Abstract
Thellungiella halophila is a salt-tolerant close relative of Arabidopsis, which is adopted as a halophytic model for stress tolerance research. We established an Agrobacterium tumefaciens-mediated transformation procedure for T. halophila. Leaf explants of T. halophila were incubated with A. tumefaciens strain EHA105 containing a binary vector pCAMBIA1301 with the hpt gene as a selectable marker for hygromycin resistance and an intron-containing β-glucuronidase gene as a reporter gene. Following co-cultivation, leaf explants were cultured on selective medium containing 10 mg l−1 hygromycin and 500 mg l−1 cefotaxime. Hygromycin-resistant calluses were induced from the leaf explants after 3 weeks. Shoot regeneration was achieved after transferring the calluses onto fresh medium of the same composition. Finally, the shoots were rooted on half strength MS basal medium supplemented with 10 mg l−1 hygromycin. Incorporation and expression of the transgenes were confirmed by PCR, Southern blot analysis and GUS histochemical assay. Using this protocol, transgenic T. halophila plants can be obtained in approximately 2 months with a high transformation frequency of 26%.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Thellungiella halophila (salt cress; synonymous to Thellungiella salsuginea, Al-Shebaz et al. 1999) is an emerging model species for studies designed to elucidate molecular mechanisms of abiotic stress tolerance in recent years (Inan et al. 2004; Wang et al. 2006; Volkov and Amtmann 2006; Wong et al. 2005). The plant is a salt-tolerant relative of Arabidopsis thaliana and has high genetic and morphological similarity with Arabidopsis. T. halophila can complete its life cycle in the presence of 300 mM NaCl. It also has a short life cycle and high seed yields. More importantly, its genome size is small, its cDNAs share over 90% identity to the model plant A. thaliana (Zhu 2001). The similarity between T. halophila and A. thaliana allows for the question of differential expression of salt tolerance genes between glycophytes and halophytes to be addressed using the molecular and genetic tools available for A. thaliana. Several molecular tools have been created for T. halophila including collections of ESTs, T-DNA insertion mutants and ecotypes, as well as cDNA libraries and microarrays (Inan et al. 2004; Amtmann et al. 2005, http://www.thellungiella.org/).
Agrobacterium-mediated genetic transformation has become a powerful tool for functional genomics (Gelvin 2005). For unraveling the function of a gene, many basic works are needed. For example, the gene expression pattern, localization of specific proteins, phenotypes of the plants when a gene is over-expressed or knocked out. Most of these studies are dependent on an efficient and suitable transfromation system. Although genetic transformation in most plant species needs a regeneration procedure, Arabidopsis can be transformed with the “floral dip” method, which is performed without the need for in vitro regeneration process (Clough and Bent 1998). This protocol has been used widely for its easy to handle and high transformation efficiency, as well as reduced occurrence of somatic mutation that is generally a problem associated with procedures requiring long-term tissue culture (Zhang et al. 2006). Similarly, the “floral dip” method was applied successfully on T. halophila, and a global T-DNA insertion mutagenesis of this plant is underway by using the same method (http://www.thellungiella.org/). However, the “floral dip” method has limitations when used for flowers with male or female sterility, as in these cases there is no seed-set for the screening of transgenic plants. This could be an obstacle for study of molecular mechanisms related to flower development and sterility. For example, it is difficult to perform genetic complementation experiments in mutants which are male or female sterile by the “floral dip” method. For the above reasons, an Agrobacterium-mediated genetic transformation system based on plant regeneration from somatic cells is needed as a supplementary method to the “floral dip” method.
This is the first report of establishing an Agrobacterium-mediated transformation method for T. halophila through plant regeneration from leaf explants. This will provide an alternative method for the molecular analysis of gene functions in this plant.
Materials and methods
Tissue culture conditions
Seeds of T. halophila (ecotype Shandong) were surface-sterilized for 30 s in 70% EtOH, transferred to 2.5% sodium hypochlorite containing 1 μl of Tween 20 per 2 ml for 20 min, rinsed five times with sterile distilled water, and placed on Petri dishes containing MS (Murashige and Skoog 1962) solid medium to germinate. Seedlings were grown at 22°C in a 16 h light (35 μmol m−2 s−1)/8 h dark cycle. The same growth-room conditions were used for tissue culture procedures. All plant media were adjusted with 1 N NaOH to pH 5.8, solidified with 0.8% agar, and autoclaved at 121°C for 20 min.
Callus formation, shoot regeneration and rooting
Leaves were excised from 3-week-old seedlings and placed horizontally on callus induction medium, which is MS basal medium supplemented with 3% (w/v) sucrose and different plant growth regulators. Callus induction was conducted under darkness or light. Three weeks later, the induced calluses were transferred to fresh medium of the same composition, and incubated in the light for shoot regeneration. After another 3 weeks, the regenerated shoots were transferred to half strength MS basal medium for rooting.
Agrobacterium and plasmid
Agrobacterium tumefaciens strain EHA105 harboring the binary vector pCAMBIA 1301 was used for transformation experiments. The binary vector pCAMBIA 1301 carried both the reporter gene for β-glucuronidase (uidA) and the selectable marker gene for hygromycin phosphotransferase (hpt). The vector pCAMBIA 1301 was introduced into A. tumefaciens strain EHA105 by the liquid nitrogen freezing-thaw method. For plant transformation, a single colony of the bacteria was inoculated in YEB medium supplemented with kanamycin (50 mg l−1) and rifampicillin (50 mg l−1) and cultured overnight at 28°C. The Agrobacterium cells were collected by centrifugation at 10,000g for 30 s at 25°C and then resuspended in 20 ml (OD600 = 0.4–0.5) of liquid MS medium containing 20 mg l−1 acetosyringone (AS).
Transformation of Thellungiella halophila leaf explants
Leaves from 1-month-old seedlings were cut into 0.5 × 0.5 cm (as leaf explants) and inoculated with the Agrobacterium cells harboring pCAMBIA 1301 in 20 ml of liquid MS medium supplemented with 20 mg l−1 AS for 15 min. Subsequently, the leaf explants were blotted on sterile filter paper to remove most of the liquid medium, and transferred to agar-solidified (0.8% w/v) MS medium supplemented with 20 mg l−1 AS, 2 mg l−1 benzyladenine (BA), 0.5 mg l−1 3-indoleacetic acid (IAA) for co-cultivation. After 3 days, the explants were washed with sterile water containing 500 mg l−1 cefotaxime to remove Agrobacterium cells. Subsequently, the explants were transferred onto selection medium (MS medium supplemented with 2 mg l−1 BA, 0.5 mg l−1 IAA, 10 mg l−1 hygromycin, and 500 mg l−1 cefotaxime), and incubated in the light. Three weeks later, the leaf explants together with the calli were transferred onto the same fresh medium for shoot induction. After another 3 weeks, hygromycin-resistant shoots were separated and transferred to half strength MS basal medium supplemented with 10 mg l−1 hygromycin for rooting.
PCR and Southern bolt analysis
Genomic DNA was prepared from leaves of hygromycin resistant plants using the cetyltrimethylammoniumbromide (CTAB) method (Muhammad et al. 1994). Genomic DNA from transgenic plants (one plant from each transgenic line) was used for PCR amplification of a 529 bp of hpt fragment using the primers of HF (5′-CGA TCT TAG CCA GAC GAG CGG GTT C-3′) and HR (5′-GCT GGG GCG TCG GTT TCC ACT ATC GG-3′). To examine whether any residual Agrobacterium exist in the regenerated plants, Agrobacterium VirB1 gene-specific primers VBFw 5′ GAA GGC AAC AGG GCC GCT GTC 3′ and VBRe 5′ TCC GCC CTC CGG GGA ACG ACG C 3′ were used to amplify the 630 bp fragment from the VirB1. The reaction mixture for PCR was incubated in a DNA thermal cycler under the following conditions: 94°C for 5 min, followed by 30 cycles of 94°C for 30 s, 60°C for 30 s, and 72°C for 1 min, with a final 10 min extension at 72°C. For Southern blot analysis, 5 μg of genomic DNA was digested with the restriction enzyme EcoRI, separated on 0.8% (w/v) agarose gel, and transferred to Hybond N+ nylon membrane. The hpt probe was labeled with digoxygenin (DIG) by PCR method using the primers of HF and HR. Prehybridization, washing, and chemiluminescent detection of the blots were performed according to the manufacturer’s instructions (Roche).
Assay for uidA expression in transgenic plantlet
Histochemical staining of GUS activity was performed according to Jefferson (1987). Calluses and plantlet of transgenic lines and calluses of wild type (control) were submerged into the solution containing 5-bromo-4-chloro-3-indolyl glucuronide (X-Gluc) at 37°C for 3 h and then destained in 70% alcohol.
Results and discussion
Callus induction from leaf explants
Plant growth regulators are important factors, which can selectively influence the genes to trigger differentiation of cells in culture (Thorpe 1983). We examined the effects of different plant growth regulators on callus induction for leaf explants of T. halophila. Callus induction on MS medium supplemented with 2 mgl−1 BA combined with either IAA, α-naphthaleneacetic acid (NAA) or 2,4-dichlorophenoxyacetic acid (2,4-d) at 0.5 mg l−1 all resulted in a 100% induction frequency (Table 1). This indicates that BA is sufficient for the callus induction. When transferring the calluses to fresh medium of the same composition, we observed that shoot regeneration only occurred on the medium containing BA and IAA (Fig. 1a). The callus induced in the light are green (Fig. 1b) and showed higher shoot regeneration frequency (90%). While callus induced under darkness are pale-yellow, and the regeneration frequency is obviously lower (Table 1). These indicate that light is effective to promote shoot regeneration.
Regeneration of transgenic plants from leaf explants in Thellungiella halophila. a Induction of multiple shoot/bud clumps after a 33-day culture on MS medium containing 2 mg l−1 BA and 0.5 mg l−1 IAA. b Green callus and bud formed on a single explant. c Regenerated plantlet after a 21-day culture on rooting medium. d Hygromycin resistant shoots after a 27-day culture on selection medium. e GUS assay of the transformed callus and buds. f GUS assay of transformed plantlet. g GUS assay of the non-transformed callus and buds
We also studied the regeneration capacity of root explants, as this kind of explants were used for transformation in the earlier studies in Arabidopsis (Valvekens et al. 1988). We observed a similar high frequency of callus induction in these explants as that for leaf explants. However, shoot regeneration rate from these calluses was lower than that from leaf explants. The experiments conducted on leaf explants from seedlings during different development stages (2-, 4- and 6-week-old) did not exhibit significant differences in callus induction and shoot regeneration (data not shown). The regenerated shoots were finally transferred to half strength MS basal medium for rooting, and roots appeared from the cut end of the shoots within 3 weeks (Fig. 1c). The rooting rates ranged from 81 to 92%.
Influence of antibiotics on shoot regeneration from leaf
To find a suitable selectable marker for T. halophila transformation, we tested the sensitivity of leaf pieces of T. halophila to a number of antibiotics that are currently used as selection agent in plant transformation. We found that 50 mg of kanamycin, 10 mg of G418, or 10 mg of hygromycin per liter completely blocked shoot regeneration. Therefore, the antibiotics we tested could be used for selection of transformed cells in this plant. As the pCAMBIA1301 contains the hpt gene which confers hygromycin resistance, we selected hygromycin as selection reagent in our transformation experiments. Cefotaxime which is used for suppressing and killing Agrobacterium, was found to severely inhibit shoot regeneration from root explants of Arabidopsis (Valvekens et al. 1988). In the case of T. halophila, cefotaxime did not inhibit the shoot regeneration capacity of its leaf explants and, therefore, cefotaxime was used to kill Agrobacterium after co-cultivation with leaf explants.
Genetic transformation of leaf explants
Leaf explants were incubated with the Agrobacterium cells harboring pCAMBIA 1301 for different intervals (5, 10, 15, 20, 25, and 30 min). We observed that transient expression of GUS increased with the infection time. Considering the post infection process, an incubation of 15 min was selected in the subsequent transformation experiments. After 3 days of co-cultivation, the leaf explants were washed with distilled water and transferred onto the selection medium containing 10 mg l−1 hygromycin, and 500 mg l−1 cefotaxime. Two weeks later, nodular green calluses grew out from the cut surface of the yellowish leaf explants. On average about 32% of the leaf explants produced hygromycin resistant calluses (Table 2). When transferring the resistant calluses to fresh medium of the same composition, shoot regeneration could be obtained in more than 85% of the green calluses (Table 2, Fig. 1d). Subsequently, more than 80% of the hygromycin resistant shoots (one shoot from one individual resistant callus) could root on half strength MS medium containing 10 mg l−1 hygromycin successfully (Table 2). Histochemical GUS assay showed that transgene expressed in calluses and in different parts of the putative transgenic plantlets (Fig. 1e, f), while there is no GUS staining in the wild-type control (Fig. 1g).
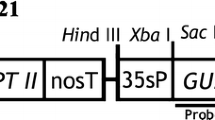
PCR and Southern blot analysis were further used to confirm the presence and the integration of T-DNA in T. halophila genome, as well as the copy number integrated. As shown in Fig. 2a, the amplification of genomic DNAs from different transgenic lines with hpt specific primer produced the expected 539 bp band, while the same band was not detected in amplification with DNA from non-transformed plants. At the same time, the VirB1 specific primers were also used to examine the presence of residual Agrobacterium in the regenerated plants. The amplification of the genomic DNA from Agrobacterium produced the 630 bp bands, while the amplification with DNAs from different transgenic lines did not give the same band (data not shown). These indicate that hygromycin resistant plants are truly transformed. Southern blot analysis using the hpt probe was performed on randomly selected transgenic lines. The results showed that all the selected lines have T-DNA integrated in their genome. As the genomic DNA was restricted with EcoRI, which cuts only once inside the T-DNA region in pCAMBIA 1301, thus, the number of the hybridization bands reflects the copy number of the T-DNA integrated into the T. halophila genome. The T-DNA copies integrated into transgenic lines ranged from 1 to 4. For example, lines 2 and 4 have single copy insertion, lines 1, 5, 6, 7, 8 and 9 have two copies, line 3 has about four copies (Fig. 2b). These results indicated that all the transgenic plants are from independent transgenic events, as they have different copy numbers of T-DNA integrated into different positions of the genome. Using this protocol, we regenerated 78 transgenic lines (each line came from one explant) in three independent experiments. On average, 26 transgenic lines (plants) were obtained from every 100 leaf explants (Table 2).
In summary, a highly efficient Agrobacterium-mediated leaf disc transformation method was established for T. halophila. This method could be used as a supplementary method to the widely used “floral dip” method, especially in complementation experiments for T. halophila mutants that are male or female sterile.
References
Al-Shebaz IA, O’Kane SLJ, Price RA (1999) Genetic placement of species excluded from Arabidopsis (Brassicaceae). Novon 9:296–307
Amtmann A, Bohnert HJ, Bressan RA (2005) Abiotic stress and plant genome evolution. Search for new models. Plant Physiol 138:127–130
Clough SJ, Bent AF (1998) A simple method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Gelvin SB (2005) Agricultural biotechnology: gene exchange by design. Nature 433:583–584
Inan G, Zhang Q, Li P, Wang Z, Cao Z, Zhang H, Zhabg C, Tanya QM, Mark GS, Zhu J, Shi H, Barbara D, Tarif C, Gong Q, Ma S, Mark F, David GW, Matthew JA, David R, Paul HM, Hans BJ, Robert JJ, Ray BA, Zhu J-K (2004) Salt cress. A halophyte and cryophyte Arabidopsis relative model system and its applicability to molecular genetic analyses of growth and development of extremophilts. Plant Physiol 135:1718–1738
Jefferson RA (1987) Assaying chimeric genes in plants: the Gus gene fusion system. Plant Mol Biol Rep 5:387–405
Muhammad AL, Ye GN, Weeden NF, Reisch BI (1994) A simple and efficient method for DNA extraction from grapevine cultivars, Vitis species and Ampelopsis. Plant Mol Biol Rep 12:6–13
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassay with tobacco tissue cultures. Physiol Plant 15:473–497
Thorpe TA (1983) Morphogenesis and regeneration in tissue culture. Beltsville Sym Agr Res 7:285–303
Valvekens D, Montagu MV, Lijsebettens MV (1988) Agrobacterium tumefaciens-mediated transformation of Arabidopsis thaliana root explants by using kanamycin selection. Proc Natl Acad Sci USA 85:5536–5540
Volkov V, Amtmann A (2006) Thellungiella halophila, a salt-tolerant relative of Arabidopsis thaliana, has specific root ion-channel features supporting K+/Na+ homeostasis under salinity stress. Plant J 48:342–353
Wang B, Davenport R, Volkov V, Amtmann A (2006) Low unidirectional sodium influx into root cells restricts net sodium accumulation in Thellungiella halophila, a salt-tolerant relative of Arabidopsis thaliana. J Exp Bot 57:1161–1170
Wong CE, Li Y, Whitty BR, Díaz-Camino C, Akhter SR, Brandle JE, Golding GB, Weretilnyk EA, Moffatt BA, Griffith M (2005) Expressed sequence tags from the Yukon ecotype of Thellungiella reveal that gene expression in response to cold, drought and salinity shows little overlap. Plant Mol Biol 5:561–574
Zhang X, Henriques R, Lin SS, Niu QW, Chua NH (2006) Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nat Protoc 1:641–646
Zhu JK (2001) Plant salt tolerance. Trends Plant Sci 6:66–71
Acknowledgments
This research was supported by the National Natural Science Foundation of China (No: 30370137) and a project from Guangdong Province (2006A20101007). We thank Dr. R.A. Jefferson, CAMBIA, Canberra, Australia, for kindly providing pCAMBIA1301. We would also like to thank the anonymous referees and the editor of this paper (Prof. Prakash P. Kumar) for their useful comments and valuable suggestions to improve the paper.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by P. Kumar.
Rights and permissions
About this article
Cite this article
Li, HQ., Xu, J., Chen, L. et al. Establishment of an efficient Agrobacterium tumefaciens-mediated leaf disc transformation of Thellungiella halophila . Plant Cell Rep 26, 1785–1789 (2007). https://doi.org/10.1007/s00299-007-0391-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-007-0391-y