Abstract
Key message
A complete set of monosomic alien addition lines of Brassica napus with one of the seven chromosomes of Isatis indigotica and the recombinant mitochondria was developed and characterized.
Abstract
Monosomic alien addition lines (MAALs) are valuable for elucidating the genome structure and transferring the useful genes and traits in plant breeding. Isatis indigotica (Chinese woad, 2n = 14, II) in Isatideae tribe of Brassicaceae family has been widely cultivated as a medicinal and dye plant in China. Herein, the intertribal somatic hybrid (2n = 52, AACCII) between B. napus cv. Huashuang 3 (2n = 38, AACC) and I. indigotica produced previously was backcrossed recurrently to parental B. napus, and 32 MAAL plants were isolated. Based on their phenotype, 5S and 45S rDNA loci and chromosome-specific SSR markers, these MAALs were classified into seven groups corresponding to potential seven types of MAALs carrying one of the seven I. indigotica chromosomes. One of the MAALs could be distinguishable by expressing the brown anthers of I. indigotica, other two hosted the chromosome with 5S or 45S rDNA locus, but the remaining four were identifiable by SSR markers. The simultaneous detection of the same SSR maker and gene locus in different MAALs revealed the paralogs on the chromosomes involved. The recombinant mitochondrial genome in MAALs was likely related with their male sterility with carpellody stamens, while the MAAL with normal brown anthers probably carried the restoring gene for the male sterility. The complete set of MAALs should be useful for exploiting the I. indigotica genome and for promoting the introgression of valuable genes to B. napus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Monosomic alien addition lines (MAALs) are plants with an alien chromosome from one donor species added to the genome of another recipient species. A complete set of MAALs which dissects the donor genome can be used for chromosomal assignment of genes and markers (Geleta et al. 2012), constructing physical maps of specific chromosomes (Kynast et al. 2004), examining alien genes expression (Cho et al. 2006) and studying syntenic correspondence among different species (Tan et al. 2005). Furthermore, MAALs provide a novel platform for studying the chromosome homoeology among different genomes and alien chromosome behavior in recipient genetic background (Wang et al. 2005; Heneen et al. 2012; McArthur et al. 2012). In crop breeding, the prime motive for producing alien addition lines is to introgress genes of interest from wild relatives into cultivated crops, and the introgressions with alien chromosome segments and target traits can be induced by homoeologous recombination, radiation or the gametocidal effect (Klindworth et al. 2012).
The common method for developing MAALs was recurrent backcrossing of the natural or synthesized amphidiploids to one parental species, and then addition lines were isolated from the resulting aneuploid progenies. MAALs can be identified among aneuploidy plants by morphological traits, chromosome-specific cytological and molecular markers, biochemical markers, and the method of molecular cytogenetics. In the family Brassicaceae, the genomes of the three cultivated Brassica diploids and the other related species have been dissected by the development of MAALs through interspecific, or intergeneric and even intertribal sexual and somatic hybridizations (see review by Prakash et al. 2009). These MAALs have enhanced the location of the useful traits and genes on particular chromosomes, such as disease resistance, important agronomic traits and fertility restoring gene for CMS (Chen et al. 1992; Cheng et al. 1994; Peterka et al. 2004; Wang et al. 2003; Wei et al. 2010; Heneen et al. 2012; Ding et al. 2013), and the investigation of chromosome homoeology (Geleta et al. 2012). However, the establishment of the whole set of MAALs for Brassica species and relatives is difficult, because of the small size of the chromosomes and lack of morphological and cytological landmarks (Budahn et al. 2008; Geleta et al. 2012; Heneen et al. 2012).
Isatis indigotica Fort. (Chinese woad) which belongs to the Isatideae tribe of the Brassicaceae family (Al-Shehbaz et al. 2006) has been widely cultivated as a medicinal and dye plant from ancient times in China, but now it is commonly used for medicinal purpose. Its root (Radix isatidis) is now the second most consumed source of plant medicine, after radix ginseng, but ginseng is mainly used as tonic. The products from Radix isatidis are usually prescribed for the indications of eruptive epidemic diseases caused by the bacteria and viruses (influenza, viral pneumonia, mumps, and hepatitis), to remove heat and eliminate toxin, to reduce heat in blood, and to soothe the sore throat. The medicine is also considered to have immune regulatory and even antitumor effects (Liu et al. 2000; Chung et al. 2011; Du et al. 2013). Some studies showed that this plant had resistance to tobacco mosaic virus (TMV) (Wang and Wang 1988), the fungus Sclerotinia sclerotiorum causing stem-rot in rapeseed (Zhao et al. 1994), and even to diamondback moth (Plutella xylostella) (Tang et al. 2010). However, the functional components of this plant with great consumption are still unclear, which largely suffers from the scarce information for its genetics and genome. In our previous study aiming to produce new germplasm for the genetic improvement of Brassica crops and to dissect the genome of I. indigotica, the intertribal somatic hybrids between Brassica napus L. (2n = 4x = 38, genomes AACC) and I. indigotica (2n = 2x = 14, II) were obtained by protoplast fusion and characterized for their phenotype and cytology (Du et al. 2009). Importantly, the hybrids had the expected sum of chromosomes from the two parents (2n = 6x = 52, AACCII) and gave rise to backcross progenies (BC1, 2n = 5x = 45, AACCI) after pollination by B. napus. In this study to continue the genetic analysis of the hybrid progenies, the complete set of B. napus monosomic additions carrying one of the seven different I. indigotica chromosomes (2n = 39, AACC + 1I1–7) is established by characterizing backcrossing progeny plants for their phenotype, cytological and molecular markers. This set of MAALs is pivotal for elucidating the genome structure of this important medicinal plant and for the introduction of useful genes and traits into Brassica crop. Specifically one new cytoplasmic male sterile (CMS) line with carpellody stamens is selected for the hybrid breeding of B. napus.
Materials and methods
Plant materials
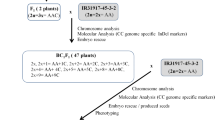
The intertribal somatic hybrid plant (As1) (2n = 6x = 52, AACCII) between Brassica napus L. cv. Huashuang 3 (2n = 4x = 38, AACC) and Isatis indigotica Fort. (2n = 2x = 14, II) was previously produced by the fusion of mesophyll protoplasts, and its BC1 plants after pollinated by Huashuang 3 had the expected chromosome complement (2n = 5x = 45, AACCI) (Du et al. 2009). Herein, these BC1 plants with the Isatis genome in haploidy state were again backcrossed to Huashuang 3 to produce BC2 plants which were derived from the embryos rescued on MS medium (Murashige and Skoog 1962). These BC2 plants with one to several chromosomes from I. indigotica were pollinated by Huashuang 3 to produce the BC3 population in which monosomic addition plants with individual chromosome of I. indigotica were identified (Fig. 1). The plants with 2n = 39 were maintained by culturing their axillary buds from main stems or branches on MS medium for further study.
Cytology and pollen viability analysis
To determine the chromosome numbers of backcross progenies, the ovaries from young flower buds were collected in the morning. After pre-treatment with 2 mM 8-hydroxyquinoline solution for 3 h at room temperature, the ovaries were fixed in Carnoy’s solution (3:1 ethanol:glacial acetic acid, v/v) overnight, and stored in 70 % ethanol at −20 °C until use. Cytogenetic observations were carried out as described by Li et al. (1995). Pollen fertility was determined as the percentage of pollen grains stained with 1 % acetocarmine.
Probe labeling and GISH/FISH analysis
Total genomic DNA was extracted and purified from young leaves according to Dellaporta et al. (1983). Genomic DNA of I. indigotica was labeled with Bio-11-dUTP (Fermentas) or digoxigenin-11-dUTP (Roche) by nick translation. The 5S rDNA and 45S rDNA isolated from tomato provided by Prof. Lijia Li, College of Biology, Wuhan University, China, were labeled with Bio-11-dUTP and digoxigenin-11-dUTP, respectively, using nick translation method. The genomic DNA of B. napus was sheared by boiling for 15 min to produce DNA fragments of 100–500 bp and used as blocks. Chromosome preparations for GISH were made according to the protocol (Ge et al. 2009). The ovaries were digested in an enzyme mixture containing 4 % cellulose Onozuka RS (Yakult, Japan), 2 % pectinase (Merck, Germany) and 2 % pectinase (Sigma-Aldrich, USA) for about 60 min at 37 °C. In situ hybridization was performed following the procedures of Cui et al. (2012). Photographs were taken using a computer-assisted fluorescence microscope (Axio Scope A1, Zeiss, Germany) with a CCD camera. Images were processed by Adobe Photoshop CS5 to adjust contrast and brightness.
SSR marker analysis
Primer sequences for I. indigotica SSR markers were kindly provided by Prof. Luqi Huang from State Key Laboratory of Dao-di Herbs, National Resource Center of Chinese Materia Medica, China Academy of Chinese Medical Sciences, Beijing, China. Of 3054 SSRs, 192 primer pairs were randomly synthesized (Table S1). These markers included 97 dinucleotide, 83 trinucleotide and 12 tetranucleotide repeats. All these 192 SSR markers were evaluated for successful PCR amplification and polymorphism by testing the genomic DNA of rapeseed and woad. PCR amplifications were performed in a volume of 10 µl containing 20 ng genomic DNA, 1 × Taq buffer, 2 mM MgCl2, 0.2 U Taq DNA polymerase (Fermentas), 2 mM dNTPs (Fermentas), 5 µM forward and reverse primer. PCR reaction program was the following: 94 °C for 5 min; 10 cycles with 94 °C for 30 s, 60 °C for 30 s, 72 °C for 45 s, with a 0.7 °C decrease in annealing temperature at each cycle; 30 cycles with 94 °C for 30 s, 55 °C for 30 s, 72 °C for 45 s, and a final extension at 72 °C for 10 min. Amplification products were separated on 6 % denaturing polyacrylamide gels. After electrophoresis, the gels were stained as previously described (Peterka et al. 2004) and photographs taken.
I. indigotica gene markers
Sequences of seven I. indigotica genes were downloaded from GenBank. These genes included IiCCoAOMT (GenBank: DQ115904), IiCDPK (GenBank: DQ482580), IiCOMT (GenBank: DQ115905), IiLTK (GenBank: DQ468350), IiMYB (GenBank: DQ468346), IiPAL (GenBank: DQ468345) and IiSDD1 (GenBank: DQ407741). PCR primers were designed by Oligo 7 software (Table S2). Reactions (10 µl) contained 1 × Taq buffer, 2 mM MgCl2, 5 mM dNTPs, 5 µM forward and reverse primer, 1 U Taq DNA polymerase and 50 ng genomic DNA. DNA fragments were amplified after a 5-min denaturation at 94 °C for 30 cycles (94 °C for 45 s, 57 °C for 30 s, 72 °C for 90 s), and a 10-min extension step at 72 °C. PCR products were separated on 0.8 % agarose gels.
Mitochondrial DNA analysis
To determine the mitochondrial genome, 21 pair primers of mitochondrial genes were designed based on the rapeseed mtDNA sequence (accession number: AP006444). Primer sequences are listed in Table S3. PCR was performed in a 10-μl reaction mixture containing 100 ng total plant DNA, 1 × Taq buffer, 2 mM MgCl2, 1 U Taq DNA polymerase, 5 mM dNTPs, 5 μM forward and reverse primer. PCR amplification was carried out with an initial denaturation step at 94 °C for 5 min followed by 30 cycles of 94 °C for 1 min, 54 °C for 1 min, 72 °C for 2 min, and a final 10-min extension at 72 °C.
Results
Development of MAALs
After BC1 plants (2n = 5x = 45, AACCI) derived from the intertribal somatic hybrids between B. napus cv. Huashuang 3 and I. indigotica were backcrossed again to Huashuang 3 as pollen parent, 22 BC2 plants were successfully obtained with the rescue of immature embryos on MS medium (Fig. 1). They had 2n = 38–43, besides one with 2n = 21, only one plant had 2n = 38 origination from B. napus, and two were monosomic additions (2n = 39), while the remaining 19 plants still kept 2–5 additional woad chromosomes (Table 1). Because of the poor male fertility of these BC2 plants, those with 2n > 38 were pollinated by Huashuang 3 to obtain BC3 progenies, and 113 plants were selected by their phenotype for the determination of chromosome numbers. Out of 113 BC3 plants, 25 ones had 2n = 39 and 18 had 2n = 40–42, while most (68) had 2n = 38, and 2 had 2n < 38. Then, 16 plants with 2n = 39 were successfully maintained clonally on MS medium by culturing their axillary buds available. In the similar way, 31 BC4 plants with 2n = 39 were identified and 25 were cloned on MS medium (Table 1). Totally, 41 plants presumably with 2n = 39 were successfully cloned to produce enough plants for further investigations. By GISH analysis of these 41 plants, 32 were confirmed to be monosomic additions with 38 chromosomes from B. napus and one from I. indigotica (Fig. 2b), but the others were actually euploid B. napus (2n = 38). In other side, several plants with 2n < 38 were found but no further study was made. One plant (2n = 20 with one alien chromosome) appeared, which likely derived from the unfertilized egg. Two plants (2n = 40) had one pair of alien woad chromosomes related to the chromosome d of I. indigotica (below).
GISH and FISH analysis of MAALs. Blue color is from the DAPI staining of chromosomes. a One root-tip cell of I. indigotica probed with the labeled 5S rDNA (green) and 45S rDNA (red). b An ovary cell (2n = 39) of one MAAL line with one labeled chromosome from I. indigotica (red). c1, c2 One ovary cell (c1) of Mf with one chromosome from I. indigotica (red) which carries 5S rDNA locus (green) (c2). Several chromosomes from B. napus also carry 5S rDNA loci. d1, d2 One ovary cell (d1) of Mg with an alien chromosome (green) which hosts 45S rDNA (red) (d2). The red signal from 45S rDNA also appears on some chromosomes from B. napus. Arrows indicate alien chromosomes. Bars 10 µm (color figure online)
The challenging task next was to classify these 32 monosomic additions into seven potential groups which likely hosted one of the seven different chromosomes from I. indigotica, according to their phenotype, chromosome-specific cytological and DNA markers (Figs. 2, 3, 4, 5; Table 2). The seven types of MAALs (designated as Ma–Mg) included 2, 2, 3, 6, 5, 13, 1 plants, respectively, Mf included much more plants (13), and Mg corresponded to only one plant (Table 2).
Cytological markers of MAALs
The chromosomes of I. indigotica had the small and similar size, and their size was even smaller than those of B. napus. Fluorescence in situ hybridization (FISH) analysis with the labeled 45S rDNA and 5S rDNA as probes identified a single locus of 5S rDNA and 45S rDNA separated on different chromosomes in I. indigotica (Fig. 2a). The locus of 5S rDNA was located near centromeres on the long arms of two chromosomes, and the locus of 45S rDNA was at the terminal parts of the short arms of other two chromosomes, which provided the chromosomal markers for the identification of two MAALs. Consequently, Mf and Mg were found to carry the alien chromosome with 5S rDNA and 45S rDNA loci, respectively (Fig. 2c, d; Table 2). Other MAALs were undistinguishable cytologically.
SSR markers of MAALs
Among 192 SSR markers from I. indigotica, 144 pairs produced the genome specific fragments, 47 pairs showed polymorphism between two parental species and one pair (ssr18) had no polymorphism (data not shown). With these woad-specific polymorphic markers to classify MAALs, most markers segregated for their presence or absence (Fig. 3), but some primers lost bands in backcross progenies or had no polymorphism between additions. According to their distribution in MAALs, these markers were divided into seven groups (a–g, Fig. 3), which was in agreement with the chromosome number of woad genome (n = 7). The number of markers in group c (19) was fewer than the other groups (23–32) (Table 2). Markers in each group were specific to one of the seven woad chromosomes (Table 2).
Five SSR markers specific to more than one chromosome of I. indigotica were also identified (Table 3). The number of duplicated SSR markers which were shared by more than one chromosome was 2, 0, 4, 3, 2, 2 and 2 for the chromosomes a–g, respectively. So no such markers appeared on chromosome b. The markers 9b, 61b and 149 were detected in two chromosomes, whereas 9a, 59b and 90 in three chromosomes. Chromosome a shared no markers with b, e and f. Chromosome c shared no markers with b. Chromosome d shared no markers with b and e. Chromosome e only shared markers with c. Chromosome f shared markers with c, d and g. Chromosome g shared markers with c, d and f. Chromosome c shared two markers with e, and similarly d and f shared two markers, suggesting a significant level of homology between c and e and between d and f.
Phenotype of MAALs
The young plants of MAALs showed a morphology biased to B. napus with some variations, but expressed some traits of the woad origin, mainly the darker green and thicker leaves. The margin of leaves was less serrated than B. napus, probably because the leaves of I. indigotica had no serrations. They grew slowly and also flowered later than B. napus. The different plants of the same MAAL line showed nearly the same phenotype. The line Ma showed weaker growth than other additions. As to flower organs, only the line Me had normal flowers with the brown anthers as I. indigotica did (Fig. 4c), and could be distinguished from the others by this trait. But other lines showed the disturbed stamen development and wrinkled petals (Fig. 4e1, f1). At the beginning of flowering, the stamens were weakly developed and some fused together (Fig. 4e1, e2), whereas the flowers of Mb did not open (Fig. 4d). The tetradynamous stamens developed into carpelloid structures and the two shorter stamens were only filaments without anthers (Fig. 4f1, f2). These feminized stamens contained stigmatoid structures at their tips and two from same side usually fused together. Sometimes, the stamens deviated to petal-like organs in few flowers (Fig. 4g). In general, these lines gave low seed sets by self-pollination, but much higher seeding rates after pollination by B. napus, except Ma with fewer seeds in each pod (Table 4).
Location of I. indigotica genes in MAALs
PCR results showed that the markers IiMYB and IiCDPK were specific to Ma (Fig. 5a; Table 2), IiSDDR1 was related to Mb (Fig. 5b). The IiPAL was amplified in four MAALs (Mc, Md, Me and Mf), respectively, showing that the gene had four homologous copies in I. indigotica (Fig. 5c; Table 2). IiCOMT was lost in BC1 plants and all MAALS, probably because of a deletion of the genomic region containing this gene in the haploidy woad genome of the respective BC1 plant (AACCI) used for backcrossing with B. napus. The marker IiLTK and IiCCoAoMT did not show polymorphism between I. indigotica and B. napus.
Recombinant mitochondrial genome in MAALs
Using a primer pair of cox2-2, one fragment of approximately 2,300 bp specific to B. napus was detected in F1, BC1 and MAALs (Fig. 6a). The cox1 in F1 and backcross progenies was the same as B. napus, but the orf 261 specific to B. napus was lost in F1 and backcross progenies. As to rps3, a larger and weaker band was amplified in B. napus by the primer pair designed based on the rapeseed mtDNA sequence, but the smaller and brighter bands in F1, BC1 and all MAALs which were the same as the one in I. indigotica (Fig. 6b). This suggested not only the sequence differences at this locus between B. napus and I. indigotica, but also the genetic recombination between the loci of these two species. No polymorphisms between B. napus and I. indigotica were detected for other 17 mitochondrial genes. These results revealed that the hybrid maintained the recombinant mitochondrial genome of two parents and was maternally transmitted to MAALs.
Discussion
As the intertribal sexual crosses between B. napus and I. indigotica only produced the unclassical hybrids with the I. indigotica chromosomes largely eliminated (Tu et al. 2010), their somatic hybridizations were made and the hybrids with the sum of the parental chromosome complements and particularly the partial fertility were successfully obtained (Du et al. 2009), which makes it feasible to dissect the genome of the important medicinal plant by developing the complete set of MAALs in this study. Using the characteristics in phenotype, chromosomes and DNA sequences specific for I. indigotica, the large number of monosomic addition plants derived from the backcrossing progenies of several generations was put into seven groups hosting one of the seven I. indigotica chromosomes in the background of B. napus (Table 2).
The distinct phenotypic traits from the donor parent are most easily recognizable for the identification of MAALs, as shown by the intergeneric or even intertribal additions of B. napus with individual chromosomes from Orychophragmus violaceus (Ding et al. 2013). But I. indigotica has few phenotypic characters which are expressed and specific for particular MAALs, except for the brown color of anthers, though it belongs to different tribe. Cytologically, the rDNA loci (5S and 18S-5.8S-25S rDNA) and their chromosomal localization have been shown to be suitable for the karyotypic characterization of species with small and similarly sized chromosomes, and thus for the recognition of certain MAALs carrying such chromosomes (Table 2; Fig. 2c, d). The 5S rDNA loci are usually localized within pericentromeric heterochromatic regions in Brassica species (Lim et al. 2005; Koo et al. 2011) and I. indigotica (this study), and most 45S rDNA occupied terminal positions in Brassicaceae species (Ali et al. 2005), but some adjacent to terminal positions (Yang and Li 2011; this study), or fewer pericentromeric regions (Mandáková and Lysak 2008). Interestingly, the 45S rDNA loci on three O. violaceus chromosomes were active and gave differential amounts of rRNA transcripts in the B. napus background, while they were completely dominant over those of B. napus in the hybrids of these two species (Ge et al. 2009). However, the expression of rRNA genes from I. indigotica was not detected in the somatic hybrids with B. napus (data not shown), and also most likely not in the addition line Mg (Table 2). This also indicated the variable expression dominance of 45S rDNA in the interspecific hybrids and allopolyploids.
A complete series of MAALs enables markers and genes to be allocated to donor chromosomes on the basis of the presence/absence of markers and genes on the chromosomes added to the recipient genome (Cho et al. 2006). In this study, 184 SSR markers and 4 genes were allocated to the seven chromosomes of I. indigotica (Table 2). On the other hand, these unique materials could confirm interchromosomal duplicate loci. For example, PAL was encoded by a multi-gene family with four members in Arabidopsis (Raes et al. 2003). In I. indigotica, IiPAL had at least four homologous copies located on chromosomes c, d, e and f, respectively (Fig. 5c). As no genetic map has now been available for woad, these SSR markers would facilitate the construction of its genetic mapping. The production of radiation hybrid lines from monosomic additions can provide valuable tools for the physical mapping of alien chromosomes (Kynast et al. 2004).
The two tribes Isatideae and Brassicaceae which were put in one of the three lineages for Brassicaceae (Beilstein et al. 2006) descended from a common ancestral karyotype, proto-Calepineae karyotype (PCK, n = 7) (Mandáková and Lysak 2008), which shared two homologous ancestral segments. SSR markers and IiPAL showed different homeology within I. indigotica chromosomes (Table 3; Fig. 5c), and the higher degree of homoeology existed between chromosomes c and e, d and f, but lower or no homoeology between chromosome b and others.
The mitochondrial genome analysis showed that the recombination of parental mtDNA which occurred in the somatic hybrid was maintained in the MAALs derived, which might be partially responsible for the male sterility shown as the carpellody stamens in these MAALs and even the euploid B. napus progeny with such cytoplasm (Fig. 6), besides other factors affecting the development of floral organs. The homeotic conversion of stamens with carpelloid structures was also presented by monosomic rapeseed (B. napus)-radish additions with Raphanus cytoplasm (Budahn et al. 2008) and Brassica napus lines with rearranged Arabidopsis mtDNA (Leino et al. 2003). This is probably associated with the reshuffling mitochondrial dysfunction and nuclear–mitochondrial interactions that affect the development of floral organs. The expression of genes responsible for stamen formation depended on proper mitochondrial function and correct nuclear–mitochondrial interaction (Teixeira et al. 2005; Carlsson et al. 2007). The male sterile phenotype was present in all but one addition lines and their euploid offspring. The normal development of stamens in the MAAL (Me) indicated that the alien chromosome carried the gene(s) for fertility restoration which could be introgressed into B. napus to establish new cytoplasmic male sterility system for hybrid production.
Despite the usage of large amount with an excellent safety record as a traditional Chinese medicine plant, the effective ingredients of I. indigotica against viruses and bacteria are still elusive. MAALs obtained here provide a feasible strategy to identify and estimate overall bioactive compounds in each one of the MAALs by integrated analysis of the metabolome and transcriptome. Preliminary result indicates that an addition plant contained chromosome d and e of I. indigotica has obvious antiviral activity (unpublished data). These MAALs are greatly useful for understanding the biosynthesis pathway of plant secondary metabolites and the related key genes. The addition lines also could serve to detect the genes for plant resistances to S. sclerotiorum and TMV.
References
Ali HBM, Lysak MA, Schubert I (2005) Chromosomal localization of rDNA in the Brassicaceae. Genome 48:341–346
Al-Shehbaz IA, Beilstein MA, Kellogg EA (2006) Systematics and phylogeny of the Brassicaceae (Cruciferae): an overview. Plant Syst Evol 259:89–120
Beilstein MA, Al-Shehbaz IA, Kellogg EA (2006) Brassicaceae phylogeny and trichome evolution. Am J Bot 93:607–619
Budahn H, Schrader O, Peterka H (2008) Development of a complete set of disomic rape-radish chromosome addition lines. Euphytica 162:117–128
Carlsson J, Lagercrantz U, Sundström J, Teixeira R, Wellmer F, Meyerowitz EM, Glimelius K (2007) Microarray analysis reveals altered expression of a large number of nuclear genes in developing cytoplasmic male sterile Brassica napus flowers. Plant J 49:452–462
Chen BY, Simonsen V, Lannér-Herrera C, Heneen WK (1992) A Brassica campestris-alboglabra addition line and its use for gene mapping, intergenomic gene transfer and generation of trisomics. Theor Appl Genet 84:592–599
Cheng BF, Chen BY, Heneen WK (1994) Addition of Brassica alboglabra Bailey chromosomes to B. campestris L. with special emphasis on seed colour. Heredity 73:185–189
Cho S, Garvin DF, Muehlbauer GJ (2006) Transcriptome analysis and physical mapping of barley genes in wheat–barley chromosome addition lines. Genetics 172:1277–1285
Chung YC, Tang FY, Liao JW, Chung CH, Jong TT, Chen SS, Tsai CH, Chiang EP (2011) Isatis indigotica induces hepatocellular cancer cell death via caspase-independent apoptosis-inducing factor translocation apoptotic pathway in vitro and in vivo. Inteqr Cancer Ther 10:201–214
Cui C, Ge XH, Gantam M, Kang L, Li ZY (2012) Cytoplasmic and genomic effects on meiotic pairing in Brassica hybrids and allotetraploids from pair crosses of three cultivated diploids. Genetics 191:725–738
Dellaporta SL, Wood J, Hicks JB (1983) A plant DNA mini preparation: version II. Plant Mol Biol Rep 1:19–21
Ding L, Zhao ZG, Ge XH, Li ZY (2013) Intergeneric addition and substitution of Brassica napus with different chromosomes from Orychophragmus violaceus: phenotype and cytology. Sci Hortic 164:303–309
Du XZ, Ge XH, Yao XC, Zhao ZG, Li ZY (2009) Production and cytogenetic characterization of intertribal somatic hybrids between Brassica napus and Isatis indigotica and backcross progenies. Plant Cell Rep 28:1105–1113
Du ZJ, Liu H, Zhang ZL, Li P (2013) Antioxidant and anti-inflammatory activities of Radix isatidis polysaccharide in murine alveolar macrophages. Int J Biol Macromol 58:329–335
Ge XH, Wang J, Li ZY (2009) Different genome-specific chromosome stabilities in synthetic Brassica allohexaploids revealed by wide crosses with Orychophragmus. Ann Bot 104:19–31
Geleta M, Heneen WK, Stoute AI, Muttucumaru N, Scott RJ, King GJ, Kurup S, Bryngelsson T (2012) Assigning Brassica microsatellite markers to the nine C-genome chromosomes using Brassica rapa var. trilocularis-B. oleracea var. alboglabra monosomic alien addition lines. Theor Appl Genet 125:455–466
Heneen WK, Geleta M, Brismar K, Xiong ZY, Pires JC, Hasterok R, Stoute AI, Scott RJ, King GJ, Kurup S (2012) Seed colour loci, homoeology and linkage groups of the C genome chromosomes revealed in Brassica rapa–B. oleracea monosomic alien addition lines. Ann Bot 109:1227–1242
Klindworth DL, Niu ZX, Chao SM, Friesen TL, Jin Y, Faris JD, Cai XW, Xu SS (2012) Introgression and characterization of a goatgrass gene for a high level of resistance to Ug99 stem rust in tetraploid wheat. G3 2:665–773
Koo DH, Hong CP, Batley J, Chung YS, Edwards D, Bang JW, Hur Y, Lim YP (2011) Rapid divergence of repetitive DNAs in Brassica relatives. Genomics 97:173–185
Kynast RG, Okagaki RJ, Galatowitsch MW, Granath SR, Jacobs MS, Stec AO, Rines HW, Phillips RL (2004) Dissecting the maize genome by using chromosome addition and radiation hybrid lines. Proc Natl Acad Sci USA 101:9921–9926
Leino M, Teixeira R, Landgren M, Glimelius K (2003) Brassica napus lines with rearranged Arabidopsis mitochondria display CMS and a range of developmental aberrations. Theor Appl Genet 106:1156–1163
Li ZY, Liu HL, Luo P (1995) Production and cytogenetics of intergeneric hybrids between Brassica napus and Orychophragmus violaceus. Theor Appl Genet 91:131–136
Lim KB, Jong HD, Yang TJ, Park JY, Kwon SJ, Kim JS, Lim MH, Kim JA, Jin M, Jin YM, Kim SH, Lim YP, Bang JW, Kim H, Park BS (2005) Characterization of rDNAs and tandem repeats in the heterochromatin of Brassica rapa. Mol Cells 19:436–444
Liu S, Chen WS, Qiao CZ, Zheng SQ, Zeng M, Zhang HM, Song ZJ (2000) Antiviral action of Radix isatidis and Folium isatidis from different germplasm against influenza A virus. Acad J Sec Mil Med Univ 21:204–206
Mandáková T, Lysak MA (2008) Chromosomal phylogeny and karyotype evolution in x = 7 Crucifer species (Brassicaceae). Plant Cell 20:2559–2570
McArthur RI, Zhu XW, Oliver RE, Klindworth DL, Xu SS, Stack RW, Wang RRC, Cai XW (2012) Homoeology of Thinopyrum junceum and Elymus rectisetus chromosomes to wheat and disease resistance conferred by the Thinopyrum and Elymus chromosomes in wheat. Chromosome Res 20:699–715
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassay with tobacco tissue cultures. Physiol Plant 15:473–479
Peterka H, Budahn H, Schrader O, Ahne R, Schütze W (2004) Transfer of resistance against the beet cyst nematode from radish (Raphanus sativus) to rape (Brassica napus) by monosomic chromosome addition. Theor Appl Genet 109:30–41
Prakash S, Bhat SR, Quiros CF, Kirti PB, Chopra VL (2009) Brassica and its close allies: cytogenetics and evolution. Plant Breed Rev 31:21–187
Raes J, Rohde A, Christensen JH, Van de Peer Y, Boerjan W (2003) Genome-wide characterization of the lignification toolbox in Arabidopsis. Plant Physiol 133:1051–1071
Tan GX, Jin HJ, Li G, He RF, Zhu LL, He GC (2005) Production and characterization of a complete set of individual chromosome additions from Oryza officinalis to Oryza sativa using RFLP and GISH analyses. Theor Appl Genet 111:1585–1595
Tang XQ, Wang KC, Luan CM, Xu RJ (2010) Effect of Isatis indigotica on feeding and growth of larvae of Plutella xylostella. Acta Agri Jiangxi 22:75–77
Teixeira RT, Farbos I, Glimelius K (2005) Expression levels of meristem identity and homeotic genes are modified by nuclear-mitochondrial interactions in alloplasmic male-sterile lines of Brassica napus. Plant J 42:731–742
Tu YQ, Sun J, Ge XH, Li ZY (2010) Production and genetic analysis of partial hybrids from intertribal sexual crosses between Brassica napus and Isatis indigotica and progenies. Genome 53:146–156
Wang QY, Wang XB (1988) A TMV-resistant material-Banlangen. Acta Agric Boreati-Sin 3:92–95
Wang YP, Sonntag K, Rudloff E (2003) Development of rapeseed with high erucic acid content by asymmetric somatic hybridization between Brassica napus and Crambe abyssinica. Theor Appl Genet 106:1147–1155
Wang YP, Zhao XX, Sonntag K, Wehling P, Snowdon RJ (2005) Behaviour of Sinapis alba chromosomes in a Brassica napus background revealed by genomic in situ hybridization. Chromosome Res 13:819–826
Wei WH, Li YC, Wang LJ, Liu SY, Yan XH, Mei DS, Li YD, Xu YS, Peng PF, Hu Q (2010) Development of a novel Sinapis arvensis disomic addition line in Brassica napus containing the restorer gene for Nsa CMS and improved resistance to Sclerotinia sclerotiorum and pod shattering. Theor Appl Genet 120:1089–1097
Yang F, Li LJ (2011) Multicolor fluorescence in situ hybridization analysis on rDNA and telomere of Isatis indigotica. Chin Tradit Herbal Drugs 42:972–975
Zhao HJ, Huang YJ, Wang YY (1994) A study on intergeneric hybridization Brassica napus L. and Isatis indigotica Fort. Sci Agric Sin 27:89–91
Acknowledgments
The study was supported by NSFC project (No. 31071451) and Specific funds of Traditional Chinese Medicine Industry (No.201407003). We thank Prof. Luqi Huang, State Key Laboratory of Dao-di Herbs, National Resource Center of Chinese Materia Medica, China Academy of Chinese Medical Sciences, for providing the sequences of SSR markers.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by K. Toriyama.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kang, L., Du, X., Zhou, Y. et al. Development of a complete set of monosomic alien addition lines between Brassica napus and Isatis indigotica (Chinese woad). Plant Cell Rep 33, 1355–1364 (2014). https://doi.org/10.1007/s00299-014-1621-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-014-1621-8