plastid-specific marker (Fig. 3c), demonstrating that DgD27 is located inside the plastids. This is in accord with plastid localization of OsD27 and AtD27 (Lin et al. 2009; Waters et al. 2012a).
Subcellular localization of DgD27. a Epifluorescence micrographs of onion epidermis transiently transformed with p35S::DgD27-GFP plasmid; b plastids are positively identified by co-expression of a protein fusion between the cRecA gene from Arabidopsis and RFP; c the merged images confirm colocalization of the two fluorescent signals; d bright field micrographs of the same field of view to provide the cell outline
DgD27 complementation of Arabidopsis d27
To confirm the biological function of DgD27, a 35S::DgD27 transgene was introduced into the Arabidopsis d27-1 mutant using a floral dip (Clough and Bent 1998). Three independent transgenic lines were selected for phenotypic appraisal, and the DgD27 expression level was checked by RT-PCR analysis. As shown in Fig. 4, all three transgenic lines had elevated levels of DgD27 mRNA expression. Expression of DgD27 in d27-1 reduced the mean number of branches, and their branching phenotypes were similar to those of wild-type plants. The expression of DgD27 in Atd27-1 plants almost completely restored the phenotype back to that of the wild type. These results confirmed DgD27 as the functional ortholog of D27.
Complementation of Arabidopsis d27-1 mutant phenotype with DgD27. a Comparison of phenotypes of wild-type (WT), d27-1 and d27-1 transformed with the 35S::DgD27 constructs. b The number of secondary rosette branches produced by WT, d27-1 and three independent homozygous lines transformed carrying 35S::DgD27. The mean number of rosette branches with a length of at least 5 mm is shown. Data are mean ± SE; n = 15. c Transcript levels of the overexpressed DgD27 for the experiment presented in b were determined by RT-PCR. Detection of UBQ transcript was used as a cDNA normalization control
Induction of DgD27 expression by auxin
To investigate auxin regulation of DgD27, qRT-PCR analysis was performed on two-bud segments cultured in a split-plate system, with or without apical application of auxin (Fig. 5b). Our results showed that, whether in bud or stem, the DgD27 expression level dramatically decreased 6 h after decapitation, but this reduction was successfully reversed by apical NAA application, and was most strongly induced in the top bud. These results indicated that auxins can induce DgD27 expression.
DgD27 expression response to decapitation, auxin, and GR24. a Plantlets were treated with 0 μM or 5 μM GR24 to the intact basal side, and an equal volume of acetone as a control. b Decapitated plants were treated with 0 μM or 5 μM NAA to the upper side (with an equal volume of ethanol as a control), and with 0 μM or 5 μM GR24 to the basal side (with an equal volume of acetone as a control). Dash vertical control, multi symbol none treated. Top buds, bottom buds and stems were collected 6 h after treatment. Detection of β-Actin was used as a normalization control. Results are means of three biological replicates analyzed by qRT-PCR, with 10–12 plants for each replicate; letters indicate significant differences (P < 0.05) between different treatments. ANOVA followed by a Duncan’s test. Error bars shown
Feedback control of DgD27 expression
We investigated the effects of SLs on DgD27 expression, using two-bud segments cultured in a split-plate system. As shown in Fig. 5a, DgD27 expression was greatly reduced in all samples upon decapitation. In the bottom bud, DgD27 expression was down-regulated by GR24 in intact plants compared with untreated controls, but this down-regulation was less than that observed upon decapitation. However, in the top bud, there was an opposite effect on DgD27 expression between GR24 treatment and decapitation: GR24 increased DgD27 expression in intact plants compared with untreated controls, and DgD27 was down-regulated upon decapitation. As shown in Fig. 5b, there was a significant difference between control and decapitated plants treated with NAA or GR24 in all samples, except the top bud with basal GR24. In the bottom bud, DgD27 expression was up-regulated by apical NAA or/with basal GR24 compared with untreated controls. Furthermore, the level of up-regulation in decapitated plants treated with both apical NAA and basal GR24 was lower than that of the decapitated plants treated with apical NAA, but higher than the decapitated plants treated with basal GR24. However in the top bud, basal GR24 had no effect on DgD27 expression, while there was a significant increase in the decapitated plants treated with both apical NAA and basal GR24 or only with apical NAA. These results indicated the presence of different feedback controls for DgD27 expression between the top and bottom buds in chrysanthemum and also a relative competitiveness between these two buds.
Expression of DgD27 with hormone treatments
We used qRT-PCR analysis to determine the expression patterns of DgD27 in intact plants given hormone treatment. As shown in Fig. 6, the results demonstrated that the expression patterns differed for each type of hormone treatment. The expression level of DgD27 was induced significantly by GA and 6-BA in buds but was induced only weakly by IAA, while it increased significantly in stems at 24 h with 6-BA treatment, and at 48 h with GA treatment. However, the expression level of DgD27 was recovered to pre-treatment levels in buds at 48 h with GA treatment, while levels remained high with a 6-BA treatment. Taken together, these data suggested that DgD27 responded to GA and CKs in both buds and stems, and exhibited different expression patterns.
Expression patterns of DgD27 in response of buds and stems to hormone treatments in chrysanthemum. a DgD27 expression was induced by treatment with 50 mg/L IAA, GA, or 6-BA in buds, respectively. b DgD27 expression was induced by treatment with 50 mg/L IAA, GA, or 6-BA in stems, respectively. Each bud and stem sample at 0 h were set to equal 1. Results are the means of three biological replicates with 10–12 plants for each replicate. Error bars shown
Effects of apical dominance on DgD27 expression
Chrysanthemum has strong apical dominance: the bud in the upper part of the plant can rapidly grow out after decapitation. To determine the effects of apical dominance on DgD27 expression levels, a classic decapitation assay was conducted. As shown in Fig. 7a, the transcript levels of DgD27 dramatically decreased in both buds 1 h after decapitation while remaining at a low level in buds 1 and 2, and increasing in buds 3 and 4 6 h after decapitation. By 48 h after decapitation, levels were high in buds 1 and 2 and had recovered almost to pre-decapitation levels in buds 3 and 4. As shown in Fig. 7b, the transcript levels of DgD27 dramatically decreased in stems 1 and 2 1 h after decapitation, but dramatically decreased in stems 3 and 4 at 6 h, and then increased 24 h after decapitation; by 48 h after decapitation, levels had recovered almost to pre-decapitation levels in all stems.
Transcript levels of DgD27 after decapitation and decapitation + NAA. a DgD27 transcript levels in buds 1–4 were analyzed 0, 1, 6, 24 and 48 h after decapitation. b DgD27 transcript levels in stems 1–4 were analyzed 0, 1, 6, 24 and 48 h after decapitation. c DgD27 transcript levels in buds 1–4 were analyzed 6 h after decapitation and after decapitation + NAA treatment. d DgD27 transcript levels in stems 1–4 were analyzed 6 h after decapitation and decapitation + NAA. Bud position was recorded basipetally. Stem position was below the numbered bud. Each bud and stem sample at 0 h were set to equal 1. Results are means of three biological replicates with 12–15 plants for each replicate. Asterisks indicate statistically significant differences (P < 0.05) calculated by Duncan’s test between the intact and decapitated plants. Error bars shown
Substitution of the decapitated apex with exogenous auxin resulted in a 2- to 20-fold increase in the expression of DgD27 in all buds and stems (Fig. 7c, d). The closer the bud is located to the exogenous auxin, the higher the expression was induced. This result indicated that induction of DgD27 expression by exogenous auxin after decapitation has a position effect, and expression consistent with the known positive effect of auxins on gene transcription in two-bud segments cultured experimentally. Our results suggested that DgD27 transcription was down-regulated rapidly by decapitation, and increased by exogenous auxin, which plays a role in the apical dominance.
Expression of DgD27 with light/dark treatments
To examine whether DgD27 can respond to light, we assessed the expression levels of DgD27 in intact plants with light/dark treatments after 5 days. As shown in Fig. 8 and Table 1, a dark treatment negatively affected plant growth, increased the height and length of internodes, and reduced the stem’s diameter and both the length and width of leaves. As shown in Fig. 9, DgD27 expression was significantly down-regulated by dark treatment in axillary buds, though it was not significantly up-regulated in stems.
Expression patterns of DgD27 in response of axillary buds and stems to light/dark treatments in chrysanthemum. Results are means of three biological replicates with 10–12 plants for each replicate. Different letters above the bars indicate significant differences (P < 0.05) calculated by a Duncan’s test
We measured the hormone levels in plants exposed to light/dark treatments after 5 days. As shown in Table 2, four hormone levels showed significant changes 5 days after dark treatments. ABA and GA levels significantly decreased both in axillary bud and stems, while the IAA level only significantly decreased in axillary buds. In contrast, the ZR level only significantly decreased in stems. These results showed that DgD27 expression in shoots has a correlation with auxins under light/dark conditions.
Response of DgD27 in shoot and root under conditions of phosphate absence
To examine whether DgD27 can respond to lack of phosphate, we assessed the expression levels of DgD27 in a hydroponic solution with no phosphate and after 2 weeks in a normal phosphate solution. As shown in Fig. 10, the level of DgD27 expression was significantly up-regulated by phosphate absence both in shoots and roots. In particular it was significantly increased at 1 h and attained higher levels in basal buds 48 h after phosphate was removed. There was a more dramatic response to phosphate absence in buds than in stems, and in addition, the level of DgD27 expression increased more rapidly in the upper buds than in the basal buds. When plantlets were treated with normal phosphate solution after a 2 weeks absence of phosphate, DgD27 expression then showed an obvious change both in shoots and roots but not in upper buds. By 48 h after the normal phosphate solution treatment, DgD27 expression levels were high in basal buds and basal stems, while they had recovered to almost T0 levels in upper bud, upper stem, and root. There was a complementary mechanism between DgD27 expression response to phosphate deficiency and recovery conditions in shoots, but not in roots. These results indicated that the response of chrysanthemum to absence of phosphate correlates with DgD27 transcript levels.
DgD27 gene expression under phosphate deficiency/recovery conditions in shoot and root. a DgD27 transcript levels in bud up were analyzed 0, 1, 10, 24 and 48 h after phosphate deficiency/recovery conditions. b DgD27 transcript levels in bud base were analyzed 0, 1, 10, 24 and 48 h after phosphate deficiency/recovery conditions. c DgD27 transcript levels in stem up were analyzed 0, 1, 10, 24 and 48 h after phosphate deficiency/recovery conditions. d DgD27 transcript levels in stem base were analyzed 0, 1, 10, 24 and 48 h after phosphate deficiency/recovery conditions. e DgD27 transcript levels in root were analyzed 0, 1, 10, 24 and 48 h after phosphate deficiency/recovery conditions. The data presented are typical of three independent biological replicates with 10–12 plants for each replicate. Error bars shown
Response of DgD27 in shoot and root under low N conditions
We investigated whether the effects of N treatment on DgD27 expression were the same as for phosphate by growing plants in a hydroponic solution with low N, and after 2 weeks transferring them to a normal N recovery solution. We found that the DgD27 expression level was highest at 1 h in leaves, but there was a mild variation up in buds, stems, and roots with a low N treatment. In addition, after 48 h of LN conditions, there was no increase in DgD27 expression in shoot, but a slight increase in root (Fig. 11). After transfer to the normal nitrogen solution treatment, DgD27 expression then showed the opposite response with low N treatment. Levels obviously increased in bud, leaf, and root, but not in stem at 48 h after the normal nitrogen solution treatment. There was also a complementary mechanism between DgD27 expression response to low N treatment and normal nitrogen solution treatment in both shoot and root. Thus, the expression of DgD27 under low N conditions was not the same as that observed under conditions of absent phosphate. These results suggested that, unlike phosphate, N does not affect DgD27 expression in buds and stems of chrysanthemum.
DgD27 gene expression under low nitrogen/nitrogen recovery conditions in shoot and root. LN low nitrogen, NR nitrogen recovery. a DgD27 transcript levels in bud were analyzed 0, 1, 10, 24 and 48 h after low nitrogen/nitrogen recovery. b DgD27 transcript levels in stem were analyzed 0, 1, 10, 24 and 48 h after nitrogen recovery/nitrogen recovery. c DgD27 transcript levels in leaf were analyzed 0, 1, 10, 24 and 48 h after low nitrogen/nitrogen recovery. d DgD27 transcript levels in root were analyzed 0, 1, 10, 24 and 48 h after nitrogen recovery/nitrogen recovery. The data presented are typical of three independent biological replicates with 10–12 plants for each replicate. Error bars shown
Response of DgBRC1 in bud under low N conditions
BRC1, a bud-specific transcription factor was previously proven to act downstream of the SL pathway (Dun et al. 2012, 2013). In chrysanthemum, P starvation induced increased endogenous SL analog levels in all parts of the plant, and DgBRC1 transcripts produced rapid responses in nodes under conditions of P starvation (Xi et al. 2015). To examine whether DgBRC1 can respond to low N conditions, we assessed the expression levels of DgBRC1 in a hydroponic solution with low N and after 2 weeks in a normal N recovery solution. As shown in Fig. 12, DgBRC1 expression showed a slight increase in buds after 1 h of LN conditions, and a slight drease after a 48 h treatment. After transfer to the normal nitrogen solution treatment, DgBRC1 expression then showed the opposite response with low N treatment. The expression levels of DgBRC1 obviously increased 48 h after the normal nitrogen solution treatment. Thus, the expression of DgBRC1 under low N conditions was the same as that of DgD27, and the results indicated that the response of chrysanthemum to low N does not correlate with DgBRC1 transcript levels.
DgDBRC1 gene expression under low nitrogen/nitrogen recovery conditions in bud. LN low nitrogen, NR nitrogen recovery. DgBRC1 transcript levels in bud were analyzed 0, 1, 10, 24 and 48 h after low nitrogen/nitrogen recovery. The data presented are typical of three independent biological replicates with 10–12 plants for each replicate. Error bars shown
Effects of phosphate deficiency/low N on shoot hormone contents
The effects of phosphate deficiency/low N on shoot hormone content were examined in hydroponically grown chrysanthemum plants after 2 weeks. As shown in Table 3, IAA levels increased in upper buds and ABA levels increased in basal buds both under phosphate deficiency and low N treatments. ZR levels decreased in basal buds only under phosphate deficiency treatments. By contrast, neither phosphate deficiency nor low N treatments affected GA levels in upper or basal buds. As shown in Table S3, ABA levels increased in upper stems under phosphate deficiency, while IAA levels and ZR levels increased under low N treatments. ZR levels decreased in basal stems only under phosphate deficiency treatments. As in buds, neither phosphate deficiency nor low N treatments affected GA levels in upper or basal stems. These results showed that multiple hormones respond to phosphate deficiency and/or low N in different parts of shoot. Thus, shoot branching was regulated by different hormonal balance responses to phosphate deficiency or low N.
Discussion
The eukaryotic sequences of D27 broadly grouped into three distinct clades. Clades 1 and 2 only contain members from land plants, while the clade 3 also contains members from chlorophyte algae and diatoms (Waters et al. 2012a). In the present study, we only isolated a D27 ortholog gene, named DgD27, which belongs to the D27 clade specific to land plants (Waters et al. 2012a). With functional complementation of d27-1 mutants, a conserved function of DgD27 was supported. The biological function of the D27-like protein family is unknown, with no known link to SLs (Delaux et al. 2012; Waters et al. 2012a).
The interactions between auxins and SLs in controlling shoot branching have been studied previously (Foo et al. 2005; Johnson et al. 2006; Crawford et al. 2010; Liang et al. 2010; Ward et al. 2013). In this study, we found that different feedback controls for DgD27 expression between the top and bottom buds were present. Similar patterns of DgD27 response to NAA/GR24 were also observed in DgCCD8 and DgD14 (Liang et al. 2010; Wen et al. 2015). However, the obvious difference was that an apical NAA treatment together with basal GR24 was not more effective at increasing DgD27 expression in the top bud. Auxins were transported basipetally down the main stem in PATS, whereas SLs were transported acropetally. DgD27 was upstream of the SLs biosynthesis pathway. Differences in apical and basal bud responses may be caused by transport over the stem segment 6 h after treatments, and this also suggested that there was relative competitiveness existing between different buds in chrysanthemum. Additionally, ABA acted downstream of the main shoot PATS and the SL pathway, while regulated bud outgrowth responses to the R:FR also suppressed lower bud outgrowth and elongation (Reddy et al. 2013; Yao and Finlayson 2015). Different ABA levels and light signaling in different buds may be another important reason leading to altered DgD27 expression patterns. In future studies, identifying the ABA levels and the responses to ABA and light treatments would help to characterize the relationship between apical and basal buds in chrysanthemum.
In this study, the response of DgD27 to GA and CKs treatments in buds was the same as that of DgD14 under the same conditions. These results indicated that GA and CKs treatments triggered the response of the SL pathway. However, the expression levels of these two genes were induced only weakly by IAA, and there was no obvious influence on any structure with IAA (Wen et al. 2015). This evidence suggested that the effects of GA and CKs treatments may be caused by the influence of apical dominance or main shoot PATS.
Use of decapitation and auxin application treatment showed that DgD27 expression responded rapidly to the release of apical dominance resulting from decapitation. Similar results were obtained in D10, CCD7, CCD8, and DgBRC1 (Ferguson and Beveridge 2009; Hayward et al. 2009; Zhang et al. 2010; Chen et al. 2013). Additionally, DgD27 expression levels in buds and stems were up-regulated after apical application of NAA, and had a positional effect: near the source of NAA, the increase of expression levels was much higher than that of the distant, further demonstrating that DgD27 as a response regulator is related to auxin regulation of bud outgrowth.
Light can affect SL levels or the signaling response to shading. Auxin signaling was required for the phytochrome B (phyB) hypobranching phenotype, SL may be acting downstream of the phyB-dependent response to both low R:FR and high R:FR (Finlayson et al. 2010). Auxin sensitivity had also been implicated in shade avoidance (Reddy and Finlayson 2014). In the current study, we found that DgD27 expression was significantly down-regulated by dark treatment in axillary buds, whereas the IAA level significantly decreased. This was consistent with the fact that auxins can induce DgD27 expression in chrysanthemum. Dark treatment is extreme shading, and differs from low/high R:FR, and it does not act as a feedback control for DgD27 expression. This may be why DgD27 expression was not up-regulated by the dark treatment, as expected. Additionally, the fact that the IAA level significantly decreased may be associated with the stage of growth, evidence of which had been provided in the research on Arabidopsis phyB (Reddy and Finlayson 2014).
SLs are able to respond to nutrient supply conditions: the expression levels of the SL pathway genes were up-regulated by conditions of P and/or N deficiency (Liu et al. 2011; Bonneau et al. 2013). In chrysanthemum, three different types of SLs were identified and levels of all three SLs showed a strong increase under absent P conditions, implying that absence of P also stimulates SL biosynthesis in chrysanthemum (Xi et al. 2015). In this study, the results showed that the expression of DgD27 rapidly increased under conditions of absent P, a result which was consistent with earlier findings in other species (Umehara et al. 2010; Liu et al. 2011). As to the expression pattern, similar patterns of DgD27 responses to P absence were also observed in DgCCD7, DgCCD8, DgMAX2, DgD14, and DgBRC1 (Wen et al. 2015; Xi et al. 2015). The expression levels of these six genes in buds had both significantly increased after 48 h in a solution without P, and these results suggested that signals triggered by P starvation had comparative continuity and cooperativity.
Meanwhile, our results showed different responses of DgD27 expression between absent P and low N conditions. The expression levels of DgD27 in buds showed that almost no difference was observed in a solution with low N. This result was also observed in DgD14 between absent P and low N conditions (Wen et al. 2015). Recent studies had indicated that the different responses of SL exudation to P or N deficiency have been shown to depend on the P level (Yoneyama et al. 2012; Czarnecki et al. 2013). In chrysanthemum, P levels were found to be significantly reduced both in shoots and roots under absent P conditions, but were not reduced in shoots under low N conditions (Wen et al. 2015). It was supposed that P did not relocate in plants exposed to low N, and this might be the reason why the SL gene was not triggered in chrysanthemum.
DgBRC1 was mainly expressed in dormant axillary buds to suppress the response of bud outgrowth (Chen et al. 2013). Expression of DgBRC1 rapidly responded to P absence and this suggested that P starvation triggered signals that facilitated bud inhibition in chrysanthemum (Xi et al. 2015). However, the response of DgBRC1 to low N conditions was the same as that of DgD27 and DgD14 under the same conditions. Thus, the present evidence may suggest that N did not regulate shoot branching directly via the SL pathway and/or DgBRC1 in chrysanthemum.
After examining the shoot hormone contents, there were found to be three reasons that may have caused the difference between the bud (stem) base and upper parts upon P deficiency and/or low N conditions: (1) Developmental status and distance from the shoot apex influence activation of axillary buds. (2) Local control and/or systemic regulation of auxin in the axillary bud. (3) ABA may play a role in regulation shoot branching upon P deficiency and/or low N conditions.
In summary, DgD27 played diverse roles in chrysanthemum regulating shoot branching by hormonal and environmental factors (e.g., light, P, N). Our results reveal new insights into the elucidation of shoot branching mechanisms via the putative SL biosynthetic component DgD27 and provide a theoretical basis for breeding cut-chrysanthemum varieties with no (or fewer) branches.
Author contribution statement
Chao Wen and Liangjun Zhao conceived and designed the study. Chao Wen, Qingcui Zhao, Jing Nie, Guoqin Liu, Lin Shen, Lin Xi, and Chenxia Cheng performed the experiments. Chao Wen wrote the paper. Liangjun Zhao, and Nan Ma reviewed and edited the manuscript. All the authors read and approved the manuscript.
Abbreviations
- CCD:
-
Carotenoid cleavage dioxygenase
- LN:
-
Low nitrogen
- P:
-
Phosphate
- PATS:
-
Polar auxin transport stream
- PBS:
-
Phosphate-buffered saline
- SL:
-
Strigolactone
- ZR:
-
Zeatin riboside
References
Abe S, Sado A, Tanaka K, Kisugi T, Asami K, Ota S, Kim HI, Yoneyama K, Xie X, Ohnishi T, Seto Y, Yamaguchi S, Akiyama K, Yoneyama K, Nomura T (2014) Carlactone is converted to carlactonoic acid by MAX1 in Arabidopsis and its methyl ester can directly interact with AtD14 in vitro. PNAS 111:18084–18089
Al-Babili S, Bouwmeester HJ (2015) Strigolactones, a novel carotenoid-derived plant hormone. Annu Rev Plant Biol 66:161–186
Alder A, Jamil M, Marzorati M, Bruno M, Vermathen M, Bigler P, Ghisla S, Bouwmeester H, Beyer P, Al-Babili S (2012) The path from β-carotene to carlactone, a strigolactone-like plant hormone. Science 335:1348–1351
Arite T, Umehara M, Ishikawa S, Hanada A, Maekawa M, Yamaguchi S, Kyozuka J (2009) d14, a strigolactone-insensitive mutant of rice, shows an accelerated outgrowth of tillers. Plant Cell Physiol 50:1416–1424
Auldridge ME, Block A, Vogel JT, Dabney-Smith C, Mila I, Bouzayen M, Magallanes-Lundback M, DellaPenna D, McCarty DR, Klee HJ (2006) Characterization of three members of the Arabidopsis carotenoid cleavage dioxygenase family demonstrates the divergent roles of this multifunctional enzyme family. Plant J 45(6):982–993
Balla J, Kalousek P, Reinöhl V, Friml J, Procházka S (2011) Competitive canalization of PIN-dependent auxin flow from axillary buds controls pea bud outgrowth. Plant J 65(4):571–577
Bennett T, Sieberer T, Willett B, Booker J, Luschnig C, Leyser O (2006) The Arabidopsis MAX pathway controls shoot branching by regulating auxin transport. Curr Biol 16:553–563
Beveridge CA, Symons GM, Turnbull CGN (2000) Auxin inhibition of decapitation-induced branching is dependent on graft-transmissible signals regulated by genes Rms1 and Rms2. Plant Physiol 123:689–697
Beveridge CA, Weller JL, Singer SR, Hofer JM (2003) Axillary meristem development. Budding relationships between networks controlling flowering, branching, and photoperiod responsiveness. Plant Physiol 131:927–934
Bonneau L, Huguet S, Wipf D, Pauly N, Truong HN (2013) Combined phosphate and nitrogen limitation generates a nutrient stress transcriptome favorable for arbuscular mycorrhizal symbiosis in Medicago truncatula. New Phytol 199:188–202
Booker J, Auldridge M, Wills S, McCarty D, Klee H, Leyser O (2004) MAX3/CCD7 is a carotenoid cleavage dioxygenase required for the synthesis of a novel plant signaling molecule. Curr Biol 14:1232–1238
Booker J, Sieberer T, Wright W, Williamson L, Willett B, Stirnberg P, Turnbull C, Srinivasan M, Goddard P, Leyser O (2005) MAX1 encodes a cytochrome P450 family member that acts downstream of MAX3/4 to produce a carotenoid-derived branch-inhibiting hormone. Dev Cell 8:443–449
Braun N, de Saint Germain A, Pillot JP, Boutet-Mercey S, Dalmais M, Antoniadi I, Li X, Maia-Grondard A, Le Signor C, Bouteiller N, Luo D, Bendahmane A, Turnbull C, Rameau C (2012) The pea TCP transcription factor PsBRC1 acts downstream of strigolactones to control shoot branching. Plant Physiol 158:225–238
Brewer PB, Dun EA, Ferguson BJ, Rameau C, Beveridge CA (2009) Strigolactone acts downstream of auxin to regulate bud outgrowth in pea and Arabidopsis. Plant Physiol 150:482–493
Brewer PB, Dun EA, Gui R, Mason MG, Beveridge CA (2015) Strigolactone inhibition of branching independent of polar auxin transport. Plant Physiol 168:1820–1829
Bruno M, Hofmann M, Vermathen M, Alder A, Beyer P, Al-Babili S (2014) On the substrate- and stereospecificity of the plant carotenoid cleavage dioxygenase 7. FEBS Lett 588:1802–1807
Challis RJ, Hepworth J, Mouchel C, Waites R, Leyser O (2013) A role for more axillary growth1 (MAX1) in evolutionary diversity in strigolactone signaling upstream of MAX2. Plant Physiol 161:1885–1902
Chatfield SP, Stirnberg P, Forde BG, Leyser O (2000) The hormonal regulation of axillary bud growth in Arabidopsis. Plant J 24:159–169
Chen XL, Zhou XY, Xi L, Li JX, Zhao RY, Ma N, Zhao LJ (2013) Roles of DgBRC1 in regulation of lateral branching in chrysanthemum (Dendranthema x grandiflora cv. Jinba). PLoS ONE 8:e61717
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Crawford S, Shinohara N, Sieberer T, Williamson L, George G, Hepworth J, Muller D, Domagalska MA, Leyser O (2010) Strigolactones enhance competition between shoot branches by dampening auxin transport. Development 137:2905–2913
Czarnecki O, Yang J, Weston DJ, Tuskan GA, Chen JG (2013) A dual role of strigolactones in phosphate acquisition and utilization in plants. Int J Mol Sci 14:7681–7701
Delaux PM, Xie X, Timme RE, Puech-Pages V, Dunand C, Lecompte E, Delwiche CF, Yoneyama K, Bécard G, Séjalon-Delmas N (2012) Origin of strigolactones in the green lineage. New Phytol 195:857–871
Domagalska MA, Leyser O (2011) Signal integration in the control of shoot branching. Nat Rev Mol Cell Biol 12(4):211–221
Dong L, Ishak A, Yu J, Zhao R, Zhao L (2013) Identification and functional analysis of three MAX2 orthologs in chrysanthemum. J Integr Plant Biol 55:434–442
Dun EA, de Saint Germain A, Rameau C, Beveridge CA (2012) Antagonistic action of strigolactone and cytokinin in bud outgrowth control. Plant Physiol 158:487–498
Dun EA, de Saint Germain A, Rameau C, Beveridge CA (2013) Dynamics of strigolactone function and shoot branching responses in Pisum sativum. Mol Plant 6:128–140
Evers JB, van der Krol AR, Vos J, Struik PC (2011) Understanding shoot branching by modelling form and function. Trends Plant Sci 16:464–467
Ferguson BJ, Beveridge CA (2009) Roles for auxin, cytokinin, and strigolactone in regulating shoot branching. Plant Physiol 149:1929–1944
Finlayson SA, Krishnareddy SR, Kebrom TH, Casal JJ (2010) Phytochrome regulation of branching in Arabidopsis. Plant Physiol 152:1914–1927
Foo E, Bullier E, Goussot M, Foucher F, Rameau C, Beveridge CA (2005) The branching gene RAMOSUS1 mediates interactions among two novel signals and auxin in pea. Plant Cell 17:464–474
Guan JC, Koch KE, Suzuki M, Wu S, Latshaw S, Petruff T, Goulet C, Klee HJ, McCarty DR (2012) Diverse roles of strigolactone signaling in maize architecture and the uncoupling of a branching-specific subnetwork. Plant Physiol 160:1303–1317
Hamiaux C, Drummond RS, Janssen BJ, Ledger SE, Cooney JM, Newcomb RD, Snowden KC (2012) DAD2 is an alpha/beta hydrolase likely to be involved in the perception of the plant branching hormone, strigolactone. Curr Biol 22:2032–2036
Hayward A, Stirnberg P, Beveridge C, Leyser O (2009) Interactions between auxin and strigolactone in shoot branching control. Plant Physiol 151:400–412
Jiang L, Liu X, Xiong G, Liu H, Chen F, Wang L, Meng X, Liu G, Yu H, Yuan Y, Yi W, Zhao L, Ma H, He Y, Wu Z, Melcher K, Qian Q, Xu HE, Wang Y, Li J (2013) DWARF 53 acts as a repressor of strigolactone signalling in rice. Nature 504:401–405
Johnson X, Brcich T, Dun EA, Goussot M, Haurogne K, Beveridge CA, Rameau C (2006) Branching genes are conserved across species. Genes controlling a novel signal in pea are coregulated by other long-distance signals. Plant Physiol 142:1014–1026
Kebrom TH, Spielmeyer W, Finnegan EJ (2013) Grasses provide new insights into regulation of shoot branching. Trends Plant Sci 18(1):41–48
Kohlen W, Charnikhova T, Liu Q, Bours R, Domagalska MA, Beguerie S, Verstappen F, Leyser O, Bouwmeester H, Ruyter-Spira C (2011) Strigolactones are transported through the xylem and play a key role in shoot architectural response to phosphate deficiency in nonarbuscular mycorrhizal host Arabidopsis. Plant Physiol 155:974–987
Kohlen W, Charnikhova T, Lammers M, Pollina T, Toth P, Haider I, Pozo MJ, de Maagd RA, Ruyter-Spira C, Bouwmeester HJ, Lopez-Raez JA (2012) The tomato CAROTENOID CLEAVAGE DIOXYGENASE8 (SlCCD8) regulates rhizosphere signaling, plant architecture and affects reproductive development through strigolactone biosynthesis. New Phytol 196:535–547
Koltai H, Cohen M, Chesin O, Mayzlish-Gati E, Bécard G, Puech V, Dor BB, Resnick N, Wininger S, Kapulnik Y (2011) Light is a positive regulator of strigolactone levels in tomato roots. J Plant Physiol 168:1993–1996
Lauressergues D, André O, Peng J, Wen J, Chen R, Ratet P, Tadege M, Mysore KS, Rochange SF (2015) Strigolactones contribute to shoot elongation and to the formation of leaf margin serrations in Medicago truncatula R108. J Exp Bot 66(5):1237–1244
Lazar G, Goodman HM (2006) MAX1, a regulator of the flavonoid pathway, controls vegetative axillary bud outgrowth in Arabidopsis. PNAS 103:472–476
Ledger SE, Janssen BJ, Karunairetnam S, Wang T, Snowden KC (2010) Modified CAROTENOID CLEAVAGE DIOXYGENASE8 expression correlates with altered branching in kiwifruit (Actinidia chinensis). New Phytol 188:803–813
Leyser O (2009) The control of shoot branching: an example of plant information processing. Plant, Cell Environ 32:694–703
Li CJ, Guevara E, Herrera J, Bangerth F (1995) Effect of apex excision and replacement by 1-naphthylacetic acid on cytokinin concentration and apical dominance in pea plants. Physiol Plant 94:465–469
Liang J, Zhao L, Challis R, Leyser O (2010) Strigolactone regulation of shoot branching in chrysanthemum (Dendranthema grandiflorum). J Exp Bot 61:3069–3078
Lin H, Wang R, Qian Q, Yan M, Meng X, Fu Z, Yan C, Jiang B, Su Z, Li J, Wang Y (2009) DWARF27, an iron-containing protein required for the biosynthesis of strigolactones, regulates rice tiller bud outgrowth. Plant Cell 21:1512–1525
Liu W, Kohlen W, Lillo A, Camp RO, Ivanov S, Hartog M, Limpens E, Jamil M, Smaczniak C, Kaufmann K, Yang WC, Hooiveld G, Charnikhova T, Bouwmeester HJ, Bisseling T, Geurtsa R (2011) Strigolactone biosynthesis in Medicago truncatula and rice requires the symbiotic GRAS-type transcription factors NSP1 and NSP2. Plant Cell 23:3853–3865
López-Bucio J, Hernández-Abreu E, Sánchez-Calderón L, Nieto-Jacobo MF, Simpson J, Herrera-Estrella L (2002) Phosphate availability alters architecture and causes changes in hormone sensitivity in the Arabidopsis root system. Plant Physiol 129:244–256
Martín AC, del Pozo JC, Iglesias J, Rubio V, Solano R, de La Peña A, Leyva A, Paz-Ares J (2000) Influence of cytokinins on the expression of phosphate starvation responsive genes in Arabidopsis. Plant J 24:559–567
Mayzlish-Gati E, De-Cuyper C, Goormachtig S, Beeckman T, Vuylsteke M, Brewer PB, Beveridge CA, Yermiyahu U, Kaplan Y, Enzer Y, Wininger S, Resnick N, Cohen M, Kapulnik Y, Koltai H (2012) Strigolactones are involved in root response to low phosphate conditions in Arabidopsis. Plant Physiol 160:1329–1341
McSteen P, Leyser O (2005) Shoot branching. Annu Rev Plant Biol 56:353–374
Minakuchi K, Kameoka H, Yasuno N, Umehara M, Luo L, Kobayashi K, Hanada A, Ueno K, Asami T, Yamaguchi S, Kyozuka J (2010) FINE CULM1 (FC1) works downstream of strigolactones to inhibit the outgrowth of axillary buds in rice. Plant Cell Physiol 51:1127–1135
Morris SE, Turnbull CG, Murfet IC, Beveridge CA (2001) Mutational analysis of branching in pea. Evidence that Rms1 and Rms5 regulate the same novel signal. Plant Physiol 126:1205–1213
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Ongaro V, Bainbridge K, Williamson L, Leyser O (2008) Interactions between axillary branches of Arabidopsis. Mol Plant 1:388–400
Pasare SA, Ducreux LJ, Morris WL, Campbell R, Sharma SK, Roumeliotis E, Kohlen W, van der Krol S, Bramley PM, Roberts AG, Fraser PD, Taylor MA (2013) The role of the potato (Solanum tuberosum) CCD8 gene in stolon and tuber development. New Phytol 198:1108–1120
Pérez-Torres CA, López-Bucio J, Cruz-Ramírez A, Ibarra-Laclette E, Dharmasiri S, Estelle M, Herrera-Estrella L (2008) Phosphate availability alters lateral root development in Arabidopsis by modulating auxin sensitivity via a mechanism involving the TIR1 auxin receptor. Plant Cell 20:3258–3272
Prusinkiewicz P, Crawford S, Smith RS, Ljung K, Bennett T, Ongaro V, Leyser O (2009) Control of bud activation by an auxin transport switch. PNAS 106:17431–17436
Reddy SK, Finlayson SA (2014) Phytochrome B promotes branching in Arabidopsis by suppressing auxin signaling. Plant Physiol 164:1542–1550
Reddy SK, Holalu SV, Casal JJ, Finlayson SA (2013) Abscisic acid regulates axillary bud outgrowth responses to the ratio of red to far-red light. Plant Physiol 163:1047–1058
Ruyter-Spira C, Kohlen W, Charnikhova T, van Zeijl A, van Bezouwen L, de Ruijter N, Cardoso C, Lopez-Raez JA, Matusova R, Bours R, Verstappen F, Bouwmeester H (2011) Physiological effects of the synthetic strigolactone analog GR24 on root system architecture in Arabidopsis: another belowground role for strigolactones? Plant Physiol 155:721–734
Schmitz G, Theres K (1999) Genetic control of branching in Arabidopsis and tomato. Curr Opin Plant Biol 2:51–55
Snowden KC, Simkin AJ, Janssen BJ, Templeton KR, Loucas HM, Simons JL, Karunairetnam S, Gleave AP, Clark DG, Klee HJ (2005) The decreased apical dominance1/Petunia hybrida CAROTENOID CLEAVAGE DIOXYGENASE8 gene affects branch production and plays a role in leaf senescence, root growth, and flower development. Plant Cell 17:746–759
Sorefan K, Booker J, Haurogne K, Goussot M, Bainbridge K, Foo E, Chatfield S, Ward S, Beveridge C, Rameau C, Leyser O (2003) MAX4 and RMS1 are orthologous dioxygenase-like genes that regulate shoot branching in Arabidopsis and pea. Genes Dev 17:1469–1474
Stirnberg P, van de Sande K, Leyser HMO (2002) MAX1 and MAX2 control shoot lateral branching in Arabidopsis. Development 129:1131–1141
Tanaka M, Takei K, Kojima M, Sakakibara H, Mori H (2006) Auxin controls local cytokinin biosynthesis in the nodal stem in apical dominance. Plant J 45:1028–1036
Umehara M, Hanada A, Magome H, Takeda-Kamiya N, Yamaguchi S (2010) Contribution of strigolactones to the inhibition of tiller bud outgrowth under phosphate deficiency in rice. Plant Cell Physiol 51:1118–1126
Varagona MJ, Schmidt RJ, Raikhel NV (1992) Nuclear localization signal(s) required for nuclear targeting of the maize regulatory protein opaque-2. Plant Cell 4:1213–1227
Wang Y, Li B, Du M, Eneji AE, Wang B, Duan L, Li Z, Tian X (2012) Mechanism of phytohormone involvement in feedback regulation of cotton leaf senescence induced by potassium deficiency. J Exp Bot 63:5887–5901
Ward SP, Salmon J, Hanley SJ, Karp A, Leyser O (2013) Using Arabidopsis to study shoot branching in biomass willow. Plant Physiol 162:800–811
Waters MT, Brewer PB, Bussell JD, Smith SM, Beveridge CA (2012a) The Arabidopsis ortholog of rice DWARF27 acts upstream of MAX1 in the control of plant development by strigolactones. Plant Physiol 159:1073–1085
Waters MT, Nelson DC, Scaffidi A, Flematti GR, Sun YK, Dixon KW, Smith SM (2012b) Specialisation within the DWARF14 protein family confers distinct responses to karrikins and strigolactones in Arabidopsis. Development 139:1285–1295
Weiler EW, Jourdan PS, Conrad W (1981) Levels of indole-3-acetic acid in intact and decapitated coleoptiles as determined by a specific and highly sensitive solid-phase enzyme immunoassay. Planta 153:561–571
Wen C, Xi L, Gao B, Wang KY, Lv SH, Kou YP, Ma N, Zhao LJ (2015) Roles of DgD14 in regulation of shoot branching in chrysanthemum (Dendranthema grandiflorum ‘Jinba’). Plant Physiol Biochem 96:241–253
Xi L, Wen C, Fang S, Chen X, Nie J, Chu J, Yuan C, Yan C, Ma N, Zhao L (2015) Impacts of strigolactone on shoot branching under phosphate starvation in chrysanthemum (Dendranthema grandiflorum cv. Jinba). Front Plant Sci 6:694
Yao C, Finlayson SA (2015) Abscisic acid is a general negative regulator of Arabidopsis axillary bud growth. Plant Physiol 169:611–626
Yoneyama K, Xie X, Kusumoto D, Sekimoto H, Sugimoto Y, Takeuchi Y, Yoneyama K (2007a) Nitrogen deficiency as well as phosphorus deficiency in sorghum promotes the production and exudation of 5-deoxystrigol, the host recognition signal for arbuscular mycorrhizal fungi and root parasites. Planta 227:125–132
Yoneyama K, Yoneyama K, Takeuchi Y, Sekimoto H (2007b) Phosphorus deficiency in red clover promotes exudation of orobanchol, the signal for mycorrhizal symbionts and germination stimulant for root parasites. Planta 225:1031–1038
Yoneyama K, Xie X, Kim HI, Kisugi T, Nomura T, Sekimoto H, Yokota T, Yoneyama K (2012) How do nitrogen and phosphorus deficiencies affect strigolactone production and exudation? Planta 235:1197–1207
Yoneyama K, Kisugi T, Xie X, Arakawa R, Ezawa T, Nomura T, Yoneyama K (2015) Shoot-derived signals other than auxin are involved in systemic regulation of strigolactone production in roots. Planta 241:687–698
Zhang S, Li G, Fang J, Chen W, Jiang H, Zou J, Liu X, Zhao X, Li X, Chu C, Xie Q, Jiang X, Zhu L (2010) The interactions among DWARF10, auxin and cytokinin underlie lateral bud outgrowth in rice. J Integr Plant Biol 52:626–638
Zhang Y, van Dijk ADJ, Scaffidi A, Flematti GR, Hofmann M, Charnikhova T, Verstappen F, Hepworth J, der Krol S, Leyser O, Smith SM, Zwanenburg B, Al-Babili S, Ruyter-Spira C, Bouwmeester HJ (2014) Rice cytochrome P450 MAX1 homologs catalyze distinct steps in strigolactone biosynthesis. Nat Chem Biol 10:1028–1033
Zhao J, Li G, Yi GX, Wang BM, Deng AX, Nan TG, Li ZH, Li QX (2006) Comparison between conventional indirect competitive enzyme-linked immunosorbent assay (icELISA) and simplified icELISA for small molecules. Anal Chim Acta 571:79–85
Zhou F, Lin Q, Zhu L, Ren Y, Zhou K, Shabek N, Wu F, Mao H, Dong W, Gan L, Ma W, Gao H, Chen J, Yang C, Wang D, Tan J, Zhang X, Guo X, Wang J, Jiang L, Liu X, Chen W, Chu J, Yan C, Ueno K, Ito S, Asami T, Cheng Z, Wang J, Lei C, Zhai H, Wu C, Wang H, Zheng N, Wan J (2013) D14-SCFD3-dependent degradation of D53 regulates strigolactone signalling. Nature 504:406–410
Acknowledgments
We would like to thank Mark T. Waters (University of Western Australia) for providing Atd27-1 mutant seeds. This project was supported by the China 863 Program (2011AA10020801) (http://www.863.gov.cn/) and the 948 Project (2011-G17) of the Ministry of Agriculture of China.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by K. K. Kamo.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wen, C., Zhao, Q., Nie, J. et al. Physiological controls of chrysanthemum DgD27 gene expression in regulation of shoot branching. Plant Cell Rep 35, 1053–1070 (2016). https://doi.org/10.1007/s00299-016-1938-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-016-1938-6