Abstract
Key Message
We show that DCN1 binds ubiquitin and RUB/NEDD8, associates with cullin, and is functionally conserved. DCN1 activity is required for pollen development transitions and embryogenesis, and for pollen tube growth.
Abstract
Plant proteomes show remarkable plasticity in reaction to environmental challenges and during developmental transitions. Some of this adaptability comes from ubiquitin-mediated protein degradation regulated by cullin-RING E3 ubiquitin ligases (CRLs). CRLs are activated through modification of the cullin subunit with the ubiquitin-like protein RUB/NEDD8 by an E3 ligase called DEFECTIVE IN CULLIN NEDDYLATION 1 (DCN1). Here we show that tobacco DCN1 binds ubiquitin and RUB/NEDD8 and associates with cullin. When knocked down by RNAi, tobacco pollen formation was affected and zygotic embryogenesis was blocked around the globular stage. Additionally, we found that RNAi of DCN1 inhibited the stress-triggered reprogramming of cultured microspores from their intrinsic gametophytic mode of development to an embryogenic state. This stress-induced developmental switch is a known feature in many important crops and leads ultimately to the formation of haploid embryos and plants. Compensating the RNAi effect by re-transformation with a promoter-silencing construct restored pollen development and zygotic embryogenesis, as well as the ability for stress-induced formation of embryogenic microspores. Overexpression of DCN1 accelerated pollen tube growth and increased the potential for microspore reprogramming. These results demonstrate that the biochemical function of DCN1 is conserved in plants and that its activity is involved in transitions during pollen development and embryogenesis, and for pollen tube growth.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant reproduction is characterized by a number of important developmental transitions, beginning with the acquisition of polarity in the globular embryo (Jenik et al. 2007). The most fundamental transition in the life cycle of higher plants is the alternation of generations in which diploid sporophytes produce haploid male and female gametophytes (reviewed by McCormick 2004; Yadegari and Drews 2004). Microgametogenesis leads to the formation of pollen, whereas megagametogenesis produces the embryo sac enclosing the egg cell (Fan et al. 2008).
Cultured microspores remarkably show both developmental plasticity and high adaptability to stress (Touraev et al. 1997). Diverse stress treatments, such as temperature shock, nutrient starvation, or chemicals, abolish the gametophytic program intrinsic to microspores (Shariatpanahi et al. 2006). Instead of developing into pollen, stressed microspores become totipotent and develop into haploid embryos and plants under non-stress conditions (Touraev et al. 2001). This process corresponds to a fundamental developmental process in plants, i.e. the transition of a gametophyte into a sporophyte, but can be seen also as the reprogramming of a cell with a restricted development fate to a totipotent state. Homozygous doubled haploids emerging after spontaneous or induced chromosome doubling in the regenerated haploids are both scientifically attractive and highly valuable for plant breeding (Touraev et al. 2001; Forster et al. 2007). However, knowledge about the molecular mechanisms controlling the developmental switch of cultured microspores towards microspore embryogenesis is still fragmentary.
The ubiquitin–proteasome system (UPS), active in all eukaryotes, mediates protein degradation through sequential enzymatic action of a ubiquitin-activating enzyme E1, a ubiquitin-conjugating protein E2, and a substrate-specific ubiquitin E3 ligase (reviewed by Hotton and Callis 2008). In plants, more than 1,300 proteins have been identified as playing a role in the UPS compared to approximately 150 proteins in Saccharomyces cerevisiae (Vierstra 2009). Approximately 1,200 or 90 % of these genes encode components of E3 ubiquitin ligases suggesting that hundreds or even thousands of proteins are regulated by the UPS. Multi-subunit protein complexes form the largest class of E3 ligases. The cullin subunit acts as platform protein, a RING H2 finger protein mediates contact to an E2 protein, and variable substrate-recognition subunits such as F-box proteins are connected to the complex by adaptors (reviewed by Schwechheimer and Calderon Villalobos 2004; Bosu and Kipreos 2008). In Arabidopsis, five canonical cullin proteins (CUL1, CUL2, CUL3A, CUL3B, and CUL4) have been shown to constitute four classes of E3 ligase complexes (Santner and Estelle 2010). Apart from protein quality control, targeted protein degradation by CRLs has been associated with photomorphogenesis, self recognition during reproduction, interplay between pathogens and their hosts, cell division, and the synthesis of and response to various plant hormones (reviewed by Santner and Estelle 2010; Dreher and Callis 2007; Hua and Vierstra 2010).
CRL activity is controlled via modification of the cullin subunit by the ubiquitin-like protein (UBL) RUB/NEDD8 in fission yeast and animals. The Arabidopsis genome contains three RUB/NEDD8 genes, two of which, RUB1 and RUB2, regulate diverse processes throughout plant development and are essential as demonstrated by embryo lethality in rub1 rub2 double mutants (Bostick et al. 2004; Parry and Estelle 2004). Derubylation is accomplished by the COP9 signalosome (Lyapina et al. 2001), whereas rubylation is mediated by an enzymatic cascade, which, in analogy to ubiquitylation, comprises three different enzymatic steps (E1, E2, and E3). In Arabidopsis, the RUB E1 and E2 functions are performed by AXR1/ECR1 and RCE1, respectively (del Pozo et al. 2002; Dharmasiri et al. 2003). Rubylation/neddylation and derubylation/deneddylation have been shown to be essential in plants, worms, and mammals, partially in fission yeast, but not in budding yeast (reviewed by Hotton and Callis 2008; Rabut and Peter 2008). They play a fundamental role in important processes such as morphogenesis (Tateishi et al. 2001), cell division (Lammer et al. 1998), signaling (del Pozo and Estelle 1999), and embryogenesis (Kurz et al. 2002, 2005). In plants, loss of AXR1 results in a variety of hormone-related phenotypes including reduced sensitivity to auxin, cytokinin, ethylene, epi-brassinolide, and jasmonic acid (Santner and Estelle 2010).
The E3 activity for rubylation/neddylation can be provided by different proteins (Scott et al. 2010; Duda et al. 2011). In mammals, the RING E3 ligase Mdm2 mediates neddylation of p53, leading to negative regulation of its transcriptional activity, and neddylation of tyrosine receptor kinases by the dual-function c-Clb E3 ligase leads to receptor recycling and signal attenuation (Watson et al. 2011). Another RUB/NEDD8 E3 is the RING H2 finger protein RBX1, which is able to recruit both ubiquitin and RUB/NEDD8 to cullins in an E3 ligase (Duda et al. 2011). In Arabidopsis, RBX1 is encoded by two genes (Gray et al. 2002). RBX1 functions synergistically with yet another RUB/NEDD8 E3, called DEFECTIVE IN CULLIN NEDDYLATION 1 (DCN1) (Higo et al. 1999), which uses a novel mechanism to bind both a cullin and the RUB E2 (Scott et al. 2010). Acting as a scaffold-like RUB/NEDD8 E3 ligase in association with RBX1 (Yang et al. 2007), DCN1/Dcn1p was proven to be required and sufficient for cullin neddylation in vitro and in vivo (Kurz et al. 2005, 2008). X-ray crystal structure analysis established that yeast DCN1 contains an N-terminal ubiquitin-binding (ubiquitin-associated or UBA) domain and a large C-terminal POTENTIATING NEDDYLATION (PONY) domain that binds NEDD8 and provides the surface for interaction with cullins and the RUB/NEDD8 E2 enzyme Ubc12 (Kurz et al. 2008). Combinatorial interactions between RBX1 and DCN1 seem to impart specificity to the multifunctional cullin-RBX1 ligase complex (Scott et al. 2010).
By using suppression subtractive hybridization of stressed embryogenic and non-stressed gametophytic microspores, we previously reported several differentially expressed genes (Hosp et al. 2007a). One of them, ntsm10 (Nicotiana tabacum stressed microspore 10), was found to be a homolog of nematode DCN-1, yeast Dcn1p, and human DCNL1 and 2 (Meyer-Schaller et al. 2009) and accordingly renamed into NtDCN1 (Nicotiana tabacum DCN1). Since plant DCN1 properties and function remained to be analyzed, we used biochemical binding assays, RNAi and overexpression to investigate its function during developmental transitions in tobacco. Here, we show that tobacco DCN1 binds ubiquitin and RUB/NEDD8 similar to other eukaryotic models and physically associates with cullin. Overexpression of DCN1 in tobacco promotes the formation of embryogenic microspores and accelerates pollen tube growth. On the other hand, we observed phenotypes in RNAi lines that show the requirement of Nicotiana tabacum NtDCN1 for developmental transitions such as from microsporogenesis to microgametogenesis, from the gametophytic to the sporophytic phase in cultured microspores, and from the globular to the heart-shaped stage in both zygotic and microspore embryogenesis.
Results
NtDCN1 Binds Ubiquitin and RUB/NEDD8, and Associates with Cullin 1
NtDCN1 encodes a 30-kDa protein comprising 259 amino acids and is represented by two almost identical paralogs in the allotetraploid tobacco genome—NtDCN1A (DQ885939) and NtDCN1B (FJ976682) (Fig. 1a). NtDCN1 has high similarity to nematode DCN-1 and yeast Dcn1p and contains a UBA and a PONY domain (Kurz et al. 2005). The closest homolog of NtDCN1 in Arabidopsis is an unknown protein (At3g12760) with 78 % identity and similar domain architecture. Another close relative, At3g28970 (ANTI-AUXIN-RESISTENT 3, AAR3) with 45 % identity but without a UBA domain, has recently been shown to regulate root responses to 2,4-dichlorophenoxyacetic acid (2,4-D), a synthetic auxin (Biswas et al. 2007). A third, still uncharacterized PONY domain-containing protein, At1g15860, shares 32 % identity with NtDCN1.
NtDCN1 is a highly conserved 30-kDa DCN1 ortholog that binds ubiquitin and RUB/NEDD8 and Associates with cullin 1. a Sequence alignment of the two DCN1 paralogs (DCN1A and DCN1B) with DCN1 orthologs from different organisms, secondary structure prediction, and domain structure. Conserved amino acids are highlighted in gray, substitutions are boxed. Regions with a high probability of α-helix formation (H), as well as UBA, PONY, and EF-hand domains are indicated (EF1, EF2, canonical EF-hands are helix-loop-helix structural domains with an affinity to calcium). b NtDCN1 binds ubiquitin and RUB/NEDD8. Recombinant NtDCN1 was incubated with beads conjugated to either ubiquitin (5 μg/μl, mix, Ubi) or RUB/NEDD8 (7–15 μg/μl, Nedd), respectively. Aliquots of wash fractions (W1–W8) and eluates (elu) were subjected to PAGE, and Western blots were probed with anti-DCN1. c Pull-down of NtDCN1 with At CUL1:His fusion protein bound to Ni–NTA beads. Aliquots of wash fractions (W1, W2) and eluates (1–3) were subjected to PAGE and Western blotting with anti-Nt DCN1. The band above 86 kDa in the first wash fraction (W1) appears to be an unspecific artifact. The association of recombinant NtDCN1 with recombinant His:AtCul1 bound to the beads is disrupted upon elution (fractions 1 and 2). The reaction was carried out with recombinant proteins in absence of Nedd8. The lower panel represents a control assay without Cul1
To demonstrate the functional properties of plant DCN1 we tested its ability to bind ubiquitin and RUB/NEDD8 in vitro. An NtDCN1-specific polyclonal antibody detected the expected 30-kD DCN1 protein on Western blots of eluted fractions resulting from incubation of recombinant NtDCN1 with ubiquitin and RUB/NEDD8 beads, respectively (Fig. 1b). Thus, NtDCN1 readily binds ubiquitin and RUB/NEDD8, similar to yeast DCN1p and nematode DCN-1 (Kurz et al. 2005). A pull-down experiment using At CUL1 shows that NtDCN1 associates with cullin 1 (Fig. 1c and Online Resource 1), confirming data obtained in yeast and nematodes (Kurz et al. 2005, 2008).
NtDCN1 is expressed throughout the plant body, and expression is differentially regulated during pollen development and microspore reprogramming in vitro
Histochemical analyses of transgenic tobacco lines harboring an NtDCN1pro:GUS construct demonstrated promoter activity throughout the plant body and indicated developmental regulation. In mature plants, the major site of GUS expression was the vascular system, as exemplified in seedlings (Fig. 2a). Strong GUS signals were also detected in meristematic and actively developing regions (Fig. 2b–e). The NtDCN1 promoter was also active in reproductive tissues, as observed in axile placentas of unfertilized ovaries (Fig. 2f) and in mature pollen (Fig. 3e). Weak GUS expression was found in early embryos and endosperms (Fig. 2g), visualized in dark-field images demonstrating mostly spotted GUS expression in these tissues (Fig. 2h). Mature seeds showed spotted GUS signals and strong GUS expression was observed at the embryonic root tip upon imbibition (Fig. 2j–l). Early seedlings had GUS signals in cotyledons and the distal part of the hypocotyl (Fig. 2l). Northern blot analyses confirmed that NtDCN1 transcripts are present in different organs of tobacco plants (Fig. 3a).
NtDCN1 is expressed throughout the plant body. Expression pattern of NtDCN1pro:GUS in seedling (a), lateral meristem (b), lateral root initial (c), apical meristem (d), first leaf primordium (e), unfertilized ovaries (f), early embryo and endosperm (g) (light microscopy, no GUS staining is visible due to weak expression), early embryo and endosperm (h) (dark-field image showing weak GUS expression as pink dots and strong signals in blue), mature seeds (j), imbibed seeds (k), early seedling (l)(color figure online)
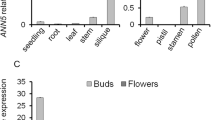
NtDCN1 is expressed during pollen development and microspore embryogenesis. a–c Expression pattern of NtDCN1 by Northern (a, b) and Western blotting (c) in vegetative tissues (a), during microspore embryogenesis (b) and pollen development (c). Northern blots were hybridized with the radiolabeled 3′ UTR of NtDCN1. Lower panels show loading controls of 18S rRNA in a, b, Coomassie staining in c infl. inflorescences, y young leaf, m intermediate leaf, o old leaf, stem stem, root root, um unicellular microspores, sm stressed microspores, em microspore-derived embryos (after 25 and 50 days), mp mature pollen, bp bicellular pollen. d Expression pattern of DCN1 detected by in situ hybridization. sm stressed microspores, dm dividing microspores, mc multicellular structures, ge globular embryos, ce cotyledon stage embryos. e Nuclear status assayed by DAPI staining and NtDCN1pro:GUS expression. um unicellular microspores, mp mature pollen, sm stressed microspores, lg late globules
Both RNA and protein blots showed that NtDCN1 is present throughout pollen development and microspore embryogenesis, in accordance with NtDCN1 promoter activity (Fig. 3b, c, e). NtDCN1 transcript levels were detected with the 3′-UTR of NtDCN1, which recognizes both NtDCN1 paralogs. They were high in unicellular microspores but decreased after the first pollen mitosis, i.e. in bicellular and mature pollen (Fig. 3b). Protein levels, in contrast, increased strongly during pollen development from unicellular microspores to mature pollen (Fig. 3c).
A slight increase in expression was detected on RNA blots between freshly isolated microspores and cultured microspores subjected to a stress treatment for 6 days for reprogramming into embryonic development (Fig. 3b), similar to protein levels (Fig. 3c). An increase in expression was detected also by in situ hybridization, by GUS assays in NtDCN1pro:GUS plants, (Fig. 3d, e), and by RT-PCR (Fig. 5b). NtDCN1 transcripts, promoter activity, and NtDCN1 protein were consistently detected in 25-day-old microspore-derived embryos (Fig. 3b–e) while a reduction of NtDCN1 expression was observed in 50-day-old, late torpedo-shaped embryos on the protein level (Fig. 3c) and at the cotyledon stage on the RNA level (Fig. 3d).
RNAi of NtDCN1 leads to severe impediment of tobacco pollen development and embryogenesis
RNAi was used to study the in vivo function of NtDCN1. A 300-bp 3′ UTR fragment corresponding to the originally identified SSH sequence was cloned into a vector downstream of the constitutive CaMV 35S promoter in sense and anti-sense orientations linked by an intron (Smith et al. 2000). Reductions in mRNA and protein levels varied from weak to strong in individual lines, as expected for RNAi knock-down plants. Both NtDCN1 mRNA levels and NtDCN1 protein levels were strongly reduced in leaves of RNAi lines (Fig. 4a). This indicated that due to their high sequence similarity both NtDCN1-paralogs were knocked down by RNA interference.
NtDCN1 is required for microspore and zygotic embryogenesis as well as for pollen development. a Northern and Western blots of three homozygous NtDCN1 RNAi lines and their respective, re-transformed homozygous promoter-silencing lines compared to the wild type. Endogenous NtDCN1 RNA levels were detected with an NtDCN1-derived probe, comprising the coding region, and double-stranded hairpin RNA levels by probing against the intron of the RNAi construct. GAPDH was used as a loading control. hp hair-pin probe, PS promoter silencing lines. b Pollen formation from cultured microspores in RNAi and promoter silencing lines. wt wild type, PS promoter silencing lines, um unicellular microspores, mp mature pollen, gp germinated pollen. Bars 30 μm. c DIC microscopy pictures of embryo development and seed formation in RNAi line L47 compared to the wild type. Early development of zygotes in both RNAi and wild-type plants was normal and synchronous, but embryos in RNAi lines were arrested around the globular stage, or showed malformed heart and cotyledon stages. z zygote, ge globular embryo, he heart stage embryo, te torpedo stage embryo. Bars 20 μm (z), 200 μm (ge, he, te). d Seed pods were smaller in RNAi lines, here shown for L47, immature seeds developed asynchronously, and only a few mature seeds developed in contrast to the wild type. is immature seeds, ms mature seeds. e Microspore embryogenesis in wild-type, RNAi (L47) and promoter silencing line PS–L47 (PS). um unicellular microspores, sm stressed microspores, ge globular embryos, ce cotyledon stage. Bars 30 μm (um, sm), 100 μm (ge), 2 mm (ce)
Nine RNAi lines with strong and six lines with medium reduction of NtDCN1 transcript abundance were analyzed for pollen and seed development. Plants with a strong reduction in RNA and protein levels appeared slightly smaller compared to wild-type plants but did not show any other morphological changes (not shown). Seed pods contained mainly stunted seeds of different sizes. Mature anthers of primary RNAi lines enclosed varying numbers of dead pollen plus mature pollen in frequencies ranging from 5 to 40 %, which correlated with the degree of NtDCN1 down-regulation.
To analyze the effect of an NtDCN1 knock-down on reproductive development, homozygous RNAi lines were produced by selfing and kanamycin selection of seeds. Homozygosity was verified by non-segregation of the kanamycin gene in the F2 progeny, and Northern and Western blots confirmed the transmission of the RNAi effect in leaves (Fig. 4a), microspores, stressed microspores, and microspore embryos (Online Resource 2). Three independent homozygous lines (L25, L35 and L47) were selected for further phenotypic analyses. The viability of microspores and early bicellular pollen grains were lower than in wild-type plants (Table 1), whereas the majority of pollen grains appeared to be dead (Fig. 4b, L47, Table 1). Microspores isolated from those plants and cultured in a pollen maturation medium (Benito Moreno et al. 1988; Touraev and Heberle-Bors 1999) developed into mature pollen at frequencies similar to microspores developing in anthers on the plants (Online Resource 3). This strongly supports the assumption that the RNAi knock-down effect operated within the microspores.
Likewise, the majority of seeds were stunted, both after self-pollination and after backcrossing with wild-type pollen, indicating reduced female fertility. Using Nomarski DIC optics we observed that the seeds were arrested at different stages, mainly at the pre-globular to globular stages (Fig. 4c, d).
To study the role of NtDCN1 in microspore embryogenesis, unicellular microspores were isolated before pollen mitosis I (PM I), i.e. prior to their in vivo death, from anthers of homozygous RNAi lines. These and corresponding wild-type microspores were subjected to a mild heat shock at 33 °C and starvation in a sugar and nitrogen-free medium for 6 days (Touraev and Heberle-Bors 1999). In the wild-type, this stress treatment as expected resulted in the formation of embryogenic microspores, and the formation of embryos (Fig. 4e) and haploid plants. In contrast, although originally viable, the vast majority of microspores isolated from all three homozygous RNAi lines died during the stress treatment. However, a few microspores (approx. 2 % of the total population) survived and dedifferentiated into embryogenic microspores. When enriched by Percoll gradient centrifugation and cultured in embryogenesis medium, these embryogenic microspores were able to form embryos. But, similar to zygotic embryos, they were either arrested at around the globular stage with irregular shapes or strongly delayed in their further development (Fig. 4e).
Alleviation of NtDCN1 RNAi by transcriptional gene silencing reconstitutes the wild-type phenotype
To prove that the observed phenotypes were caused by NtDCN1 knock-down, we used transcriptional silencing of the promoter controlling the RNAi construct to restore the wild-type phenotype. The construct used for re-transformation contained an inverted repeat of the full 35S promoter under control of the constitutively active ubiquitin promoter and lacked a poly-adenylation site, thereby switching off the 35S promoter by RNA-directed DNA methylation (Mette et al. 2000; Sijen et al. 2001). The hygromycin phosphotransferase (HPT) gene was included in the construct to be able to select double transformants on hygromycin and kanamycin. Both NtDCN1 mRNA and protein levels recovered in the double-transgenic lines, exemplified by line PS-L47 in Fig. 4a, and hyper-methylation of the 35S promoter was confirmed (Online Resource 4). As anticipated, plant and flower size, pollen development, seed set, and zygotic embryogenesis, as well as microspore embryogenesis, were restored in a manner and in frequencies similar to wild-type plants in all lines with reconstituted NtDCN1 expression, represented by PS-L47 (Fig. 4b, e). We thus conclude that the phenotypes seen in homozygous NtDCN1 RNAi lines were indeed caused by a knock-down of the NtDCN1 gene.
Overexpression of NtDCN1 accelerates pollen tube growth and promotes reprogramming of microspores
Transgenic plants were created that harbored the NtDCN1 full-length cDNA under control of the DC3 promoter, shown to be highly active during male gametophyte development and microspore embryogenesis (Wilde et al. 1988; Touraev et al. 1995). Out of 25 independent transgenic lines overexpressing DCN1 in young leaves, two lines with strong overexpression in microspores (ox14 and ox21, 4A, B) were self-pollinated, and homozygous offspring were selected. These plants did not show an apparent phenotype, including in seed formation.
Pollen viability and germination, as measured by FDA staining and in vitro germination test, did not differ much between wild-type plants and overexpressing lines (Table 2) and the difference between overexpressing lines depended on the degree of overexpression. Significant differences, however, were detected in pollen tube growth rates (Fig. 5d). With 10.22 μm/min for ox14 and 7.95 μm/min for ox21, overexpressor pollen tubes in GK-medium grew at about twice the speed of wild-type pollen tubes (4.38 μm/min, Table 2). Wild-type tobacco pollen, in turn, grew at more than double the speed reported by (Certal et al. 2008), probably due to a more appropriate culture medium, while ox14 pollen tube growth rates approached in vivo growth rates of wild-type tobacco pollen (25 μm/min, Cheung et al. 2000).
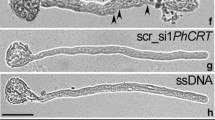
Overexpression of NtDCN1 accelerates pollen tube growth and promotes reprogramming of microspores in vitro. a Overexpression of NtDCN1 in microspores determined by Western blotting. ctrl recombinant NtDCN1, wt wild type, ox14 overexpressing line 14, ox21 overexpressing line 21. In the lower panel, the Coomassie-stained gel is shown as a loading control. b RT-PCR showing the time-course of NtDCN1 expression during the 6-day stress treatment leading to the reprogramming of microspores. NtDCN1 expression is elevated in unstressed and stressed microspores of overexpressing lines ox14 and ox21 compared to the wild type. u unstressed microspores, 2 2-day-stressed microspores, 4 4-day-stressed microspores, 6 6-day-stressed microspores. c In the overexpressing lines, 2 days of stress treatment were sufficient to induce embryogenesis at a high frequency compared to wild-type plants. Pictures of embryos in microspore cultures prepared from ox14-plants were taken after 2 weeks (globular stage) and 6 weeks (cotyledon stage). Bars 300 μm. d Pollen tube growth in wild type and overexpressing lines. Pictures were taken after 30′ and 120′, respectively. wt wild type, ox14 overexpressing line 14, ox21 overexpressing line 21. Bars 30 μm
To evaluate whether NtDCN1 is not only necessary but sufficient for stress-induced reprogramming of microspores towards embryogenesis, we incubated microspores from wild-type and overexpressor plants under reprogramming conditions, followed by culturing them in embryogenesis medium for embryo formation. NtDCN1 transcript levels, measured throughout the 6-day stress treatment, during which reprogramming occurs in the wild-type, increased continuously in both wild-type and NtDCN1-overexpressing microspores, and clearly higher levels were detected in overexpressing microspores already during the first days of culture (Fig. 5b). Approximately 80 % of wild-type microspores dedifferentiated into embryogenic microspores after a stress treatment of 6 days (Table 3), similar to published data (Touraev et al. 1999). After 2 weeks in embryogenesis medium most of the reprogrammed microspores had developed into multi-cellular structures, and after 6 weeks, these had formed well-developed globular to torpedo-shaped embryos. A shorter, 2-day, exposure to the stress treatment decreased the frequency of embryogenic microspores to 40 %, and only 10–12 % of the surviving cells developed into multicellular structures and embryos (Table 3; Fig. 5c). In the overexpressing lines, the frequency of embryogenic microspores after 2 days of stress treatment reached almost 70 %, similar to wild-type microspores after a 6-day stress treatment, and the majority of these embryogenic microspores developed into multicellular structures and embryos (Table 3; Fig. 5c).
When we completely omitted the stress treatment and cultured ox14 and ox21 microspores directly in embryogenesis medium, no embryos formed. Cytological investigations of embryogenic cultures, however, revealed that microspores from both overexpressing lines developed into multi-cellular structures faster and at a higher frequency than wild-type microspores (Table 4).
Discussion
NtDCN1 shares biochemical features with other eukaryotic DCN1 proteins
We have found that NtDCN1 binds cullin 1, ubiquitin, and RUB, similar to DCN1 proteins of other eukaryotes and consistent with the features of a potential RUB E3 ligase (Kurz et al. 2008). Ubiquitin binding is particular to DCN1 and distinguishes the DCN1 gene from DCN1-like genes such as AAR3, the only member of the DCN gene family characterized in plants to date (Biswas et al. 2007). Unfortunately, due to the lack of suitable cullin and RUB antibodies, we were not able to show differences in cullin rubylation. Further experiments will have to show whether cullins are DCN1-mediated rubylation targets in the developmental transitions that we have described, and if so, which specific CRLs they constitute. These could be SCF-like complexes similar to those involved in cell cycle regulation (Lammer et al. 1998; Vodermaier 2004; Inze and De Veylder 2006) or in the signal transduction of hormones (Lechner et al. 2006; Dreher and Callis 2007). Apart from the CUL1 subunit of SCFs, CUL3a, CUL3b, and CUL4 are also known to be modified by RUB (Dreher and Callis 2007). In addition, non-cullin proteins may be rubylated, as the list of neddylated proteins is increasing (Rabut and Peter 2008; Watson et al. 2011).
NtDCN1 is necessary for pollen development while overexpression promotes pollen tube growth
During pollen development, NtDCN1 transcript levels quickly decreased after PM I and remained low in mature pollen grains, similar to the expression of its Arabidopsis ortholog At3g12760 (Honys and Twell 2004), while protein levels strongly increased. This phenomenon has been described for a large number of pollen-expressed genes (Holmes-Davis et al. 2005) A feedback mechanism may be operating in developing pollen grains, down-regulating NtDCN1 transcription when the protein has accumulated beyond a certain threshold level. NtDCN1 thus seems to be stored in mature pollen grains, presumably to be used during pollen germination and tube growth.
A loss of function of NtDCN1 in pollen, induced by RNAi-mediated knock-down, led to a developmental arrest after PM I in early bicellular pollen. This stage marks the developmental phase change from microsporogenesis to microgametogenesis, i.e. when genes required for microgametogenesis are activated after PM I (Ma 2005). Pollen defects have also been found in other components of the rubylation and ubiquitylation pathways. Pollen of insertion mutants in the AtDCN1-like AAR3 gene failed to transmit the auxin-resistant phenotype to the next generation when used to pollinate wild-type Arabidopsis plants (Biswas et al. 2007). Similarly, mutants in the AXR1 gene, encoding a RUB E1 (del Pozo et al. 2002), produced significantly less pollen (Lincoln et al. 1990), and phytotoxin-resistant mutants in CORONATINE-INSENSITIVE 1 (COI1), which is part of an SCFCOI1-complex involved in jasmonate signaling (Xu et al. 2002), had a male-sterility phenotype (Feys et al. 1994).
Pollen lethality has been observed in Arabidopsis thaliana RNAi plants to be unrelated to the loss of function of particular genes and to be a general feature of RNAi knock-down in this species (Xing and Zachgo 2007). Unlike in Arabidopsis, however, the frequencies of dead pollen were much higher in our tobacco RNAi knock-down plants, while microspores were viable, and we observed defects not only in pollen but also in seed set and embryonic development. In addition, our experiments to alleviate the RNAi knock-down of NtDCN1 by transcriptional silencing showed that complementation restored the RNAi-affected functions, i.e. pollen lethality as well as zygotic and microspore embryogenesis, arguing for a specific effect of RNAi on NtDCN1.
In our NtDCN1 overexpressing lines, pollen development per se was not significantly affected, presumably because of an already high amount of NtDCN1 protein present in mature wild-type pollen (see above). Overexpression of NtDCN1 resulted in a clear acceleration of pollen tube growth compared to wild-type plants, possibly due to an enhanced turnover of stored proteins. Proteome analyses have verified that pollen grains store synthesized proteins for later use (Holmes-Davis et al. 2005; Noir et al. 2005; Dai et al. 2007) and pollen function appears to depend on selective protein degradation by the 26S proteasome (Sheng et al. 2006; Dai et al. 2007). Indeed, the proteolytic machinery is required during virtually all stages of pollen development and germination (Capron et al. 2003; Liu et al. 2008; Scoccianti et al. 1999; Yang et al. 1999; Zhao et al. 2001; Doelling et al. 2007) including the control of self-incompatibility (Qiao et al. 2004). Our results suggest that NtDCN1 is an important player in some of these processes and that rubylation may activate ubiquitin E3 ligases specific for repressor proteins, e.g. such controlling translation of stored mRNAs, or specific for ubiquitin E3 ligase growth factor receptors regulating pollen tube growth. Alternatively, overexpression of NtDCN1 may promote pollen tube growth by stimulating actin turnover. In human cells, inhibition of neddylation caused abnormalities in the actin cytoskeleton by affecting the accumulation of the small GTPase RhoA, a recently identified CRL substrate (Leck et al. 2010).
NtDCN1 is necessary for zygotic embryogenesis
In our RNAi lines, both male reproductive and zygotic development were severely affected, indicating that NtDCN1 is an essential gene. The developmental arrest at the globular stage of embryogenesis found in NtDCN1 RNAi lines resembled that in Arabidopsis cullin3 mutants (Thomann et al. 2005; Figueroa et al. 2005) and in apc2 mutants (Capron et al. 2003). APC2 is a distant member of the cullin protein family and encodes a subunit of the E3 ligase anaphase-promoting complex or cyclosome (APC/C). Similarly, DCN1 RNAi caused a complete developmental arrest in embryos of nematodes (Kurz et al. 2005) while the same study showed that Dcn1p is not essential in budding yeast. DCN1 knockout mice (SCCRO −/− mice) have reduced viability and severe developmental defects (Kim et al. 2008) and, similar to our NtDCN1 knock-down tobacco plants, were smaller and showed male infertility with dysfunctional spermatogenesis (Kaufman et al. 2007). The other components of the neddylation/rubylation pathway, i.e. RUB/NEDD8 E1 and E2, are essential in both animals and plants (Hotton and Callis 2008; Bosu and Kipreos 2008) while with growth reduction and gametophyte or embryonic lethality, antisense suppression of RBX1 in Arabidopsis (Gray et al. 2002) resulted in a phenotype comparable to our NtDCN1 RNAi plants.
Apart from the described phenotype and smaller size, our RNAi plants looked normal, thus proving that NtDCN1 is not essential for viability. Rubylation of target proteins in the absence of NtDCN1 may have accounted for this. Given the recent discovery that RBX1 and DCN1 interact in neddylation and that the function of DCN1 is to impart specificity to the cullin-RBX1 ligase complex (Scott et al. 2010), it appears that in plants with down-regulated DCN1 RBX1 is still able to rubylate target proteins to a degree sufficient to maintain viability and vegetative development. The specificity provided by DCN1 seems to be more important for reproductive processes. In in vitro experiments, in fact, cullin neddylation has been shown to occur without DCN1 activity through direct neddylation of cullins by RBX1, albeit less efficiently (Kurz et al. 2005; Yang et al. 2007; Kim et al. 2008). Alternatively, other members of the DCN gene family may have substituted for DCN1 in vegetative processes, similar to the human DCN1-like proteins DCNL 3 and 5, which are able to substitute for DCN1 in dcn1Δ HeLa cells (Meyer-Schaller et al. 2009). In fact, in Western blots using an anti-RUB/NEDD8 antibody we were able to detect high-molecular-weight complexes and an excess of free RUB in leaves (not shown). Third, the NtDCN1 knock-down may have been incomplete, and residual expression of NtDCN1 in the RNAi plants may have been sufficient to maintain vegetative processes.
Unlike the antisense suppression phenotype of RBX1 in Arabidopsis, the NtDCN1 RNAi trait was transmitted to the offspring through a few functional gametes. Incomplete penetrance and variable expressivity have been reported also for the Arabidopsis cul1 and cul3 mutant phenotypes (Shen et al. 2002; Thomann et al. 2005; Figueroa et al. 2005) and the rhf1a rhf2a (RING-H2 group F) RING-type E3 ligase double mutant phenotype (Liu et al. 2008). In the latter, only a fraction of embryo sacs, pollen, and embryos aborted while other embryos with a mutated gene developed normally.
NtDCN1 is necessary and increases the potential for the reprogramming of microspores towards totipotency
Despite its scientific attractiveness and the economic importance of microspore-derived doubled haploid plants for plant breeding (Forster et al. 2007), the molecular mechanisms controlling this process have remained largely unknown. Transition of microspores into the embryogenic state involves the degradation of earlier synthesized RNAs and proteins (Garrido et al. 1993; Joosen et al. 2007; Malik et al. 2007), particularly of gametophyte-specific proteins and metabolites (discussed in Hosp et al. 2007b). The isolation of a number of protease and ubiquitin-interacting genes from embryogenic microspores in independent studies supports this assumption (Hosp et al. 2007b; Maraschin et al. 2005). However, no key regulators involved in microspore reprogramming have been identified until now.
Our RNAi experiments showed that NtDCN1 is necessary for the stress-induced transition from microsporogenesis to microspore embryogenesis. Most microspores died during the stress treatment. Like in pollen development and zygotic embryogenesis, the RNAi phenotype showed incomplete penetrance and variable expressivity which allowed enriching the surviving embryogenic microspores by density centrifugation and culturing them to produce embryos. These embryos were arrested in their majority at the globular stage while a few continued with embryonic development and developed into plants.
As compared to freshly isolated microspores and unlike in normal pollen development, the NtDCN1 gene was slightly up-regulated by the reprogramming stress treatment on the RNA and protein level. RNA and protein levels remained high in microspore-derived embryos until the globular stage, but were lower in late torpedo-shaped embryos. We conclude that NtDCN1 function is required until the globular stage of development, in line with the developmental arrest at the globular stage found in microspore-derived and zygotic embryos of NtDCN1 RNAi lines.
The gain-of-function phenotype obtained by overexpressing NtDCN1 with the DC3 promoter indicated that NtDCN1 is not only necessary but also increases the potential for the phase change from gametophytic to sporophytic development of cultured microspores. Overexpression reduced the time of the stress treatment required to reprogram microspores. Interestingly, constitutive overexpression of DCN1 in mice resulted in embryo-lethality while in conditional mutants overexpression of DCN1 contributed to tumor formation (Broderick et al. 2010).
In cultured microspores that were not subjected to any stress treatment, the number of multi-cellular structures (early embryos) as well as the number of cells in these embryos were increased as compared to the wild type. However, these multicellular structures did not develop into advanced embryos, possibly due to an insufficient amount of conditioning factors produced by these embryos, a phenomenon frequently observed in microspore cultures of different species. Alternatively, a full priming of microspore totipotency may require the action of multiple reprogramming genes, similar to the genes that are required for the direct reprogramming of differentiated mammalian cells (Takahashi and Yamanaka 2006).
Materials and methods
Plant materials and microscopy
Nicotiana tabacum cv. “Petit Havana” SR1 plants were grown in at 25 °C with a 16-h day. Microspores were isolated and cultured for maturation into pollen and for microspore embryogenesis according to Touraev and Heberle-Bors (1999).
Ovules were prepared according to Figueroa et al. (2005). After gently squeezing the fertilized ovules, the embryos were released and observed under a Zeiss Axioplan microscope equipped with Nomarski (DIC) optics. FDA staining and pollen germination assays were performed as described (Touraev and Heberle-Bors 1999). Pollen tube length was measured from images projected on a digitizer by using SIGMA-SCAN software V3.90 (Jandel Scientific, now http://www.systat.com, Ylstra et al. 1996). Histochemical GUS assays were performed according to Jefferson et al. (1987).
Cloning
Full-length cloning of NtDCN1 was carried out according to the instructions of the SMART RACE cDNA amplification kit (Clontech, http://www.clontech.com) using adapter and gene-specific primers. RACE products were T/A-cloned (Invitrogen, http://www.invitrogen.com) and sequenced. The UNIVERSAL GENOME WALKER Kit (Clontech) was used for promoter isolation. The largest products were cloned into pCR2.1 and sequenced. The promoter region was inserted into the plant transformation vector pBI101.1. NtDCN1 RNAi construct: A 270-bp fragment of the 3′ end of DCN1 was cloned into pART69 (Gleave 1992) in antisense and sense orientation, separated by the adh Y5 intron, and flanked by the 35S promoter and the ocs terminator, respectively. This cassette was cloned into pBI101.1 for plant transformation. ProSi (CaMV 35S promoter silencing) construct: A 1.4-kb fragment of the 35S promoter from pART69 was cloned into the same vector in antisense orientation. This cassette was fused to a Ubi promoter fragment from pAHC25 and cloned into the pBIN:Hyg:TX vector (NCBI acc. no. Z37515). NtDCN1 overexpression construct: Restriction sites were added to the 5′ and 3′ ends of NtDCN1 by PCR using specific primers. The product was T/A-cloned (Invitrogen), sequenced, and cloned into pBI101.1 containing the DC3 promoter.
In silico sequence analyses
Similarity searches were performed using BLAST (Altschul et al. 1997) at http://www.ncbi.nlm.nih.gov/blast/Blast.cgi applying default parameters and non-redundant databases. Promoter analyses were done at http://www.dna.affrc.go.jp/PLACE/ (Higo et al. 1999) and http://www-bimas.cit.nih.gov/molbio/proscan/, on-line analyses using the ExPASy pI/Mw tool (Wilkins et al. 1999). Domain and motif analyses were performed with InterProScan (Zdobnov and Apweiler 2001) at http://www.ebi.ac.uk/interpro/, SMART at http://smart.embl-heidelberg.de/ (Schultz et al. 1998), and at http://www.predictprotein.org/ (Rost et al. 2004).
Transformation of tobacco and PCR analysis of transgenic tobacco plants
The constructs were transformed into Agrobacterium strain LBA4404 by electroporation. Tobacco leaf disks were transformed according to Curtis et al. (1995) and Horsch et al. (1985). Transformants were selected on MS medium containing antibiotics, rooted and transferred to soil. DNA was isolated using the CTAB method (Doyle and Doyle 1990). The presence of the construct was verified by PCR, applying standard conditions.
In situ and RNA gel blot hybridization analyses
In situ hybridization was done as described in Dornelas et al. (1999). Tissue sections were obtained using a microtome. Probes of a 5′-fragment (500 bp) of NtDCN1 were prepared using the DIG RNA Labelling Kit (Amersham Biosciences, now https://www2.gehealthcare.com). For RNA gel blot analyses, five μg of total RNA was loaded per lane. PCR-generated probes were labeled with [α-32P]dCTP using the RADPRIME DNA Labeling System (Invitrogen). RNA blots on Hybond-N membranes (Amersham Biosciences) were prepared and hybridized at 65 °C according to standard procedures under high stringency. Hybridization was visualized on BIOMAX MR X-ray films (Kodak, http://www.kodak.com). For expression analysis by multiple tissue Northerns, a PCR-derived fragment of DCN1, corresponding to the original SSH-derived sequence, was applied. A PCR-generated 697 bp 5′-fragment of DCN1 was used for screening endogenous mRNA levels of RNAi plants, 431 bp of the Y5 intron of the pART69 vector served as a probe for screening the ectopic expression of the RNAi construct, and GAPDH and 18S rRNA were used as internal controls for normalizing RNA loads.
Recombinant proteins
A pUC:NtDCN1 full-length clone was used as a template for PCR and the product was cloned into pGEM-4T-1 (Promega, http://www.promega.com). After sequencing and transformation into E. coli ED3, expression and harvest of recombinant NtDCN1 were done following standard procedures. For protein purification glutathione sepharose 4B (Amersham Biosciences) was used according to the manufacturer’s instructions. A rabbit antiserum was produced by Sanova Diagnostik (http://www.sanova.at). The AtCUL1:His-tag vector was kindly provided by M. Estelle. Expression and purification were performed as recommended by the Ni–NTA purification system handbook (Invitrogen).
Protein gel blot analyses
Proteins from 100 mg of plant tissue were isolated by standard protocols. The protein content was measured with a Bradford assay (BioRad, http://www.bio-rad.com). Ten to twenty μg of protein was separated in 10–12.5 % SDS-PAGE. Equal protein load on nitrocellulose membrane was verified by Ponceau S staining after transfer or Coomassie-staining of the gel. Membranes were blocked in PBS-T with 5 % milkpowder and incubated with Protein A purified polyclonal rabbit antiserum against recombinant NtDCN1 (1:2,500). AP-labeled anti-rabbit IgG (1:5,000, Sigma-Aldrich, http://www.sigma.com) and CDPStar reagent (Amersham Biosciences) were used for photodetection on HYPERFILM ECL (Amersham Biosciences).
Ubiquitin/NEDD8 binding assay and pull-down
The binding assay was performed with ubiquitin-agarose (Sigma-Aldrich) and NEDD8-agarose (Boston Biochem. http://www.bostonbiochem.com), essentially as described in Kurz et al. (2005). Initial binding was performed with 120 μg recombinant NtDCN1 and 5 μl (50 μg) of agarose-immobilized ubiquitin (7–15 μg/μl) or NEDD8 (5 μg/μl). Aliquots of starting reaction, washes, and eluates were separated using 12.5 % SDS-PAGE. Western detection was done as described above. For the pull-down, recombinant AtCUL1:His protein (kindly provided by Mark Estelle) and recombinant NtDCN1 were incubated with Ni–NTA beads (Qiagen, http://www1.qiagen.com). All washing and elution steps were performed according to user’s instructions.
Accession numbers
Sequence data from this article can be found in the EMBL/GenBank data libraries under accession numbers DQ885939 (DCN1A), FJ976682 (DCN1B) and DQ885938 (promoter).
References
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucl Acids Res 25(17):3389–3402
Benito Moreno RM, Macke F, Alwen A, Heberle-Bors E (1988) In situ seed production after pollination with in vitro matured, isolated tobacco pollen. Planta 176:145–148
Biswas KK, Ooura C, Higuchi K, Miyazaki Y, Van Nguyen V, Rahman A, Uchimiya H, Kiyosue T, Koshiba T, Tanaka A, Narumi I, Oono Y (2007) Genetic characterization of mutants resistant to the antiauxin p-chlorophenoxyisobutyric acid reveals that AAR3, a gene encoding a DCN1-like protein, regulates responses to the synthetic auxin 2,4-dichlorophenoxyacetic acid in Arabidopsis roots. Plant Physiol 145(3):773–785
Bostick M, Lochhead SR, Honda A, Palmer S, Callis J (2004) Related to ubiquitin 1 and 2 are redundant and essential and regulate vegetative growth, auxin signaling, and ethylene production in Arabidopsis. Plant Cell 16(9):2418–2432
Bosu DR, Kipreos ET (2008) Cullin-RING ubiquitin ligases: global regulation and activation cycles. Cell Div 3:7
Broderick SR, Golas BJ, Pham D, Towe CW, Talbot SG, Kaufman A, Bains S, Huryn LA, Yonekawa Y, Carlson D, Hambardzumyan D, Ramanathan Y, Singh B (2010) SCCRO promotes glioma formation and malignant progression in mice. Neoplasia 12(6):476–484
Capron A, Serralbo O, Fulop K, Frugier F, Parmentier Y, Dong A, Lecureuil A, Guerche P, Kondorosi E, Scheres B, Genschik P (2003) The Arabidopsis anaphase-promoting complex or cyclosome: molecular and genetic characterization of the APC2 subunit. Plant Cell 15(10):2370–2382
Certal AC, Almeida RB, Carvalho LM, Wong E, Moreno N, et al (2008) Exclusion of a proton ATPase from the apical membrane is associated with cell polarity and tip growth in Nicotiana tabacum pollen tubes. Plant Cell 20:614–634
Cheung AY, Wu H-M, Di Stilio V, Glaven R, Chen C, et al (2000) Pollen-pistil interactions in Nicotiana tabacum. Ann Bot 85:29–37
Curtis IS, Davey MR, Power JB (1995) Leaf disk transformation. Methods Mol Biol 44:59–70
Dai S, Wang T, Yan X, Chen S (2007) Proteomics of pollen development and germination. J Proteome Res 6(12):4556–4563
del Pozo JC, Estelle M (1999) The Arabidopsis cullin AtCUL1 is modified by the ubiquitin-related protein RUB1. Proc Natl Acad Sci USA 96(26):15342–15347
del Pozo JC, Dharmasiri S, Hellmann H, Walker L, Gray WM, Estelle M (2002) AXR1-ECR1-dependent conjugation of RUB1 to the Arabidopsis Cullin AtCUL1 is required for auxin response. Plant Cell 14(2):421–433
Dharmasiri S, Dharmasiri N, Hellmann H, Estelle M (2003) The RUB/Nedd8 conjugation pathway is required for early development in Arabidopsis. EMBO J 22(8):1762–1770
Doelling JH, Phillips AR, Soyler-Ogretim G, Wise J, Chandler J, Callis J, Otegui MS, Vierstra RD (2007) The ubiquitin-specific protease subfamily UBP3/UBP4 is essential for pollen development and transmission in Arabidopsis. Plant Physiol 145(3):801–813
Dornelas MC, Wittich P, von Recklinghausen I, van Lammeren A, Kreis M (1999) Characterization of three novel members of the Arabidopsis SHAGGY-related protein kinase (ASK) multigene family. Plant Mol Biol 39(1):137–147
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Dreher K, Callis J (2007) Ubiquitin, hormones and biotic stress in plants. Ann Bot (Lond) 99(5):787–822
Duda DM, Scott DC, Calabrese MF, Zimmerman ES, Zheng N, Schulman BA (2011) Structural regulation of cullin-RING ubiquitin ligase complexes. Curr Opin Struct Biol 21(2):257–264. doi:10.1016/j.sbi.2011.01.003
Fan YF, Jiang L, Gong HQ, Liu CM (2008) Sexual reproduction in higher plants I: fertilization and the initiation of zygotic program. J Integr Plant Biol 50(7):860–867
Feys B, Benedetti CE, Penfold CN, Turner JG (1994) Arabidopsis mutants selected for resistance to the phytotoxin coronatine are male sterile, insensitive to methyl jasmonate, and resistant to a bacterial pathogen. Plant Cell 6(5):751–759
Figueroa P, Gusmaroli G, Serino G, Habashi J, Ma L, Shen Y, Feng S, Bostick M, Callis J, Hellmann H, Deng XW (2005) Arabidopsis has two redundant Cullin3 proteins that are essential for embryo development and that interact with RBX1 and BTB proteins to form multisubunit E3 ubiquitin ligase complexes in vivo. Plant Cell 17(4):1180–1195
Forster BP, Heberle-Bors E, Kasha KJ, Touraev A (2007) The resurgence of haploids in higher plants. Trends Plant Sci 12(8):368–375
Garrido D, Eller N, Heberle-Bors E, Vicente O (1993) De novo transcription of specific mRNAs during the induction of tobacco pollen embryogenesis. Sex Plant Reprod 6:40–45
Gleave AP (1992) A versatile binary vector system with a T-DNA organisational structure conducive to efficient integration of cloned DNA into the plant genome. Plant Mol Biol 20(6):1203–1207
Gray WM, Hellmann H, Dharmasiri S, Estelle M (2002) Role of the Arabidopsis RING-H2 protein RBX1 in RUB modification and SCF function. Plant Cell 14(9):2137–2144
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database: 1999. Nucl Acids Res 27(1):297–300
Holmes-Davis R, Tanaka CK, Vensel WH, Hurkman WJ, McCormick S (2005) Proteome mapping of mature pollen of Arabidopsis thaliana. Proteomics 5(18):4864–4884
Honys D, Twell D (2004) Transcriptome analysis of haploid male gametophyte development in Arabidopsis. Genome Biol 5(11):R85
Horsch RB, Fry JE, Hoffmann NL, Eichholtz D, Rogers SG, Fraley RT (1985) A simple and general method for transferring genes into plants. Science 277:1229–1231
Hosp J, de Faria Maraschin S, Touraev A, Boutilier K (2007a) Functional genomics of microspore embryogenesis. Euphytica 158(3):275–285
Hosp J, Tashpulatov A, Roessner U, Barsova E, Katholnigg H, Steinborn R, Melikant B, Lukyanov S, Heberle-Bors E, Touraev A (2007b) Transcriptional and metabolic profiles of stress-induced, embryogenic tobacco microspores. Plant Mol Biol 63(1):137–149
Hotton SK, Callis J (2008) Regulation of cullin RING ligases. Annu Rev Plant Biol 59:467–489
Hua Z, Vierstra RD (2010) The cullin-RING ubiquitin-protein ligases. Annu Rev Plant Biol 62:299–334. doi:10.1146/annurev-arplant-042809-112256
Inze D, De Veylder L (2006) Cell cycle regulation in plant development. Annu Rev Genet 40:77–105
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6(13):3901–3907
Jenik PD, Gillmor CS, Lukowitz W (2007) Embryonic patterning in Arabidopsis thaliana. Annu Rev Cell Dev Biol 23:207–236
Joosen R, Cordewener J, Supena ED, Vorst O, Lammers M, Maliepaard C, Zeilmaker T, Miki B, America T, Custers J, Boutilier K (2007) Combined transcriptome and proteome analysis identifies pathways and markers associated with the establishment of rapeseed microspore-derived embryo development. Plant Physiol 144(1):155–172
Kaufman AJ, Kim A, Ryan R, Huryn L, Conway A, Manova K, Morris P, Hunnicutt G, Ramanathan Y, Singh B (2007) 171: creation of a SCCRO (DCN-1) knockout mouse results in decreased CUL3 neddylation and male infertility. J Surg Res 137(2):224
Kim AY, Bommeljé CC, Lee BE, Yonekawa Y, Choi L, Morris LG, Huang G, Kaufman A, Ryan RJ, Hao B, Ramanathan Y, Singh B (2008) SCCRO (DCUN1D1) is an essential component of the E3 complex for neddylation. J Biol Chem 283(48):33211–33220
Kurz T, Pintard L, Willis JH, Hamill DR, Gonczy P, Peter M, Bowerman B (2002) Cytoskeletal regulation by the Nedd8 ubiquitin-like protein modification pathway. Science 295(5558):1294–1298
Kurz T, Ozlu N, Rudolf F, O’Rourke SM, Luke B, Hofmann K, Hyman AA, Bowerman B, Peter M (2005) The conserved protein DCN-1/Dcn1p is required for cullin neddylation in C. elegans and S. cerevisiae. Nature 435(7046):1257–1261
Kurz T, Chou YC, Willems AR, Meyer-Schaller N, Hecht ML, Tyers M, Peter M, Sicheri F (2008) Dcn1 functions as a scaffold-type E3 ligase for cullin neddylation. Mol Cell 29(1):23–35
Lammer D, Mathias N, Laplaza JM, Jiang W, Liu Y, Callis J, Goebl M, Estelle M (1998) Modification of yeast Cdc53p by the ubiquitin-related protein rub1p affects function of the SCFCdc4 complex. Genes Dev 12(7):914–926
Lechner E, Achard P, Vansiri A, Potuschak T, Genschik P (2006) F-box proteins everywhere. Curr Opin Plant Biol 9(6):631–638
Leck YC, Choo YY, Tan CY, Smith PG, Hagen T (2010) Biochemical and cellular effects of inhibiting Nedd8 conjugation. Biochem Biophys Res Commun 398(3):588–593. doi:10.1016/j.bbrc.2010.06.128
Lincoln C, Britton JH, Estelle M (1990) Growth and development of the axr1 mutants of Arabidopsis. Plant Cell 2(11):1071–1080
Liu J, Zhang Y, Qin G, Tsuge T, Sakaguchi N, Luo G, Sun K, Shi D, Aki S, Zheng N, Aoyama T, Oka A, Yang W, Umeda M, Xie Q, Gu H, Qu LJ (2008) Targeted degradation of the cyclin-dependent kinase inhibitor ICK4/KRP6 by RING-type E3 ligases is essential for mitotic cell cycle progression during Arabidopsis gametogenesis. Plant Cell 20(6):1538–1554
Lyapina S, Cope G, Shevchenko A, Serino G, Tsuge T, Zhou C, Wolf DA, Wei N, Deshaies RJ (2001) Promotion of NEDD-CUL1 conjugate cleavage by COP9 signalosome. Science 292(5520):1382–1385
Ma H (2005) Molecular genetic analyses of microsporogenesis and microgametogenesis in flowering plants. Annu Rev Plant Biol 56:393–434
Malik MR, Wang F, Dirpaul JM, Zhou N, Polowick PL, Ferrie AM, Krochko JE (2007) Transcript profiling and identification of molecular markers for early microspore embryogenesis in Brassica napus. Plant Physiol 144(1):134–154
Maraschin SF, de Priester W, Spaink HP, Wang M (2005) Androgenic switch: an example of plant embryogenesis from the male gametophyte perspective. J Exp Bot 56(417):1711–1726
McCormick S (2004) Control of male gametophyte development. Plant Cell 16(Suppl):S142–S153
Mette MF, Aufsatz W, van der Winden J, Matzke MA, Matzke AJ (2000) Transcriptional silencing and promoter methylation triggered by double-stranded RNA. EMBO J 19(19):5194–5201
Meyer-Schaller N, Chou YC, Sumara I, Martin DD, Kurz T, Katheder N, Hofmann K, Berthiaume LG, Sicheri F, Peter M (2009) The human Dcn1-like protein DCNL3 promotes Cul3 neddylation at membranes. Proc Natl Acad Sci U S A 106: 12365–12370
Noir S, Brautigam A, Colby T, Schmidt J, Panstruga R (2005) A reference map of the Arabidopsis thaliana mature pollen proteome. Biochem Biophys Res Commun 337(4):1257–1266. doi:10.1016/j.bbrc.2005.09.185
Parry G, Estelle M (2004) Regulation of cullin-based ubiquitin ligases by the Nedd8/RUB ubiquitin-like proteins. Semin Cell Dev Biol 15(2):221–229
Qiao H, Wang H, Zhao L, Zhou J, Huang J, Zhang Y, Xue Y (2004) The F-box protein AhSLF-S2 physically interacts with S-RNases that may be inhibited by the ubiquitin/26S proteasome pathway of protein degradation during compatible pollination in Antirrhinum. Plant Cell 16(3):582–595
Rabut G, Peter M (2008) Function and regulation of protein neddylation. Protein modifications: beyond the usual suspects’ review series. EMBO Rep 9(10):969–976
Rost B, Yachdav G, Liu J (2004) The predictprotein server. Nucl Acids Res 32(Web Server issue):W321–326
Santner A, Estelle M (2010) The ubiquitin-proteasome system regulates plant hormone signaling. Plant J 61:1029–1040
Schultz J, Milpetz F, Bork P, Ponting CP (1998) SMART, a simple modular architecture research tool: identification of signaling domains. Proc Natl Acad Sci USA 95(11):5857–5864
Schwechheimer C, Calderon Villalobos LI (2004) Cullin-containing E3 ubiquitin ligases in plant development. Curr Opin Plant Biol 7(6):677–686
Scoccianti V, Speranza A, Crinelli R, Calzoni GL, Biasi R, Altamura MM, Bagni N (1999) Development-related changes of protein ubiquitination in pollen from male and female kiwifruit (Actinidia deliciosa). Physiol Plant 107:128–135
Scott DC, Monda JK, Grace CR, Duda DM, Kriwacki RW, Kurz T, Schulman BA (2010) A dual E3 mechanism for Rub1 ligation to Cdc53. Mol Cell 39(5):784–796. doi:10.1016/j.molcel.2010.08.030
Shariatpanahi ME, Bal U, Heberle-Bors E, Touraev A (2006) Stresses applied for the re-programming of plant microspores towards in vitro embryogenesis. Physiol Plant 127(4):519–534
Shen WH, Parmentier Y, Hellmann H, Lechner E, Dong A, Masson J, Granier F, Lepiniec L, Estelle M, Genschik P (2002) Null mutation of AtCUL1 causes arrest in early embryogenesis in Arabidopsis. Mol Biol Cell 13(6):1916–1928
Sheng X, Hu Z, Lu H, Wang X, Baluska F, Samaj J, Lin J (2006) Roles of the ubiquitin/proteasome pathway in pollen tube growth with emphasis on MG132-induced alterations in ultrastructure, cytoskeleton, and cell wall components. Plant Physiol 141(4):1578–1590
Sijen T, Vijn I, Rebocho A, van Blokland R, Roelofs D, Mol JN, Kooter JM (2001) Transcriptional and posttranscriptional gene silencing are mechanistically related. Curr Biol 11(6):436–440
Smith NA, Singh SP, Wang MB, Stoutjesdijk PA, Green AG, Waterhouse PM (2000) Total silencing by intron-spliced hairpin RNAs. Nature 407(6802):319–320
Takahashi K, Yamanaka S (2006) Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126(4):663–676
Tateishi K, Omata M, Tanaka K, Chiba T (2001) The NEDD8 system is essential for cell cycle progression and morphogenetic pathway in mice. J Cell Biol 155(4):571–579
Thomann A, Brukhin V, Dieterle M, Gheyeselinck J, Vantard M, Grossniklaus U, Genschik P (2005) Arabidopsis CUL3A and CUL3B genes are essential for normal embryogenesis. Plant J 43(3):437–448
Touraev A, Heberle-Bors E (1999) Microspore embryogenesis and in vitro pollen maturation in tobacco. Methods Mol Biol 111:281–291
Touraev A, Lezin F, Heberle-Bors E, Vicente O (1995) Maintenance of gametophytic development after symmetrical division in tobacco microspore culture. Sex Plant Reprod 8:70–76
Touraev A, Vicente O, Heberle-Bors E (1997) Initiation of microspore embryogenesis by stress. Trends Plant Sci 2:285–303
Touraev A, Ilham A, Vicente O, Heberle-Bors E (1999) Stress-induced microspore embryogenesis in tobacco: an optimized system for molecular studies. Plant Cell Rep 15:561–565
Touraev A, Pfosser M, Heberle-Bors E (2001) The microspore: a haploid multipurpose cell. Adv Bot Res 35:53–109
Vierstra RD (2009) The ubiquitin-26S proteasome system at the nexus of plant biology. Nat Rev Mol Cell Biol 10(6):385–397. doi:10.1038/nrm2688
Vodermaier HC (2004) APC/C and SCF: controlling each other and the cell cycle. Curr Biol 14(18):R787–R796
Watson IR, Irwin MS, Ohh M (2011) NEDD8 pathways in cancer. Sine Quibus Non Cancer Cell 19(2):168–176. doi:10.1016/j.ccr.2011.01.002
Wilde H, Nelson W, Booij H, de Vries S, Thomas TL (1988) Gene-expression programs in embryonic and non-embryonic carrot cultures. Planta 176:205–211
Wilkins MR, Gasteiger E, Bairoch A, Sanchez JC, Williams KL, Appel RD, Hochstrasser DF (1999) Protein identification and analysis tools in the ExPASy server. Methods Mol Biol 112:531–552
Xing S, Zachgo S (2007) Pollen lethality: a phenomenon in Arabidopsis RNA interference plants. Plant Physiol 145:330–333
Xu L, Liu F, Lechner E, Genschik P, Crosby WL, Ma H, Peng W, Huang D, Xie D (2002) The SCF(COI1) ubiquitin-ligase complexes are required for jasmonate response in Arabidopsis. Plant Cell 14(8):1919–1935
Yadegari R, Drews GN (2004) Female gametophyte development. Plant Cell 16(Suppl):S133–S141
Yang M, Hu Y, Lodhi M, McCombie WR, Ma H (1999) The Arabidopsis SKP1-LIKE1 gene is essential for male meiosis and may control homologue separation. Proc Natl Acad Sci USA 96(20):11416–11421
Yang X, Zhou J, Sun L, Wei Z, Gao J, Gong W, Xu RM, Rao Z, Liu Y (2007) Structural basis for the function of DCN-1 in protein Neddylation. J Biol Chem 282(34):24490–24494
Ylstra B, Muskens M, Van Tunen AJ (1996) Flavonols are not essential for fertilization in Arabidopsis thaliana. Plant Mol Biol 32:1155–1158
Zdobnov EM, Apweiler R (2001) InterProScan—an integration platform for the signature-recognition methods in InterPro. Bioinformatics 17(9):847–848
Zhao J, Morozova N, Williams L, Libs L, Avivi Y, Grafi G (2001) Two phases of chromatin decondensation during dedifferentiation of plant cells: distinction between competence for cell fate switch and a commitment for S phase. J Biol Chem 276(25):22772–22778
Acknowledgments
We thank Kristina Belogradova and Svetlana Akimcheva for excellent management of transgenic lines, and Maria Kalyna for assistance with DIC and GUS assays. We are grateful to Ortrun Mittelsten Scheid and Barbara Hohn for commenting on the manuscript, to Marjori Matzke for helpful remarks, and Irina Sadovnik and Håvard Nyhagen Henriksen for critical reading. Fatima Touraeva and Maria Granilshikova are gratefully acknowledged for plant care. Mark Estelle kindly provided the AtCul1-His-tag construct. J.H. was supported by a DOC scholarship granted from the Austrian Academy of Sciences.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by X. S. Zhang.
J. Hosp and A. Ribarits made equal contributions.
Electronic supplementary material
Below is the link to the electronic supplementary material.
299_2014_1609_MOESM1_ESM.jpg
Online Resource 1 Pull-down of Recombinant NtDCN1 with At CUL1:His Bound to Ni–NTA Beads. Aliquots of wash fractions (W1, W2) and eluates (1-4) were subjected to PAGE and Coomassie staining. The band at 86 kDa in fractions 1 to 3 corresponds to co-eluting His:AtCul1 while DCN1 manifests above 26 kDa. (JPEG 1488 kb)
299_2014_1609_MOESM2_ESM.tif
Online Resource 2 Expression of NtDCN1 in Microspores of NtDCN1 RNAi Plants. Western blot with protein isolated from wt and RNAi line 25, respectively; LU late unicellular microspores, str stressed microspores, gl globular embryos, T/C torpedo/cotyledon stage embryos. (TIFF 406 kb)
299_2014_1609_MOESM4_ESM.jpg
Online Resource 4 Hyper-Methylation of the 35S Promoter during Transcriptional Silencing of the NtDCN1 RNAi Construct in “Knock-Up” Double Transformants. (a) Schemes of the 35S promoter region of RNAi and promoter-silencing constructs. Black bars indicate the localisation of the probe used for Southern blotting; M/H indicates restriction sites for the methylation-sensitive enzyme HpaII (the isoschizomer of methylation-insensitive MspI); the schemes are not drawn to scale. (b) Southern blot performed with HpaII-digested DNA of NtDCN1 RNAi lines 35 and 47 and derived retransformed, promoter-silenced lines (35PS1, 47PS1, 47PS2, 47PS3); plasmids of the RNAi construct (RNAi) and the 35S TGS (ProSi) construct with partially digested DNA, respectively, are shown as size control.(JPEG 217 kb)
Rights and permissions
About this article
Cite this article
Hosp, J., Ribarits, A., Retzer, K. et al. A tobacco homolog of DCN1 is involved in pollen development and embryogenesis. Plant Cell Rep 33, 1187–1202 (2014). https://doi.org/10.1007/s00299-014-1609-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-014-1609-4