Abstract
Plants express many calmodulins (CaMs) and calmodulin-like (CML) proteins that sense and transduce different Ca2+ signals. Previously, we reported divergent soybean (Glycine max) CaM isoforms (GmCaM4/5) with differential abilities to activate CaM-dependent enzymes. To elucidate biological functions of divergent CaM proteins, we isolated a cDNA encoding a CML protein, AtCML8, from Arabidopsis. AtCML8 shows highest identity with GmCaM4 at the protein sequence level. Expression of AtCML8 was high in roots, leaves, and flowers but low in stems. In addition, the expression of AtCML8 was induced by exposure to salicylic acid or NaCl. AtCML8 showed typical characteristics of CaM such as Ca2+-dependent electrophoretic mobility shift and Ca2+ binding ability. In immunoblot analyses, AtCML8 was recognized only by antiserum against GmCaM4 but not by GmCaM1 antibodies. Interestingly, AtCML8 was able to activate phosphodiesterase (PDE) but did not activate NAD kinase. These results suggest that AtCML8 acts as a CML protein in Arabidopsis with characteristics similar to soybean divergent GmCaM4 at the biochemical levels.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In plants, intracellular Ca2+ concentration is one of most important second messengers in the regulation of cellular events such as developmental cues and environmental biotic and abiotic stimuli (Sanders et al. 2002). Calcium is decoded by many calcium-binding proteins in calcium mediated signaling (DeFalco et al. 2010). Calmodulins (CaMs), representative Ca2+ binding proteins, are highly conserved and ubiquitous proteins in plants (McCormack et al. 2005). CaM also mediates Ca2+ signals in dependence on developmental and environmental cues and transmits signals to various target enzymes and signaling proteins, including metabolic enzymes, transcription factors, ion channels, protein kinases/phosphatases and structural proteins (Snedden and Fromm 2001; Bouché et al. 2005; Kim et al. 2009).
In addition to CaMs, plants also possess a large repertoire of CaM-like proteins (CMLs) that encode potential calcium sensors and exhibit significant structural divergence from the typical CaM (McCormack and Braam 2003; McCormack et al. 2005; Ranty et al. 2006; Boonburapong and Buaboocha 2007). Based on sequence analysis of CML proteins, amino acids have been substituted in the EF-hand loop motif, which binds a single Ca2+ ion. Recent data indicated that CaM and CML proteins differ in their Ca2+ affinity and target-binding activities (Hua et al. 2003; Lee et al. 2000; Popescu et al. 2007). However, the biochemical and physiological functions of CMLs are still largely unknown in plants.
Previously, we cloned five genes specifying CaM isoforms from soybean designated as GmCaM1 to GmCaM5, and divided them into two groups, the conserved group (GmCaM1 through 3) and a divergent group (GmCaM4 and 5) on the basis of primary structural diversity (Lee et al. 1995). In biochemical analyses, we reported that a divergent GmCaM isoform (GmCaM4) showed a different CaM-dependent enzyme regulation pattern. Although a conserved CaM isoform (GmCaM1) activated both PDE and NAD kinase, which are well-characterized CaM-dependent enzymes, the divergent CaM isoform (GmCaM4) activated PDE but not NAD kinase (Lee et al. 1995, 1997). In addition, CaM isoforms reciprocally regulated NOS and calcineurin. GmCaM4 activated mammalian NOS, while GmCaM1 serves as a competitive antagonist of this activation. In contrast, GmCaM1 activated the mammalian protein phosphatase calcineurin (CaN), while GmCaM4 competitively antagonized its activation (Cho et al. 1998; Lee et al. 2000).
In physiological analyses, two divergent GmCaM isoforms (GmCaM4 and GmCaM5) were involved in plant disease resistance responses in a salicylic acid independent manner in tobacco and Arabidopsis (Heo et al. 1999; Park et al. 2004). These CaM isoforms are believed to serve as mediators of intracellular Ca2+ signals that are transiently increased by pathogen attack in plants. In addition, transgenic plants over-expressing GmCaM4 showed enhanced resistance against salt stress in Arabidopsis through the up-regulation of a MYB transcription factor that regulated salt and dehydration responsive genes (Yoo et al. 2005). The biochemical and physiological differences distinguishing the two CaM sub-families may be due to altered primary structures between the divergent CaM isoforms and the highly conserved CaM isoforms.
The presence of GmCaM4 homologs at the protein level has previously been predicted by immunoblot analysis in Arabidopsis total protein extracts with GmCaM4 specific antibody (Lee et al. 1995). Recently, it was reported that divergent CaMs from Arabidopsis, including AtCaM8 (AtCML8), were able to functionally complemented a yeast calmodulin null mutant (Zielinski 2002). In addition, AtCaM7 in this divergent sub-family is a transcriptional regulator that directly interacts with the promoters of light-inducible genes and promotes photomorphogenesis (Kushwaha et al. 2008; Kim et al. 2009). It has also been reported that AtCML9 is responsive to diverse stimuli, and that the loss of function of AtCML9 results in altered responses to abiotic stress and ABA (Magnan et al. 2008). However, the biochemical activity of the CML proteins in Arabidopsis has not yet been determined. In order to understand the biochemical role of the CML proteins, we isolated one member of this group, AtCML8, which is most closely aligned with GmCaM4/5 in the model species Arabidopsis. Here, we describe the biochemical properties of AtCML8 which resemble those of a soybean divergent CaM (GmCaM4) isoform with respect to antigenicity and CaM-dependent enzyme activation.
Materials and methods
Phylogenetic analysis of AtCML8
For phylogenetic analysis of AtCML8, 13 CaMs and AtCML8 were aligned with CLUSTAL-W program (http://www-igbmc.u-strasbg.fr/BioInfo/ClustalW/clustalw.html). GenBank accession numbers for the analysis are as follows: AtCaM1, NM123137; AtCaM2, NM180013; AtCaM3, NM115539; AtCaM4, NM105313; AtCaM5, NM179766; AtCaM6, NM180529; AtCaM7, NM114249; AtCML8, NM117546; GmCaM1, L01430; GmCaM2, L01431; GmCaM3, L01432; GmCaM4, L01433; GmCaM5, L19359 and Human CaM, AAH08437, respectively.
Production of recombinant AtCML8 protein in E. coli
A T7 expression vector, pET-3d, was used for production of AtCML8 protein in E. coli BL21 (DE3) pLysS. The full-length AtCML8 cDNA clone was amplified by PCR with a forward (5′) primer containing a NcoI site (5′-CCATGGAAGAAACAGCACTGAC-3′) and a reverse (3′) primer containing a BamHI site (5′-GGATCCTCAGTCAATGTTGATCATCAT-3′). The PCR product was cloned in pGEM-T Easy Vector (Promega, Madison, WI) and sequenced to verify no PCR mistake. The cDNA was sub-cloned NcoI and BamHI site of the pET-3d expression vector (NOVAGEN). Recombinant AtCML8 protein was expressed in E. coli and purified by phenyl-Sepharose column chromatography (Amersham Bioscience) as described (Harmon et al. 1984). Protein concentration was determined using Protein assay kit (Bio-Rad) with bovine serum albumin and bovine calmodulin (σ) as standards.
Reverse transcription (RT)-PCR analysis
RT–PCR analysis was carried out as previously described (Hwang et al. 2009). cDNA was prepared from each tissue; roots, rosette leaves, stem, cauline leaves, and flowers. Primers for the PCR were: 5′-ATGGAAGAAACAGCACTGACAAAA-3′ and 5′-TCAGTCAATGTTGATCATCATCTT-3′ for AtCML8, and 5′-CCAACAACGTGAAATCGACAG-3′ and 5′-TCTTGGTATTGCTGGTACTCT-3′ for the internal standard tubulin2 (GenBank accession number, M84700). The sizes predicted from amplified products were 456 bp for AtCML8 and 243 bp for internal standard tubulin2. PCR amplifications were carried out in conditions of a 1 min denaturation at 94°C, followed by 25 cycles of 94°C for 40 s, 55°C for 30 s, and 72°C for 40 s. After PCR, 10 μl of each reaction was removed and electrophoresed in 1% agarose gel.
For quantitative real-time PCR, total RNAs were isolated from 10-day-old Arabidopsis plants treated with 0.5 mM SA (salicylic acid) and 100 mM NaCl. RNA samples were treated with DNase I at 37°C for 30 min. cDNA synthesis was performed using the SuperScript III Reverse Transcriptase (Invitrogen) according to the manufacturer’s instructions. The following primers were used for the quantitative real-time PCR: AtCML8: 5′-CAACAGTGATCCGTTCGTTG-3′ and 5′-TCTTCCTCCGCATCACTTTC-3′; tubulin2: 5′-TGGCATCAACTTTCATTGGA-3′ and 5′-ATGTTGCTCTCCGCTTCTGT-3′. The tubulin2 was used as a control to normalize the expression data. Quantitative real-time RCRs were performed in triplicate with a Bio-Rad CFX96™ Real-time system (http://www.bio-rad.com/). Data were analyzed with Bio-Rad CFX MANAGER software (2−ΔΔCt method).
45Ca2+ binding assay
GST and CaMs purified by phenyl-Sepharose column chromatography were electrophoresed on two independent SDS-denaturing gels. One gel was stained and the second gel was blotted onto PVDF membrane (Millipore). The blot was incubated in binding buffer (60 mM KCl, 5 mM MgCl2, and 10 mM imidazole–HCl, pH 6.8) containing 1 μCi 45Ca2+ (Maruyama et al. 1984). After washing, the blot was exposed and visualized by X-ray film.
Immunoblot analysis
To determine antibody cross-reactions among AtCML8, GmCaM1, GmCaM4 and bovine brain CaM, purified anti-GmCaM1 and anti-GmCaM4 antibodies were used in immunoblot experiments. Purified AtCML8 protein, GmCaM proteins, and bovine brain CaM were electrophoresed on 13.5% SDS–polyacrylamide gels in the presence of either 5 mM EGTA or 5 mM CaCl2 in the SDS sample buffer. Proteins were transferred onto a PVDF membrane (Millipore) and incubated either with anti-GmCaM1 or with anti-GmCaM4 antibody. Protein bands were detected using the ECL system (Amersham Pharmacia Biotech) after incubating with horseradish peroxidase-conjugated mouse anti-goat IgG antibody (Pierce).
Calmodulin-dependent enzyme assays
Phosphodiesterase (PDE) activity was assayed using commercially available bovine heart CaM-deficient phosphodiesterase (Roche Applied Science) as described (Schechtele and Marme 1988). NAD kinase was partially purified from pea seedlings by successive protamine sulfate precipitation, polyethylene glycol precipitation, and DEAE-Sephacel column chromatography as previously described (Muto and Miyachi 1977). Effluents from the DEAE-Sephacel column were used for NAD kinase assays without further purification. NAD kinase assay was done as described (Harmon et al. 1984) with varying amount of activator CaMs. As controls, NAD kinase activation was examined with reaction mixtures either in the presence of 5 mM EGTA or absence of exogenous activator CaM to verify that our NAD kinase preparation was free of endogenous CaM contamination and that the activation of NAD kinase was a calcium-dependent process. To determine activation parameters, curve fittings were done by the aid of the GraFit software through a non-linear regression analysis with a modified Hill equation (Lee et al. 1995).
where v is the observed rate, V max is the maximal activity, [CaM] is the concentration of added CaM, K act is the concentration of CaM required for half-maximal activity, and n is the Hill coefficient.
Results
Comparison of the AtCML8 amino acid sequence
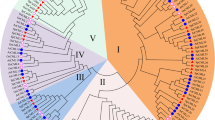
To understand the biological significance and molecular genetic characteristics of CaM-like protein, we cloned a CaM-like gene cDNA from Arabidopsis by polymerase chain reaction. Recently, the cDNA clone had been renamed as AtCML8 (McCormack and Braam 2003). The AtCML8 cDNA encodes 150 amino acids, whereas other conserved CaMs have 148 amino acid residues. AtCML8 has been shown to exhibit high amino acid sequence identity of 85.4% with a CaM-like protein, AtCML11 from Arabidopsis (McCormack and Braam 2003), and amino acid identity of 81% with GmCaM4 from soybean (Lee et al. 1995). Because the biochemical activity of AtCML11 has not been characterized as CaMs, the AtCML8 protein sequence was aligned with those of well-characterized CaMs such as AtCaM2, GmCaM1, and GmCaM4 (Fig. 1a). In order to visualize the primary sequence relationship of AtCML8 with other known CaMs, a phylogenetic analysis with 13 other CaMs was performed (Fig. 1b). The phylogenetic tree shows a close relationship of AtCML8 with GmCaM4 and GmCaM5 among the plant CaMs.
Comparison of the deduced amino acid sequences of AtCML8 and other CaMs. a Sequence alignment of AtCML8 with other divergent soybean CaMs (GmCaM4/5) and conserved Arabidopsis CaM (AtCaM2). Sequences are arranged to illustrate the relationships among the four Ca2+-binding domains. Identical amino acids are indicated by a dash (−). Residues marked with asterisks (*) correspond to ones that act as Ca2+-binding motifs, EF-hands, in conserved CaM proteins. b Phylogenetic relationships of AtCML8 and CaM proteins. Thirteen CaMs and CML8 amino acid sequences from Arabidopsis, soybean, and human were compared to construct a phylogenetic tree by neighbor joining method
Gene expression of AtCML8 in various tissues and in response to environmental stresses
Several reports have explored the expression patterns of CaMs and CaM-like proteins in different tissues and in the response to environmental stresses (Takezawa et al. 1995; Yang et al. 1998; Lee et al. 1999; McCormack et al. 2005). In order to observe the relative expression level of AtCML8, RT–PCR analysis was performed. The amplified products by RT–PCR were 450 bp for AtCML8 and 300 bp for Tubulin2, respectively. As shown in Fig. 2a, the AtCML8 gene was highly expressed in root (RO), leaves (RL and CL) and flower (FL). In stems (ST), however, the transcripts of AtCML8 were detected only at very low level. In order to investigate whether expression of the AtCML8 gene could be induced in response to biotic or abiotic stimuli, we examined the expression patterns in salicylic acid (SA, inducer of PR proteins) and NaCl-treated Arabidopsis plants by quantitative RT–PCR analysis. As shown in Fig. 2b and c, treatments of the plants with SA or NaCl resulted in increased transcripts of the tCML8 gene. Especially, SA treatment showed high increase in relative mRNA levels (Fig. 2b). Therefore, these results imply that AtCML8 may be involved in response to pathogen attack and NaCl stress, as it is known for GmCaM4/5 in soybean.
Expression analysis of AtCML8 gene in Arabidopsis tissues and in response to environmental stresses. a Expression patterns of AtCML8 gene in Arabidopsis tissues. The cDNAs were synthesized from total RNAs of each tissue, RO root, RL rosette leaves, ST stem, CL cauline leaves, and FL flower of 3 weeks grown plants. Subsequently, polymerase chain reactions (PCRs) were performed using gene-specific oligonucleotides primers. Amplified products were separated on a 1% agarose gel. Amplified products of the tubulin2 were used for quantitative controls. The sizes of PCR products are shown on the left. b qRT-PCR analysis of AtCML8 gene expression during 0.5 mM salicylic acid treatment for 0, 2, 6, and 12 h. c qRT-PCR analysis of AtCML8 gene expression during 100 mM NaCl treatment for 0, 2, 6, and 12 h. In b and c, tubulin2 was used as an internal control to normalize the expression data. Data represent the average of three independent experiments ± SD
Basic characteristics of AtCML8 as a calmodulin-like protein
In order to determine the characteristics of an Arabidopsis CML8, we expressed AtCML8 in E. coli, and purified the protein to homogeneity by Ca2+-dependent hydrophobic interaction chromatography. During the purification, the AtCML8 protein behaved in a fashion similar to that observed with other CaM isoforms with respect to heat stability and elution profiles on a phenyl-Sepharose column. To test the characteristic Ca2+-binding pattern of the calmodulin-like protein, we investigated the electrophoretic mobility shift of the AtCML8 protein upon Ca2+ binding. Purified proteins were electrophoretically separated in SDS gels in the presence of either 5 mM CaCl2 or 5 mM EGTA in sample buffer (Fig. 3a). AtCML8 protein showed a Ca2+-dependent electrophoretic mobility shift, although the extent of this shift was different among calmodulin isoforms. In order to investigate further the Ca2+-binding ability of AtCML8, 45Ca2+-binding assays were carried out (Fig. 3b). Purified proteins were electrophoresed on an SDS gels and stained or blotted onto membranes. The membranes were overlaid with a buffer containing 45Ca2+ and subsequently washed and exposed to X-ray film. As a result, calcium was bound to AtCML8 and GmCaM1 but not bound to GST.
Ca2+-binding ability of AtCML8. a Ca2+-dependent electrophoretic mobility shifts of AtCML8. AtCML8, GmCaM1, and four proteins were produced in E. coli using a T7 expression vector system and purified to homogeneity using phenyl-Sepharose column chromatography as described under “Materials and methods.” 2 μg of purified CaMs, CML protein and bovine brain calmodulin (σ) was electrophoresed on a 13.5% SDS–PAGE either in the presence of 5 mM CaCl2 or 5 mM EGTA in sample buffers. Protein bands were visualized by Coomassie Brilliant Blue staining. b 45Ca2+ binding assay. Protein samples were separated by SDS–PAGE, and either stained with Coomassie Brilliant Blue (right panel) or overlaid and autoradiographed with 45Ca2+ (left panel). GmCaM-1 and GST are a positive control and negative control for 45Ca2+ binding assay, respectively
To investigate antigenic characteristics of AtCML8, we performed western blot analysis by using anti-GmCaM1 and anti-GmCaM4 antibodies. Previously, we had shown antigenic differences between conserved type CaM (GmCaM1 and bovine CaM) and divergent type CaM (GmCaM4) by using specific goat antisera raised against GmCaM1 and GmCaM4 (Lee et al. 1995). As shown in Fig. 4, anti-GmCaM4 antiserum recognized GmCaM-4 and AtCML8 as expected, but not GmCaM1 and bovine CaM. In contrast, anti-GmCaM1 antiserum recognized GmCaM1 and bovine CaM, but not GmCaM4 and AtCML8. These results suggest that AtCML8 has similar antigenic characteristic as the divergent CaM isoform, GmCaM4 in Arabidopsis.
Immunoblot analysis of purified AtCML8 protein with antibodies raised against GmCaM1 or GmCaM4 proteins. Equal amount of GmCaM1/4 proteins, bovine CaM, and AtCML8 (25 ng per lane) was electrophoresed on a 13.5% SDS–PAGE. Fractionated proteins were blotted onto a PVDF membrane and incubated with purified either anti-GmCaM1 or anti-GmCaM4 antisera. The immune complexes were visualized by using the ECL system after incubating with horseradish peroxidase-conjugated anti-goat IgG
Differential activation of phosphodiesterase and NAD kinase by AtCML8
To assess the biochemical ability of the AtCML8 to activate CaM-dependent enzymes, two representative CaM-dependent enzymes, phosphodiesterase (PDE) and NAD kinase, were tested. In PDE enzyme assay, not only GmCaM1 and GmCaM4 but also AtCML8 activated bovine brain PDE well (Fig. 5a). The half-maximal activation value (K act) of AtCML8 was 90.01 nM, which is higher than those of GmCaM1 and 4. However, the maximal activation value (V max) of two GmCaM isoforms and AtCML8 for PDE were not significantly different from each other. In the presence of EGTA, PDE enzyme did not show activity regardless of the presence of activator, AtCML8 (data not shown). This result indicates that the activation of the enzyme by AtCML8 is a calcium-dependent process.
Activation of calmodulin-dependent enzymes by CaMs and AtCML8. a Activation of phosphodiesterase (PDE) by CaMs and AtCML8. Dose/response curves of PDE are shown. Data points represent means of three independent assays for PDE, and error bars represent standard deviations. Fitted curves are drawn from the Hill equation as described under “Materials and methods.” PDE assay was done using CaM-deficient bovine heart PDE and CaMs or CML (GmCaM1/4 and AtCML8) as activators. Activity of PDE was monitored with varying amounts of CaMs or AtCML8 and expressed as a relative activity to that of PDE in the presence of 100 nM activator CaMs or CML (GmCaM1/4 and AtCML8). b Differential activation of NAD kinase by CaMs or AtCML8. Dose/Response curves of NAD kinase are shown. Data points represent means of three independent assays for NAD kinase, and error bars represent standard deviations. The activity of pea NAD kinase is expressed as the activity relative to that of NAD kinase in the presence of 80 nM of isolated CaMs or CML (GmCaM1/4, and AtCML8)
In NAD kinase assays, we observed GmCaM1-activated NAD kinase with a K act (8.07 nM) and Hill coefficient (1.86) in the presence of Ca2+ but not in the absence of Ca2+. However, the divergent AtCML8 did not activate NAD kinase, similar to the behavior of GmCaM4, even at 500-fold higher concentration than that of GmCaM1 for a maximal activation of NAD kinase (Fig. 5b). The results strongly indicated that AtCML8 is a functional CML protein and may be an ortholog of GmCaM4 at the biochemical level in Arabidopsis.
Discussion
Calmodulins are well known as highly conserved calcium-binding proteins in eukaryotes (Chin and Means 2000; DeFalco et al. 2010). Many studies in plants revealed the presence of multiple calmodulin isoforms and CML proteins (McCormack et al. 2005). Although the Ca2+/calmodulin mediated signal transduction mechanism of plants is very similar to that of animals, the presence of multiple CaMs and CMLs implicates that the plant proteins may have unique features (Snedden and Fromm 2001; Sanders et al. 2002; McCormack and Braam 2003; Bouché et al. 2005). One group of conserved plant CaMs shows identities of 95% or higher when compared with animal CaMs. However, a set of divergent plant CaMs shows less than 75% identity with the conserved CaMs in both animals and plants. In this study, the AtCML8 cDNA was shown to encode a CML protein in Arabidopsis when it was isolated and characterized at the biochemical level.
The comparison of amino acid sequences of CaMs in Arabidopsis showed that all CaMs except AtCML8, AtCML9 (At3g51920) and other CMLs exhibit higher than 90% identity to each other. In contrast, AtCML8 shares less than 72% identity with the AtCaMs of the highly conserved type. When comparing AtCML8 and AtCaM2, one difference is that Tyr99 of AtCML8 was substituted by Phe99 of AtCaM2 in the third Ca2+-binding EF-hand domain (Fig. 1a). This Tyr99 residue was only found in the plant divergent CaMs, GmCaM4, GmCaM5 and NtCaM13 (Yamakawa et al. 2001). Like bovine and human CaMs, this Tyr99 is thought to be important for the regulation of the calmodulin by phosphorylation. Other telltale substitutions were changed from Gly40 in AtCaM2 to Asp40 in AtCML8, and from Ser81 in AtCaM2 to Ala81 in AtCML8. These residues are known as essential residues promoting helix bending (Marmé and Dieter 1983). Although the amino acid sequences of AtCML8 diverge considerably from that of AtCaM2, the positions of strong hydrophobic residues in the protein are highly conserved (Zielinski 2002).
Zielinski (2002) reported that AtCaM-8 (AtCML8) was identified as a CML protein based on sequence comparisons, when expressed in yeast conferred reduced rates of growth in Ca2+-depleted media to the yeast cells. In addition, AtCaM-8 (AtCML8) showed different target specificities when compared with those of highly conserved AtCaMs (Köhler and Neuhaus 2000; Charpenteau et al. 2004). However, no direct evidence was provided if AtCML8 constituted a functional CML protein. Here, the AtCML8 protein is identified as a genuine Ca2+-binding sensor protein by providing evidence not only through the observation of the Ca2+-dependent electrophoretic mobility shift assay but also by 45Ca2+-binding assays (Fig. 3). Also, the 17 kDa protein in Arabidopsis was recognized immunologically by a GmCaM4 specific antibody (Lee et al. 1995). We show that AtCML8 is recognized by a GmCaM4 specific antibody but not by GmCaM1 specific antibody (Fig. 4).
CaMs and CMLs could have different functions based on target specificity, distinct subcellular localization, and different Ca2+ affinity (Hoeflich and Ikura 2002; McCormack and Braam 2003). The binding of Ca2+ to CaM is well known to evoke a conformational change of CaM that results in a more compact structure. The structural change of CaM by Ca2+ exposes its hydrophobic surface that serves to interact with and alter the activities of target proteins. We previously reported that CaM isoforms exhibited differences in the Ca2+ dependence of activation of target enzymes and activated differentially and/or reciprocally. Two soybean CaM isoforms (GmCaM1/4) exhibited the opposite activation and competitive inhibition of CaM target enzymes (Lee et al. 1995; Lee et al. 2000).
Recently, Magnan et al. (2008) reported that AtCML9 gene was induced in the response to various environmental stimuli. Specifically, the protein was shown to participate in salt stress tolerance through its effects on ABA-mediated pathways. To identify the physiological function of AtCML8, we investigated not only AtCML8 T-DNA insertion mutants but also AtCML8 over-expressing transgenic plants. However, we could not detect any phenotype either during normal growth or under stressed conditions. These results imply that, in Arabidopsis, other CML isoforms are able to compensate the function of AtCML8 through gene redundancy, or that the AtCML8 protein is controlled by fine-tuning of its activation but not by its quantity.
Here, we report that the biochemical activity of AtCML8 from Arabidopsis is similar to that of GmCaM4, a known divergent CaM from soybean. The characterization of divergent CaM isoforms in Arabidopsis is a requisite for identifying its biological functions in plant Ca2+-signaling pathways. The model character of Arabidopsis thaliana and the available transgenic and mutational tools will help in studying biological functions in calcium signaling of the CaMs that have evolved into a plant-specific function.
References
Boonburapong B, Buaboocha T (2007) Genome-wide identification and analyses of the rice calmodulin and related potential calcium sensor proteins. BMC Plant Biol 7:4
Bouché N, Yellin A, Snedden WA, Fromm H (2005) Plant-specific calmodulin-binding proteins. Annu Rev Plant Biol 56:435–466
Charpenteau M, Jaworski K, Ramirez BC, Tretyn A, Ranjeva R, Ranty B (2004) A receptor-like kinase from Arabidopsis thaliana is a calmodulin-binding protein. Biochem J 379:841–848
Chin D, Means AR (2000) Calmodulin: a prototypical calcium sensor. Trends in Cell Biol 10:322–328
Cho MJ, Vaghy PL, Kondo R, Lee SH, Davis JP, Rehl R, Heo WD, Johnson JD (1998) Reciprocal regulation of mammalian nitric oxide synthase and calcineurin by plant calmodulin isoforms. Biochemistry 37:15593–15597
DeFalco TA, Bender KY, Snedden WA (2010) Breaking the code: Ca2+ sensors in plant signaling. Biochem J 425:27–40
Harmon AC, Jarrett HW, Cormier MJ (1984) An enzymatic assay for calmodulins based on plant NAD kinase activity. Anal Biochem 141:168–178
Heo WD, Lee SH, Kim MC, Kim JC, Chung WS, Chun HJ, Lee KJ, Park CY, Park HC, Choi JY, Cho MJ (1999) Involvement of specific calmodulin isoforms in salicylic acid-independent activation of plant disease resistance responses. Proc Natl Acad Sci USA 96:766–771
Hoeflich KP, Ikura M (2002) Calmodulin in action: diversity in target recognition and activation mechanisms. Cell 108:739–742
Hua W, Liang S, Lu YT (2003) A tobacco (Nicotiana tabaccum) calmodulin-binding protein kinase, NtCBK2, is regulated differentially by calmodulin isoforms. Biochem J 376:291–302
Hwang JE, Hong JK, Je JH, Lee KO, Kim DY, Lee SY, Lim CO (2009) Regulation of seed germination and seedling growth by an Arabidopsis phytocystatin isoform, AtCYS6. Plant Cell Rep 28:1623–1632
Kim MC, Chung WS, Yun D-J, Cho MJ (2009) Calcium and calmodulin-mediated regulation of gene expression in plants. Mol Plant 2:13–21
Köhler C, Neuhaus G (2000) Characterisation of calmodulin binding to cyclic nucleotide-gated ion channels from Arabidopsis thaliana. FEBS lett 4710:133–136
Kushwaha R, Singh A, Chattopadhyay S (2008) Calmodulin7 plays an important role as transcriptional regulator in Arabidopsis seedling development. Plant Cell 20:1747–1759
Lee SH, Kim JC, Lee MS, Heo WD, Seo HY, Yoon HW, Hong JC, Lee SY, Bahk JD, Hwang IH, Cho MJ (1995) Identification of novel divergent calmodulin isoform from soybean which has differential ability to activate calmodulin-dependent enzymes. J Biol Chem 270:21806–21812
Lee SH, Seo HY, Kim JC, Heo WD, Chung WS, Lee KJ, Kim MC, Cheong YH, Choi JY, Lim CO, Cho MJ (1997) Differential activation of NAD kinase by plant calmodulin isoforms: the critical role of domain I. J Biol Chem 272:9252–9259
Lee SH, Kim MC, Heo WD, Kim JC, Chung WS, Park CY, Park HC, Cheong YH, Kim CY, Lee KJ, Bahk JD, Lee SY, Cho MJ (1999) Competitive binding of calmodulin isoforms to calmodulin-binding proteins: implication for the function of calmodulin isoforms in plants. Biochim Biophys Acta 1433:56–67
Lee SH, Johnson JD, Walsh MP, Van Lierop JE, Sutherland C, Xu A, Snedden WA, Kosk-Kosicka D, Fromm H, Narayanan N, Cho MJ (2000) Differential regulation of Ca2+/calmodulin-dependent enzymes by plant calmodulin isoforms and free Ca2+ concentration. Biochem J 350:299–306
Magnan F, Ranty B, Charpenteau M, Sotta B, Galaud J-P, Aldon D (2008) Mutations in AtCML9, a calmodulin-like protein from Arabidopsis thaliana, alter plant responses to abiotic stress and abscisic acid. Plant J 56:575–589
Marmé D, Dieter P (1983) Role of calcium and calmodulin in plants. In: Cheung WY (ed) Calcium and cell function, vol 4, Academic Press, New York, p 263
Maruyama K, Mikawa T, Ebshi S (1984) Detection of calcium binding proteins by 45Ca autoradiography on nitrocellulose membrane after sodium dodecyl sulfate gel electrophoresis. J Biol Chem 95:511–519
McCormack E, Braam J (2003) Calmodulins and related potential calcium sensors of Arabidopsis. New Phytol 159:585–598
McCormack E, Tsai Y-C, Braam J (2005) Handling calcium signaling: Arabidopsis CaMs and CMLs. Trends Plant Sci 10:383–389
Muto S, Miyachi S (1977) Properties of a protein activator of NAD kinase from plants. Plant Physiol 59:55–60
Park CY, Heo WD, Yoo JH, Lee JH, Kim MC, Chun HJ, Moon BC, Kim IH, Park HC, Choi MS, Ok HM, Cheong MS, Lee SM, Kim HS, Lee KH, Lim CO, Chung WS, Cho MJ (2004) Pathogenesis-related gene expression by specific calmodulin isoforms is dependent on NIM1, a key regulator of systemic acquired resistance. Mol Cells 18:207–213
Popescu SC, Popescu GV, Bachan S, Zhang Z, Seay M, Gerstein M, Snyder M, Dinesh-Kumar SP (2007) Differential binding of calmodulin-related proteins to their targets revealed through high-density Arabidopsis protein microarrays. Proc Natl Acad Sci USA 104:4730–4735
Ranty B, Aldon D, Galaud J-P (2006) Plant calmodulins and calmodulin-related proteins: multifaceted relays to decode calcium signals. Plant Signal Behav 1:96–104
Sanders D, Pelloux J, Brownlee C, Harper JF (2002) Calcium at the crossroads of signaling. Plant Cell 14:S401–S417
Schechtele C, Marme D (1988) Calcium-binding proteins. In: Thompson M (ed), vol 1 CRC Press, Florida, p 83–96
Snedden WA, Fromm H (2001) Calmodulin as a versatile calcium signal transducer in plants. New Phytol 151:35–66
Takezawa D, Liu ZH, An G, Poovaiah BW (1995) CaM gene family in potato: developmental and touch induced expression of the mRNA encoding a novel isoform. Plant Mol Biol 27:693–703
Yamakawa H, Mitsuhara I, Ito N, Seo S, Kamada H, Ohashi Y (2001) Transcriptionally and post-transcriptionally regulated response of 13 calmodulin genes to tobacco mosaic virus-induced cell death and wounding in tobacco plant. Eur J Biochem 268:3916–3929
Yang T, Lev-Yadun S, Feldman M, Fromm H (1998) Developmentally regulated organ-, tissue-, and cell-specific expression of calmodulin genes in common wheat. Plant Mol Biol 37:109–120
Yoo JH, Park CY, Kim JC, Heo WD, Cheong MS, Park HC, Kim MC, Moon BC, Choi MS, Kang YH, Lee JH, Kim HS, Lee SM, Yoon HW, Lim CO, Yun D-J, Lee SY, Chung WS, Cho MJ (2005) Direct interaction of a divergent CaM isoform and the transcription factor, MYB2, enhances salt tolerance in Arabidopsis. J Biol Chem 280:3697–3706
Zielinski RE (2002) Characterization of three new members of the Arabidopsis thaliana calmodulin gene family: conserved and highly diverged members of the gene family functionally complement a yeast calmodulin null. Planta 214:446–455
Acknowledgments
We thank Dr. Hans J. Bohnert for critical reading and insightful comments. This work was supported by grants from World Class University Program (R32-10148) funded by MOEST and by BioGreen 21 Program (20080401034023) funded by RDA. KEK was supported by scholarship from the BK21 program of MOEST.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by J. R. Liu.
Rights and permissions
About this article
Cite this article
Park, H.C., Park, C.Y., Koo, S.C. et al. AtCML8, a calmodulin-like protein, differentially activating CaM-dependent enzymes in Arabidopsis thaliana . Plant Cell Rep 29, 1297–1304 (2010). https://doi.org/10.1007/s00299-010-0916-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-010-0916-7