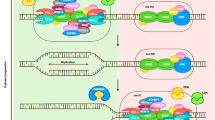
Abstract
The spatial and temporal organization of genome duplication, also referred to as the replication program, is defined by the distribution and the activities of the sites of replication initiation across the genome. Alterations to the replication profile are associated with cell fate changes during development and in pathologies, but the importance of undergoing S phase with distinct and specific programs remains largely unexplored. We have recently addressed this question, focusing on the interplay between the replication program and genome maintenance. In particular, we demonstrated that when cells encounter challenges to DNA synthesis, the organization of DNA replication drives the response to replication stress that is mediated by the ATR/Rad3 checkpoint pathway, thus shaping the pattern of genome instability along the chromosomes. In this review, we present the major findings of our study and discuss how they may bring new perspectives to our understanding of the biological importance of the replication program.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
DNA replication is an essential and conserved process that is carried out by a complex and highly regulated machinery. DNA synthesis initiates at sites distributed along the chromosomes called replication origins, which are activated at different times during S phase and are used with particular frequencies, or efficiencies, in a population of cells. The timings and efficiencies of origin usage, together with their genome-wide distribution, delineate the program of DNA replication. This program is altered in response to intrinsic and extrinsic signals, with signature patterns described for distinct cell types and growth conditions (Hiratani et al. 2008, 2010; Pope et al. 2010; Ryba et al. 2011; Wu and Nurse 2014; Perrot et al. 2018; Singh and Wu 2018). However, although a large body of work has been dedicated to characterizing such profiles and investigating their regulation, it remains an open question whether and how these programs contribute to cellular function.
One context in which the control of DNA replication may be particularly critical is in situations of replication stress: challenges to DNA synthesis can lead to the generation of lesions and errors in the genetic material (Hills and Diffley 2014; Mazouzi et al. 2014; Kotsantis et al. 2018). To preserve genome integrity, cells activate checkpoint pathways that not only regulate cell cycle progression and gene expression but also modulate DNA synthesis, and in particular origin activation (Saldivar et al. 2017). Our recent study explores the interplay between the replication program, the checkpoint inhibition of origin firing, and genome instability in replication stress conditions (Gómez-Escoda and Wu 2018). Using the fission yeast Schizosaccharomyces pombe as a model system, we uncovered a critical role for the organization of DNA replication in delineating the landscape of genetic instability, shedding new light on how this architecture may modulate both genome maintenance and evolution.
The spatiotemporal program of DNA replication
The organization of DNA replication in chromosomal domains of timing and efficiency has been observed in diverse organisms, suggesting that such configurations are a common feature of replication in eukaryotes. Interestingly, replication programs are conserved between related species of yeast as well as between mouse and human cells (Ryba et al. 2010; Yaffe et al. 2010; Muller and Nieduszynski 2012). At the same time, these programs are flexible, and signature profiles have been associated with distinct developmental states (Hiratani et al. 2010). For instance, there is a clear relationship between the replication program and cell fate, as conserved replication timing domains have been identified in mouse and human embryonic stem cells, constituting a shared feature of pluripotence (Hiratani et al. 2008; Palou et al. 2017). In addition, development and differentiation are accompanied by alterations in replication patterns in organisms ranging from worms to frogs to humans (Hyrien et al. 1995; Hiratani et al. 2008, 2010; Desprat et al. 2009; Pope et al. 2010; Rodríguez-Martínez et al. 2017). Finally, the profile of origin usage is sensitive to nutritional conditions, as found in fission yeast cells that have been temporarily starved of nitrogen (Wu and Nurse 2014). Therefore, the replication program displays a dynamic specificity in response to internal and external signals.
Why might cells use different replication programs in particular contexts? One possibility is that these profiles simply arise as a consequence of the physiological changes that are occurring in distinct environments. However, the conservation of specific organizations of DNA replication described above suggests that it may be important for cellular function apart from simply ensuring the duplication of the genome. For example, links between replication and gene expression have been found in multiple systems. In mammalian cells, early replicating regions are correlated with transcriptional activity, and alterations in replication timing during differentiation are associated with changes in chromosome architecture and transcription (Rivera-Mulia et al. 2015, 2018; Pourkarimi et al. 2016; Rodríguez-Martínez et al. 2017). Along the same lines, a direct role for replication timing in the control of gene expression has been demonstrated in the budding yeast, where this parameter has been implicated in regulating the level of histone transcription (Müller and Nieduszynski 2017). Furthermore, we have previously shown that the profile of origin usage in the fission yeast determines the distribution of meiotic recombination along the chromosomes (Wu and Nurse 2014). While these observations hint toward replication being involved in diverse processes, we are only beginning to understand the biological roles of the organization of DNA replication.
Although the genetic material is particularly fragile during its duplication, little is known about whether and how the replication program contributes to genome integrity. Intriguingly, genetic analyses have established correlations between replication timing and mutation frequencies in budding yeast (Lang and Murray 2011), and whole-genome sequencing of normal and cancer cell lines has revealed differences in the types and distributions of mutations that are found in early vs. late replicating chromosomal regions (Koren et al. 2012; Liu et al. 2013; Sima and Gilbert 2014; Tomkova et al. 2018). These findings thus hint at an intimate coupling between the replication program and genome maintenance.
Challenges to DNA synthesis: causes and consequences
During DNA replication, the genetic material is particularly vulnerable to damage and to acquiring errors. On one hand, even during an “unperturbed” S phase, cells encounter a variety of insults that impede DNA synthesis. For instance, markers of genotoxic stress are detected during replication in embryonic stem cells (Ahuja et al. 2016). On the other hand, replication stress can be induced by extrinsic DNA damaging agents, and it can also arise due to intracellular challenges such as limiting levels of nucleotides and replication factors, difficult to replicate sites with secondary DNA structures, chromatin accessibility, or collisions between the replication and transcription machineries (Zeman and Cimprich 2014; Mazouzi et al. 2014; Tubbs and Nussenzweig 2017). These challenges can then lead to the slowing, stalling, or collapsing of replication forks. If not properly managed, replication stress may, therefore, result in DNA breaks as well as incomplete DNA synthesis with cells entering mitosis prior to completing replication.
Notably, the genome instability that arises from replication stress is considered to be a key contributor to cancer. Genetic alterations are a fundamental feature of cancer cells and a driving force in tumorigenesis. Recent studies have identified thousands of coding mutations that are heterogeneous within and between tumors (McGranahan and Swanton 2017), with mutations in established cancer genes present in only a subset of cells within a tumor. This suggests that tumor fitness may be modulated by the numerous other genetic alterations that are present, which may be generated in part by replication stress-associated instability (Negrini et al. 2010). These findings thus highlight how our understanding of cancer progression will benefit from investigating the cellular response to replication challenges and the mechanisms that modulate these pathways.
The checkpoint response to replication stress
Eukaryotic cells utilize a number of mechanisms to protect the genome from endogenous and exogenous sources of DNA damage (Palou et al. 2017; Saldivar et al. 2017). Challenges to DNA synthesis that disrupt the coordination between replicative helicase and DNA polymerase lead to the formation of single-stranded DNA and the activation of the S phase/ATR checkpoint (Byun et al. 2005). The subsequent modification of a number of substrates by checkpoint kinases brings about specific changes in gene expression, cell cycle arrest, replication fork protection and subsequent restart, and inhibition of replication initiation (Saldivar et al. 2017; Mikolaskova et al. 2018; Villa-Hernández and Bermejo 2018). These mechanisms are conserved throughout eukaryotes, and the importance of much of this regulation has been previously described. However, the impact of the checkpoint control of origin firing, first reported over 20 years ago (Santocanale and Diffley 1998; Shirahige et al. 1998), has remained surprisingly elusive. This inhibition of origins occurs through modulation of the CDK (cyclin-dependent kinase) and DDK (Dbf4-dependent kinase) pathways (Lopez-Mosqueda et al. 2010; Zegerman and Diffley 2010), and initial studies in budding yeast suggested that inefficient and late-firing origins are specifically targeted for inhibition (Santocanale and Diffley 1998; Shirahige et al. 1998; Santocanale et al. 1999). However, the selection of these target origins, the extent of inappropriate initiation in the absence of the checkpoint, and the significance of this regulation remained contested. As the conservation of the checkpoint regulation of origin activation from yeast to human indicates a central role for this mechanism in the DNA damage response, we have taken advantage of this process to investigate the interplay between the organization of DNA replication and genome maintenance (Gómez-Escoda and Wu 2018).
Interplay between the replication program and the response to replication stress
In the fission yeast Schizosaccharomyces pombe, replication stress activates the conserved Rad3/ATR checkpoint kinase, which then signals through Cds1/CHK1 to modify a number of substrates that include replication factors and cell cycle regulators (Bentley et al. 1996; Lindsay et al. 1998; Edwards et al. 1999; Labib and De Piccoli 2011; Willis et al. 2016). Previous work in this organism reported differing results for the checkpoint inhibition of origin firing, identifying 2–28% of the origins in the genome as sensitive to this control (Feng et al. 2006; Heichinger et al. 2006; Hayashi et al. 2007; Mickle et al. 2007). For our study, we therefore began by evaluating origin activity in Rad3/ATR-deleted cells following exposure to the ribonucleotide reductase inhibitor hydroxyurea (HU), which triggers replication stress through depletion of nucleotide pools. As we surmised that the detection of checkpoint-inhibited initiation sites in earlier studies may have been impeded by the use of high HU concentrations, we applied a moderate level of the drug that permits slow progression through S phase. Our results showed that in rad3Δ cells treated with 6 mM HU, origin firing is significantly deregulated at a subset of initiation sites in the genome. Specifically, we identified 176 deregulated sites out of 876 total origins that are increased by up to 25% efficiency in these conditions. In contrast to the previous findings, these checkpoint-inhibited origins (CIOs) span a broad spectrum of activities. However, regardless of their individual characteristics, they are clearly clustered in late-firing and inefficient regions of the genome, with a large portion of the origins in such domains (up to ~ 70%) being deregulated in HU-treated rad3Δ cells. Together with this signature distribution of CIOs, we established the quantitative profile of the extent of inappropriate firing in HU-treated rad3Δ cells. This then allowed us to probe how the organization of genome duplication impacts genome maintenance.
First, we evaluated the consequences of deregulated origin firing for genome instability, using the single-stranded DNA (ssDNA) binding protein Ssb1/RPA and the DNA repair protein Rad52 as molecular markers. Our data demonstrated abnormal ssDNA formation and high levels of Rad52 recruitment at CIOs. We further found that upon HU exposure, the changes in RPA and Rad52 binding between rad3Δ vs. wild type were correlated with the extent of origin deregulation: the increase in activity of a CIO in rad3Δ was directly proportional to the increase in RPA and Rad52 occupancy at this site. We then evaluated the relationship between the genome-wide profiles of RPA and Rad52 with respect to inappropriate replication initiation. Taking into account all origins as well as all RPA- and Rad52-binding sites in HU-treated cells, our analysis revealed a clear correspondence between the domains of origin deregulation vs. the densities of RPA and Rad52 loci along the chromosomes in a rad3Δ mutant (Fig. 1, middle and bottom panels). Our results thus suggested unscheduled origin firing as a critical determinant of the overall pattern of genome instability. Strikingly, the profile of origin deregulation in rad3Δ cells under replication stress showed a strong negative correlation with the wild-type program of origin efficiencies, consistent with our observation that origins in late-firing and inefficient regions of the genome are inhibited by the checkpoint (Fig. 1, compare top panel with middle and bottom panels). Taken together, these results led us to propose that the replication program defines the checkpoint regulation of origin firing in replication stress conditions, with consequences for the landscape of genome instability.
Schematic representation of the relationship between the replication program, origin deregulation, and genome instability. Top: replication program in wild-type cells. Origin activity (dashed black line) represents the efficiency of origin usage, which is measured by the frequency of initiation at a given origin in a population of cells. Middle: change in origin efficiency (dashed red line) between rad3Δ and wild-type cells in replication stress conditions. Bottom: density of DNA damage hotspots (dashed blue line), as assayed by Rad52 binding, in rad3Δ cells exposed to replication stress. x-axis: chromosome coordinates, y-axis: origin efficiency (top), changes in origin usage (middle), and density of DNA damage hotspots (bottom). The replication program shows strong negative correlations with origin deregulation and the level of genome instability. Inappropriate initiation and DNA damage hotspots are clustered in inefficient and late-firing replication domains
To test this model, we took advantage of the rif1Δ genetic background in which replication initiation is altered genome-wide (Hayano et al. 2012; Cornacchia et al. 2012; Yamazaki et al. 2012). We first determined the effect of this modified replication program on origin deregulation in the absence of rad3 and found a complete redistribution of checkpoint-inhibited origins: these rif1-CIOs are more uniformly positioned along the chromosomes and no longer clustered in distinct domains (Fig. 2). We then assessed whether the changes in checkpoint inhibition are accompanied by alterations in Rad52 binding. Our data showed that the correlation between origin deregulation and Rad52 accumulation observed in HU-treated checkpoint-defective cells is maintained in the rif1Δ rad3Δ genetic background. Strikingly, Rad52 sites in rif1Δrad3Δ cells were no longer clustered in the same genomic domains as in rad3Δ, revealing a reorganization of Rad52 loci. Collectively, these findings support our model that the replication program governs the checkpoint regulation of origin firing and the profile of genome instability when cells are challenged for DNA synthesis.
Alteration of the replication program has direct consequences for the checkpoint regulation of origin firing. Left: schematic representation of the replication program in wild-type (top) vs. rif1Δ (bottom) cells. Origin activity (dashed black lines) represents the efficiency of origin usage. x-axis: chromosome coordinates, y-axis: origin efficiency. The distinct replication domains apparent in wild-type cells are lost in rif1Δ. Right: Inappropriate initiation in rad3Δ (top, dashed red line) and rif1Δ rad3Δ (bottom, dashed green line) cells. Consistent with the changes in the replication program, the signature pattern of origin deregulation in rad3Δ is reorganized in rif1Δ rad3Δ. This is accompanied by changes in the profiles of DNA damage hotspots (not shown). x-axis: chromosome coordinates, y-axis: changes in origin efficiency
Perspectives for genome organization and evolution
Our results demonstrate that the control of origin usage by the Rad3/ATR pathway in replication stress conditions is a key aspect of its function in genome maintenance. Why might genome duplication need to be organized for the response to such challenges? One possibility is that the use of particular replication programs may promote elevated local concentrations of checkpoint factors that may result in a more efficient response, either for checkpoint signaling or for DNA repair (Rivera-Mulia et al. 2015). Related to this idea, particular regions may be more accessible to the recombination machinery or for the search for homology, which has been proposed to be affected by the spatial organization of the genome (Misteli and Soutoglou 2009; Nagai et al. 2010). Moreover, evolution may have favored an arrangement in which potential sites of DNA damage are enriched in regions, where mutations have a lower likelihood to be deleterious. The replication program may also be important in the context of an “unperturbed” cell cycle, where cells are in fact constantly exposed to endogenous challenges to replication (Zeman and Cimprich 2014). In support of this idea, ATR function is essential in vertebrate cells (Eykelenboom et al. 2013), and embryonic stem cells display markers of replication stress that are dependent on ATR (Ahuja et al. 2016). Therefore, how the genome is organized for its duplication may ultimately have consequences for its integrity in the context of normal and challenging situations.
While our study has assessed the immediate consequences of origin deregulation, it is also interesting to consider the long-term impacts of the replication program. For instance, our work may provide insight into how this aspect of genome architecture may contribute to genome evolution. Indeed, our demonstration that replication origins are hotspots of DNA damage when they are fired inappropriately in stress conditions is consistent with comparative genomic analyses and laboratory evolution experiments in budding yeast that indicate a co-localization of chromosomal breakpoints with origins (Di Rienzi et al. 2009; Gordon et al. 2009). The coupling between the replication program and genome instability that we uncovered may also help us understand the correlations between replication timing and the differences in the types and frequencies of mutations that are found in different genomic regions (Lang and Murray 2011; Koren et al. 2012; Liu et al. 2013; Lu et al. 2014; Polak et al. 2015; Tomkova et al. 2018). Furthermore, our results may be pertinent in the context of natural situations in which checkpoints are inefficient or inactive. This is, for instance, the case in the rapid and synchronous embryonic cell cycles observed in many metazoa (Kermi et al. 2017). Finally, checkpoint deficiencies, replication stress, and altered patterns of origin usage have also all been identified as features of cancer cells (Halazonetis et al. 2008; Ciccia and Elledge 2010; Bester et al. 2011; Donley and Thayer 2013). Our results may therefore have implications for how the replication program may critically contribute to heterogeneity, somatic evolution, and cancer development. All together, given the flexibility of the organization of DNA replication in response to internal and external stimuli, it is tempting to speculate that the way cells duplicate their genetic material plays crucial roles in adaptation and evolution in the long term.
Conclusion
The organization of DNA replication is a conserved feature of eukaryotic genomes. Its remarkable similarity between related species highlights potential evolutionary constraints and suggests a biological importance for this architecture. Future studies in this area of research will open exciting frontiers in our understanding of how the essential process of DNA replication may not only serve to copy the genome but also to modulate cellular pathways and shape genome evolution.
References
Ahuja AK, Jodkowska K, Teloni F et al (2016) A short G1 phase imposes constitutive replication stress and fork remodelling in mouse embryonic stem cells. Nat Commun 7:10660. https://doi.org/10.1038/ncomms10660
Bentley NJ, Holtzman DA, Flaggs G et al (1996) The Schizosaccharomyces pombe rad3 checkpoint gene. EMBO J 15:6641–6651
Bester AC, Roniger M, Oren YS et al (2011) Nucleotide deficiency promotes genomic instability in early stages of cancer development. Cell 145:435–446. https://doi.org/10.1016/j.cell.2011.03.044
Byun TS, Pacek M, Yee M-C et al (2005) Functional uncoupling of MCM helicase and DNA polymerase activities activates the ATR-dependent checkpoint. Genes Dev 19:1040–1052. https://doi.org/10.1101/gad.1301205
Ciccia A, Elledge SJ (2010) The DNA damage response: making it safe to play with knives. Mol Cell 40:179–204. https://doi.org/10.1016/j.molcel.2010.09.019
Cornacchia D, Dileep V, Quivy J-P et al (2012) Mouse Rif1 is a key regulator of the replication-timing programme in mammalian cells. EMBO J 31:3678–3690. https://doi.org/10.1038/emboj.2012.214
Desprat R, Thierry-Mieg D, Lailler N et al (2009) Predictable dynamic program of timing of DNA replication in human cells. Genome Res 19:2288–2299. https://doi.org/10.1101/gr.094060.109
Di Rienzi SC, Collingwood D, Raghuraman MK, Brewer BJ (2009) Fragile genomic sites are associated with origins of replication. Genome Biol Evol 1:350–363. https://doi.org/10.1093/gbe/evp034
Donley N, Thayer MJ (2013) DNA replication timing, genome stability and cancer: late and/or delayed DNA replication timing is associated with increased genomic instability. Semin Cancer Biol 23:80–89. https://doi.org/10.1016/j.semcancer.2013.01.001
Edwards RJ, Bentley NJ, Carr AM (1999) A Rad3-Rad26 complex responds to DNA damage independently of other checkpoint proteins. Nat Cell Biol 1:393–398. https://doi.org/10.1038/15623
Eykelenboom JK, Harte EC, Canavan L et al (2013) ATR activates the S-M checkpoint during unperturbed growth to ensure sufficient replication prior to mitotic onset. Cell Rep 5:1095–1107. https://doi.org/10.1016/j.celrep.2013.10.027
Feng W, Collingwood D, Boeck ME et al (2006) Genomic mapping of single-stranded DNA in hydroxyurea-challenged yeasts identifies origins of replication. Nat Cell Biol 8:148–155. https://doi.org/10.1038/ncb1358
Gómez-Escoda B, Wu P-YJ (2018) The organization of genome duplication is a critical determinant of the landscape of genome maintenance. Genome Res. https://doi.org/10.1101/gr.224527.117
Gordon JL, Byrne KP, Wolfe KH (2009) Additions, losses, and rearrangements on the evolutionary route from a reconstructed ancestor to the modern Saccharomyces cerevisiae genome. PLoS Genet 5:e1000485. https://doi.org/10.1371/journal.pgen.1000485
Halazonetis TD, Gorgoulis VG, Bartek J (2008) An oncogene-induced DNA damage model for cancer development. Science 319:1352–1355. https://doi.org/10.1126/science.1140735
Hayano M, Kanoh Y, Matsumoto S et al (2012) Rif1 is a global regulator of timing of replication origin firing in fission yeast. Genes Dev 26:137–150. https://doi.org/10.1101/gad.178491.111
Hayashi M, Katou Y, Itoh T et al (2007) Genome-wide localization of pre-RC sites and identification of replication origins in fission yeast. EMBO J 26:1327–1339. https://doi.org/10.1038/sj.emboj.7601585
Heichinger C, Penkett CJ, Bähler J, Nurse P (2006) Genome-wide characterization of fission yeast DNA replication origins. EMBO J 25:5171–5179. https://doi.org/10.1038/sj.emboj.7601390
Hills SA, Diffley JFX (2014) DNA replication and oncogene-induced replicative stress. Curr Biol 24:R435–R444. https://doi.org/10.1016/j.cub.2014.04.012
Hiratani I, Ryba T, Itoh M et al (2008) Global reorganization of replication domains during embryonic stem cell differentiation. PLoS Biol 6:e245. https://doi.org/10.1371/journal.pbio.0060245
Hiratani I, Ryba T, Itoh M et al (2010) Genome-wide dynamics of replication timing revealed by in vitro models of mouse embryogenesis. Genome Res 20:155–169. https://doi.org/10.1101/gr.099796.109
Hyrien O, Maric C, Méchali M (1995) Transition in specification of embryonic metazoan DNA replication origins. Science 270:994–997
Kermi C, Furno Lo E, Maiorano D (2017) Regulation of DNA replication in early embryonic cleavages. Genes (Basel). https://doi.org/10.3390/genes8010042
Koren A, Polak P, Nemesh J et al (2012) Differential relationship of DNA replication timing to different forms of human mutation and variation. Am J Hum Genet 91:1033–1040. https://doi.org/10.1016/j.ajhg.2012.10.018
Kotsantis P, Petermann E, Boulton SJ (2018) Mechanisms of oncogene-induced replication stress: jigsaw falling into place. Cancer Discov 8:537–555. https://doi.org/10.1158/2159-8290.CD-17-1461
Labib K, De Piccoli G (2011) Surviving chromosome replication: the many roles of the S-phase checkpoint pathway. Philos Trans R Soc B Biol Sci 366:3554–3561. https://doi.org/10.1098/rstb.2011.0071
Lang GI, Murray AW (2011) Mutation rates across budding yeast chromosome VI are correlated with replication timing. Genome Biol Evol 3:799–811. https://doi.org/10.1093/gbe/evr054
Lindsay HD, Griffiths DJ, Edwards RJ et al (1998) S-phase-specific activation of Cds1 kinase defines a subpathway of the checkpoint response in Schizosaccharomyces pombe. Genes Dev 12:382–395
Liu L, De S, Michor F (2013) DNA replication timing and higher-order nuclear organization determine single-nucleotide substitution patterns in cancer genomes. Nat Commun 4:1502. https://doi.org/10.1038/ncomms2502
Lopez-Mosqueda J, Maas NL, Jonsson ZO et al (2010) Damage-induced phosphorylation of Sld3 is important to block late origin firing. Nature 467:479–483. https://doi.org/10.1038/nature09377
Lu J, Li H, Hu M et al (2014) The distribution of genomic variationsin human iPSCs is related to replication-timing reorganization during reprogramming. Cell Rep 7:70–78. https://doi.org/10.1016/j.celrep.2014.03.007
Mazouzi A, Velimezi G, Loizou JI (2014) DNA replication stress—causes, resolution and disease. Exp Cell Res 329:85–93. https://doi.org/10.1016/j.yexcr.2014.09.030
McGranahan N, Swanton C (2017) Clonal heterogeneity and tumor evolution: past, present, and the future. Cell 168:613–628. https://doi.org/10.1016/j.cell.2017.01.018
Mickle KL, Ramanathan S, Rosebrock A et al (2007) Checkpoint independence of most DNA replication origins in fission yeast. BMC Mol Biol 8:112. https://doi.org/10.1186/1471-2199-8-112
Mikolaskova B, Jurcik M, Cipakova I et al (2018) Maintenance of genome stability: the unifying role of interconnections between the DNA damage response and RNA-processing pathways. Curr Genet 64:971–983. https://doi.org/10.1007/s00294-018-0819-7
Misteli T, Soutoglou E (2009) The emerging role of nuclear architecture in DNA repair and genome maintenance. Nature 10:243–254. https://doi.org/10.1038/nrm2651
Muller CA, Nieduszynski CA (2012) Conservation of replication timing reveals global and local regulation of replication origin activity. Genome Res 22:1953–1962. https://doi.org/10.1101/gr.139477.112
Müller CA, Nieduszynski CA (2017) DNA replication timing influences gene expression level. J Cell Biol 216:1907–1914. https://doi.org/10.1083/jcb.201701061
Nagai S, Heun P, Gasser SM (2010) Roles for nuclear organization in the maintenance of genome stability. Epigenomics 2:289–305. https://doi.org/10.2217/epi.09.49
Negrini S, Gorgoulis VG, Halazonetis TD (2010) Genomic instability—an evolving hallmark of cancer. Nat Rev Mol Cell Biol 11:220–228. https://doi.org/10.1007/BF01882039
Palou R, Palou G, Quintana DG (2017) A role for the spindle assembly checkpoint in the DNA damage response. Curr Genet 63:275–280. https://doi.org/10.1007/s00294-016-0634-y
Perrot A, Millington CL, Gómez-Escoda B et al (2018) CDK activity provides temporal and quantitative cues for organizing genome duplication. PLoS Genet 14:e1007214. https://doi.org/10.1371/journal.pgen.1007214
Polak P, Karlić R, Koren A et al (2015) Cell-of-origin chromatin organization shapes the mutational landscape of cancer. Nature 518:360–364. https://doi.org/10.1038/nature14221
Pope BD, Hiratani I, Gilbert DM (2010) Domain-wide regulation of DNA replication timing during mammalian development. Chromosome Res 18:127–136. https://doi.org/10.1007/s10577-009-9100-8
Pourkarimi E, Bellush JM, Whitehouse I (2016) Spatiotemporal coupling and decoupling of gene transcription with DNA replication origins during embryogenesis in C. elegans. Elife. https://doi.org/10.7554/eLife.21728
Rivera-Mulia JC, Buckley Q, Sasaki T et al (2015) Dynamic changes in replication timing and gene expression during lineage specification of human pluripotent stem cells. Genome Res 25:1091–1103. https://doi.org/10.1101/gr.187989.114
Rivera-Mulia JC, Dimond A, Vera D et al (2018) Allele-specific control of replication timing and genome organization during development. Genome Res. https://doi.org/10.1101/gr.232561.117
Rodríguez-Martínez M, Pinzón N, Ghommidh C et al (2017) The gastrula transition reorganizes replication-origin selection in Caenorhabditis elegans. Nat Struct Mol Biol 24:290–299. https://doi.org/10.1038/nsmb.3363
Ryba T, Hiratani I, Lu J et al (2010) Evolutionarily conserved replication timing profiles predict long-range chromatin interactions and distinguish closely related cell types. Genome Res 20:761–770. https://doi.org/10.1101/gr.099655.109
Ryba T, Hiratani I, Sasaki T et al (2011) Replication timing: a fingerprint for cell identity and pluripotency. PLoS Comput Biol 7:e1002225. https://doi.org/10.1371/journal.pcbi.1002225
Saldivar JC, Cortez D, Cimprich KA (2017) The essential kinase ATR: ensuring faithful duplication of a challenging genome. Nat Rev Mol Cell Biol 18:622–636. https://doi.org/10.1038/nrm.2017.67
Santocanale C, Diffley JF (1998) A Mec1- and Rad53-dependent checkpoint controls late-firing origins of DNA replication. Nature 395:615–618. https://doi.org/10.1038/27001
Santocanale C, Sharma K, Diffley JF (1999) Activation of dormant origins of DNA replication in budding yeast. Genes Dev 13:2360–2364
Shirahige K, Hori Y, Shiraishi K et al (1998) Regulation of DNA-replication origins during cell-cycle progression. Nature 395:618–621. https://doi.org/10.1038/27007
Sima J, Gilbert DM (2014) Complex correlations: replication timing and mutational landscapes during cancer and genome evolution. Curr Opin Genet Dev 25:93–100. https://doi.org/10.1016/j.gde.2013.11.022
Singh B, Wu P-YJ (2018) Regulation of the program of DNA replication by CDK: new findings and perspectives. Curr Genet. https://doi.org/10.1007/s00294-018-0860-6
Tomkova M, Tomek J, Kriaucionis S, Schuster-Böckler B (2018) Mutational signature distribution varies with DNA replication timing and strand asymmetry. Genome Biol 19:129. https://doi.org/10.1186/s13059-018-1509-y
Tubbs A, Nussenzweig A (2017) Endogenous DNA damage as a source of genomic instability in cancer. Cell 168:644–656. https://doi.org/10.1016/j.cell.2017.01.002
Villa-Hernández S, Bermejo R (2018) Cohesin dynamic association to chromatin and interfacing with replication forks in genome integrity maintenance. Curr Genet 64:1005–1013. https://doi.org/10.1007/s00294-018-0824-x
Willis NA, Zhou C, Elia AEH et al (2016) Identification of S-phase DNA damage-response targets in fission yeast reveals conservation of damage-response networks. Proc Natl Acad Sci USA 113:E3676–E3685. https://doi.org/10.1073/pnas.1525620113
Wu P-YJ, Nurse P (2014) Replication origin selection regulates the distribution of meiotic recombination. Mol Cell 53:655–662. https://doi.org/10.1016/j.molcel.2014.01.022
Yaffe E, Farkash-Amar S, Polten A et al (2010) Comparative analysis of DNA replication timing reveals conserved large-scale chromosomal architecture. PLoS Genet 6:e1001011. https://doi.org/10.1371/journal.pgen.1001011
Yamazaki S, Ishii A, Kanoh Y et al (2012) Rif1 regulates the replication timing domains on the human genome. EMBO J 31:3667–3677. https://doi.org/10.1038/emboj.2012.180
Zegerman P, Diffley JFX (2010) Checkpoint-dependent inhibition of DNA replication initiation by Sld3 and Dbf4 phosphorylation. Nature 467:474–478. https://doi.org/10.1038/nature09373
Zeman MK, Cimprich KA (2014) Causes and consequences of replication stress. Nat Cell Biol 16:2–9. https://doi.org/10.1038/ncb2897
Acknowledgements
We thank Damien Coudreuse for critical reading of the manuscript. This work was supported by funding from the Institut National du Cancer (INCA, PLBIO 15-043) and the Région Bretagne. We apologize to any authors whose work was not cited due to space restrictions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by M. Kupiec.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Singh, B., Wu, PY.J. Linking the organization of DNA replication with genome maintenance. Curr Genet 65, 677–683 (2019). https://doi.org/10.1007/s00294-018-0923-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-018-0923-8