Abstract
Key message
A high-quality rice activation tagging population has been developed and screened for drought-tolerant lines using various water stress assays. One drought-tolerant line activated two rice glutamate receptor-like genes. Transgenic overexpression of the rice glutamate receptor-like genes conferred drought tolerance to rice and Arabidopsis.
Abstract
Rice (Oryza sativa) is a multi-billion dollar crop grown in more than one hundred countries, as well as a useful functional genetic tool for trait discovery. We have developed a population of more than 200,000 activation-tagged rice lines for use in forward genetic screens to identify genes that improve drought tolerance and other traits that improve yield and agronomic productivity. The population has an expected coverage of more than 90 % of rice genes. About 80 % of the lines have a single T-DNA insertion locus and this molecular feature simplifies gene identification. One of the lines identified in our screens, AH01486, exhibits improved drought tolerance. The AH01486 T-DNA locus is located in a region with two glutamate receptor-like genes. Constitutive overexpression of either glutamate receptor-like gene significantly enhances the drought tolerance of rice and Arabidopsis, thus revealing a novel function of this important gene family in plant biology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rice (Oryza sativa) is a staple food crop for more than half of the world’s population. Increasing rice productivity can significantly address the global food security challenge (Zhang 2007). In addition to being an important food crop grown in more than one hundred countries, rice is also a useful genetic tool for gene discovery due to its high throughput genetic transformation, relatively small genome size, complete genome sequence and syntenic relationship with other cereals, and its diverse genetic resources (Zhang et al. 2007; Krishnan et al. 2009). The discovery of rice traits that impart higher yield per acre and allow cultivation under biotic and abiotic stress conditions can positively impact global food production.
Advanced genomic technologies are enabling discovery of genes that can increase crop productivity in a sustainable manner. The use of forward (phenotype to gene) and reverse (gene to phenotype) genetic approaches is an effective way to identify trait genes. Various techniques have been used to facilitate the functional analysis of rice and other plant genes, including generation of loss-of-function mutations from chemical and physical mutageneses (Krishnan et al. 2009; Satoh et al. 2010; Suzuki et al. 2010), from insertions of endogenous or exogenous DNA, such as T-DNA (Hsing et al. 2007) or from transposable elements (Park et al. 2007; Ayliffe and Pryor 2009). Compared with transposons, T-DNA insertions appear to be relatively randomly distributed in a plant’s genome (Kolesnik et al. 2004). Thus, loss-of-function populations have been good sources from which to discover rice genes, given a useful assay for the genes of interest (Ryu et al. 2004; Tani et al. 2004; An et al. 2005a, b; Krishnan et al. 2009; Tsuchida-Mayama et al. 2010). Their main limitation is that genes are masked if they act redundantly, if they are compensated for by alternative metabolic or regulatory circuits, or if they are essential for plant growth and development (Weigel et al. 2000; Tani et al. 2004). To overcome these disadvantages, gain-of-function strategies have been employed, which include activation-tagging and full-length cDNA overexpressing gene strategies (Hakata et al. 2010; Tsuchida-Mayama et al. 2010; Higuchi-Takeuchi et al. 2013). Activation tagging is a powerful approach which increases the expression of adjacent genes using transcriptional enhancers randomly inserted in the genome (Weigel et al. 2000; Gou and Li 2012). This method has been successfully used to generate mutant pools in rice (Jeong et al. 2002; An et al. 2005a; Wan et al. 2009), Arabidopsis (Pogorelko et al. 2008; Wang et al. 2012), populus (Fladung and Polak 2012; Trupiano et al. 2013), and tomato (Mathews et al. 2003). About 25,000 rice T-DNA insertional mutant lines were generated using the vector pCAS04, which has both promoter-trapping and activation-tagging functions. About 40 % of the lines had single-copy insertions, and the average T-DNA insertion number was 2.3 by Southern blot analysis (Ma et al. 2009). Over 100,000 T-DNA insertion lines were developed in japonica rice (Yi and An 2013), 107,169 insertion sites were identified in this population from 58,943 T-DNA tagged lines (An et al. 2005a; Jeong et al. 2006), and 172,500 flanking sequence tags have been reported in rice (Jung and An 2013). Recently, a transposon-based activation-tagging system has been developed (Guiderdoni and Gantet 2012; Carter et al. 2013; Harb and Pereira 2013). Activation-tagged mutant populations have proven to be a powerful genetic resource for trait discovery and for understanding plant biology (Tani et al. 2004), with populations successfully screened for seed germination (Zhao et al. 2011), plant immune responses (Bao et al. 2013), plant growth and development (Shao et al. 2012; Park et al. 2013), disease resistance (Xia et al. 2004; Grant et al. 2003), and abiotic stress tolerance (Park et al. 2011, 2013).
Abiotic stress is a primary cause of crop losses, causing average yield losses of more than 50 % for major crops (Boyer 1982; Bray et al. 2000). Among the various abiotic stresses, drought is the major factor that limits crop productivity worldwide. Exposure of plants to a water-limiting environment during developmental stages appears to activate cascades of physiological and developmental changes (Xoconostle-Cázares et al. 2010). Although many reviews on molecular mechanisms of abiotic stress responses and genetic regulatory networks of drought stress tolerance have been published (Xiong and Zhu 2001; Wang et al. 2003; Shinozaki et al. 2003; Chaves and Oliveira 2004; Vinocur and Altman 2005; Yamaguchi-Shinozaki and Shinozaki 2005; Valliyodan and Nguyen 2006), it remains a major challenge to fully understand the basic biochemical and molecular mechanisms for drought stress perception, transduction and tolerance. Genetic research has shown that drought tolerance is a quantitative trait, controlled by many genes. Molecular marker-assisted breeding has led to improved drought tolerance in crops. However, marker accuracy and breeding efficiency remain problematic (Ashraf 2010). Transgenic approaches to engineering drought tolerance in crops have made some progress (Vinocur and Altman 2005; Lawlor 2013).
Glutamate receptors exist in both animal and plant cells (Dean 2002; Davenport 2002). In animals, ionotropic glutamate receptors (iGluRs) represent a class of cation-selective, ligand-gated ion channels mediating excitatory synaptic transmission in the central nervous system (Dean 2002). Plant GLR genes were first identified from Arabidopsis where they were shown to play a role in light-mediated signal transduction (Lam et al. 1998). In the last decade, various other functions have been attributed to plant GLR genes including the regulation of root growth and branching (Liu et al. 2006), the response to aluminum (Sivaguru et al. 2003), ion transport (Kim et al. 2001; Dean 2002; Turano et al. 2002; McAinsh and Pittman 2009), and metabolic and signaling pathways (Lam et al. 1998; Tapken and Hollmann 2008). Suppression of AtGLR1.1 expression indicated that AtGLR1.1 functions as a regulator of carbon/nitrogen metabolism, a key step in ABA biosynthesis, and the control of seed germination (Kang and Turano 2003). Overexpression of the AtGLR3.1 glutamate receptor homolog led to impairment in external Ca2+-induced stomatal closure and imposed Ca2+ oscillation-induced long-term stomatal closure without affecting the anion channel activity or the Ca2+ oscillation kinetics (Cho et al. 2009). Recent studies demonstrated that AtGLR3.2, AtGLR3.3, and AtGLR3.6 attenuated wound-induced surface potential changes (Mousavi et al. 2013), and the loss of function of certain members of the glutamate receptor-like family of ion-channel proteins affected wound-induced signal generation (Christmann and Grill 2013). A new family of 13 SlGLR genes has been identified in tomato and represents the first comprehensive list of GLRs in a plant other than Arabidopsis (Aouini et al. 2012). Li et al. (2006) identified a rice GLR gene which may be involved in cell proliferation and cell survival in the root apical meristem at an early seedling stage. In summary, GLR-like genes have multiple functions in plants; no clear role in water stress tolerance has been attributed to rice GLR proteins.
In this study, we report the construction of a population of >200,000 rice activation-tagged lines with advanced molecular features. By screening the population using greenhouse assays, we identified a tagged line exhibiting enhanced drought tolerance. We demonstrated that the drought tolerance phenotype was caused by activation of a GLR gene, and that overexpression of either GLR gene, OsGLR1 and OsGLR2, significantly improved drought tolerance in both rice and Arabidopsis. These results demonstrate that GLR genes play important roles in plant abiotic stress biology.
Materials and methods
Plant materials and transformation
Four japonica (Oryza sativa ssp. japonica) varieties, Zhonghua 11, Chaoyou 1, Taichong 65 and Nipponbare, were used for the creation of a rice activation-tagging population using Agrobacterium-mediated transformation, with the majority of the lines from Zhonghua 11. The transgenic events for trait recapitulation were generated using Zhonghua 11. A 16.3-Kb T-DNA-based binary construct, as shown in Fig. 1, was used to generate the activation-tagging population, which contains four tandem copies of the enhancer of Cauliflower mosaic virus (CaMV) 35S promoter (corresponding to sequences from −341 to −64, as defined by Odell et al. 1985). The binary vector (Construct A) was created by modifying the activation tagging vector of Arabidopsis, pHSbarENDs (Aukerman and Sakai 2003), in which pNOS-bar-NOS terminator was replaced by the cassette containing LTP2-DsRed2-PIN terminator–UBI-HygR-NOS terminator. The final construct contained vector sequences (pUC9) and a poly-linker for plasmid rescue, kanamycin resistance for transforming Agrobacterium, DsRed for selecting transgenic seeds, ampicillin resistance for plasmid rescue of the T-DNA-flanking sequences, and hygromycin resistance for selecting transgenic plants for screening. Also, the Ds maize transposal element was embedded in the vector to separate 35S enhancer elements from promoters in T-DNA, maximizing the chance of enhancing promoters in the vicinity of T-DNA insertion sites.
Functional regions of T-DNA in construct A which was used to generate the rice activation-tagged population. Arrowheads indicate direction of transcripts. RB and LB indicate the right border and left border of T-DNA, respectively. Four tandem copies of the CaMV 35S enhancer were inserted near to the RB. LTP2 barley lipid transfer protein 2 promoter; DsRed Discosoma red fluorescent protein gene; TPin Pin2 terminator; UBI maize ubiquitin promoter; HYG hygromycin resistance gene; TNos Nos terminator; Ds maize transposal element; pUC9 pUC9 plasmid skeleton including ampicillin resistance gene
Rice transformation essentially followed the Agrobacterium-mediated method as described by Lin and Zhang (2005), except for the use of Agrobacterium tumefaciens strain AGL0 and that the concentrations of the Agrobacterium-harboring plasmid constructs for callus infection were OD600 <0.1. The transgenic seedlings (T0) were rescued and T1 seeds were further advanced to the T2 generation in the field, and then stored in 4 °C.
Arabidopsis thaliana ecotype Columbia was grown in a growth chamber for 1 week under 8-h light/16-h dark and then for 3 weeks at 10-h light/14-h dark at 22 °C day/20 °C night and 70 % relative humidity conditions. Arabidopsis was transformed using well-known standard methodology as described by Clough and Bent (1998).
Genetic analysis of T-DNA insertion loci number
A total of 986 random tagging lines were used to estimate the DsRed segregation ratio and T-DNA insertion locus number. Two hundred seeds from each line were used in the experiments and the DsRed+ and DsRed– seed numbers were counted under green fluorescent lights. Statistical method \( \chi_{\text{C}}^{2} = \mathop \sum \nolimits \frac{{(\left| {{\text{observed}} - {\text{expected}}} \right| - 1/2)^{2} }}{\text{expected}} \) was used to test whether the DsRed seed segregation ratio (DsRed+:DsRed–, r = A/a) of each line was 3:1 or 15:1. If \( \chi_{\text{C}}^{2} < \chi_{0.05,1}^{2} = 3.8415 \), if was 2.2 ≤ A/a ≤ 4.3 or 9.1 ≤ A/a ≤ 36.8; we accepted that the segregation fitted the ratio of 3:1 or 15:1 at 0.05 levels when n = A + a = 200. According to the law of segregation and independent assortment, if the segregation ratio was 3:1 or 15:1, the T-DNA insertion loci number was 1 or 2, respectively.
Greenhouse drought tolerance assay
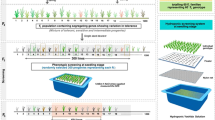
Transgenic T2 (or T1) seeds of the tagging lines which were red under green fluorescent lights were used for greenhouse drought tolerance screens. Seeds were sterilized with 800 ppm carbendazol for 8 h at 32 °C and washed 3–5 times, soaked in water for 16 h at 32 °C, and then germinated overnight at 35–37 °C in incubator. The germinated seeds were sowed in a tray filled with a mixture of organic soil (FangJie soil from Beijing HuiYeShengDa Center), vermiculite (Beijing QingYuanShiJi Garden Center) and sand (Beijing Shuntun Construction Material Market) (V:V:V = 3:3:2). Twenty uniform seedlings from each line, one positive control (a drought-tolerant variety, Mianhui 501), one negative control (a drought-sensitive variety, Dongbeiyin 2), and one tissue-cultured Zhonghua11line were included in the drought tolerance screens. The seedlings were grown under normal greenhouse conditions (sodium lamp and metal halide lamp at 1:1 ratio to provide long days of 16-h/8-h light/dark, 20–35 °C, and 30–90 % relative humidity) in an IRRI solution (Gregorio et al. 1997). For each line, 20 plants were grown in different positions of the tray. After the seedlings grew to the 3-leaf stage, watering was stopped until the leaves became curved (approximately 9–15 days, depending on the season). Water was added below the trays to enable seedling recovery in 5–7 days, and the plants were scored for their recovery degree. The following scoring system was used: if more than half of a stem was green = 1; if more than two-thirds of a leaf was green = 1; if less than two-thirds but more than one-third of a leaf was green = 0.5; if less than one-third of a leaf was green = 0.2; if no part of a leaf or less than half of a stem was green = 0. The recovery degree was the sum of the scores of the green tissues of a plant, and the data were statistically analyzed using SAS PROC MIXED. The lines, which showed significant differences from the controls (p < 0.05) and other mutant plants, were considered to be the primary positive lines. The primary positives were screened again with 32 seedlings. In the third screen, 60 seedlings were included in the experiments under the same condition as the primary screen. The screening data were analyzed using SAS PROC MIXED (p < 0.05).
Survival rate (percentage of surviving plants over the total plant number) was also used as a parameter in identifying drought-tolerant tagged lines. Only the lines that passed the third round of screens were considered drought-tolerant lines and were moved into the candidate gene cloning step for recapitulating the trait. The drought tolerance assay for gene recapitulation was carried out similar to the screening of the tagging lines. Nine to 12 events from each construct and 8–16 seedlings from each event at T1 or T2 generations were tested along with the transgenic events from control vector (DP0005 or DP0009) or event null. The results were analyzed by SAS PROC MIXED.
In the Arabidopsis drought assays, T1 seeds were sowed in planting soil under normal watering conditions for 4 weeks. The planting soil was watered to saturation before the drought treatment, and the Arabidopsis plants grew without watering for about 13 days. When the relative water content of the soil had decreased to about 3–4 %, the Arabidopsis plants were re-watered for 4 days, and the plant survival rate then measured. The results were analyzed by SAS PROC MIXED.
T-DNA-flanking sequence and identification of activation-tagged genes
Genomic DNA was isolated from 3-week-old seedling leaf tissues of the tagging lines (AH01486 and other tagging lines) using the CTAB method (Murray and Thompson 1980) to identify the T-DNA flanking sequence by the plasmid rescue method (Behringer and Medford 1992). Ten microgram of genomic DNA was digested overnight with 2 μL each of BglII/HindIII/XhoI (NEB, Ipswich, MA) in a 200-μl reaction solution. The digested genomic DNA was self-ligated overnight at 16 °C in a 400-μl reaction solution. The self-ligation sample was transformed into competent E. coli DH5α via electroporation. The rescued clones were sequenced using the primer of M13R-46 and the right border-flanking sequences of the T-DNA were determined from the rice genomic annotation in the US National Center for Biotechnology Information of the National Library of Medicine, National Institutes of Health, and the website for the rice genome annotation project at Michigan State University. Based on the T-DNA insertion locus information, the T-DNA insertion site on both right border (RB) and left border (LB) were defined through PCR using a pair of primer complementing the T-DNA and the genomic DNA nearby the T-DNA insertion site, respectively.
GLR gene cloning and vector constructions
To clone the cDNA of rice glutamate receptor-like 1 gene (OsGLR1, LOC_Os09g26144.1), two primers were designed based on the sequence information of LOC_Os09g26144.1. The primers were as follows: 56-26144-F: 5′-TCTGCCTGCAGGATGGCTGGTCACACCCCGAATC-3′ and 56-26144-R: 5′-TAACCTGCAGGTTACTGGCCTCTGCCATTCTCT-3′. The SbfI site was designed in both primers. PCRs were carried out using these primers and pooled cDNA from leaf, stem and root tissues of Zhonghua 11 plants as a template. The PCR product with the expected size of ~2.8 Kb was purified and then digested by SbfI. The DNA fragment was ligated with PstI-digested vector DP0009 (pCAMBIA1300-35s-NOS-Hyg-deleteAsRED). The resultant construct with the sense orientation was selected as OsGLR1 overexpression vector DP0025. The OsGLR1 gene sequences and orientation in DP0025 were confirmed by sequencing.
The rice glutamate receptor-like 2 (OsGLR2, LOC_Os09g26160.1) gene was amplified from Zhonghua11 cDNA pool using the following primers which were designed based on the sequences of LOC_Os09g26160.1. The primers were 56-26160-F: 5′-TTTAGGATCCTTGTCCATGGAAGCTG-3′ and 56-26160-R: 5′-ACTCTCGAGTTATTGCCCCCTCTCTGAATG-3′. A BamHI restriction site was added in primer 56-26160-F, and an XhoI site in primer 56-26160-R. An expected ~2.8-Kb PCR fragment was purified and digested by BamHI and XhoI. This DNA fragment was ligated with BglII-SalI-digested DP0005 (pCAMBIA1300-35s-NOS-Hyg) and the OsGLR2 overexpression vector DP0015 was obtained. The nucleotide sequences and orientation of the OsGLR2 gene in DP0015 were confirmed by sequencing.
T-DNA was inserted in the second intron of OsGLR1 in AH01486 mutant. To understand whether the first two 5′ exons of OsGLR1 in the contributed to the enhanced drought tolerance in AH01486, we cloned the 5′ genomic DNA fragment of OsGLR1 by PCR, using rice Zhonghua11 genomic DNA as a PCR template. The primers were 56-26144F: 5′-TCTGCCTGCAGGATGGCTGGTCACACCCCGAATC-3′ and 56-26144R: 5′-CGGCGTTAAACATTTTGAGATGGAGG-3′. The expected ~1.7-Kb PCR product was purified and then digested by SbfI. This DNA fragment was ligated with DP0009 which had been digested by PstI +NruI to obtain the overexpression vector DP0011. The nucleotide sequence and orientation of the 5′-part of OsGLR1 in DP0011 were confirmed by sequencing.
RT-PCR analyses
Total RNA was extracted from about 50 mg leaf, or stem, or root tissues of plants at the 4-leaf stage and cultured in IRRI solution using RNAiso Plus kit (TaKaRa) according to the manufacturer’s instructions. The cDNA was prepared by RevertAidTM First Strand cDNA Synthesis Kit (Fermentas) from 500 ng of total RNA. The real-time PCR (RT-PCR) was performed based on the normal procedure using SYBRRPremix Ex TaqTM Kit (TaKaRa) and 7500-Fast real-time PCR equipment (ABI) according to the manual. Gene expression was normalized by the EF1α mRNA levels. The primers used in the RT-PCR analysis for the OsGLR1 gene and OsGLR2 gene were as below: OsGLR1-3′RT-F: 5′-GACGCCGTACCTGAGGATCTTC-3′; OsGLR1-3′RT-R: 5′- CACTTGCGCTCGATGAGGTTC-3′; OsGLR1-5′RT-F: 5′-GTCAAGGTCCGGCAAGAAG-3′; OsGLR1-5′RT-R: 5′-CTTCCCTCTGTCCATGATGTTC-3′; OsGLR2-RT-F: 5′-CTCCTAAAGATATTCCACG-3′; OsGLR2-RT-R: 5′-ACGCTCTCCACATCACTGC-3′.
Sequence alignments
Alignments of the DNA and protein sequences were performed using the Clustal V method (Higgins and Sharp 1989) with the default parameters (GAP PENALTY = 10, GAP LENGTH PENALTY = 10). Default parameters for pairwise alignments and calculation of percent identity of amino acid sequences using the Clustal V method are KTUPLE = 1, GAP PENALTY = 3, WINDOW = 5 and DIAGONALS SAVED = 5. For nucleic acids, these parameters are KTUPLE = 2, GAP PENALTY = 5, WINDOW = 4 and DIAGONALS SAVED = 4.
Results
The rice activation-tagging population
Several laboratories have developed rice activation-tagging populations and demonstrated the utility of the tagging technology for gene discovery (Jeong et al. 2002; Tani et al. 2004; An et al. 2005a, b; Krishnan et al. 2009; Ma et al. 2009). We have developed a Japonica population of more than 200,000 activation-tagging lines (ATLs) using a modified T-DNA which contains tetramerized CaMV 35S enhancers near the RB (Fig. 1). The number of insertion lines statistically required to tag every rice gene was estimated to be between 180,689 and 460,000 (Hirochika et al. 2004). By RT-PCR analyses of a number of tagged lines, we observed that the tetramerized 35S enhancers as present in the tested activation-tagging construct A could activate rice genes within a region 23 Kb from the RB and 17 Kb from the LB, and that, on average, this region (40 Kb) contains 4–5 rice genes. The activated gene expression levels depend upon the genes, their locations, and the tissues (data not shown). We have observed more than 80 types of visible phenotypes associated with root, shoot, leaf, panicle, etc. as indicated in Fig. 2. These phenotypes can be resulted from gene activation or knock-off by the inserted T-DNA or transformation-caused mutations. We are in the process of confirming the phenotypes and studying if the phenotypes are linked with T-DNA insertion. We have demonstrated that a semi-sterile mutant (Fig. 2e) dominantly co-segregated with the T-DNA insertion. These results confirm that the CaMV 35S enhancers in the tested construct configuration can function in rice tissues and activate genes located bi-directionally from the enhancers, within a large genomic region. Based on the present rice genomic data, this rice tagging population is estimated to cover more than 90 % of the rice genes.
The pUC9 plasmid construction in the T-DNA is useful for plasmid rescue of the T-DNA flanking sequences as indicated in the “Materials and methods”. The color marker (DsRed gene) is useful for identifying the transgenic tissues and seeds. We selected the transgenic seeds under a green fluorescent light in experiments, and this non-destructive process was significantly simple and cost-effective compared to using antibiotics or molecular techniques to identify transgenic seeds.
Molecular features of the population
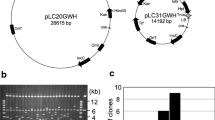
Segregation of the color marker can be used to estimate the number of T-DNA loci in the genome. We analyzed DsRed-positive and DsRed-negative seeds of pooled T1 seeds from 986 ATLs which showed that overexpressed DsRed could be assigned as a qualitative trait. The statistical analysis indicates that about 74 % (14.81 + 59.13 %) of the tagged lines segregate as a single T-DNA insertion locus (p < 0.05), 15 % of the lines have two T-DNA loci, and only 3 % of the lines have more than three T-DNA insertion loci. About 8 % of the lines have either one or two T-DNA insertion loci (Fig. 3a). Plasmid rescue and sequence analyses of 172 tagging lines show that 80 % of the 172 lines harbor only a single T-DNA insertion in the genome, 16 % of the lines have two loci, and less than 4 % of the lines have three or more T-DNA insertion loci (Fig. 3b). Thus, the majority of the tagged lines have only one or two T-DNA insertion loci. This is a significant attribute of population since the presence of single T-DNA insertions simplifies the identification of candidate genes.
The number of T-DNA insertion loci in rice activation tagging population. a Statistical analysis of DsRed-positive and DsRed-negative seeds from 986 tagging lines. Two hundred seeds from each line were counted and analyzed by Latin-square model. b T-DNA insertion numbers of 172 tagging lines analyzed by plasmid rescue and sequence analysis. (0, 2.2), 0 and <2.2; [2.2, 4.3), ≥2.2 and ≤4.3; (4.3, 9.1], >4.3 and ≤9.1; (9.1, 36.8], >9.1 and ≤36.8; (36.8, 199], >36.8 and ≤199
To further characterize the molecular features of the population, we analyzed the locations of the T-DNA insertion sites in the rice genome. The 172 tagging lines harbor a total of 213 T-DNA insertion sites which are randomly distributed across the 12 rice chromosomes (data not shown). As shown in Table 1, 59 % of the T-DNAs had been inserted in promoter or terminator regions, and more than 19 % of the T-DNA had been inserted between genes; these are advantageous locations for the gain-of-function mechanism of enhancer-based activation tagging. Notably, about 21 % of the T-DNAs had been inserted into the coding region, inclusive of introns and exons. These types of insertions may also cause loss-of-function and provide a knock-out approach to identifying gene function.
Line AH01486 exhibited significantly enhanced drought tolerance under greenhouse conditions
AH01486 was identified in greenhouse drought tolerance screens. This line showed a rapid and dramatic recovery in 10 of 13 plants (77 %) after reintroduction of water following water stress. In contrast, only 33 % of the tissue-cultured Zhonghua 11 control plants survived this treatment. These results indicate that AH01486 plants have significantly enhanced tolerance to water deficit when compared with control plants.
To confirm this observation, we carried out the drought assays using T2 seeds of AH01486. In the first round of screens, 85 % of AH01486 plants survived, while only 35 % of the control survived. The average recovery degree of AH01486 plants was very significantly higher than that of the tissue-cultured Zhonghua11 controls (Table 2, p < 0.01). In repeated screens under the same conditions, both the survival rate and the recovery degree of AH01486 plants were consistently and significantly enhanced for drought tolerance compared to the controls (Table 2, p < 0.05). These results demonstrate that the AH01486 plants have enhanced drought tolerance under greenhouse conditions at the seedling stage.
T-DNA insertion locus and flanked candidate genes in AH01486
To identify the gene(s) responsible for the drought tolerance phenotype in AH01486, we cloned the T-DNA-flanking fragments using plasmid rescue. The RB- and LB-flanking sequences are shown in Supplemental Fig. 1. Sequence analyses and Blast searches revealed one T-DNA insertion locus in AH01486; its RB was inserted at the 15,760,153 bp and the other border at the 15,760,233 bp of Chromosome 9 (Blasted by MUS 6.1 rice genome database) (Fig. 4). The T-DNA insertion resulted in a 79-bp deletion in rice genomic DNA (based on the rice genome database available in June–July, 2009). The T-DNA in AH01486 has a RB–LB–LB connection. The T-DNA has a 22-bp deletion at the RB and a 65-bp deletion at the 3′ end of the Ubi1 promoter at the LB. The T-DNA was inserted in the second intron of OsGLR1 (LOC_Os09g26144.1). Upstream of the RB, there are two retrotransposon genes. Downstream of the T-DNA LB, there is another glutamate receptor gene OsGLR2 (LOC_Os09g26160.1) and a MYB family transcription factor gene (Fig. 4).
Map of the insertion position of the T-DNA in AH01486 genome and the nearby genes. The right border (RB) at the 15,760,153 bp and the left border (LB) at the 15,760,233 bp of Chromosome 9, respectively, based on Blast search using MUS 6.1 rice genome database. The T-DNA was inserted into the second intron of glutamate receptor-like 1 gene (OsGLR1). OsGLR2, rice glutamate receptor-like 2 gene
A functional activation-tagged allele should result in up-regulation of the candidate gene in tissues where it is normally expressed, ectopic expression in tissues that do not normally express that gene, or both. To understand which genes are activated in AH01486, RT-PCR analyses were carried out. The semi-quantitative RT-PCR assays indicated that expression of the MYB transcription factor was unchanged (data not shown). In contrast, as shown in Fig. 5, both the OsGLR1 (5′ part) and OsGLR2 genes were significantly activated in root, stem and leaf tissues of AH01486 plants compared to the control. In leaf tissues, OsGLR1 and OsGLR2 gene expression levels were increased about 10- and 20-fold, respectively, compared to the control. Since the T-DNA was inserted into the second intron of OsGLR1, the activated expression level is from the 5′ part of OsGLR1 (the first two exons of OsGLR1). Based on these results, our phenotype recapitulation efforts focused on these two GLR genes.
Activated expression levels of OsGLR1 (a) and OsGLR2 (b) genes in various tissues of AH01486 plants measured by real-time PCR assay. Zhonghua 11-TC, tissue-cultured Zhonghua 11; L, leaves; R, roots; S, stems. The numbers on the top of the columns are the fold-increase compared with the corresponding control tissues
Both OsGLR1 and OsGLR2 genes were cloned using PCR primers based on the information in the rice genome sequence database, and using the rice cDNA pool as the template. Their DNA and protein sequences are listed in Supplemental Fig. 2, 3. The 5′-part of OsGLR1 was cloned from the genomic DNA and its sequences are shown in Supplemental Fig. 4. There is 37 % identity between OsGLR1 and OsGLR2 at the amino acid sequence level across the whole peptides, and 46 % identical amino acid residues in the conserved C-terminal domain. DP0011, DP0015, and DP0025 vectors were constructed to overexpress the 5′ part of OsGLR1, full-length of OsGLR2 and OsGLR1 under the control of the CaMV 35S promoter (Fig. 6) to understand the contribution of the genes to the improved drought tolerance of AH01486.
Both OsGLR1 and OsGLR2 conferred enhanced drought tolerance in rice
The overexpression vectors (DP0011, DP0015, and DP0025; Fig. 6) and the control vectors (DP0005 and DP0009, Supplemental Fig. 5) were transformed into Zhonghua 11. More than 60 independent transgenic events were generated for each vector. The greenhouse drought tolerance assays were carried out at the T1 and/or T2 generations. The transgenic plants were identified by hygromycin selection, and only hygromycin-resistant seedlings were used in the drought assays. Overexpression of either OsGLR1 or OsGLR2 genes in rice did not alter plant growth and development, and the transgenic plants exhibited same phenotypes as the controls before stop-watering. The greenhouse drought tolerance assays showed that there were no statistical differences among tissue-cultured Zhonghua 11 control, DP0005-transgenic events, DP0009-transgenic events, and event null (data not shown).
As shown in Tables 3 and 4, constitutive (CaMV 35S-driven) overexpression of OsGLR2 genes resulted in a significant enhancement of drought tolerance under greenhouse conditions. At the T1 stage, overexpression of the OsGLR2 gene enhanced survival rate and the average recovery degree after water deficit; 67 % of OsGLR2-transgenic rice survived the drought treatment, while only 30 % of the control survived this treatment; the average recovery degree was 1.04 for OsGLR2-transgenic rice and 0.3 for the control (Table 3). This indicates that the OsGLR2-transgenic plants have enhanced drought tolerance. The transgenic events were further evaluated at the T2 generation. The results in Table 4 and Fig. 7d show that overexpression of OsGLR2 increased drought tolerance in rice, with the improved tolerance correlated to the transgene expression level (Fig. 7b). These results demonstrate that activated OsGLR2 expression contributed to the enhanced drought tolerance of AH01486.
Transgene expression levels of OsGLR1 (a) and OsGLR2 (b) in the leaves of transgenic events measured by real-time PCR assay. The base level of expression in CK (Control, tissue-cultured Zhonghua 11) event was set at 1.00, and the numbers on the top of the columns are the fold-increase compared with the corresponding control. Phenotypes of DP0025.15 and control plants (CK) (c) and DP0015.10 (the two plants on the right side of d) and its segregated null plants (the two plants on the left side of d) 5 days after re-watering. The data in a and c were from T1 plants, and the results in b and d were from T2 plants
As shown in Table 5, the T1 OsGLR1-transgenic rice plants show a higher survival rate and average recovery degree compared to the control under moderate drought stress in the greenhouse assays. These results show that overexpression of OsGLR1 also enhanced the drought tolerance of the OsGLR1-transgenic rice plants, and the improved tolerance is correlated to the transgene expression level (Fig. 7a, c). To further understand if the new polypeptide encoded by the truncated OsGLR1 gene contributed to the enhanced drought tolerance of AH01486, the DP0011 vector was transformed into Zhonghua 11. Overexpression of this truncated OsGLR1 under CaMV 35S promoter had no effect on drought tolerance in multiple rice transgenic events (Table 6, p > 0.1). These results show that the truncated OsGLR1 did not contribute to the improved drought tolerance of AH01486.
OsGLR genes enhanced drought tolerance in Arabidopsis
Arabidopsis GLR studies demonstrated that plant GLR proteins contain “three-plus-one” transmembrane domains (M1–M4) and putative ligand-binding domains (GlnH1 and GlnH2) (Lam et al. 1998), and sequence analysis indicated that the rice OsGLR1 and OsGLR2 have the same domains as Arabidopsis homologs (data not shown). To understand if rice GLR genes could improve drought tolerance in dicot plants, vectors expressing OsGLR1 (construct DP0025) and OsGLR2 genes (construct DP0015) were transformed into Arabidopsis.
Overexpression of either OsGLR1 or OsGLR2 genes in Arabidopsis did not alter plant growth and development, and the transgenic plants exhibited same phenotypes as the controls before stop-watering. During the water withdrawal, the wilting levels of control plants (generated from construct DP0009) were more apparent than those of the AtDP0015 and AtDP0025 events, which grew better than the controls after re-watering. On the fourth day after re-watering, 30–70 % of OsGLR1- or OsGLR2-transgenic plants survived, whereas the corresponding survival rates were <15–20 % for control plants (Figs. 8a, c, 9a, c). The increased survival rates were closely related to the transgene expression levels (Figs. 8b, 9b).
Overexpression of OsGLR1 increased drought tolerance in Arabidopsis. a The survival rate of three DP0025-transgenic events and one control event. b Transgene expression levels of OsGLR1 in the transgenic leaf tissues. c Phenotypes of three DP0025-transgenic events and a control event 4 days after re-watering. Data are expressed as mean ± SD at least three independent replications (n = 3). Single asterisk and double asterisk indicate the difference between the transgenic events and the control at significant (p < 0.05) and very significant (p < 0.01) level, respectively
Overexpression of OsGLR2 increased drought tolerance in Arabidopsis. a Survival rate of three DP0015-transgenic events and one control event. b Transgene expression levels of OsGLR2 in the transgenic leaf tissues. c Phenotypes of three DP0015-transgenic events and a control event 4 days after re-watering. Data are expressed as mean ± SD at least three independent replications (n = 3). Single asterisk and double asterisk indicate the difference between the transgenic events and the control at significant (p < 0.05) and very significant (p < 0.01) level, respectively
These results are consistent with the results from rice (Tables 3, 4, 5) and demonstrate that overexpression of OsGLR1 or OsGLR2 gene can confer enhanced drought tolerance in rice and Arabidopsis. In addition, these results further show that a monocot gene (rice GLR genes) can function in a dicot plant (Arabidopsis), demonstrating the cross-validation of drought tolerance across species.
Discussion
Activation tagging has several advantages over inactivation approaches for uncovering gene functions in rice, Arabidopsis, and tomato (Tani et al. 2004). Several rice activation tagging populations have been developed (Jeong et al. 2002; Tani et al. 2004; An et al. 2005a, b; Krishnan et al. 2009). Herein, we report a japonica rice population of 200,000 ATL with several advantages. Theoretically, this size of the population covers >90 % of the rice genome if each tagged line has a single gene activated (Hirochika et al. 2004). However, we found that a single tetramer of the CaMV 35S enhancer activates 4–5 rice genes on average within a 40-Kb region. Therefore, we expect that this population is likely to cover most rice genes. This population has several advantageous features, such as 73–80 % of the tagging lines having a single T-DNA insertion locus, about a quarter of the population having a T-DNA insertion within a gene, the T-DNA being randomly distributed along the 12 rice chromosomes, and a number of lines exhibiting various visible phenotypes.
Activation tagging has been used to study plant growth and development, disease resistance gene discovery, metabolic pathway dissection, and signal transduction (Tani et al. 2004). Herein, we further demonstrated the utility of the activation-tagging technology in agronomic trait gene discovery and stress biology.
GLR proteins are found in both animal and plant cells. This gene family comprises diverse signaling pathways and processes, including excitatory synaptic transmission in the central nervous system (Dean 2002), light-signal transduction (Lam et al. 1998), and ion channel function in plants (Tapken and Hollmann 2008). We discovered that two rice GLR genes confer significantly enhanced drought tolerance in rice and Arabidopsis (Tables 3, 4, 5; Figs. 7, 8, 9). These results demonstrate that GLR genes have important roles in response to acute drought stress and in water-use efficiency in plants. Drought tolerance is a complicated trait which requires coordination of multiple responsive pathways (Xiong and Zhu 2001; Shinozaki et al. 2003; Wang et al. 2003; Chaves and Oliveira 2004; Vinocur and Altman 2005; Yamaguchi-Shinozaki and Shinozaki 2005; Valliyodan and Nguyen 2006). Although the mode of action of GLR is unknown, the role in drought tolerance may be related to regulation of carbon/nitrogen metabolism and ABA biosynthesis (Kang and Turano 2003) and Ca2+-induced stomatal closure (Cho et al. 2009). The observation that a truncated OsGLR1 gene could not impact drought tolerance in rice (Table 6) suggests an important role for the C-terminal transmembrane domain (Lam et al. 1998).
Rice transgenic events with high levels of GLR expression exhibited healthy and normal architecture (data not shown). The native rice OsGLR1 and OsGLR2 genes functioned well in Arabidopsis to improve vegetative growth performance under drought stress. These results suggest that ectopic overexpression of rice GLR genes can be used to improve agronomic performance of both monocot and dicot crops.
Abbreviations
- ATL:
-
Activation tagging line
- CaMV:
-
Cauliflower mosaic virus
- CTAB:
-
Cetyltrimethyl ammonium bromide
- CDS:
-
Coding sequence
- DsRed:
-
Discosoma red fluorescent protein
- GLR:
-
Glutamate receptor-like
- RNAi:
-
RNA interference
- RT-PCR:
-
Reverse-transcription polymerase chain reaction
- T-DNA:
-
Transfer DNA
- UTR:
-
Untranslated region
References
An G, Lee S, Kim SH, Kim SR (2005a) Molecular genetics using T-DNA in rice. Plant Cell Physiol 46:14–22
An G, Jeong DH, Jung KH, Lee S (2005b) Reverse genetic approaches for functional genomics of rice. Plant Mol Biol 59:111–123
Aouini A, Matsukura C, Ezura H, Asamizu E (2012) Characterisation of 13 glutamate receptor-like genes encoded in the tomato genome by structure, phylogeny and expression profiles. Gene 493:36–43
Ashraf M (2010) Inducing drought tolerance in plants: recent advances. Biotechnol Adv 28:169–183
Aukerman MJ, Sakai H (2003) Regulation of flowering time and floral organ identity by a microRNA and its APETALA2-like target genes. Plant Cell 15:2730–2741
Ayliffe MA, Pryor AJ (2009) Transposon-based activation tagging in cereals. Functional Plant Biol 36:915–921
Bao Z, Yang H, Hua J (2013) Perturbation of cell cycle regulation triggers plant immune response via activation of disease resistance genes. Proc Natl Acad Sci USA 110:2407–2412
Behringer FJ, Medford JI (1992) A plasmid rescue technique for the recovery of plant DNA disrupted by T-DNA insertion. Plant Mol Biol Rep 10:190–198
Boyer JS (1982) Plant productivity and environment. Science 218:443–448
Bray EA, Bailey-Serres J, Weretilnyk E (2000) Responses to abiotic stresses. In: Gruissem W, Buchannan B, Jones R (eds) Biochemistry and molecular biology of plants. American Society of Plant Physiologists, Rockville, pp 1158–1249
Carter JD, Pereira A, Dickerman AW, Veilleux RE (2013) An active Ac/Ds transposon system for activation tagging in tomato cultivar M82 using clonal propagation. Plant Physiol 162:145–156
Chaves MM, Oliveira MM (2004) Mechanisms underlying plant resilience to water deficits: prospects for water-saving agriculture. J Exp Bot 55:2365–2384
Cho D, Kim SA, Murata Y, Lee S, Jae SK, Nam HG, Kwak JM (2009) De-regulated expression of the plant glutamate receptor homolog AtGLR3.1 impairs long-term Ca2+-programmed stomatal closure. Plant J 58:437–449
Christmann A, Grill E (2013) Plant biology: electric defence. Nature 500:404–405
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Davenport R (2002) Glutamate receptors in plants. Ann Bot 90:549–557
Dean RM (2002) The structure and function of glutamate receptor ion channels. Nat Rev Neurosci 3:91–101
Fladung M, Polak O (2012) Ac/Ds-transposon activation tagging in poplar: a powerful tool for gene discovery. BMC Genomics 13:61
Gou X, Li J (2012) Activation tagging. Methods Mol Biol 876:117–133
Grant JJ, Chini A, Basu D, Loake GJ (2003) Targeted activation tagging of the Arabidopsis NBS-LRR gene, ADR1, conveys resistance to virulent pathogens. Mol Plant Microbe Interact 16:669–680
Gregorio GB, Senadhira D, Mendoza RD (1997) Screening rice for salinity tolerance. IRRI Discussion paper Series No. 22
Guiderdoni E, Gantet P (2012) Ac–Ds solutions for rice insertion mutagenesis. Methods Mol Biol 859:177–187
Hakata M, Nakamura H, Iida-Okada K, Miyao A, Kajikawa M, Imai-Toki N, Pang JH, Amano K, Horikawa A, Tsuchida-Mayama T, Song JY, Igarashi M, Kitamoto HK, Ichikawa T, Matsui M, Kikuchi S, Nagamura Y, Hirochika H, Ichikawa H (2010) Production and characterization of a large population of cDNA-overexpressing transgenic rice plants using Gateway-based full-length cDNA expression libraries. Breed Sci 60:575–585
Harb A, Pereira A (2013) Activation tagging using the maize En–I transposon system for the identification of abiotic stress resistance genes in Arabidopsis. Methods Mol Biol 1057:193–204
Higgins DG, Sharp PM (1989) Fast and sensitive multiple sequence alignments on a microcomputer. Comput Appl Biosci 5:151–153
Higuchi-Takeuchi M, Mori M, Matsui M (2013) High-throughput analysis of rice genes by means of the heterologous full-length cDNA overexpressor (FOX)-hunting system. Int J Dev Boil 57:517–523
Hirochika H, Guiderdoni E, An G, Hsing YI, Eun MY, Han CD, Upadhyaya N, Ramachandran S, Zhang Q, Pereira A, Sundaresan V, Leung H (2004) Rice mutant resources for gene discovery. Plant Mol Biol 54:325–334
Hsing YI, Chern CG, Fan MJ, Lu PC, Chen KT, Lo SF, Sun PK, Ho SL, Lee KW, Wang YC, Huang WL, Ko SS, Chen S, Chen JL, Chung CI, Lin YC, Hour AL, Wang YW, Chang YC, Tsai MW, Lin YS, Chen YC, Yen HM, Li CP, Wey CK, Tseng CS, Lai MH, Huang SC, Chen LJ, Yu SM (2007) A rice gene activation/knockout mutant resource for high throughput functional genomics. Plant Mol Biol 63:351–364
Jeong DH, An S, Kang HG, Moon S, Han JJ, Park S, Lee HS, An K, An G (2002) T-DNA insertional mutagenesis for activation tagging in rice. Plant Physiol 130:1636–1644
Jeong DH, An S, Park S, Kang HG, Park GG, Kim SR, Sim J, Kim YO, Kim MK, Kim SR, Kim J, Shin M, Jung M, An G (2006) Generation of a flanking sequence-tag database for activation-tagging lines in japonica rice. Plant J 45:123–132
Jung KH, An G (2013) Functional characterization of rice genes using a gene-indexed T-DNA insertional mutant population. Methods Mol Biol 956:57–67
Kang J, Turano FJ (2003) The putative glutamate receptor 1.1 (AtGLR1.1) functions as a regulator of carbon and nitrogen metabolism in Arabidopsis thaliana. Proc Natl Acad Sci USA 100:6872–6877
Kim SA, Kwak JM, Jae SK, Wang MH, Nam HG (2001) Overexpression of the AtGluR2 gene encoding an Arabidopsis homolog of mammalian glutamate receptors impairs calcium utilization and sensitivity to ionic stress in transgenic plants. Plant Cell Physiol 42:74–84
Kolesnik T, Szeverenyi I, Bachmann D, Kumar CS, Jiang S, Ramamoorthy R, Cai M, Ma ZG, Sundaresan V, Ramachandran S (2004) Establishing an efficient Ac/Ds tagging system in rice: large-scale analysis of Ds flanking sequences. Plant J 37:301–314
Krishnan A, Guiderdoni E, An G, Hsing YI, Han CD, Lee MC, Yu SM, Upadhyaya N, Ramachandran S, Zhang Q, Sundaresan V, Hirochika H, Leung H, Pereira A (2009) Mutant resources in rice for functional genomics of the grasses. Plant Physiol 149:165–170
Lam HM, Chiu J, Hsieh MH, Meisel L, Oliveira IC, Shin M, Coruzzi GM (1998) Glutamate receptor genes in plants. Nature 396:125–126
Lawlor DW (2013) Genetic engineering to improve plant performance under drought: physiological evaluation of achievements, limitations, and possibilities. J Exp Bot 64:83–108
Li J, Zhu S, Song X, Shen Y, Chen H, Yu J, Yi K, Liu Y, Karplus VJ, Wu P, Deng XW (2006) A rice glutamate receptor-like gene is critical for the division and survival of individual cells in the root apical meristem. Plant Cell 18:340–349
Lin YJ, Zhang QF (2005) Optimising the tissue culture conditions for high efficiency transformation of indica rice. Plant Cell Rep 23:540–547
Liu PW, Liu LH, Remans T, Tester M, Forde BG (2006) Evidence that L-glutamate can act as an exogenous signal to modulate root growth and branching in Arabidopsis thaliana. Plant Cell Physiol 47:1045–1057
Ma Y, Liu L, Zhu C, Sun C, Xu B, Fang J, Tang J, Luo A, Cao S, Li G, Qian Q, Xue Y, Chu C (2009) Molecular analysis of rice plants harboring a multi-functional T-DNA tagging system. J Genet Genomics 36:267–276
Mathews H, Clendennen SK, Caldwell CG, Liu XL, Connors K, Matheis N, Schuster DK, Menasco DJ, Wagoner W, Lightner J, Wagner DR (2003) Activation tagging in tomato identifies a transcriptional regulator of anthocyanin biosynthesis, modification, and transport. Plant Cell 15:1689–1703
McAinsh MR, Pittman JK (2009) Shaping the calcium signature. New Phytol 181:275–294
Mousavi SAR, Chauvin A, Pascaud F, Kellenberger S, Farmer EE (2013) Glutamate receptor-like genes mediate leaf-to-leaf wound signalling. Nature 500:422–426
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4326
Odell JT, Nagy F, Chua NH (1985) Identification of DNA sequences required for activity of the cauliflower mosaic virus 35S promoter. Nature 313:810–812
Park SH, Jun NS, Kim CM, Oh TY, Huang J, Xuan YH, Park SJ, Je BI, Piao HL, Park SH, Cha YS, Ahn BO, Ji HS, Lee MC, Suh SC, Nam MH, Eun MY, Yi G, Yun DW, Han CD (2007) Analysis of gene-trap Ds rice populations in Korea. Plant Mol Biol 65:373–384
Park MY, Kang JY, Kim SY (2011) Overexpression of AtMYB52 confers ABA hypersensitivity and drought tolerance. Mol Cells 31:447–454
Park MY, Kim SA, Lee SJ, Kim SY (2013) ATHB17 is a positive regulator of abscisic acid response during early seedling growth. Mol Cells 35:125–133
Pogorelko GV, Fursova OV, Ogarkova OA, Tarasov VA (2008) A new technique for activation tagging in Arabidopsis. Gene 414:67–75
Ryu CH, You JH, Kang HG, Hur J, Kim YH, Han MJ, An K, Chung BC, Lee CH, An G (2004) Generation of T-DNA tagging lines with a bidirectional gene trap vector and the establishment of an insertion-site database. Plant Mol Biol 54:489–502
Satoh H, Matsusaka H, Kumamaru T (2010) Use of N-methyl-N-nitrosourea treatment of fertilized egg cells for saturation mutagenesis of rice. Breed Sci 60:475–485
Shao J, Liu X, Wang R, Zhang G, Yu F (2012) The Over-expression of an arabidopsis B3 transcription factor, ABS2/NGAL1, leads to the loss of flower petals. PLoS One 7:11
Shinozaki K, Yamaguchi-Shinozaki K, Seki M (2003) Regulatory network of gene expression in the drought and cold stress responses. Curr Opin Plant Biol 6:410–417
Sivaguru M, Pike S, Gassmann W, Baskin TI (2003) Aluminum rapidly depolymerizes cortical microtubules and depolarizes the plasma membrane: evidence that these responses are mediated by a glutamate receptor. Plant Cell Physiol 44:667–675
Suzuki T, Moriguchi K, Tsuda K, Eiguchi M, Kumamaru T, Satoh H, Kurata N (2010) Neighboring nucleotide bias around MNU induced mutations in rice. Rice Genet Newsl 25:90–91
Tani H, Chen X, Nurmberg P, Grant JJ, SantaMaria M, Chini A, Gilroy E, Birch PR, Loake GJ (2004) Activation tagging in plants: a tool for gene discovery. Funct Integr Genomics 4:258–266
Tapken D, Hollmann M (2008) Arabidopsis thaliana glutamate receptor ion channel function demonstrated by ion pore transplantation. J Mol Biol 383:36–48
Trupiano D, Yordanov Y, Regan S, Meilan R, Tschaplinski T, Scippa GS, Busov V (2013) Identification, characterization of an AP2/ERF transcription factor that promotes adventitious, lateral root formation in Populus. Planta 238:271–282
Tsuchida-Mayama T, Nakamura H, Hakata M, Ichikawa H (2010) Rice transgenic resources with gain-of-function phenotypes. Breed Sci 60:493–501
Turano FJ, Muhitch MJ, Felker FC, McMahon MB (2002) The putative glutamate receptor 3.2 from Arabidopsis thaliana (AtGLR3.2) is an integral membrane peptide that accumulates in rapidly growing tissues and persists in vascular-associated tissues. Plant Sci 163:43–51
Valliyodan B, Nguyen HT (2006) Understanding regulatory networks and engineering for enhanced drought tolerance in plants. Curr Opin Plant Biol 9:189–195
Vinocur B, Altman A (2005) Recent advances in engineering plant tolerance to abiotic stress: achievements and limitations. Curr Opin Biotechnol 16:123–132
Wan S, Wu J, Zhang Z, Sun X, Lv Y, Gao C, Ning Y, Ma J, Guo Y, Zhang Q, Zheng X, Zhang C, Ma Z, Lu T (2009) Activation tagging, an efficient tool for functional analysis of the rice genome. Plant Mol Biol 69:69–80
Wang W, Vinocur B, Altman A (2003) Plant responses to drought, salinity and extreme temperatures: towards genetic engineering for stress tolerance. Planta 218:1–14
Wang M, Liu X, Wang R, Li W, Rodermel S, Yu F (2012) Overexpression of a putative Arabidopsis BAHD acyltransferase causes dwarfism that can be rescued by brassinosteroid. J Exp Bot 63:5787–5801
Weigel D, Ahn JH, Blázquez MA, Borevitz JO, Christensen SK, Fankhauser C, Ferrándiz C, Kardailsky I, Malancharuvil EJ, Neff MM, Nguyen JT, Sato S, Wang ZY, Xia Y, Dixon RA, Harrison MJ, Lamb CJ, Yanofsky MF, Chory J (2000) Activation tagging in Arabidopsis. Plant Physiol 122:1003–1013
Xia Y, Suzuki H, Borevitz J, Blount J, Guo Z, Patel K, Dixon RA, Lamb C (2004) An extracellular aspartic protease functions in Arabidopsis disease resistance signaling. EMBO J 23:980–988
Xiong L, Zhu JK (2001) Abiotic stress signal transduction in plants: molecular and genetic perspectives. Physiol Plant 112:152–166
Xoconostle-Cázares B, Ramírez-Ortega FA, Flores-Elenes L, Ruiz-Melrano R (2010) Drought tolerance in crop plant. Am J Plant Physiol 5:241–256
Yamaguchi-Shinozaki K, Shinozaki K (2005) Organization of cis-acting regulatory elements in osmotic- and cold-stress-responsive promoters. Trends Plant Sci 10:88–94
Yi J, An G (2013) Utilization of tagging lines in rice. J Plant Biol 56:85–90
Zhang QF (2007) Strategies for developing green super rice. Proc Natl Acad Sci USA 104:16402–16409
Zhang J, Guo D, Chang Y, You C, Li X, Dai X, Weng Q, Zhang J, Chen G, Li X, Liu H, Han B, Zhang Q, Wu C (2007) Non-random distribution of T-DNA insertions at various levels of the genome hierarchy as revealed by analyzing 13804 T-DNA flanking sequences from an enhancer-trap mutant library. Plant J 49:947–959
Zhao T, Zeng Y, Kermode AR (2011) Identification of seed dormancy mutants by activation tagging. Methods Mol Biol 773:185–198
Acknowledgments
We thank Li Xin, Guimin Zhang, Jing Zhang, Chunxia Liu, Guangwu Chen, Xiaocui Huang, Yingbin Chen, Cong Li, Chengfeng Du, Huanming Sun, Xiuping Meng, Qiaoyan Pan, Guowen Zhang, Tom Greene, Bo Shen, and James Zhou, et al. for their contributions and supports to the research program. Thanks to the employees of Beijing Kaituo DNA Biotech Research Center, Co., Ltd., Beijing Weiming Kaituo Crop Co., Ltd., and DuPont Pioneer for their various supports. Thanks also to Binbin Lei and Shufen Li for their critical review of the manuscript. This research was supported by DuPont Pioneer ABT collaboration program.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by K. Chong.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lu, G., Wang, X., Liu, J. et al. Application of T-DNA activation tagging to identify glutamate receptor-like genes that enhance drought tolerance in plants. Plant Cell Rep 33, 617–631 (2014). https://doi.org/10.1007/s00299-014-1586-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-014-1586-7