Abstract
Self-incompatibility (SI) restricts fertilisation and fruit setting in many tree fruit crops. In apple, we have produced transgenic trees harbouring extra copies of the endogenous S-gene controlling SI. Two independent transgenic genotypes were characterised in detail. Controlled self- and cross-pollination of the flowers of trees from both genotypes over a 3-year-period showed that the transgenic lines produced normal levels of fruit and seeds after selfing. In contrast, the controls produced much less fruit following self- compared to cross-pollination. Fruit set data correlated with the results of microscopic evaluation of pollen tube growth through the pistil, which revealed inhibition after selfing in the controls but not in the transgenic lines. The self-fertile phenotype was associated with the complete absence of pistil S-RNase proteins, which are the products of the targeted S-gene. These results confirm that self-fertility was due to inhibition of expression of the S-RNase gene in the pistil, resulting in un-arrested self-pollen tube growth, and fertilisation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fruit production in many tree fruit crops is dependent on cross-pollination between cultivars. This is due to the existence of a self-incompatibility (SI) mechanism, which is a widespread intraspecific system to prevent self-fertilisation that is controlled by a single S-locus (de Nettancourt 2001). Cross-pollination between compatible cultivars depends on insects as pollen vectors during flowering, and their activity is impaired by inclement weather. Suboptimal pollination efficiencies are one of the factors contributing to low annual fruit crops that may occur during certain years in commercial orchards (Goldway et al. 1999). There is a strong interest in the self-fertile character in many fruit and nut tree crops because self-pollination could ensure more consistently high production yields compared to cross-pollination. This has been apparent in sweet cherry and almond, where cultivars with dysfunctional SI genes have been obtained through mutagenesis and interspecific crosses, respectively (Godini et al. 1998). Although the severity of the SI reaction varies between cultivars and pollination conditions, true self-fertile apple cultivars are commercially non-existent. Moreover, because high crop loads often create lower quality fruit and stimulate a biennial bearing tendency, it is unclear whether self-fertility would be beneficial for apple production. For a scientifically sound evaluation of the impact of self-fertility in apple, isogenic self-incompatible and self-compatible genotypes should be compared. We have, therefore, used a transgenic approach for inactivating the SI mechanism in apple to obtain self-fertile trees. These trees have been analysed under contained greenhouse conditions and may further be used to investigate if self-fertility would be advantageous in the commercial production of apple fruit.
As in many other species displaying gametophytic SI, the SI mechanism in apple is controlled through the action of cytotoxic pistillar proteins, the S-RNases, which are the products of an S-locus gene (Broothaerts et al. 1995). In the case of an incompatible interaction, the S-RNases act on elongating pollen tubes, entering their cytoplasm and degrading the pollen RNA (Golz et al. 2001). Several functionally different S-RNases operate in the species, each encoded by a different allele of the S-gene. In model Solanaceous plant species, it has been shown through both gain-of-function and loss-of-function studies that the presence of the S-RNase in the pistil was sufficient and necessary, respectively, for the development of a SI response (Lee et al. 1994; Murfett et al. 1994).
Several alleles of the S-gene in apple have been identified and cloned (reviewed in Broothaerts 2003). In 1994, we transformed apple with constructs targeted at knocking out S-gene expression in the pistil. The resulting trees were analysed for self-fertilisation behaviour under controlled growth conditions during the 3 years following their maturation (1999–2001). In this paper, we report on the results obtained for two transgenic lines and one control line that were studied in detail at the molecular and phenotypic level.
Materials and methods
Construction of the binary vector
The S-gene silencing construct was derived from the S3 cDNA obtained from a style cDNA library (Broothaerts et al. 1995). PCR was used to amplify the coding sequence from the cloned S3 cDNA from start to stop codon, using primers OWB149 (5′-TCTCTAGAGCTCTTGAACAAACATTATTC-3′) and OWB 150 (5′-ACTCTAGATGAGCTCTTAATACTG-3′). After verifying the amplified product by cloning and sequence analysis, the fragment was ligated in a sense orientation into the SacI site of pFF19, between the enhanced 35S promoter and terminator sequences of cauliflower mosaic virus (CaMV) (Timmermans et al. 1991). The resulting chimaeric expression cassette was subsequently ligated into the HindIII/EcoRI restriction sites of binary vector pGPTV-KAN (Becker et al. 1992), containing the nptII expression cassette near the left T-DNA border, in the opposite direction to the p35S::S3 gene cassette. The recombinant binary vector was electroporated into Agrobacterium tumefaciens strain EHA105.
Apple transformation
Apple leaves from in vitro shoots of cv. Elstar were co-cultivated with the A. tumefaciens strain harboring the binary vector and transgenic calli and shoots were selected on 100 mg/l kanamycin. Upon their appearance 8–16 weeks following transformation, the new shoots were elongated and propagated in vitro, followed by their transfer to ex vitro conditions by grafting the shoots directly onto the stem of an untransformed M.9 rootstock, grown in the greenhouse. Further propagation of the trees occurred by chipbudding. The potted plants were grown to maturity in a greenhouse without supplemental light or heat, except for the first growing season. Transgene insertion was verified by multiplex PCR using primers for the endogenous (intron-containing) S3-allele and both nptII and the p35S::S3 gene fusion (Broothaerts et al. 2001). In the latter case, primers were designed within the S3 cDNA sequence and in the 35S promoter region.
Pollination assays
Greenhouse-grown trees were covered with nylon cages before flower anthesis and flowers were hand-thinned to three flowers per flower cluster. Immediately before pollination, the petals and anthers were removed from the flowers. The pollen used for pollination was derived from either untransformed ‘Elstar’ trees (self-pollination) or from Delbard Jubilé, a cross-compatible variety (cross-pollination). Seven days after pollination, some flowers were removed, fixed in FAA and analysed for pollen tube growth by fluorescence microscopy following softening in Na2SO3 and staining with aniline blue. Fruit set was determined in July (after June drop) and ripe fruit was picked in September, and dissected to determine seed number. Fruit set data were calculated using a minimum of five flowering trees per genotype in 2000 and one or two trees during the other years. Mean values for the 3 years are weighed, taking into account the number of flowers used. Control trees were potted untransformed ‘Elstar’ trees that were either derived from the same tissue culture used for transformation or were obtained as 2-year-old trees from a local supplier.
Western blot analysis
Protein extracts were prepared from five styles using the following extraction buffer: 10 mM thiourea, 100 mM Tris-HCl pH 8.5, 0.5% Tween-20, 2% polyvinylpolypyrrolidone, 14 mM β-mercaptoethanol. Proteins were separated by SDS-PAGE, followed by electroblotting onto a nylon membrane (Hybond N; Amersham, Piscataway, N.J.). Immunodetection was performed by conventional techniques, using a chicken antibody raised against the apple S-RNase conserved peptide sequence CKDPPDKLFT (kindly received from H. Kokko, University of Kuopio, Finland). The secondary antibody was an anti-chicken IgG, coupled to alkaline phosphatase (Sigma, St. Louis, Mo.), and detection used the colorimetric substrate BM purple (Boehringer-Mannheim, Germany).
Results
Production of transgenic apple plants bearing a sense S3-allele construct
The apple variety ‘Elstar’ displays a strong SI response, as reflected by the low fruit and seed set following selfing. Previous investigations had shown that ‘Elstar’ bears the alleles S3 and S5, and cDNAs for both S-alleles have been cloned (Broothaerts et al. 1995; Janssens et al. 1995). S3 is by far the most common S-allele within the domesticated apple species and the presence of its gene product in the pistil has been confirmed by in situ immuno-localization studies, applying an S3-peptide-specific antibody (Certal et al. 1999). Based on the S3 cDNA, a co-suppression construct was prepared for the transformation of ‘Elstar’. This construct contained the full-length sense S3 cDNA sequence driven by the double CaMV 35S promoter (Fig. 1). From 1,500 in vitro leaf explants co-cultivated with A. tumefaciens, 13 independent transgenic plants were produced. These lines were shown to contain an intact S-transgene and nptII gene by PCR. Based on preliminary pollination experiments (see below), transformation events 81 and 102 were chosen for further analysis. Copy number analysis by Southern blotting revealed the integration of T-DNA copies at six and two genomic locations, respectively (data not shown). We previously found the in vitro grown shoots of these two transgenic lines to weakly transcribe the transgene S3 mRNA, as well as producing hybridisation signals of a much larger size (Janssens 1997). Note that the targeted endogenous S-gene is expressed only in the pistil of the flower and not in any other tissues (Broothaerts et al. 1995).
Construction of the binary vector used for the genetic transformation of ‘Elstar’ (see Materials and methods). The S3 coding region was PCR-amplified and cloned into the SacI site of pFF19; the whole expression cassette was then transferred to binary vector pGPTV-KAN. 35S-prom, 35S-ter Duplicated 35S promoter and terminator, respectively, of cauliflower mosaic virus (CaMV); Nos-prom, Ag7-ter promoter of nopaline synthase gene and terminator of gene 7, respectively, both from Agrobacterium tumefaciens; NptII Escherichia coli neomycin phosphotransferase gene conferring resistance to kanamycin; RB, LB right and left border of the T-DNA, respectively
Controlled pollination tests
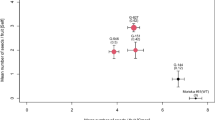
Plants from transgenic lines 81 and 102 were grown to maturity in the greenhouse (Fig. 2A) and were shown to bear normal fertile flowers (Fig. 2B). These plants were analyzed for SI by hand-pollination. Some of the flowers were self-pollinated (using pollen from control ‘Elstar’ trees), and some were cross-pollinated with a compatible pollen donor variety (Delbard Jubilé, bearing the S-alleles S2 and S22; Broothaerts 2003). This experiment was repeated during the following 2 years and the results are summarised in Table 1. In control ‘Elstar’ trees, the mean fruit set following self-pollination during 1999–2001 was 4%, while fruit setting after cross-pollination increased to 30%. In line 81, the mean fruit set following both self- and cross-pollination was identical (31–32%), indicating that both pollen sources were equally efficient at fertilisation. Similar results were obtained for line 102. Using transgenic pollen from either transgenic line for pollination of the control or the transgenic flowers gave comparable results to those obtained when using pollen from untransformed trees. This indicates that the transformation has not affected the SI behaviour of the pollen, i.e. they were arrested in an incompatible style and grew without restriction in a compatible (transgenic) style. Some variation in the fruit setting was observed from year to year, which was likely caused by differences in tree maturity or vigour and in the environmental conditions in the greenhouse, e.g. in 2000 temperatures were generally higher during and after flowering, which promoted fruit setting. The fruits produced after selfing the transgenic genotypes were visually indistinguishable from the ones produced after crossing (Fig. 2C) and the number of seeds was similar in both cases (mean of seven seeds per fruit). In contrast, fruit resulting from selfing contained roughly only half the number of seeds per fruit compared to fruit resulting from cross-pollination on the control trees. This is further evidence that the SI mechanism was not operational in the transgenic trees, while it functioned in the control trees, albeit not completely effectively.
Transgenic apple trees revealing the self-fertility phenotype (line 81). A Mature flowering tree (upper part) growing in the greenhouse. B Flower close-up revealing the anthers (with pollen grains) and the five styles composing the pistil. C Mature fruit developed as a result of self-pollination (two upper fruits) or cross-pollination (lower left fruit)
Further evidence for the inactivation of the SI mechanism also came from fluorescence microscopic analysis of pollen tube growth through the pistil (Fig. 3). In control trees, self-pollen was never observed near the base of the pistil, while cross-pollen was abundantly present in that region at the time of analysis (7 days after hand pollination). In the transgenic plants, both self- and cross-pollen grew efficiently through the styles, and pollen tubes were found at the stylar bases in both cases. This indicates that pollen tube growth following self-pollination in the transgenic trees was apparently not restricted by the SI mechanism.
Pollen tube growth through the style of ‘Elstar’ flowers following self-pollination in control trees (A–C) and in transgenic line 81 (D–F). Fluorescence microscopy pictures of squash preparations of styles, stained for pollen tubes (particularly callose seen along the tube wall and in plugs). A, D Top of the style just below the stigma; B, E halfway down the style; C, F basal part of the style above the ovary (note the weakly fluorescing trichomes at the pistillar base in C)
S-RNase expression in the pistil
To investigate if the absence of a functional SI mechanism in the transgenic trees was the result of inhibition of S-RNase expression, protein extracts of styles were separated by electrophoresis and blotted. An anti-S-RNase antibody raised against a conserved peptide was used to detect the apple S-RNases. The control styles clearly showed the presence of the pistil S-RNases, whereas the transgenic pistils were completely devoid of them (Fig. 4). As the separation method employed did not discriminate between the two ‘Elstar’ S-RNases (S3 and S5), this result reveals that the transgene employed not only inhibited the expression of the targeted S3-allele, but also that of the S5-allele. From native-PAGE and segregation experiments, we know that both S-RNases are expressed in wild-type ‘Elstar’ pistils. S-allele-specific PCR analysis of segregating seedlings following self-pollination of the transgenic lines confirmed that fertilisation resulted from both S3 and S5 pollen grains (data not shown).
Discussion
Genetic engineering of fruit tree species for traits expressed at tree maturity is strongly hampered by long juvenile periods. To our knowledge, this is the first report of a transgenic approach to alter the reproductive phenotype of a fruit tree species. The first true self-fertile trees obtained in apple were shown to stably express self-fertility over several years, without having any obvious adverse effects on tree growth or fruit appearance. The strategy required the inhibition of a single key gene of the SI system, whose function is to specifically prevent self-pollen from fertilising the egg cells. We used a co-suppression approach to achieve S-RNase gene silencing. The presence of RNA transcripts derived from the transgene on RNA blots (Janssens 1997) provided evidence that S-gene downregulation occurred post-transcriptionally. We have detected several copies of the T-DNA construct integrated in the genome of both transgenic lines and we are now in the process of analysing the integration patterns in more detail. This may reveal the presence of (direct or inverted) repeated structures, which have been related to gene silencing in other plants (Muskens et al. 2000). Obviously, the silencing constructs employed in this work are now outdated and may in future work be replaced by (hairpin) constructs that induce gene silencing at a higher frequency (Smith et al. 2000). Our findings further show that, also in the Rosaceae family, expression of the S-RNase genes in the pistil is necessary for the development of a SI response. This is further evidence for the similarity between the SI mechanisms in Rosaceous and Solanaceous plants, despite their divergence several million years ago. According to a recent phylogenetic reconstruction, RNase-based SI systems originated from a common ancestor that predated the divergence of most Angiosperm dicot families (Igic and Kohn 2001). A self-compatible mutant of Japanese pear was previously shown to lack the genomic sequence spanning the S-RNase gene (Sassa et al. 1997), further substantiating the central role of this gene in regulating SI in fruit crops.
This paper describes results obtained on two selected transgenic lines and one control line. Several other transgenic lines were produced in the same transformation experiment. Some of these lines reveal the self-fertile trait, while others are indistinguishable from the controls or show an intermediate phenotype. The detailed analysis of these lines is in progress and will be described elsewhere.
As shown by Western blot staining, the S-RNase signal was completely absent from lanes loaded with protein extract from the transgenic pistils. This indicates that the transgene has resulted in a complete, or at least significant, downregulation of S3-RNase expression. Moreover, because of their apparent similarity, the gel analysis system employed did not resolve the S3- and S5-RNases. The absence of any S-RNase signal on the blot, therefore, indicated that both S-RNase alleles were rendered nonfunctional, despite the use of a cDNA transgene derived from the S3-RNase gene only. The S3 and S5-allele sequence are 77% identical in their coding region. Presumably, this homology has caused the silencing of both endogenous alleles in the transgenic apple trees analysed. In three out of six silenced Petunia inflata plants, both S-alleles had been rendered non-functional, while in the remaining three plants only the S-allele corresponding to the targeted allele was affected (Lee et al. 1994). In these experiments, an antisense S-allele sequence that was driven by its own promoter was used. The homology between the S-alleles affected in the transgenic Petunia plants approximated 80% for the full coding region, but the antisense transgene was only composed of approximately 70% of this region.
Following on from the extensive research on the molecular basis of gametophytic SI during the past 20 years, this report describes the first application of the findings in an economically important, edible crop. Further work should study the behaviour of the transgenic trees under natural growth conditions to find out to what extent self-fertility affects crop yield under various environmental conditions. For the first time, this issue can be addressed appropriately by performing field studies comparing self-fertile and self-incompatible trees in an identical genetic background. The self-fertile trees may furthermore be used as a tool to study inbreeding depression, or for the development of homozygous breeding lines.
References
Becker D, Kemper E, Schell J, Materson R (1992) New plant binary vectors with selectable markers located proximal to the left T-DNA border. Plant Mol Biol 20:1195–1197
Broothaerts W (2003) New findings in apple S-genotype analysis resolve previous confusion and request the re-numbering of some S-alleles. Theor Appl Genet 106:703–714
Broothaerts W, Janssens GA, Proost P, Broekaert WF (1995) cDNA cloning and molecular analysis of two self-incompatibility alleles from apple. Plant Mol Biol 27:499–511
Broothaerts W, Wiersma PA, Lane WD (2001) Multiplex PCR combining transgene and S-allele control primers to simultaneously confirm cultivar identity and transformation in apple. Plant Cell Rep 20:349–353
Certal AC, Sanchez AM, Kokko H, Broothaerts W, Oliveira MM, Feijó JA (1999) S-RNases in apple are expressed in the pistil along the pollen tube growth path. Sex Plant Reprod 12:94–98
Godini A, Palasciano M, Cozzi G, Petruzzi G (1998) Role of self-pollination and horticultural importance of self-compatibility in cherry. Acta Hortic 468:567–574
Goldway M, Shai O, Yehuda H, Matityahu A, Stern RA (1999) ‘Jonathan’ apple is a lower-potency pollenizer of ‘Topred’ than ‘Golden Delicious’ due to partial S-allele incompatibility. J Hortic Sci Biotechnol 74:381–385
Golz JF, Oh H-Y, Su V, Kusaba M, Newbigin E (2001) Genetic analysis of Nicotiana pollen-part mutants is consistent with the presence of an S-ribonuclease inhibitor at the S-locus. Proc Natl Acad Sci USA 98:15372–15376
Igic B, Kohn JR (2001) Evolutionary relationships among self-incompatibility RNases. Proc Natl Acad Sci USA 98:13167–13171
Janssens G (1997) Molecular analysis of the self-incompatibility (S-) gene in apple (Malus x domestica Borkh.). S-genotyping of apple cultivars and (anti)sense suppression of the S-gene. PhD dissertation no. 350, Fac Agric Appl Biol Sci, Katholieke Universiteit Leuven, Belgium
Janssens GA, Goderis IJ, Broekaert WF, Broothaerts W (1995) A molecular method for S-allele identification in apple based on allele-specific PCR. Theor Appl Genet 91:691–698
Lee H-S, Huang S, Kao T-h (1994) S-proteins control rejection of incompatible pollen in Petunia inflata. Nature 367:560–563
Murfett J, Atherton TL, Mou B, Gasser CS, McClure BA (1994) S-RNase expressed in transgenic Nicotiana causes S-allele-specific pollen rejection. Nature 367:563–566
Muskens MWM, Vissers APA, Mol JNM (2000) Role of inverted DNA repeats in transcriptional and post-transcriptional gene silencing. Plant Mol Biol 43:243–260
Nettancourt D de (2001) Incompatibility and incongruity in wild and cultivated plants. Springer, Berlin Heidelberg New York
Sassa H, Hirano H, Nishio T, Koba T (1997) Style-specific self-compatible mutation caused by deletion of the S-RNase gene in Japanese pear (Pyrus serotina). Plant J 12:223–227
Smith NA, Singh SP, Wang M-B, Stoutjesdijk PA, Green AG, Waterhouse PM (2000) Total silencing by intron-spliced hairpin RNAs. Nature 407:319–320
Timmermans MCP, Maliga P, Vieira J, Messing J (1991) The pFF plasmids: cassettes utilizing CaMV sequences for expression of foreign genes in plants. J Biotechnol 14:333–344
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by P. Debergh
Rights and permissions
About this article
Cite this article
Broothaerts, W., Keulemans, J. & Van Nerum, I. Self-fertile apple resulting from S-RNase gene silencing. Plant Cell Rep 22, 497–501 (2004). https://doi.org/10.1007/s00299-003-0716-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-003-0716-4