Abstract
RNA interference (RNAi) is a potent trigger for specific gene silencing of expression in a number of organisms and is an efficient way of shutting down gene expression. 1-Aminocyclopropane-1-carboxylate (ACC) oxidase catalyzes the oxidation of ACC to ethylene, a plant growth regulator that plays an important role in the tomato ripening process. In this research, to produce double-stranded (ds)RNA of tomato ACC oxidase, we linked the sense and antisense configurations of DNA fragments with 1,002-bp or 7-nt artificially synthesized fragments, respectively, and then placed these under the control of a modified cauliflower mosaic virus 35S promoter. The dsRNA expression unit was successfully introduced into tomato cultivar Hezuo 906 by Agrobacterium tumefaciens-mediated transformation. Molecular analysis of 183 transgenic plants revealed that the dsRNA unit was integrated into the tomato genome. With respect to the construct with the 1,002-bp linker, the severity of phenotypes indicated that 72.3% of the transformed plants had non-RNA interference, about 18.1% had semi-RNA interference, and only 9.6% had full-RNA interference. However when the construct with the 7-nt linker was used for transformation, the results were 13.0%, 18.0%, and 69.0%, respectively, indicating that the short linker was more efficient in RNAi of transgenic tomato plants. When we applied this fast way of shutting down the ACC oxidase gene, transgenic tomato plants were produced that had fruit which released traces of ethylene and had a prolonged shelf life of more than 120 days. The RNA and protein analyses indicated that there was non-RNA interference, semi-RNA interference and full-RNA interference of ACC oxidase in the transgenic tomato plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Double-stranded RNA (dsRNA) is a powerful signal for specific gene silencing in a number of organisms, from Caenorhabditis elegans (Fire et al. 1998; Boutla et al. 2002), Drosophila (Alder et al. 2003) to plants (Chuang and Meyerowitz 2000; Klahre et al. 2002) and animals (Paddison et al. 2002; Zeng and Cullen 2003). This dsRNA-mediated gene silencing now is referred to as RNA interference (RNAi) and is closely related to post-transcriptional gene silencing (PTGS) in plants (Fire 1999; Sharp and Zamore 2000; Sijen and Kooter 2000). There is strong evidence of dsRNA having a key role as an inducer of PTGS in plants and of it acting as a trigger of RNA-mediated virus resistance (Vance and Vaucheret 2001). One of the advantages of RNAi over conventional gene knockout or knockdown is that only 20–30 bp to a few 100 bp of a gene sequence are sufficient to induce RNAi in plants and animals. The discovery of RNAi has changed our understanding of how organisms guard their genomes and has lead to the development of new strategies to block gene function. However, some inconsistent results have been observed (Chuang and Meyerowitz 2000; de Wit et al. 2002).
The phytohormone ethylene—olefin in its simplest form—plays an important role in many aspects of plant growth, development, and environmental responses (Abeles et al. 1992; Alexander and Grierson 2002). The ethylene biosynthetic pathway has been well-characterized (Yang and Hoffman 1984): S-adenosyl-l-Met is converted to 1-aminocyclopropane-1-carboxylate (ACC) by ACC synthase (Olson et al. 1991), then ACC is converted to ethylene by ACC oxidase (Barry et al. 1996). In fruits, both ACC synthase and ACC oxidase are induced during ripening and contribute to both the regulation of ethylene production and the ripening process. In tomato, the ACC oxidase gene family consists of three members, withACO1 being the only predominant ACC oxidase in fruit (Barry et al. 1996).
During the ripening of climacteric fruit, such as tomato, the burst of autocatalytic ethylene coordinates and accelerates the ripening process (Alexander and Grierson 2002). Both antisense mRNA and sense mRNA strategies have been used in tomato to prolong shelf life (Ye et al. 1996; Liu et al. 1998; Yao et al. 1999; Xiong et al. 2003). However, since RNAi can shut down the gene expression specifically, dsRNA may be much more effective than either the sense or antisense RNA. In the study reported here, we introduced a unit of tomato ACC oxidase dsRNA into tomato (Lycopersicon esculentum Mill) cv. Hezuo 906 by Agrobacterium tumefaciens-mediated transformation. Fruit of the transgenic plants exhibited a prolonged shelf life, and the ethylene production rate of the ripened fruits and leaves was significantly inhibited.
Materials and methods
Plants and bacteria
Tomato (Lycopersicon esculentum Mill) cv. Hezuo 906 seeds were obtained from the New-Changzheng Tomato Institute of Shanghai. Plants were grown in the glasshouse under supplementary lighting. Agrobacterium tumefaciens strain LBA4404 was used for transformation. Fruits were picked at various developmental stages defined as follows: (1) immature green; small, with expanding hairy fruit; (2) mature green; fruits that had fully expanded and lost the hairs from the surface but did not show visible signs of ripening; (3) breaker; fruit in which the first visible color change could be observed; (4) breaker+3, +8, and +100; fruit 3, 8, and 100 days following the onset of color change.
A. tumefaciens-mediated transformation of tomato plants
A. tumefaciens-mediated transformation of tomato stem explants was carried out as described Jones et al. (1998).
Construction of the transformation vector
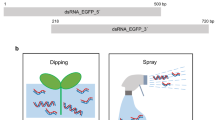
Total RNA was isolated from ripened fruits of tomato as described by Jones et al. (1998). Sense and antisense DNA fragments 71–571 of the ACC oxidase gene were obtained using the reverse transcription (RT)-PCR. The primers used were the oligonucleotides Acos1, Acos2; Acoa1, and Acoa2 (Table 1) and were synthesized using a Oligo 1000M DNA synthesizer (Beckman, Fullerton, Calif., www.beckmancoulter.com). The cycling parameters of the PCR were: one cycle at 94°C for 10 min [during which 2.0 U Taq polymerase (Promega, Madison, Wis., www.promega.com) was added], then 30 cycles of 94°C for 30 s (denaturing), 65°C for 40 s (annealing), and 72°C for 1 min (extension). The amplified products were cloned directly into the TA cloning vector pMD 18-T (TaKaRa, Japan, www.takara.com.cn). Two artificially synthesized fragments, the 1,002-bp crtW+crtY fragment (Genbank no. AY345165) and the 7-nt (TCAAGAG) fragment, were used as linkers between gene-specific fragments in the sense and antisense orientations, respectively. We obtained the dsRNA unit OACCRi1 (containing the 1,002-bp linker) and OACCRi2 (containing the 7-nt linker) (Fig. 1). DNA was sequenced by the di-deoxy chain-termination method using an autofluorescence sequencer (Model 377; ABI, Foster City, Calif., www.appliedbiosystems.com).
The double-stranded (ds)RNA of the 1-aminocyclopropane-1-carboxylate (ACC) oxidase gene constructs. The dsRNA unit OACCRi1. ACC oxidase gene-specific sequences (open boxes with arrows indicating the orientation) in the antisense and sense orientations were linked with a 1,002-bp fragment or a 7-nt (TCAAGAG) fragment (hatched box) and controlled by the D35S Omega promoter (solid arrow)
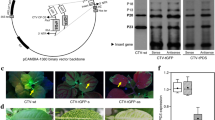
In order to improve the efficiency of gene transformation in tomato, we modified the plant binary vector pCAMBIA1201 (Genbank no. AF234293). Our new binary vector, pOACCRi, used the tobacco Ubi-U4 promoter (Plesse et al. 1997) instead of the cauliflower mosaic virus (CaMV) 35S promoter to control the reporter intron-β-gluronidase (GUS) gene expression. The hygromycin resistance gene was replaced by a modified neomycin phosphotransferase II (nptll) gene (Peng et al. 2001), which contained the catalase intron in the coding sequence. The OACCRNAi unit was placed under the control of a modified CaMV 35S promoter fragment with a duplicated enhancer. The unit was terminated by the termination sequences of the nopaline synthase (nos) gene from A. tumefaciens. DNA cloning was performed according to standard procedures (Sambrook et al. 1989).
Southern blotting
DNA for Southern blotting was extracted following the nuclei lysis hexadecyl trimethylammonium bromide (CTAB) extraction method (Richards et al. 1998). Tomato genomic DNA was digested with PstI. Fragments were separated on a 0.8% (w/v) agarose gel (in 40 mM Tris-acetate, 1 mM EDTA and 0.5 μg/ml ethidium bromide) at 40 V for 4 h. The digested tomato genomic DNA was transferred onto a nylon membrane (Amersham Pharmacia Biotech, Piscataway, N.J., www4.amersham -biosciences.com) and hybridized with the probe, a PCR-derived nptII fragment. The probe was labeled with α-[32P]-dCTP, which was purchased from Amersham Pharmacia Biotech, using a PCR labeling kit (TaKaRa). Hybridization was carried out using a standard protocol (Sambrook et al. 1989).
Northern blotting
Tomato fruit total RNA was prepared as detailed by Lu et al. (2001). The same total RNA extract was used for high- and low-molecular-weight RNA gel blot analysis. RNA polyacrylamide gel electrophoresis (PAGE), blotting, and hybridization was done according to the method of Silhavy et al. (2002). For Northern blots, 10 μg of total RNA was separated by gel electrophoresis on a 1.2% agarose gel and blotted onto a nylon membrane (Amersham Pharmacia Biotech). The RNA was fixed to the membrane using a UV cross-linker (Stratagene, La Jolla, Calif., www.stratagene.com). α-[32P]-CTP-labeled ACC oxidase RNA probes were synthesized with Riboprobe in vitro Transcription Systems purchased from Promega. Hybridization was done as described by Sambrook et al. (1989). Membranes were rehybridized with a cDNA probe corresponding to 18S rRNA to confirm equal loading of the gels.
Western blotting
Polyclonal antibodies were raised in a rabbit against denatured ACC oxidase, which was expressed in Escherichia coli and purified according to the QIAexpress Protein Purification System (Qiagen, Valencia, Calif., www.qiagen.com). Protein was extracted from tomato fruit according to the protocol of Meyer et al. (1988). Samples (2 μg protein) were electrophoresed on 12% sodium dodecyl sulfate (SDS) polyacrylamide gels and subsequently blotted onto transfer buffer (25 mM Tris pH 8.3, 192 mM glycine, 20% methanol). Western blotting was as described by Sambrook et al. (1989). Antibodies against ACC oxidase and goat-anti-rabbit alkaline phosphyatase conjugate were used at a dilution of 1:5,000.
ACC oxidase enzyme activity assay
ACC oxidase enzyme activity was determined using the procedure described by Barry et al. (1996). Three samples were analyzed each time and the assay repeated three times.
Ethylene measurements
The samples of fruits and leaves were placed in a sealed container at 25°C for 1 h. One-milliliter gas samples were withdrawn for ethylene measurement by gas chromatography as described by Picton et al. (1993). Three samples were analyzed each time and the analysis repeated three times.
Results
ACC oxidase dsRNA-mediated genetic interference and phenotype of the transgenic plants
We obtained 188 transgenic explants that showed GUS activity, of which 183 developed into plants and survived after being transplanted to soil. Of these 183 plants, 83 and 100 were transformed with the OACCRi1 and the OACCRi2 construct, respectively. The transgenic and control plants blossomed in the spring, and the fruits matured in May in the experimental field plot.
The fruits of the transgenic and control plants showed various phenotypes. The fruits of eight transgenic OACCRi1 and 69 transgenic OACCRi2 plants had a significantly prolonged shelf life with respect to those from the other transgenic plants (Fig. 2a). While the former fruits (transgenic OACCRi1 and OACCRi2 plants) slowly changed color from green to buff, those of the control tomato plants changed from breaker (+day) to pink and then to red in only 8 days (Fig. 2b). Fruits of 15 transgenic OACCRi1 plants and 18 transgenic OACCRi2 plants had a longer shelf life than the control fruits, while the fruits of 60 transgenic OACCRi1 plants and 13 transgenic OACCRi2 plants had the same shelf life as the control fruit.
The colors of tomato fruits of transgenic and nontransgenic plants at breaker+8 days. a Fruit of transgenic plant tomato showed full-RNAi. b Fruit of nontransgenic plant Hezuo 906. c Changes in the colors of tomato fruits of cv. Hezuo 906 plants from breaker+1 day to breaker+120 days, semi-RNAi and full-RNAi transgenic plants of ACC oxidase: a fruit of the control tomato plant changed to red at breaker+5 days, b semi-RNAi transgenic tomato fruit changed to red at breaker+30 days, c full-RNAi transgenic tomato fruit changed first to buff and then to a weak-red color very slowly; at breaker+120 days, the fruit was rosy in color
Three tomato fruits were plucked from a control plant and each of 183 transgenic plants at the breaker stage. Fruits from the control plants changed from green to red within 5 days and rotted after 10–15 days under conventional storage conditions (room temperature) (Fig. 2c, part a). Fruits of 72.3% of the OACCRi1 transgenic tomato plants showed a color change and had a shelf life similar to those of control plants (non-RNAi); fruits of 18.1% of the OACCRi1 transgenic plants changed color from green to red within 30 days and had a prolonged shelf life of over 50 days before rotting (Fig. 2c, part b) (semi-RNAi); fruits of the remaining 9.6% OACCRi1 transgenic plants gradually changed color from green to buff and then slowly to light red slowly, a process that took over 120 days (Fig. 2c, part c) (full-RNAi). In OACCRi2 transgenic plants, 13.0%, 18.0%, and 69.0% showed non-RNAi, semi-RNAi and full-RNAi characteristics, respectively.
Most of the fruits of the control tomato cultivar Hezuo 906 rotted within 10 days of being picked, while most of those of transgenic tomato, which showed non-RNAi of the ACC oxidase gene, rotted within 15 days. Almost 95% of the fruits showed semi-RNAi, and these took 50–60 days to rot. Most of the fruits that showed full-RNAi rotted after 120 days (Fig. 3).
DNA and RNA analysis
Six transgenic plants, two of each showing full-RNAi, semi-RNAi, and non-RNAi, respectively, were selected for Southern blot analysis. The results indicated that the OACCRi unit was integrated into the tomato genome of the six transgenic tomato plants (Fig. 4).
Southern blot analysis of the OACCRi unit in six transgenic plants. Genomic DNA was isolated, digested, and separated on a 0.8% agarose gel, transferred to a nylon membrane and hybridized with a α-[32P]-dCTP-labeled probe specific for the PCR-derived nptII fragment. Lanes: pRi Plasmid of pOACCRi, WT wild-type tomato plant, 1 OACCRi1 transgenic plants (full-RNAi), 2 OACCRi2 transgenic plants (full-RNAi), 3 OACCRi1 transgenic plants (semi-RNAi), 4 OACCRi2 transgenic plants (semi-RNAi), 5 OACCRi1 transgenic plants (non-RNAi), 6 OACCRi2 transgenic plants (non-RNAi)
In order to determine the pattern of RNA accumulation and RNA interference, we extracted RNA samples from fruit at different stages of ripening—from mature green to breaker+3, +8, and +10—for RNA gel blot analysis. The level of ACC oxidase mRNA in fruit of the control Hezuo 906 plant increased dramatically from a low level in the mature green fruit to a peak at the breaker+3 stage before declining at breaker+8. At no stage were small RNAs detected in the control (Fig. 5a). Figure 5b,c shows the levels of ACC oxidase in the fruit of plants transformed with the OACCRi1 and OACCRi2 constructs. ACC oxidase mRNA was not detected from the mature green stage to breaker+10. However, the 21- to 23-nt RNA levels were slightly higher. These results indicate that dsRNA mediated ACC oxidase RNAi in transgenic OACCRi1 and OACCRi2 plants. The level of ACC oxidase mRNA in the fruit of plants displaying semi-RNAi at different stages is shown in Fig. 5d. There was some increase in the mRNA level of ACC oxidase at different stages, with some blot signals around 21- to 23-nt RNA, but these blot signals were weaker than those in plants that showed full-RNAi. Figure 5e shows that the increased ACC oxidase mRNA level in plants that showed non-RNAi was the same as that in the control plants and that there was no sign of 21- to 23-nt RNA.
ACC oxidase RNA gel blot analysis and a comparison of fruit RNA abundance in control and transgenic tomato plants at different stages of fruit development. Total RNA was separated by gel electrophoresis on a 1.2% agarose gel, blotted onto a nylon membrane, fixed to the membrane using a UV cross-linker, and hybridized with α-[32P]-CTP-labeled ACC oxidase RNA probes. Lanes: M Mature green, B+3 breaker+3 days, B+8 breaker+8 days, B+10 breaker+10 days. Equal loading was confirmed by rehybridization with a probe for 18s rRNA. a Control tomato fruit (Hezuo 906), b OACCRi1, full-RNAi, c OACCRi2, full-RNAi, d OACCRi2, semi-RNAi, e OACCRi2, non-RNAi
ACC oxidase enzyme activity and protein abundance
Fruits ranging from the mature-green to fully ripened stages were used to analyze ACC oxidase activity (Table 2). In the fruits of control plant Hezuo 906 and transgenic OACCRi1 and OACCRi2 plants showing non-RNAi we detected ACC oxidase activity at a low basal level in the green fruits; this level increased substantially from the breaker stage to the fully ripened stage, peaking at breaker+3 and then declining. ACC oxidase activity was very low at all stages in transgenic fruits that showed full-RNAi (Table 2), while in transgenic fruits showing semi-RNAi, ACC oxidase activity showed the same trend as that observed in the control fruit except that the activity was fivefold lower than in the latter, resulting in a slower ripening process (Table 2).
We also determined the level of ACC oxidase protein in the fruits of the control plant and in those showing ACC oxidase non-RNAi, semi-RNAi, and full-RNAi at the mature-green, breaker+3, breaker+8, and breaker+10 stages. The accumulation of ACC oxidase protein was found to be correlated with the degree of prolonged ripening and RNA abundance. The greatest abundance of protein was found in control Hezuo 906 fruit, where it increased from a low level in mature-green fruit, peaked at the breaker+3 stage before declining at breaker+8 (Fig. 6a). ACC oxidase protein was weakly expressed in fruits showing semi-RNAi (Fig. 6c) and was not detected at levels above background in tomato fruits exhibiting full-RNAi (Fig. 6b). The level of ACC oxidase protein in fruits of non-RNAi transgenic plants was the same as in those of the control plants (Fig. 6d).
Western blot analysis of the ACC oxidase protein accumulation. Protein samples were electrophoresed on 12% SDS-polyacrylamide gels and transferred to a nitrocellulose membrane. The blots were blocked with non-fat dry milk, washed, and then incubated with polyclonal anti-ACC oxidase antibodies. M Mature green, B+3 breaker+3 days, B+8 breaker+8 days, B+10 breaker+10 days. a Control tomato fruit (Hezuo 906), b ACC oxidase full-RNAi tomato fruit, c ACC oxidase semi-RNAi tomato fruit, d ACC oxidase non-RNAi tomato fruit
Ethylene production
Ethylene levels were measured during ripening (Fig. 7a). In control fruits, ethylene production peaked at the breaker+3 stage and then declined. In full-RNAi fruits, ethylene was produced at a very low rate continuously but was only 1% of the level observed in the control. In semi-RNAi fruits, ethylene production showed the same trend as that in the control but at a level nearly threefold lower. Ethylene production in non-RNAi fruits was similar to that of the control (Fig. 7a).
Ethylene levels were also measured in wounded leaves (Fig. 7b). In the wounded leaves of the control plant, ethylene production was rapidly induced within 30 min, peaked at about 2 h after wounding, and then declined. This trend was repeated in wounded leaves showing non-RNAi. In leaves that exhibited full-RNAi ethylene production was only detectable 2 h after wounding, and at only 0.5% of that of the control plant. In wounded leaves showing semi-RNAi, ethylene production was only 20–30% of that observed in the control leaves at different times following wounding (Fig. 7b). These results indicate that the ethylene production rate of ripened fruits and leaves was significantly inhibited in the ACC oxidase semi-RNA interference plant, and it was almost completely repressed as a result of ACC oxidase RNA interference in those plants expressing RNA interference.
Discussion
During the last decade, various groups have shown that cells are capable of shutting down or silencing genes in a number of novel ways. Two types of anti-mRNA strategies can be distinguished: one is single-stranded sense-mRNA (Di Serio et al. 2001) with antisense-mRNA (D’Aoust et al. 1999; Hackett et al. 2000; Kurreck 2003), and the other is RNA interference induced by dsRNA (Bernstein et al. 2001; Sharp 2001; Zamore 2001). When RNA-induced silencing was first discovered in transgenic plants, it was termed PTGS (Bosher and Labouesse 2000; Hutvagner et al. 2000). It has since been found to be closely related to the phenomenon of RNAi in other organisms (Klahre et al. 2002; Boutla et al. 2002).
In the investigation reported here, we examined the efficacy of employing the RNAi strategy to knockout and knockdown the ACC oxidase gene, which is involved in the last step of the biosynthesis of ethylene, a key inducer of the ripening process of climacteric fruits like tomato. The knockout or knockdown of ACC oxidase should reduce or block ethylene biosynthesis and, consequently, prolong the shelf life of tomato. We obtained transgenic tomatoes that showed a full spectrum of RNAi (non-, semi-, and full-RNAi) on ACC oxidase gene expression at the level of mRNA, protein, protein activity, ethylene production, ripening, and fruit shelf life; the latter ranged from a normal 15 days (control, non-RNAi) to a prolonged 120 days. Part of the wide variation of the RNAi effect is related to the constructs, or the linker length between the sense and antisense fragment of the ACC oxidase gene. In transgenic OACCRi1 tomato plants, in which dsRNA is linked with a long, 1,002-bp fragment, most of the fruit showed a non-RNAi effect, some showed a semi-RNAi effect, while less than 10% showed a full-RNAi effect. This result indicates that only in 10% of the transgenic tomato plants was the ACC oxidase gene shut down, even though the other 90% of the transgenic tomato plants were confirmed to have transformed the dsRNA expression units into the genome DNA. However, when we used another dsRNA construction, OACCRi2, in which the dsRNA was linked to a short fragment (7 nt), we obtained opposite results. Most of the transgenic plants exhibited a full-RNAi effect, some showed a semi-RNAi effect, and only 13% of transgenic plants expressed a non-RNAi effect. In a study by Wesley et al. (Wesley et al. 2001), a 560-nt GUS link was used, but in our research the link was a 1,002-bp artificial sequence bias of the typical plant codon usage. Our results suggest that the short linker is more efficient than the long linker in producing the RNAi effect in transgenic plants.
The wide spectrum of RNAi effect generated by the same construct, from very weak to very strong, suggests that there are other components which are either involved in or influence the RNAi, or PTGS. It has been shown that almost 85% (Wesley et al. 2001) or 100% (Smith et al. 2000) of the plants transformed with an intron-containing hairpin RNA construct can be silenced. However, in our investigation, only 9.6–69.0% of the tomato plants transformed with a dsRNA construct showed silencing. One explanation for the difference in percentage of silencing may be that in our constructs a very highly modified 35S promoter fragment with a duplicated enhancer was used to drive the expression of the OACCRi unit. Another 35S promoter, which drove the modified nptII resistance gene, may have caused some interference with the 35S promoter that controls the OACCRi unit, depending on the position of the insertion and interaction with chromatins. It may also be possible that tomato cultivar Hezuo 906 could partially resist RNAi as shown in some other plants (Fire 1999; Chuang and Meyerowitz 2000).
It is well known that ethylene plays a crucial role in the initiation and development of ripening in tomato fruits. Manipulation of ethylene production—in particular, reducing its production—has been widely used in tomato to delay senescence and prolong the shelf life of the fruit. Previous researchers have genetically engineered tomato with genes like the antisense ACC oxidase gene (Ye et al. 1996), the antisense ACC synthase (Yao et al. 1999), and the double-antisense ACC oxidase and ACC synthase fusion gene (Xiong et al. 2003), and some low ethylene-producing tomato lines whose fruit have an extended shelf life have been obtained. We have demonstrated here a RNAi-mediated gene silencing for the ACC oxidase gene and have successfully generated transgenic tomato plants that produce trace levels of ethylene and exhibit a prolonged shelf life, with similar levels of total soluble sugar, titratable acid, amino acids, and total soluble solids as the control plants.
References
Abeles FB, Morgan PW, Saltveit ME (1992) Ethylene in plant biology, 2nd edn. Academic, San Diego, pp 264–296
Alder MN, Dames S, Gaudet J, Mango SE (2003) Gene silencing in Caenorhabditis elegans by transitive RNA interference. RNA 9:25–32
Alexander L, Grierson D (2002) Ethylene biosynthesis and action in tomato: a model for climacteric fruit ripening. J Exp Bot 53:2039–2055
Barry CS, Blume B, Bouzayen M, Cooper W, Hamilton AJ, Grierson D (1996) Differential expression of the 1-aminocyclopropane-1-carboxylate oxidase gene family of tomato. Plant J 9:525–535
Bernstein E, Denli AM, Hannon GJ (2001) The rest is silence. RNA 7:1509–1521
Bosher JM, Labouesse M (2000) RNA interference: genetic wand and genetic watchdog. Nat Cell Biol 2:E31–E36
Boutla A, Kalantidis K, Tavernarakis N, Tsagris M, Tabler M (2002) Induction of RNA interference in Caenorhabditis elegans by RNAs derived from plants exhibiting post-transcriptional gene silencing. Nucleic Acids Res 30:1688–1694
Chuang CF, Meyerowitz EM (2000) Specific and heritable genetic interference by double-stranded RNA in Arabidopsis thaliana. Proc Natl Acad Sci USA 97:4985–4990
D’Aoust MA, Yelle S, Nguyen-Quoc B (1999) Antisense inhibition of tomato fruit sucrose synthase decreases fruit setting and the sucrose unloading capacity of young fruit. Plant Cell 11:2407–2418
Di Serio F, Schob H, Iglesias A, Tarina C, Bouldoires E, Meins F (2001) Sense- and antisense-mediated gene silencing in tobacco is inhibited by the same viral suppressors and is associated with accumulation of small RNAs. Proc Natl Acad Sci USA 98:6506–6510
Fire A (1999) RNA-triggered gene silencing. Trends Genet 15:358–363
Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC (1998) Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391:806–811
Hackett RM, Ho CW, Lin Z, Foote HC, Fray RG, Grierson D (2000) Antisense inhibition of the Nr gene restores normal ripening to the tomato Never-ripe mutant, consistent with the ethylene receptor-inhibition model. Plant Physiol 124:1079–1086
Hutvagner G, Mlynarova L, Nap JP (2000) Detailed characterization of the posttranscriptional gene-silencing-related small RNA in a GUS gene-silenced tobacco. RNA 6:1445–1454
Jones, CG, Scothern GP, Lycett GW, Tucker GA (1998) Co-ordinated gene silencing by a single chimeric gene construct. Planta 204:499–505
Klahre U, Crete P, Leuenberger SA, Iglesias VA, Meins F (2002) High molecular weight RNAs and small interfering RNAs induce systemic posttranscriptional gene silencing in plants. Proc Natl Acad Sci USA 99:11981–11986
Kurreck J (2003) Antisense technologies. Improvement through novel chemical modifications. Eur J Biochem 270:1628–1644
Liu CY, Tian YC, Shen QG, Jiang H, Ju R, Yan T, Liu CD, Mang KQ (1998) Cloning of ACC synthase cDNA and its inhibition of fruit ripening by its antisense RNA in transgenic tomato plants. Chin J Biotechnol 14:139–146
Lu C, Zainal Z, Tucker GA, Lycett GW (2001) Developmental abnormalities and reduced fruit softening in tomato plants expressing an antisense Rab11 GTPase gene. Plant Cell 13:1819–1833
Meyer Y, Grosset J, Chartier Y, Cleyet-Marel JC (1988) Preparation by two-dimensional electrophoresis of proteins for antibody production: antibodies against proteins whose synthesis is reduced by auxin in tobacco mesophyll protoplasts. Electrophoresis 9:704–712
Olson DC, White JA, Edelman L, Harkins RN, Kende H (1991) Differential expression of two genes for 1-aminocyclopropane-1-carboxylate synthase in tomato fruits. Proc Natl Acad Sci USA 88:5340–5344
Paddison PJ, Caudy AA, Hannon GJ (2002) Stable suppression of gene expression by RNAi in mammalian cells. Proc Natl Acad Sci USA 99:1443–1448
Peng RH, Huang XM, LI X, Shun AJ, Yao QH, Peng YL (2001) Construction of plant binary expression vector containing intron-kanamycin gene and transformation in Nicotiana tabacum. Sheng Wu Hua Xue Yu Sheng Wu Wu Li Xue Bao 27:55–60
Picton S, Gray J, Barton S, AbuBakar U, Lowe A, Grierson D (1993) cDNA cloning and characterisation of novel ripening-related mRNAs with altered patterns of accumulation in the ripening inhibitor (rin) tomato ripening mutant. Plant Mol Biol 23:193–207
Plesse B, Durr A, Marbach J, Genschik P, Fleck J (1997) Identification of a new cis-regulatory element in a Nicotiana tabacum polyubiquitin gene promoter. Mol Gen Genet 254:258–266
Richards E, Reichardt M, Rogers S (1998) Preparation of plant DNA using CTAB. In: Ausubel JA, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl KS (eds) Current protocol in molecular biology. Current Protocol, Boston, pp 233–237
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Sharp PA (2001) RNA interference—2001. Genes Dev 15:485–490
Sharp PA, Zamore PD (2000) Molecular biology. RNA interference. Science 287:2431–2433
Sijen T, Kooter JM (2000) Post-transcriptional gene-silencing: RNAs on the attack or on the defense? Bioessays 22:520–531
Silhavy D, Molnar A, Lucioli A, Szittya G, Hornyik C, Tavazza M, Burgyan J (2002) A viral protein suppresses RNA silencing and binds silencing-generated, 21- to 25-nucleotide double-stranded RNAs. EMBO J 21:3070–3080
Smith NA, Singh SP, Wang MB, Stoutjesdijk PA, Green AG, Waterhouse PM (2000) Total silencing by intron-spliced hairpin RNAs. Nature 407:319–320
Vance V, Vaucheret H (2001) RNA silencing in plants—defense and counterdefense. Science 292:2277–2280
Wesley SV, Helliwell CA, Smith NA, Wang MB, Rouse DT, Liu Q, Gooding PS, Singh SP, Abbott D, Stoutjesdijk PA, Robinson SP, Gleave AP, Green AG, Waterhouse PM (2001) Construct design for efficient, effective and high-throughput gene silencing in plants. Plant J 27:581–590
de Wit T, Grosveld F, Drabek D (2002) The tomato RNA-directed RNA polymerase has no effect on gene silencing by RNA interference in transgenic mice. Transgenic Res 11:305–310
Xiong AS, Yao QH, Li X, Fan HQ, Peng RH (2003) Double-antisense ACC oxidase and ACC synthase fusion gene introduced into tomato by agrobacterium-mediated transformation and analysis the ethylene production of transgenic plants. Acta Biol Exp Sin 36:35–41
Yang S, Hoffman N (1984) Ethylene biosynthesis and its regulation in higher plants. Annu Rev Plant Physiol 84:520–525
Yao QH, Peng RH, Huang XM, Fan HQ, Zhang DB, Jiang L, Yang YM (1999) Construction of a fruit-specific expression vector of antisense tomato ACC synthase gene. Acta Agric Shanghai 15[Suppl]:1–5
Ye ZB, Li HX, Zheng YL, Liu HL (1996) Inhibition of introducing antisense ACC oxidase gene into tomato genome on expression of its endogeous gene. J Huazhong Agric Univ 15:305–309
Zamore PD (2001) RNA interference: listening to the sound of silence. Nat Struct Biol 8:746–750
Zeng Y, Cullen BR (2003) Sequence requirements for micro RNA processing and function in human cells. RNA 9:112–123
Acknowledgements
We are grateful to Prof. Yi Li (Transgenic Plant Research Center, University of Connecticut, USA) and Prof. Zong-ming Cheng (Department of Plant Sciences and Landscapes Systems, University of Tennessee, USA) for their careful reading and useful comments on the manuscript. We sincerely thank Prof. Miao-quan Wang (New-Changzheng Tomato Institute of Shanghai) for providing the seeds of Hezuo 906. This work was supported by Shanghai Municipal Committee of Agriculture (No. 99-1-8). We deeply appreciate the cooperation of the Shanghai YongYe Bio-Engineering Co. in providing the greenhouse facilities and technical help.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by I.S. Chung
Rights and permissions
About this article
Cite this article
Xiong, AS., Yao, QH., Peng, RH. et al. Different effects on ACC oxidase gene silencing triggered by RNA interference in transgenic tomato. Plant Cell Rep 23, 639–646 (2005). https://doi.org/10.1007/s00299-004-0887-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-004-0887-7