Abstract
Yeasts create multicellular structures of varying complexity, such as more complex colonies and biofilms and less complex flocs, each of which develops via different mechanisms. Colony biofilms originate from one or more cells that, through growth and division, develop a complicated three-dimensional structure consisting of aerial parts, agar-embedded invasive parts and a central cavity, filled with extracellular matrix. In contrast, flocs arise relatively quickly by aggregation of planktonic cells growing in liquid cultures after they reach the appropriate growth phase and/or exhaust nutrients such as glucose. Creation of both types of structures is dependent on the presence of flocculins: Flo11p in the former case and Flo1p in the latter. We recently showed that formation of both types of structures by wild Saccharomyces cerevisiae strain BR-F is regulated via transcription regulators Tup1p and Cyc8p, but in a divergent manner. Biofilm formation is regulated by Cyc8p and Tup1p antagonistically: Cyc8p functions as a repressor of FLO11 gene expression and biofilm formation, whereas Tup1p counteracts the Cyc8p repressor function and positively regulates biofilm formation and Flo11p expression. In addition, Tup1p stabilizes Flo11p probably by repressing a gene coding for a cell wall or extracellular protease that is involved in Flo11p degradation. In contrast, formation of BR-F flocs is co-repressed by the Cyc8p–Tup1p complex. These findings point to different mechanisms involved in yeast multicellularity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Cyc8p and Tup1p in transition between colony and biofilm mode of multicellularity
Two major types of colony structures are formed by Saccharomyces cerevisiae strains: smooth colonies created by the majority of laboratory strains and structured colony biofilms formed by various wild strains isolated from natural environment (Palkova and Vachova 2016; Palkova et al. 2014; Vachova and Palkova 2018). Wild strains possess the ability to flip multicellular structure formation between smooth colony and structured biofilm modes, depending on nutrient availability. In addition, wild strains can undergo stable switching under rich nutritive conditions during which they turn off biofilm-specific traits and begin to form smooth colonies, independently of nutrient sources. This process, called domestication, is reversible (Stovicek et al. 2014). So far identified biofilm-specific traits that are switched off during domestication include cell adhesion to and invasion of semisolid material (such as agar), cell–cell adhesion via extracellular fibers, dependent on the presence of Flo11p adhesin and production of hydratable extracellular matrix (Stovicek et al. 2010, 2014; Vachova et al. 2011).
Recently, we discovered an important role of transcription regulators Cyc8p and Tup1p in regulation of biofilm-specific traits and thus in modulating colony complexity between smooth and biofilm modes (Nguyen et al. 2018). Cyc8p (Ssn6p) and Tup1p are conserved in eukaryotic organisms including mammals and have mainly been known as a transcriptional corepressor, consisting of four molecules of Tup1p and one molecule of Cyc8p (Smith and Johnson 2000; Varanasi et al. 1996). Later, Wong and Struhl (2011) proposed that Cyc8p–Tup1p repression occurs by masking activation domains and blocking activator-mediated recruitment of coactivators. Accordingly, Cyc8–Tup1p binding to DNA is mediated via its interaction with sequence-specific DNA-binding interacting proteins, both repressors and activators (Kliewe et al. 2017; Smith and Johnson 2000). By studying multicellular structures formed by S. cerevisiae strains with modified level of Cyc8p and Tup1p, we found that these transcription regulators antagonistically regulate the level of Flo11p and colony biofilm structure (Fig. 1a) (Nguyen et al. 2018). Overexpression of CYC8 completely blocks FLO11 expression and formation of organized extracellular fibers among cells, prevents cell adhesion to and penetration into agar and results in the formation of smooth colonies. Unexpectedly, the same cell behavior was identified when the TUP1 gene was deleted, implying opposing effects of Cyc8p and Tup1p on biofilm-specific traits and the development of complex biofilms. Further, in-depth analyses of changes at different FLO11 expression levels (mRNA and protein) in strains in which the TUP1 and CYC8 genes were controlled by inducible promoters, revealed that Cyc8p is the major repressor of FLO11, while Tup1p counteracts Cyc8p function and thus induces, probably indirectly, FLO11 expression (Nguyen et al. 2018). As Tup1p and Cyc8p are able to form a co-repressor complex that apparently has a different function (see below), relative levels of Cyc8p and Tup1p can fine-tune the level of free Cyc8p, needed for FLO11 gene repression. The level of Cyc8p repressor could also be influenced by formation of Cyc8p-prion, an inactive form of Cyc8p, as hypothesized by Patel et al. (2009). However, counteracting the Cyc8p repressor function is not the only role of Tup1p in colony biofilm formation. Tup1p is also essential for regulating levels of Flo11p molecules by protecting them against degradation, possibly via Tup1p-dependent repression of a cell wall or extracellular protease that degrades Flo11p (Fig. 1a) (Nguyen et al. 2018). Such a protease is yet to be identified but we can speculate that it may be involved in Flo11p shedding and its subsequent degradation and, thus in balancing mutual levels of cell wall attached and free/released Flo11p within the colony/biofilm. Such a protease could also degrade Flo11p in those parts of colonies where its presence is not required. Further studies of the dynamics of Flo11p accumulation within the colony structure are needed to clarify this point. Differences in Flo11p processing were found in a strain defective in the serine protease Kex2p (Karunanithi et al. 2010), but Flo11p does not contain prominent Kex2p cleavage sites (Bader et al. 2008) and, therefore, is probably not a direct target of Kex2p. So, potentially, a protease that cleaves Flo11p could itself be processed via Kex2p. In addition, studies of Tup1p orthologues showed that Tup1p represses production of secreted aspartyl proteinases (SAPs) in Candida albicans (Naglik et al. 2004) and extracellular proteases in Aspergillus nidulans (Schachtschabel et al. 2013).
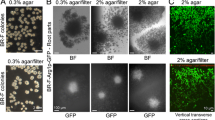
Scheme of Tup1p and Cyc8p regulation of multicellular processes in S. cerevisiae. a Antagonistic regulation by Tup1p and Cyc8p of biofilm and domesticated (colony) lifestyles. Cyc8p represses expression of FLO11 gene encoding the major player in biofilm formation; Tup1p positively regulates Flo11p formation by antagonizing Cyc8p-mediated repression and by stabilization of Flo11p by repressing a protease involved in Flo11p degradation. Aerial views and side views obtained by two-photon excitation confocal microscopy of biofilm/colony cross sections stained with Calcofluor white (false green color) are shown, together with respective schemes. Adapted from Nguyen et al. (2018). b Synergistic regulation by the Tup1p–Cyc8p co-repressor of multicellular floc formation. Co-repressor complex of four Tup1p and one Cyc8p molecules represses formation of flocculin Flo1p. Absence of either Tup1p or Cyc8p or both results in the formation of Flo1p, the presence of which causes cell flocculation and formation of multicellular aggregates (Fleming et al. 2014; Nguyen et al. 2018). Bright field microscopic images of planktonic cells and cells forming flocs are adapted from Nguyen et al. (2018)
Cyc8p and Tup1p in yeast adhesion and invasion in S. cerevisiae and yeast/fungal pathogens
Cell adhesion to and invasion into a semisolid support are important processes in the formation of multicellular structures and are also related to virulence of pathogenic yeast/fungi. The involvement of Tup1p and Cyc8p proteins in these processes has been demonstrated in various yeast/fungi, but the findings are rather inconsistent and imply differences in Tup1p/Cyc8p functions in particular species (or even strains), growth conditions and/or stages of filamentous cell formation.
In S. cerevisiae, similar to inhibition of biofilm colony formation, cell adhesion and pseudohyphae invasion of the wild strain are also completely blocked by Cyc8p overexpression and TUP1 deletion and thus antagonistically regulated by Cyc8p and Tup1p (Nguyen et al. 2018). On the other hand, deletion of TUP1 stimulated adhesion to plastic and to agar and increased FLO1 and FLO11 mRNA levels in laboratory strain BY4741 (S288c background), which is impaired in haploid adhesion, biofilm formation and diploid pseudohyphal growth, partially due to non-functional FLO8 (Fichtner et al. 2007).
In C. albicans, filamentous growth and hypha-specific genes were repressed by Tup1p. Accordingly, deletion of TUP1 increased C. albicans filamentation and surface invasion and the strains formed wrinkled invasive colonies (Braun and Johnson 1997; Garcia-Sanchez et al. 2005, Su et al. 2018). Ssn6p (a Cyc8p orthologue) was also reported to be a repressor of filamentation and of wrinkled colony morphology under particular conditions and some of these repressive effects were enhanced by deletion of the gene, coding for histone deacetylase Rpd31p (Lee et al. 2015). However, other reports implicate Ssn6p in different types of phenotypic switching rather than in filamentous growth. ssn6 strains displayed an unstable phenotype and switched between different forms, with partial changes in cell and colony morphology. It was, therefore, concluded that Ssn6p may act as a repressor of phenotypic switching (Garcia-Sanchez et al. 2005). In addition, roles were identified for both Ssn6p and Tup1p in white-opaque switching of C. albicans: Ssn6p being a negative regulator of the opaque transcription program in white cells and of the white transcription program in opaque cells (Hernday et al. 2016) and Tup1p as a repressor of the opaque state, which, together with its negative regulator Wor1p, controls the opaque switch under different circumstances (Alkafeef et al. 2018). In summary, both Tup1p and Ssn6p seem to contribute to repression of C. albicans filamentation, at least partially independently of one another and dependently on growth conditions and dependently on the functions of other interacting regulators. Hence, Kim et al. (2015) hypothesize that the interplay between Ssn6p and Tup1p, either in a complex or as individual proteins, and other regulatory proteins (e.g., Nrg1p or Rpd3p) differently regulates hyphal-specific genes under different stages of the dimorphic transition from yeast to hyphal form (yeast state, hyphal induction and hyphal maintenance and extension).
As to other yeast/fungi, Tup1p is indispensable for yeast to hypha transition in the maize pathogen Ustilago maydis; accordingly TUP1 deletion reduces filamentation and virulence of this fungus (Elias-Villalobos et al. 2011). Similarly, in Penicillium marneffei, TupA is required for cell filamentation, and in Cryptococcus neoformans for the formation of dikaryotic hyphae and virulence (Lee et al. 2005, 2009; Todd et al. 2003). MoTUP1 was recently identified in Magnaporthe oryzae (a rice pathogen), and its deletion caused decreased production of aerial hyphae and decreased pathogenicity of the fungus (Chen et al. 2015; Li et al. 2017). Hence, contrary to C. albicans and in agreement with S. cerevisiae wild strains, Tup1p seems to function as an activator of filamentation in these yeast/fungi.
Cyc8p and Tup1p in yeast flocculation
Cell flocs, mostly studied in the yeast S. cerevisiae, represent a different type of yeast multicellular population (Smukalla et al. 2008). Flocs are formed by flocculation of planktonic cells within liquid yeast cultures depending not only on the presence of flocculin Flo1p (the major flocculin), but also on other Ca2+-dependent lectin-type flocculins such as Flo10p (Verstrepen and Klis 2006). Cell flocculation of flocculent strains is dependent on nutrient availability as it can be induced by nutrient limitation of mainly sugars and nitrogen sources (Soares 2011; Teunissen et al. 1995). It is, therefore, usually initiated at the end of exponential or during stationary phase. Formation of multicellular flocs is regulated by Tup1p and Cyc8p differently from regulation of biofilm formation (Fig. 1) (Lipke and Hullpillsbury 1984; Nguyen et al. 2018; Stratford 1992; Teunissen et al. 1995). Impairment of any of these regulators induces floc formation even under non-flocculation conditions and increases, by ~ five times, the flocculation of wild strains (Fig. 1b) (Nguyen et al. 2018). A model was proposed, in which the Tup1p–Cyc8p co-repressor complex binds to a distinct site at the FLO1 promoter (Chen et al. 2017; Fleming et al. 2014). This subsequently brings to the promoter region the histone deacetylases Hda1p and Rpd3p, which deacetylate histones in nucleosomes over the FLO1 promoter and upstream regions and thus repress FLO1 expression. In the absence of Cyc8p–Tup1p, chromatin structure over the FLO1 promoter is remodeled, leading to FLO1 de-repression and subsequent cell flocculation. Regulation of flocculation in other yeast is less well studied; speculations exist that flocculation of C. albicans cells is repressed by Sfl1p in concert with Tup1p and Nrg1p (Bauer and Wendland 2007).
In summary, Cyc8p and Tup1p play important pleiotropic roles in yeast multicellularity, being important regulators of key processes, contributing to different aspects of/stages in the formation of various structured populations (Nguyen et al. 2018). Differences in Cyc8p/Tup1p functions observed between different species/strains may reflect either evolutionary functional divergence of these regulators, or, more probably, the fact that the multicellular structures are complex and plastic and processes contributing to their formation, such as filamentation, are not uniform. For example, various types of filaments can be distinguished among the species and nutritive conditions, which can play different roles in different processes, such as cell invasion involved in attachment of a structure to a substrate or cell extension from a structure, involved in occupation of new niches (Vopalenska et al. 2010). Moreover, different stages of these processes have to be controlled. Cyc8p and Tup1p have been described as interacting and forming complexes with numerous additional regulators and chromatin remodeling proteins, each of which could modify the functions of Cyc8p and/or Tup1p proteins in respect to a particular process under distinct circumstances. In this regard, more information on distinct Cyc8p and/or Tup1p complexes and, in particular, on their functional dynamics during structured population development is required to resolve the complex regulatory networks among these proteins and their roles in yeast multicellularity.
References
Alkafeef SS, Yu C, Huang L, Liu H (2018) Wor1 establishes opaque cell fate through inhibition of the general co-repressor Tup1 in Candida albicans. PLoS Genet 14:e1007176
Bader O, Krauke Y, Hube B (2008) Processing of predicted substrates of fungal Kex2 proteinases from Candida albicans, C. glabrata, Saccharomyces cerevisiae and Pichia pastoris. BMC Microbiol 8:116
Bauer J, Wendland J (2007) Candida albicans Sfl1 suppresses flocculation and filamentation. Eukaryot Cell 6:1736–1744
Braun BR, Johnson AD (1997) Control of filament formation in Candida albicans by the transcriptional repressor TUP1. Science 277:105–109
Chen Y, Zhai S, Sun Y, Li M, Dong Y, Wang X, Zhang H, Zheng X, Wang P, Zhang Z (2015) MoTup1 is required for growth, conidiogenesis and pathogenicity of Magnaporthe oryzae. Mol Plant Pathol 16:799–810
Chen X, Poorey K, Carver MN, Muller U, Bekiranov S, Auble DT, Brow DA (2017) Transcriptomes of six mutants in the Sen1 pathway reveal combinatorial control of transcription termination across the Saccharomyces cerevisiae genome. PLoS Genet 13:e1006863
Elias-Villalobos A, Fernandez-Alvarez A, Ibeas JI (2011) The general transcriptional repressor Tup1 is required for dimorphism and virulence in a fungal plant pathogen. PLoS Pathog 7:e1002235
Fichtner L, Schulze F, Braus GH (2007) Differential Flo8p-dependent regulation of FLO1 and FLO11 for cell-cell and cell-substrate adherence of S. cerevisiae S288c. Mol Microbiol 66:1276–1289
Fleming AB, Beggs S, Church M, Tsukihashi Y, Pennings S (2014) The yeast Cyc8-Tup1 complex cooperates with Hda1p and Rpd3p histone deacetylases to robustly repress transcription of the subtelomeric FLO1 gene. Biochim Biophys Acta 1839:1242–1255
Garcia-Sanchez S, Mavor AL, Russell CL, Argimon S, Dennison P, Enjalbert B, Brown AJ (2005) Global roles of Ssn6 in Tup1- and Nrg1-dependent gene regulation in the fungal pathogen, Candida albicans. Mol Biol Cell 16:2913–2925
Hernday AD, Lohse MB, Nobile CJ, Noiman L, Laksana CN, Johnson AD (2016) Ssn6 defines a new level of regulation of white-opaque switching in Candida albicans and is required for the stochasticity of the switch. mBio 7:e01565–e01515
Karunanithi S, Vadaie N, Chavel CA, Birkaya B, Joshi J, Grell L, Cullen PJ (2010) Shedding of the mucin-like flocculin Flo11p reveals a new aspect of fungal adhesion regulation. Curr Biol 20:1389–1395
Kim J, Lee JE, Lee JS (2015) Histone deacetylase-mediated morphological transition in Candida albicans. J Microbiol 53:805–811
Kliewe F, Engelhardt M, Aref R, Schuller HJ (2017) Promoter recruitment of corepressors Sin3 and Cyc8 by activator proteins of the yeast Saccharomyces cerevisiae. Curr Genet 63:739–750
Lee H, Chang YC, Kwon-Chung KJ (2005) TUP1 disruption reveals biological differences between MATa and MATalpha strains of Cryptococcus neoformans. Mol Microbiol 55:1222–1232
Lee H, Chang YC, Varma A, Kwon-Chung KJ (2009) Regulatory diversity of TUP1 in Cryptococcus neoformans. Eukaryot Cell 8:1901–1908
Lee JE, Oh JH, Ku M, Kim J, Lee JS, Kang SO (2015) Ssn6 has dual roles in Candida albicans filament development through the interaction with Rpd31. FEBS Lett 589:513–520
Li Y, Zhang X, Hu S, Liu H, Xu JR (2017) PKA activity is essential for relieving the suppression of hyphal growth and appressorium formation by MoSfl1 in Magnaporthe oryzae. PLoS Genet 13:e1006954
Lipke PN, Hullpillsbury C (1984) Flocculation of Saccharomyces cerevisiae tupl mutants. J Bacteriol 159:797–799
Naglik J, Albrecht A, Bader O, Hube B (2004) Candida albicans proteinases and host/pathogen interactions. Cell Microbiol 6:915–926
Nguyen PV, Hlavacek O, Marsikova J, Vachova L, Palkova Z (2018) Cyc8p and Tup1p transcription regulators antagonistically regulate Flo11p expression and complexity of yeast colony biofilms. PLoS Genet 14:e1007495
Palkova Z, Vachova L (2016) Yeast cell differentiation: lessons from pathogenic and non-pathogenic yeasts. Semin Cell Dev Biol 57:110–119
Palkova Z, Wilkinson D, Vachova L (2014) Aging and differentiation in yeast populations: elders with different properties and functions. FEMS Yeast Res 14:96–108
Patel BK, Gavin-Smyth J, Liebman SW (2009) The yeast global transcriptional co-repressor protein Cyc8 can propagate as a prion. Nat Cell Biol 11:344–349
Schachtschabel D, Arentshorst M, Nitsche BM, Morris S, Nielsen KF, van den Hondel CA, Klis FM, Ram AF (2013) The transcriptional repressor TupA in Aspergillus niger is involved in controlling gene expression related to cell wall biosynthesis, development, and nitrogen source availability. PloS One 8:e78102
Smith RL, Johnson AD (2000) Turning genes off by Ssn6-Tup1: a conserved system of transcriptional repression in eukaryotes. Trends Bioch Sci 25:325–330
Smukalla S, Caldara M, Pochet N, Beauvais A, Guadagnini S, Yan C, Vinces MD, Jansen A, Prevost MC, Latge JP, Fink GR, Foster KR, Verstrepen KJ (2008) FLO1 is a variable green beard gene that drives biofilm-like cooperation in budding yeast. Cell 135:726–737
Soares EV (2011) Flocculation in Saccharomyces cerevisiae: a review. J Appl Microbiol 110:1–18
Stovicek V, Vachova L, Kuthan M, Palkova Z (2010) General factors important for the formation of structured biofilm-like yeast colonies. Fungal Genet Biol 47:1012–1022
Stovicek V, Vachova L, Begany M, Wilkinson D, Palkova Z (2014) Global changes in gene expression associated with phenotypic switching of wild yeast. BMC Genom 15:136
Stratford M (1992) Yeast flocculation: reconciliation of physiological and genetic viewpoints. Yeast 8:25–38
Su C, Yu J, Lu Y (2018) Hyphal development in Candida albicans from different cell states. Curr Genet. https://doi.org/10.1007/s00294-018-0845-5
Teunissen AW, van den Berg JA, Steensma HY (1995) Transcriptional regulation of flocculation genes in Saccharomyces cerevisiae. Yeast 11:435–446
Todd RB, Greenhalgh JR, Hynes MJ, Andrianopoulos A (2003) TupA, the Penicillium marneffei Tup1p homologue, represses both yeast and spore development. Mol Microbiol 48:85–94
Vachova L, Palkova Z (2018) How structured yeast multicellular communities live, age and die? FEMS Yeast Res 18:foy033
Vachova L, Stovicek V, Hlavacek O, Chernyavskiy O, Stepanek L, Kubinova L, Palkova Z (2011) Flo11p, drug efflux pumps, and the extracellular matrix cooperate to form biofilm yeast colonies. J Cell Biol 194:679–687
Varanasi US, Klis M, Mikesell PB, Trumbly RJ (1996) The Cyc8 (Ssn6)-Tup1 corepressor complex is composed of one Cyc8 and four Tup1 subunits. Mol Cell Biol 16:6707–6714
Verstrepen KJ, Klis FM (2006) Flocculation, adhesion and biofilm formation in yeasts. Mol Microbiol 60:5–15
Vopalenska I, St’ovicek V, Janderova B, Vachova L, Palkova Z (2010) Role of distinct dimorphic transitions in territory colonizing and formation of yeast colony architecture. Environ Microbiol 12:264–277
Wong KH, Struhl K (2011) The Cyc8-Tup1 complex inhibits transcription primarily by masking the activation domain of the recruiting protein. Genes Dev 25:2525–2539
Acknowledgements
We thank Derek Wilkinson for proofreading of the manuscript. ZP was supported by LQ1604 NPU II provided by Ministry of Education, Youth and Sports; LV by RVO61388971 from Czech Academy of Sciences; and the research was performed in BIOCEV supported by CZ.1.05/1.1.00/02.0109 BIOCEV provided by European Regional Development Fund and Ministry of Education, Youth and Sports.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Rights and permissions
About this article
Cite this article
Váchová, L., Palková, Z. Diverse roles of Tup1p and Cyc8p transcription regulators in the development of distinct types of yeast populations. Curr Genet 65, 147–151 (2019). https://doi.org/10.1007/s00294-018-0883-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-018-0883-z