Abstract
Key message
Sixty-three proteins were identified to be differentially accumulated due to iron deficiency in shoot and root. The importance of these proteins alterations on shoot physiology is discussed.
Abstract
Iron (Fe) is an essential micronutrient for plant growth and its accumulation affects the quality of edible plant organs. To investigate the adaptive mechanism of a Chinese rice variety grown under iron deficiency, proteins differentially accumulated in leaves and roots of Yangdao 6, an indica cultivar, under Fe deficiency growth condition, were profiled using a two-dimensional electrophoresis (2-DE) and matrix-assisted laser desorption/ionization time of flight mass spectrometry (MALDI-TOF/MS). The accumulations of seventy-three proteins were detected to be increased or decreased upon iron deficiency, and sixty-three of them were successfully identified. Among the sixty-three proteins, a total of forty proteins were identified in rice leaves, and twenty-three proteins were in roots. Most of these proteins are involved in photosynthesis, C metabolism, oxidative stress, Adenosine triphosphate synthesis, cell growth or signal transduction. The results provide a comprehensive way to understand, at the level of proteins, the adaptive mechanism used by rice shoots and roots under iron deficiency.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Iron is an essential mineral nutrient in plants, functioning in a variety of physiological and metabolic processes, such as chlorophyll synthesis, respiration, redox reactions and electron transfer. Though iron is abundant in the geosphere, most of it is in the forms that are not bioavailable for plants. Thus iron deficiency is a major abiotic stress limiting plant growth rates and crop yield (Chen and Barak 1982; Schmidt 2003). Low accumulation of iron in edible portions of plants also poses negative nutritional problems for humans and animals.
Plants evolved two distinct Fe absorption strategies, e.g. reduction and chelation. Dicotyledons and non-graminaceous monocots utilize the reduction strategy (strategy I). It is comprised of three steps: release of H+ via H+-ATPase to increase the solubility of Fe, reduction of Fe3+ to Fe2+ by ferric reductase oxidase and transport of Fe2+ into roots by Fe-regulated transporter 1 (IRT1) (Colangelo and Guerinot 2004; Dinneny et al. 2008). The strategy II plants, mainly graminaceous species, use chelation, in which the phytosiderophores (PSs), such as mugineic acids (MAs) are released into the rhizosphere to directly chelate Fe3+ (Curie et al. 2001, 2009). Rice uses mixed strategies to take up iron from the soil (Ishimaru et al. 2006).
MA is an important chelator in strategy II plants, and its synthesis in roots and secretion into the rhizosphere are induced under Fe-deficient conditions. MA is synthesized from S-adenosylmethionine (SAM). Nicotianamine synthase (NAS) first catalyses SAM to Nicotianamine (NA), NA is then converted to a 3′-oxo intermediate by Nicotianamine amino-transferase (NAAT), and 2′-deoxymugineic acid is synthesized by the subsequent action of a reductase. Other kinds of MAs may also be formed in this process. In the above biosynthetic pathway, NAS and NAAT are key rate-limiting enzymes. In addition, MA has also been identified in the xylem and phloem where it is suggested to play a role in long distance Fe transport in plants.
In long distance transport, iron needs to be bound by chelating compounds, such as NA or citrate. In the xylem, Fe(III)-citrate is the major form. In the phloem, NA has been suggested to be the major chelator to bind both Fe2+ and Fe3+. In a tomato mutant (Curie et al. 2009; Haydon and Cobbett 2007), loss of NA led to chlorosis, a symptom of iron deficiency (Ling et al. 1999). In the phloem, Fe-NA is the main existing form in which yellow stripe-like (YSL) family members are thought to transport the Fe-NA complex (Curie et al. 2009; Haydon and Cobbett 2007). In rice, OsYSL2 and OsYSL15 are both up-regulated in response to Fe deficiency. They may be involved in the long distance Fe transport from root to shoot to seed, via the phloem (Inoue et al. 2009).
As described above, previous research has revealed some aspects of plants’ adaptation mechanisms to iron deficiency. To map the metabolic changes induced by iron deficiency in a holistic way, 2-DE proteomic profiling techniques have been used to study root proteome changes in different plant species. But all of the studies were conducted in strategy I plant roots. Proteomic studies in Strategy II plants have only been conducted in the root plasma membrane of maize (Hopff et al. 2013). The interaction between shoot and root in most of these studies also had been ignored. In this study, we used two-dimensional gel electrophoresis coupled with MS to analyze differential protein accumulation profiles in rice leaves and roots under normal and deficiency growth conditions. The metabolic protein changes induced by Fe deficiency which we found in rice can be compared to those reported by others in Strategy I plants.
Materials and methods
Plant materials
Rice (Oryza sativa L. spp. Indica var. Yangdao 6) seeds were soaked in water for 72 h in dark to induce germination. Germinated seeds were transferred to plastic containers filled with moisturized quartz sand. After 2 weeks, the seedlings were transferred into 7-L plastic containers filled with half-strength Yoshida’s rice nutrient solution (10 mg L−1 NaH2PO4·2H2O, 40 mg L−1 CaCl2, 40 mg L−1 K2SO4, 40 mg L−1 MgSO4·7H2O, 0.5 mg L−1 MnCl2·4H2O, 0.05 mg L−1 (NH4)6·Mo7O24·2H2O, 0.2 mg L−1 H3BO3, 0.01 mg L−1 ZnSO4·7H2O, 0.01 mg L−1 CuSO4·5H2O, and 2.0 mg L−1 Fe(II)-EDTA and 10 mg L−1 NH4NO3) (Yoshida et al. 1976). After 10 days in this condition, the plants were transferred to either Fe-deficient (0) or Fe-sufficient (2.0 mg L−1) conditions. The pH was adjusted to 5.0 ± 0.2 with HCl. Leaf and root samples were collected 2 weeks after this treatment. The samples were snap frozen in liquid nitrogen and stored at −80 °C.
Measurement of Fe content
Shoots and roots were dried at 105 °C for 30 min and then at 80 °C until a constant weight was achieved. Shoots (500–600 mg) and roots (500–600 mg) were soaked in 10 ml HNO3:HClO4 (3:1, v/v) for 2 h, and then boiled until the liquid became transparent. Fe content was measured by atomic absorption spectrophotometry (TAS-986; Beijing, China).
The experimental results are presented as the mean ± standard deviation (SD) from three independent experiments. The results were subjected to ANOVA and the Tukey test using the statistical package SPSS 16.0. Differences between treatments were tested by the least significant difference (LSD) test at 0.05 probability level.
Protein extraction
Protein was extracted following the traditional method (Ding et al. 2011). Rice samples were thoroughly ground in a mortar with liquid N2 and the protein was extracted in ice-cold acetone that contained 10 % trichloroacetic acid (TCA), 10 mM dithiothreitol (DTT), and 1 mM phenylmethylsulfonyl fluoride (PMSF) incubated at −20 °C for 1 h, followed by centrifugation at 20,000g for 30 min at 4 °C. After being air dried, the pellet was rinsed three times with ice-cold acetone that contained 10 mM DTT and 1 mM PMSF and incubated at −20 °C for 1 h before being centrifuged at 20,000g for 15 min at 4 °C. The pellet was dried under vacuum and resuspended in 200 μL lysis buffer (7 M urea, 2 M thiourea, 4 % 3-[(3-cholamidopropyl) dimethylammonio] propanesulfonic acid (CHAPS), 0.5 % (v/v) immobilized pH gradient (IPG) buffer (pH 4–7 NL, Amersham Biosciences), and distilled water). The mixed sample was incubated at 4 °C overnight, then centrifuged at 40,000g for 15 min at 20 °C, and the supernatant was used for 2-DE analysis. Protein concentrations were determined with the standard Bradford assay, using bovine serum albumin as the standard.
Two-dimensional gel electrophoresis and protein quantification
Isoelectric focusing (IEF) and SDS-PAGE were described previously by Ding, using 100 μg protein per gel (Ding et al. 2011). Gels were stained with silver nitrate (Yan et al. 2000).
Protein visualization, image analysis and quantification followed the method by Zhao et al. (2012). The stained gels were scanned using a VersaDoc4000 image system (Bio-Rad), followed by analysis of the protein spots with PDQUEST 8.0 software (Bio-Rad). For each sample, at least three 2-DE gels were used for analysis. Spots showing significant variation in terms of intensity between the control and other treatments were selected according to the results of paired Student’s t tests (P ≤ 0.05). Protein spots were manually excised from the gels and in-gel digested by trypsin. Gel digestion and protein identification followed Ding et al. (2011). Only significant hits based on MASCOT probability analysis (P < 0.05) were accepted.
Results
Reduction of Fe and chlorophyll content and variation of shoot/root ratio under iron deficiency
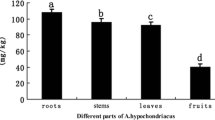
The Fe content of leaves under iron deficiency was 21.4 % less than that in rice under Fe-sufficient growth conditions; that in roots was 50.3 % less (Fig. 1).
Iron is a key component in the biosynthesis of chlorophyll. In this study, Chlorophyll a, chlorophyll b, chlorophyll a + b and carotenoid content were measured (Table 1). Compared to those measured from rice grown under Fe-sufficient condition, they were decreased by 49.1, 50, 47.8 and 36.8 %, respectively. As expected, all the leaves of plants growing under iron deficiency showed chlorosis.
Deficiency of micronutrients can affect the shoot/root ratio in some plant species. Our data showed the shoot-to-root ratio increased from 3.23 under normal condition to 5.15 under a deficiency condition in rice (Table 1).
Changes in protein profiles under iron deficiency
The protein profiles of rice leaves and roots were investigated from iron deficient and sufficient conditions. Approximately one thousand protein spots per gel were consistently detected in a triplicate experiment (Fig. 2). Among them, seventy-three differentially accumulated proteins were analyzed, forty-five and twenty-eight were identified from leaves and roots, respectively (Fig. 3a, b).
Functional classification of identified proteins
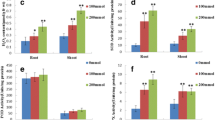
To obtain a comprehensive understanding of the functionalities of these differentially accumulated proteins under iron deficiency, we used BlastP algorithm in NCBI to classify them by the analysis of conserved domains (Altschul et al. 1990). Figure 4a, b showed the differentially abundance changed proteins’ classification in leaves and roots, respectively. Forty out of the forty-five changed proteins were successfully identified in leaves (Table 2). These proteins were mainly involved in carbon metabolism, photosynthesis, stress, energy metabolism, N metabolism, cell growth and signal transduction (Fig. 4a). In roots, twenty-three out of twenty-eight changed proteins were identified (Table 3). These proteins were involved in carbon metabolism, oxidative stress, cell growth, secondary metabolism and protein metabolism (Fig. 4b). When compared to normal Fe condition, the spots showed significant changes in their abundance with mean ratios above 2.0 or below −0.5 under Fe deficiency, and the detailed fold changes were listed in Tables 2 and 3.
Proteins showing changes in rice leaves
C metabolism
Among the forty abundance changed proteins, eight of them were involved in carbon metabolism, including glycolysis, Citric acid pathway (TCA) cycle and the Calvin Benson pathway.
Glycolysis fermentation
In the leaf of rice grown under deficiency, two glycolytic enzymes: phosphoglycerate kinase (spot 1303) and chloroplastic aldolase (spot 3302), their abundance was increased. It might indicate increased efficiency in glycolysis. It was reported that glycolysis can provide the reducing power and ATP needed in the up-regulated Fe acquisition mechanisms (López-Millán et al. 2013).
Under anaerobic conditions glycolysis leads to fermentation and the production of ethanol, thereby continuing ATP production. A degree of anaerobiosis, caused by increased consumption of O2 under iron deficiency (López-Millán et al. 2009) is suggested by our finding of an increase in TASSELSEED2 (spot 7104), a short chain alcohol dehydrogenase (DeLong et al. 1993).
Calvin Benson pathway
The accumulation of Ribulose bisphosphate carboxylase (RuBisCO) (spots 5005, 8105, 8106 and 8202), a key enzyme of the Calvin cycle, was increased in iron-deficient leaves. Among the compounds produced by the Calvin cycle is citric acid, which can be used as a Fe chelator in xylem transport.
Citric acid cycle
Nicotinamide adenine dinucleotide phosphate (NADP) dependent malic enzyme (spot 7704), a key enzyme in the TCA cycle, catalyzing the formation of oxaloacetate from malate, was increased under iron deficiency. This result suggested that the citric acid cycle might be enhanced in response to iron deficiency.
Taxadien-5-alpha-ol O-acetyltransferase (spot 6503), which catalyzes the first acylation step of taxol biosynthesis, was increased compared to that in iron sufficient leaves. During the chemical reaction it catalyzes acetyl-CoA, the substrate of the TCA cycle is produced. Iron starvation may induce more substrates for the TCA cycle.
Nitrogen metabolism
Iron deficiency also has an impact on N metabolism. Four proteins involved in this pathway changed in iron-deficient leaf. Ferredoxin-nitrite reductase (spots 3006, 3010, 7506, 7709, 7714), an enzyme containing Fe, was decreased under iron deficiency. Glutamine synthetase (spot 4307) and putative isoflavone reductase (spot 6203) participating in the assimilation of nitrogen showed an increase. Increased abundance of these two proteins can probably help the plant complement a decrease in N metabolism caused by lower ferredoxin-nitrate reductase.
Methionine is the synthetic precursor of nicotianamine and mugineic acid, chelators that are involved in the solubility and movability of iron (Ling et al. 1996, 1999; Li and Kaplan 2004; Takahashi et al. 2003). In our study, methylthioribulose-1-phosphate dehydratase (spot 6604) catalyzing the final step of Methionine recycling was increased. The increased accumulation of this chelator synthesis-related protein is probably related to maintenance of solubility and supply of Fe under deficiency condition in rice leaves.
Hypothetical protein OsI_38931 (spot 5001), involved in adenine salvage was increased in abundance in iron-deficient leaves. In barley iron-deficient roots, the biosynthesis of MAs is related to adenine salvage in the methionine cycle and activity of adenine phosphoribosyl transferase, which has a possible role in phytosiderophore production (Itai et al. 2000). Thus we speculated that the physiological responses mentioned in barley might also exist in rice grown in deficiency condition.
Stress and defense
Eleven of the abundance changed proteins were involved in stress and/or defense metabolism. These accounted for 28 % of the total accumulations changed proteins in leaves under iron deficiency. This might point to a strong impact of iron deficiency with the stress and defense system.
Glutathione-ascorbate pathway
Ascorbic acid, associated with chloroplasts, plays a role in ameliorating the oxidative stress of photosynthesis. This compound is an electron donor in cell metabolism, playing a pivotal role in development and defense systems, especially in the oxidative-regulated pathway (Zaharieva and Abadia 2003; Kiddle et al. 2003; Noctor and Foyer 1998). GDP-mannose 3, 5-epimerase (spots 6507, 7502), a key enzyme involved in biosynthesis of ascorbate, was increased compared to the control. Ascorbate peroxidase utilizes ascorbic acid as its specific electron donor to reduce H2O2 to water. In our study, ascorbate peroxidase (spot 2107, 4003) was decreased in iron-deficient leaves. This is not surprising, since iron is an indispensable component in the heme reaction domain in peroxidases. Furthermore, monodehydroascorbate reductase (spot 3502, 5503), hydroxyacylglutathione hydrolase (spot 5102) and glutathione S-transferase (spot 4007), playing key roles in ascorbate regeneration, were all increased. All these results which might indicate the alteration of glutathione-ascorbate pathway could be one of the strategies adopted by plants for reductive detoxification in iron-deficient leaves.
Other stress-related proteins
Other stress-induced proteins, i.e. drought-induced S-like ribonuclease (spot 1008) and late embryogenesis abundant protein (LEA; spot 7405), were also increased in iron-deficient rice leaves. LEA-like protein was previously seen to be induced by drought stress, functioning in abiotic stress tolerance by minimizing the negative effects of oxidation (Shinozaki and Yamaguchi-Shinozak 2007; Mowla et al. 2006). In addition to conferring cellular protection upon stress (Chatelain et al. 2012), it has been found that LEA-like protein also participates the long distance transport of iron in phloem (Kruger et al. 2002).
Chitinase 8 (spot 3002) is often produced in higher plants as a general defense response after wounding or pathogenic attack (Witmer et al. 2003). It was increased under iron deficiency. This demonstrated the defense system was induced in rice leaves under iron deficiency.
ATP synthesis
In iron-deficient rice leaves, the abundance of energy-related proteins was increased. ATP synthase in mitochondria resides in two regions: the F0 portion is within the membrane; the F1 portion of the ATP synthase inside the matrix of the mitochondria, above the membrane. The F0 portion (spot 6608) and F1 portion (spot 6710, 6711, 7703), were both increased. This indicated that under iron deficiency in rice leaves, some improvement of energy supply might occur, replacing some of that lost due to lowered photosynthesis.
Signal transduction
Plant 14-3-3 protein (spot 115) is well known for playing an indispensable role in activation of the plasma membrane H+-ATPase. This protein might also be involved in iron mobilization (Xu and Shi 2006). The abundance of this protein was increased under iron deficiency.
DEAD-box ATP-dependent RNA helicase 56 (spot 5602) showed increased abundance under iron deficiency. It is involved in pre-mRNA splicing and required for the export of mRNA out of the nucleus. We note that RNA helicases of the DEAD-box protein family possess an ATPase activity that is dependent on, ATPase is usually induced by iron deficiency (Cordin et al. 2006).
Protein phosphatase 2A (spot 1704), its abundance was increased in iron-deficient leaves. It is a major intracellular protein phosphatase that associated with multiple aspects of cell growth and metabolism, such as DNA replication, transcription, signal transduction and intermediary metabolism. Its activity is also associated with growth inhibition and cell cycle arrest (Schönthal 1998; Mumby and Walter 1993; Hubbard and Cohen 1993; Virshup et al. 1993).
Heme-binding protein (spot 6204, 6704) can bind divalent iron to store iron atoms for physiological utilization in plants. Its abundance was increased in iron-deficient rice leaves. Hemes play a critical role in diverse biological processes in eukaryotic cells, including respiration, protein targeting, transcription and translation regulation, ion channel regulation and signaling, microRNA processing, and protein degradation (Severance and Hamza 2009; Mochizuki et al. 2010).
Protein metabolism
Two proteins induced by iron deficiency were related with protein metabolism. Elongation factor 1 beta’ (spot 109) is the workhorse of protein synthesis on the ribosome. It assists in elongating the newly synthesized polypeptide chain (Andersen et al. 2003). Preglutelin (spot 6102), a primary protein storage protein was also increased under iron deficiency.
Differentially expressed proteins in rice roots
C metabolism
Glycolysis
Glyceraldehyde phosphate dehydrogenase (spot 5704) catalyzes the transition of phosphoglyceraldehyde into 1, 3-bisphosphoglycerate, one of the energy-yielding steps in glycolysis pathway. The abundance of the protein was increased in iron-deficient rice roots. Similar to that in leaves, the glycolysis metabolic pathway might be enhanced in roots in adaptation to iron deficiency.
Pentose phosphate pathway
Pentose phosphate pathway, an alternative pathway to glycolysis, is a process that generates NADPH and pentoses. Ribose-5-phosphate isomerase (spot 3105) and xylulokinase (spot 3610) involved in the pentose phosphate pathway were both increased their accumulations in iron-deficient roots. The products glyceraldenyde-3-phosphate and fructose 6-phosphate in the pentose phosphate pathway can be reused by glycolysis. Thus the up-regulation of the pentose phosphate pathway can facilitate glycolysis.
Calvin Benson pathway
The Calvin cycle is known to be the major pathway for CO2 fixation. Ribulose-1, 5-bisphosphate carboxylase/oxygenase large subunit (spot 3807) is a key CO2 assimilation enzyme of the Calvin cycle (Watson and Tabita 1997). Its increase in iron-deficient roots, which do not perform photosynthese, was a surprise and no function for it can be suggested at present.
Citric acid cycle
Three enzymes, aconitate hydratase (spot 5904), malate dehydrogenase (spot 5316), and NADP dependent malic enzyme (spot 5701, 6706) involved in the TCA cycle, were found to be increased in iron-deficient roots. Furthermore, acetyl-CoA synthetase (spot 4818), whose main function is to convey carbon atoms within the acetyl group to the citric acid cycle to be oxidized for energy production, was also increased in response to iron deficiency. Acetyl-CoA can participate in many other biochemical reactions in addition to the TCA cycle. One of the pathways is to catalyze the formation of fatty acid starting with an esterase. The esterase enzyme (spot 2321) was decreased under iron deficiency. It may indicate that under iron deficiency, acetyl-CoA has priority to enter the TCA cycle, rather than the fatty acid pathway. These results suggest that the TCA cycle efficiency might be enhanced under iron deficiency in rice roots.
Amino acid metabolism
Isopropylmalate synthase (spot 4713) is the key enzyme involved in the synthesis of leucine (Nelson and Cox 2000). In plants and microorganisms, leucine acts as metabolic signal and is involved in amino acid biosynthesis, the assimilation of ammonia, amino acid degradation, nutrient transport, and the formation of pili (Kohlhaw 2003). Its responsive regulatory protein can control glutamate synthase expression. When levels of leucine-responsive regulatory protein are high, glutamate synthase expression is also high (Calvo and Matthews 1994). In our study, α- isopropylmalate synthase showed increased abundance under iron deficiency. We speculate that glutamate might also have been increased, offering more substrates to the MAs synthesis cycle.
Stress and defense
The abundance of ascorbate peroxidase (spot 2001) was decreased due to iron deficiency, similar to the response in leaves.
Under a stress environment, plants will activate their defense systems to remove abnormal and unwanted intracellular proteins. 26S proteasome regulatory particle triple-A ATPase subunit 3 (spot 4520), which was increased in iron deficiency, participates in the stress response by removing abnormal proteins. It also participates in a number of diverse developmental processes, presumably by regulating the levels of one or more short-lived regulators/enzymes. For plants in particular, this pathway has been implicated in cell-cycle progression, photomorphogenesis, hormone responses, leaf, floral and xylem differentiation, and pathogen resistance (Fu et al. 1999).
Secondary metabolism
UDP-glucose is produced by catalysis of UDP-glucose pyrophosphorylase (spot 4507) and sucrose synthase. It was increased under iron deficiency which was consistent with the situation in P-deficient rice roots (Wasaki et al. 2003). Sucrose synthase activity increased more than twice in—P roots of bean (Ciereszko et al. 1998). This may imply more sucrose synthase is generated in rice iron-deficient root to supply the energy needs.
UDP-glucose 6-dehydrogenase (spot 6606) was decreased in our study. This protein exhibited an increase due to heat stress (Xu and Huang 2008) and P deficiency (Wasaki et al. 2003), but the mechanisms are unclear.
Cell growth and development
Typical root morphology changes in response to iron deficiency in plants included an increased density of root hairs, lateral roots, and transfer cells (Ling et al. 2002). Plant root hairs are important organs for the uptake of nutrients and water from the soil. In our study, the protein coded by ROOT HAIR DEFECTIVE 3 (spot 4816) involved in cell and tissue development, showed increased accumulation under iron deficiency. The phylogenic analysis of the gene encoding this enzyme with other homologs and the functional analysis of this gene may shed more light on the mechanism of physiological adaptation in roots during Fe deficiency.
Caffeoyl-CoA 3-O-methyltransferase (spot 1103, 1111) is an enzyme related to the biosynthesis of lignin. Lignin is a major structural component of secondarily thickened plant cell walls. Strong down regulation of caffeoyl-CoA 3-O-methyltransferase can decrease the lignin level in alfalfa (Guo et al. 2001). Therefore, our observed decrease of this protein might indicate iron deficiency disturbing cell generation in roots.
Beta-glucan synthase (spot 5722) catalyzes synthesis of the major structural component of yeast cell walls (Mol et al. 1994). It showed increased abundance in Fe-deficient roots.
Ribose-5-phosphate pyrophosphokinase (spot 7321) catalyzes the transfer of phosphate (phosphoryl or pyrophosphoryl transfer) from ATP to a second substrate. It was also known to be implicated in the synthesis of glycocalyx, suggesting that the formation of glycocalyx might be involved in the root response to iron deficiency (Ouyang et al. 2013).
Protein metabolism
Elongation factors are a set of proteins that are used in protein synthesis in the cell, promoting the GTP-dependent translocation of the nascent protein chain from the A-site to the P-site of the ribosome. In roots, an elongation factor (spot 4304) showed decreased abundance compared to the control.
OSJNBa0039C07.11 (spot 4714) exhibited increased abundance in response to iron deficiency in rice roots. It participates in protein modification related to N metabolism (Suyal et al. 2014).
N-ethylmaleimide–sensitive fusion protein (spot 4817) is a cytosolic ATPase required for many intracellular vesicle fusion reactions. It is a general component of the intracellular fusion machinery (Whiteheart et al. 1994), and its abundance was increased due to Fe deficiency in the roots.
Discussion
In this study, we analyzed the differential accumulation of protein in leaves and roots of rice grown under iron deficiency condition. Sixty-three proteins’ accumulations were observed to be differentially changed, and forty and twenty-three were identified in leaves and roots, respectively. To our knowledge, this is the first proteomic research that incorporated leaves from a plant grown under Fe deficiency conditions in strategy II plants (López-Millán et al. 2013). We classified the identified proteins according to the metabolic pathway they participated. Even though some of the enzymes involved in the pathway are not rate-limiting steps, we assumed their abundance change might be the marker indicating the changed efficiency of the pathway. Missing identification of key enzymes may be due to the low resolution of the 2-DE method.
Both in roots and leaves, some enzymes relevant to energy metabolism (i.e. ATP synthesis, glycolysis and the TCA cycle) were increased (Fig. 5a, b). These up-regulated pathways were in harmony with the proteomic results in Arabidopsis and sugar beet (López-Millán et al. 2000; Thimm et al. 2001). In iron-deficient sugar beet roots, the ATP concentration increased five-fold over the control. A similar situation was found in tomato roots, in which the protein profiling showed increased abundance of mitochondrial ATP synthase under iron deficiency (Li et al. 2008). In cucumber, iron starvation induced activation of metabolism leading to the consumption of stored carbohydrates to produce NAD(P)H, ATP and phosphoenolpyruvate. Activation of catabolic pathways was supported by the enhancement of glycolytic enzymes and concentrations. Another reason for the possible enhancement of ATP levels may be due to accelerated H+-ATPase under iron deficiency; specifically the root plasma membrane one that secretes protons increasing the solubility of soil iron (Santi and Schmidt 2008). In a previous study (Santi and Schmidt 2008), a correlation was found between the rate of ATP formation and the level of H+-ATPase. In iron-deficient rice leaves, the F0 and F1 portion of ATP synthase increased. Plant 14-3-3 protein and ATP-dependent helicase that can activate H+-ATPase were also increased in leaves. The N-ethylmaleimide sensitive fusion protein related to ATPase in roots increased under iron deficiency. It seems probable that, under iron deficiency, roots and shoots may need more energy for uptake and transport of low levels of iron, as well as accomplishing the whole plant’s physiological process.
a Metabolic pathway networks in rice leaves. The proteins in dashed box means it was decreased compared to control, the other means it was increased under iron deficiency. b Metabolic pathway networks in rice roots. The proteins in dashed box means it was decreased compared to control, the other means it was increased under iron deficiency
The usual reduction of chlorophyll content and decrease in photosynthesis rate in rice plants grown under Fe deficiency were observed in this study. However, to our surprise, the accumulation of Rubisco, the marker enzyme for photosynthesis, was increased. This discrepancy between our result and others, in which the enzymatic activity of Rubisco was decreased during Fe deficiency condition in sugar beet (Winder and Nishio 1995) which might be due to the different tissue collection timing points and species specificity. We collected the leaf tissue just 14 days after the induction of deficiency, while in most other studies, the collections of tissues were at period of time in which a severe chlorosis symptom could be observed. Another explanation could be that we measured absolute protein quantity rather than enzymatic activity. It is possible that only part of the Rubisco accumulated during Fe deficiency condition was enzymatically active. Previous studies have shown that Rubisco can also serve as a storage protein in addition to being a key enzyme in photosynthesis (Warren et al. 2004). Future studies on the measurement of real enzyme activity will be especially needed. If much of the accumulated Rubisco protein is only a pool of N storage, it will indicate that the metabolism or export of resources from source leaf to sink tissues is inhibited.
Thirteen stress-related proteins showed changes in response to iron deficiency (Tables 2, 3). Three out of five proteins in the ascorbate cycle were identified to be up-regulated. Ascorbate cycle is a metabolic pathway that can detoxify hydrogen peroxide (H2O2) generated under stress conditions. Previous studies have found differing results on the expressions of ascorbate synthesis enzymes. In our study the accumulation of this enzyme was reduced, consistent with the study on roots of tomato under Fe deficiency (Li et al. 2008). Fe is an indispensable component for the synthesis of the ascorbate enzyme, thus, the lower content of the enzyme may simply be due to the lack of substrate. In leaves, drought-induced proteins were up-regulated. The interaction of Fe deficiency and drought stress physiology is unknown at this stage.
It has been long recognized that some chelators are indispensable for the movement of Fe in both xylem and phloem. In this study, we found that the accumulations of proteins related to MAs synthesis, and TCA cycle, in which malate and citrate can be generated, were increased during deficiency conditions. We also found that the accumulation of an LEA protein was increased in rice grown under Fe deficiency. In addition to being a drought stress-related protein, LEA protein has also be found to be involved in the transport of Fe in phloem. The increased content of this protein may be able to accelerate the mobility of Fe in phloem in rice plants.
Root architecture can have adaptive changes, e.g. formation of new root hairs, under Fe deficiency condition, and this information was mainly obtained from the studies on dicots. The importance of this process in monocots during Fe deficiency condition is largely unknown. In this study, four proteins involved in tissue development were increased under iron deficiency (Table 3), indicating a possibly similar mechanism in monocots as in dicots. We found that the abundance of an elongation factor in leaves was increased, however, in roots the abundance of the protein was decreased, indicating the importance of cell reprogramming on the de novo protein synthesis or turn over in plants grown under stress conditions.
Another interesting characteristic discovered from this study is the increased shoot-to-root ratio caused by the deficiency of Fe. It is well known that the shoot-to-root ratios can be changed under the deficiency of nutrients. However, this has been very rarely studied in plant species under Fe deficiency. The increased ratio suggested that the phloem export of resources, especially carbon, may have been inhibited. But the actual cause is unknown.
Fe and Mg are both important chemicals for cellular metabolism and indispensable components for the structure of chlorophyll. Previous studies have shown that under Mg deficiency condition, the rate of photosynthesis was decreased, phloem loading and transport of carbon were inhibited, and the shoot-to-root ratio was increased (Cakmak and Kirkby 2008). The increased shoot-to-root ratio was caused by decreased export of carbon from shoot tissues. Further work is needed to see if this situation occurs during Fe deficiency in rice. The need for this work illustrates the importance of the incorporation of shoot tissue in studying responses to Fe deficiency in plants.
In conclusion, the analysis of the responses at protein levels in both shoot and root provided a comprehensive and unique way to understand the adaptive mechanisms used by rice grown under Fe deficiency condition. Apparently, rice responded to the lack of Fe at the whole plant level and an integrative strategy was used by the plant to cope up with the stress. Further functional characterization of the identified proteins, especially those involved in the interaction between shoot and root will certainly shed more light on the understanding of the mechanism used by rice grown under Fe deficiency.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Andersen GR, Nissen P, Nyborg J (2003) Elongation factors in protein biosynthesis. Trends Biochem Sci 28:434–441
Cakmak I, Kirkby EA (2008) Role of magnesium in carbon partitioning and alleviating photooxidative damage. Physiol Plant 133:692–704
Calvo JM, Matthews RG (1994) The leucine-responsive regulatory protein, a global regulator of metabolism in Escherichia coli. Microbiol Rev 58:466–490
Chatelain E, Hundertmark M, Leprince O, Le Gall S, Satour P, Deligny-Penninck S, Rogniaux H, Buitink J (2012) Temporal profiling of the heat-stable proteome during late maturation of Medicago truncatula seeds identifies a restricted subset of late embryogenesis abundant proteins associated with longevity. Plant Cell Environment 35:1440–1455
Chen Y, Barak P (1982) Iron nutrition of plants in calcareous soils. In: Brady NC (ed) Advance in agronomy. Rehovot, Israel, pp 217–240
Ciereszko I, Zambrzycka A, Rychter A (1998) Sucrose hydrolysis in bean roots (Phaseolus vulgaris L.) under phosphate deficiency. Plant Sci 133:139–144
Colangelo EP, Guerinot ML (2004) The essential bHLH protein FIT1 is required for the iron deficiency response. Plant Cell 16:3400–3412
Cordin O, Banroques J, Tanner NK, Linder P (2006) The DEAD-box protein family of RNA helicases. Gene 367:17–37
Curie C, Panaviene Z, Loulergue C, Dellaporta SL, Briat JF, Walker EL (2001) Maize yellow stripe1 encodes a membrane protein directly involved in Fe(III) uptake. Nature 409:346–349
Curie C, Cassin G, Counch D, Divol F, Higuchi K, Jean ML, Misson J, Schikora A, Czernic P, Mari S (2009) Metal movement within the plant: contribution of nicotianamine and yellow stripe 1-like transporters. Ann Bot 103:1–11
DeLong A, Calderon-Urrea A, Dellaporta SL (1993) Sex determination gene TASSELSEED2 of maize encodes a short-chain alcohol dehydrogenase required for stage-specific floral organ abortion. Cell 74:757–768
Ding CQ, You J, Liu ZH, Rehmani MIA, Wang SH, Li GH, Wang QS, Ding YF (2011) Proteomic analysis of low nitrogen stress-responsive proteins in roots of rice. Plant Mol Biol Rep 29:618–625
Dinneny JR, Long TA, Wang JY, Jung JW, Mace D, Pointer S, Barron C, Brady SM, Schiefelbein J, Benfey PN (2008) Cell identity mediates the response of Arabidopsis roots to abiotic stress. Science 320:942–945
Fu HY, Doelling JH, Rubin DM, Vierstra RD (1999) Structural and functional analysis of the six regulatory particle triple-A ATPase subunits from the Arabidopsis 26S proteasome. Plant J 18:529–539
Guo D, Chen F, Inoue K, Blount JW, Dixon RA (2001) Downregulation of caffeic acid 3-O-methyltransferase and caffeoyl CoA 3-O-methyltransferase in transgenic alfalfa. impacts on lignin structure and implications for the biosynthesis of G and S lignin. Plant Cell 13:73–88
Haydon MJ, Cobbett CS (2007) Transporters of ligands for essential metal ions in plants. New Phytol 174:499–506
Hopff D, Wienkoop S, Lüthje S (2013) The plasma membrane proteome of maize roots grown under low and high iron conditions. J Proteomics. doi:10.1016/j.jprot.2013.01.006
Hubbard MJ, Cohen P (1993) On target with a new mechanism for the regulation of protein phosphorylation. Trends Biochem Sci 18:172–177
Inoue H, Kobayashi T, Nozoye T, Takahashi M, Kakei Y, Suzuki K, Nakazono M, Nakanishi H, Mori S, Nishizawa NK (2009) Rice OsYSL15 is an Fe-regulated Fe(III)-deoxymugineic acid transporter expressed in the roots and is essential for Fe uptake in early growth of the seedlings. J Biol Chem 284:3470–3479
Ishimaru Y, Suzuki M, Tsukamoto T, Suzuki K, Nakazono M, Kobayashi T, Wada Y, Watanabe S, Matsuhashi S, Takahashi M, Nakanishi H, Mori S, Nishizawa NK (2006) Rice plants take up Fe as an Fe3+-phytosiderophore and as Fe2+. Plant J 45:335–346
Itai R, Suzuki K, Yamaguchi H, Nakanishi H, Nishizawa NK, Yoshimura E, Mori S (2000) Induced activity of adenine phosphoribosyltransferase (APRT) in iron-deficient barley roots: a possible role for phytosiderophore production. J Exp Bot 51:1179–1188
Kiddle G, Pastori GM, Bernard S, Pignocchi C, Antoniw J, Verrier PJ, Foyer CH (2003) Effects of leaf ascorbate content on defense and photosynthesis gene expression in Arabidopsis thaliana. Antioxid Redox Signal 5:23–32
Kohlhaw GB (2003) Leucine biosynthesis in fungi: entering metabolism through the back door. Microbiol Mol Biol Rev 67:1–15
Kruger C, Berkowitz O, Stephan UW, Hell R (2002) A metal-binding member of the late embryogenesis abundant protein family transports iron in the phloem of Ricinus communis L. J Biol Chem 227:25062–25069
Li L, Kaplan J (2004) A mitochondrial vacuolar signaling pathway in yeast that affects iron and copper metabolism. J Biol Chem 279:33653–33661
Li J, Wu XD, Hao ST, Wang XJ, Ling HQ (2008) Proteomic response to iron deficiency in tomato root. Proteomics 8:2299–2311
Ling HQ, Pich A, Scholz G, Ganal MW (1996) Genetic analysis of two tomato mutants affected in the regulation of iron metabolism. Mol Gen Genet 252:87–92
Ling HQ, Koch G, Baumlein H, Ganal MW (1999) Map-based cloning of chloronerva, a gene involved in iron uptake of higher plants encoding nicotianamine synthase. Proc Natl Acad Sci 96:7098–7103
Ling HQ, Bauer P, Bereczky Z, Keller B, Ganal M (2002) The tomato fer gene encoding a bHLH protein controls iron-uptake responses in roots. Proc Natl Acad Sci 99:13938–13943
López-Millán AF, Morales F, Andaluz S, Gogorcena Y, Abadía A, De Rivas Las J, Abadía J (2000) Responses of sugar beet roots to iron deficiency. Changes in carbon assimilation and oxygen use. Plant Physiol 124:885–897
López-Millán AF, Morales F, Gogorcena Y, Abadía A, Abadía J (2009) Metabolic responses in iron deficient tomato plants. J Plant Physiol 166:375–384
López-Millán AF, Grusak MA, Abadía A, Abadía J (2013) Iron deficiency in plants: an insight from proteomic approaches. Front Plant Sci 4:254
Mochizuki N, Tanaka R, Grimm B, Moulin M, Masuda T, Smith AG, Tanaka A, Terry MJ (2010) The cell biology of tetrapyrroles: a life and death struggle. Trends Plant Sci 15:488–498
Mol PC, Park HM, Mullins JT, Cabib E (1994) A GTP-binding protein regulates the activity of (1→3)-beta-glucan synthase, an enzyme directly involved in yeast cell wall morphogenesis. J Biol Chem 269:31267–31274
Mowla SB, Cuypers A, Driscoll SP, Kiddle G, Thomson J, Foyer CH, Theodoulou FL (2006) Yeast complementation reveals a role for an Arabidopsis thaliana late embryogenesis abundant (LEA)-like protein in oxidative stress tolerance. Plant J 48:743–756
Mumby MC, Walter G (1993) Protein serine/threonine phosphatases: structure, regulation, and functions in cell growth. Physiol Rev 73:673–699
Nelson DL, Cox MM (2000) Lehninger. Principles of Biochemistry, New York
Noctor G, Foyer CH (1998) Ascorbate and glutathione: keeping active oxygen under control. Annu Rev Plant Physiol Plant Mol Biol 49:249–279
Ouyang JP, Guo WB, Li B, Gu L, Zhang HJ, Chen XH (2013) Proteomic analysis of differential protein expression in Acidithiobacillus ferrooxidans cultivated in high potassium concentration. Microbiol Res 168:455–460
Santi S, Schmidt W (2008) Laser microdissection-assisted analysis of the functional fate of iron deficiency-induced root hairs in cucumber. J Exp Bot 59:697–704
Schmidt W (2003) Fe solutions: acquisition strategies and signaling pathway in plants. Trends Plant Sci 8:188–193
Schönthal AH (1998) Role of PP2A in intracellular signal transduction pathways. Front Biosci 3:1262–1273
Severance S, Hamza I (2009) Trafficking of heme and porphyrins in metazoa. Chem Rev 109:4596–4616
Shinozaki K, Yamaguchi-Shinozak K (2007) Gene networks involved in drought stress response and tolerance. J Exp Bot 58:221–227
Suyal DC, Yadav A, Shouche Y, Goel R (2014) Differential proteomics in response to low temperature diazotrophy of Himalayan psychrophilic nitrogen fixing Pseudomonas migulae S10724 strain. Curr Microbiol 68:543–550
Takahashi M, Terada Y, Nakai I, Nakanishi H, Yoshimura E, Mori S, Nishizawa NK (2003) Role of nicotianamine in the intracellular delivery of metals and plant reproductive development. Plant Cell 15:1263–1280
Thimm O, Essigmann B, Kloska S, Altmann T, Buckhout TJ (2001) Response of Arabidopsis to iron deficiency stress as revealed by microarray analysis. Plant Physiol 127:1030–1043
Virshup DM, Cegielska A, Russo A, Kelly TJ, Shaffer S (1993) The initiation of SV40 DNA replication is controlled by a phosphorylation-dephosphorylation cycle. Adv Protein Phosphatases 7:271–293
Warren CR, Livingston NJ, Turpin DH (2004) Photosynthetic responses and N allocation in Douglas-fir needles following a brief pulse of nutrients. Tree Physiol 24:601–608
Wasaki J, Yonetani R, Kuroda S, Shinano T, Yazaki J et al (2003) Transcriptomic analysis of metabolic changes by phosphorus stress in rice plant roots. Plant Cell Environ 26:1515–1523
Watson GMF, Tabita FR (1997) Microbial ribulose 1, 5-bisphosphate carboxylase/oxygenase: a molecule for phylogenetic and enzymological investigation. FEMS Microbiol Lett 146:13–22
Whiteheart SW, Rossnagel K, Buhrow SA, Brunner M, Jaenicke R, Rothman JE (1994) N-ethylmaleimide-sensitive fusion protein: a trimeric ATPase whose hydrolysis of ATP is required for membrane fusion. J Cell Biol 126:945–954
Winder TL, Nishio JN (1995) Early iron deficiency stress response in leaves of sugar beet. Plant Physiol 108:1487–1494
Witmer X, Nonogaki H, Beers EP, Bradford KJ, Welbaum GE (2003) Characterization of chitinase activity and gene expression in muskmelon seeds. Seed Science Research 13:167–178
Xu CP, Huang BR (2008) Root proteomic responses to heat stress in two Agrostis grass species contrasting in heat tolerance. J Exp Bot 59:4183–4194
Xu WF, Shi WM (2006) Expression Profiling of the 14-3-3 gene family in response to salt stress and potassium and iron Deficiencies in young tomato (Solanum lycopersicum) roots: analysis by real-time RT-PCR. Ann Bot 98:965–974
Yan JX, Wait R, Berkelman T, Harry RA, Westbrook JA, Wheeler CH, Dunn MJ (2000) A modified silver staining protocol for visualization of proteins compatible with matrix-assisted laser desorption/ionization and electrospray ionization-mass spectrometry. Electrophoresis 21:3666–3672
Yoshida S, Forno DA, Cock JH, Gomez KA (1976) Laboratory manual for physiological studies of rice. Los Baños
Zaharieva TB, Abadia J (2003) Iron deficiency enhances the levels of ascorbate, glutathione, and related enzymes in sugar beet roots. Protoplasma 221:269–275
Zhao XF, Ding CQ, Chen L, Wang SH, Wang QS, Ding YF (2012) Comparative proteomic analysis of the effects of nitric oxide on alleviating Cd-induced toxicity in rice (Oryza sativa L.). Plant Omics J 5:604–614
Acknowledgments
This work was supported by the Ministry of National Science and Technology of China (Project No. 2012BAD04B00) and the Science and Technology Department of Henan Province (Project No. 2013BAD07B00). We thank Professor Andre Jagendorf for the language editing.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Wendy Harwood.
Rights and permissions
About this article
Cite this article
Chen, L., Ding, C., Zhao, X. et al. Differential regulation of proteins in rice (Oryza sativa L.) under iron deficiency. Plant Cell Rep 34, 83–96 (2015). https://doi.org/10.1007/s00299-014-1689-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-014-1689-1