Abstract
Homologous recombination repairs discontinuities in DNA including single-strand gaps (SSGs) and double-strand breaks (DSBs). This commentary describes how the RecBCD and RecF pathways might be exchangeable for the repair of their respective DSB and SSG canonical substrates. In particular, I will discuss how the RecBCD pathway could engage in the repair of an SSG even when the latter is not associated with a DSB.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Homologous recombination (HR) is an essential mechanism involved in DNA damage repair. In the bacterium Escherichia coli, RecA protein polymerizes around single-stranded DNA (ssDNA) to form a nucleoprotein filament that will invade a homologous duplex DNA, allowing the exchange of DNA strands (for a review, see Kuzminov 1999).
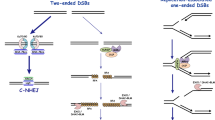
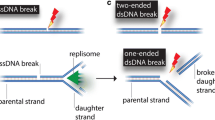
Two types of DNA damages are substrates for HR: (1) single-strand gaps (SSGs, also named daughter-strand gaps) arise during replication, when DNA synthesis is blocked by a lesion on one of the strands; and (2) double-strand breaks (DSBs) which are induced by ionizing radiation, or can arise during replication through an unrepaired SSG.
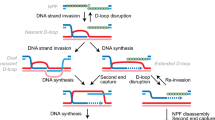
While RecA is the key protein involved in the repair of these damages by homologous recombination, two distinct recombination pathways have been originally described (Clark 1973): the RecF and the RecBCD pathways. Both are involved in the pre-synapsis stage of HR, which consists in assembling the nucleofilament of RecA onto ssDNA. The RecF pathway is involved in SSG repair where it mediates the loading RecA protein onto SSB-coated ssDNA that is already present at the gap (Morimatsu and Kowalczykowski 2003). Since no extensive ssDNA is initially present at a DSB, the RecBCD helicase/nuclease complex processes the ends by resection and concomitantly loads RecA on the generated ssDNA overhangs (Anderson and Kowalczykowski 1997, for a review, see Dillingham and Kowalczykowski 2008). The following steps of HR are common to both pathways: search for homology and exchange of DNA strands occur during the synapsis stage. Branch migration and resolution of the Holliday junction completes the HR process leading to the recombinant molecule (Kuzminov 1999; Spies and Kowalczykowski 2005).
The conventional view of two separate specialized pathways for SSG and DSB repair has changed after several studies reported that RecF is also involved in DSB repair. In normal situations, RecBCD accounts for about 99 % of DSB repair, as measured by conjugational recombination. However, a combined inactivation of RecBCD and exonuclease I results in repair of DSBs by RecF with wild-type efficiency (Horii and Clark 1973; Lloyd et al. 1987). In vitro studies by Handa et al. (2009) showed that the RecJ nuclease and RecQ helicase can substitute for RecBCD at DSBs, leading to resection and RecF-mediated loading of RecA on ssDNA. The RecF pathway is therefore not restricted to SSG repair, but can also participate to DSB repair. Similarly, in bacteria devoid of RecB, such as Deinococcus radiodurans, the RecF pathway is employed to repair DSB (Bentchikou et al. 2010). In contrast, until recently, no evidence pointed towards a role for the RecBCD pathway in repairing SSGs, unless the gaps were converted to DSBs (Wang and Smith 1983).
The processing of DSBs and its genetic requirements have been widely explored through the study of biological processes such as conjugation or transduction since these processes initiate at DSBs. Moreover, the recent possibility to generate a unique DSB at a specific locus on the bacterial chromosome helped to clarify the mechanisms involved (Lesterlin et al. 2013; Cockram et al. 2015). But, our understanding of how SSGs are processed has remained relatively incomplete.
Our group recently developed a system to insert a single replication-blocking lesion into the chromosome of a living bacteria (Pagès et al. 2012; Naiman et al. 2014), thus allowing to study the repair of the SSG generated at the damaged site. By means of genetic markers placed at both the damaged and non-damaged strand, we were able to directly monitor sister chromatid strand exchange in the vicinity of the lesion site (Laureti et al. 2015). Using this system, we explored the repair of the SSG generated at a single UV lesion by monitoring the exchange of genetic markers between the damaged strand and its non-damaged homologous sister chromatid. We named this process homology-directed gap repair (HDGR).
We found that 90 % of cells repaired these SSGs by HDGR and these frequencies dropped to less than 10 % in the absence of RecA (Fig. 1). As expected, the inactivation of RecF led to a strong decrease of HDGR frequencies, with the majority of the colonies arising from “damaged chromatid loss” events in this strain (Laureti et al. 2015). Surprisingly, we observed a decrease in HDGR frequencies in a recB strain, suggesting an involvement of the RecBCD pathway in SSG repair.
The graph represents the partition of DNA damage-tolerance mechanisms for the TT(6-4) photoproduct in our parental strain (uvrA, mutS), and the different mutants impaired in homologous recombination: recA, recF, recB, and recF recB. The bars in blue represent the extent of SSG repair, also named homology-directed gap repair (HDGR). Adapted from Laureti et al.: the graph represents the average of the bypass of the lesion located on the leading and lagging strands
It appears that the action of RecB does not occur through the conversion of the SSG into DSB. Indeed, if a DSB was formed and processed by RecBCD, it would lead to extended resection of the DNA end. The helicase activity of RecBCD unwinds an average of 30 kbp in vitro (Roman et al. 1992), and recent in vivo studies involving the generation of a single DSB confirmed a 30-kbp region of ssDNA coated by RecA (Cockram et al. 2015). Such resection would remove the markers we have placed at a short distance (between 100 and 800 bp) upstream of the lesion. Instead, the markers persist in the presence of RecBCD, and we were able to visualize the exchange of genetic markers between the damaged and non-damaged chromatid, evidencing genuine HDGR. Therefore, the RecBCD pathway could directly be involved in SSG repair. Interestingly, the inactivation of RecB leads to a decrease in HDGR even in the presence of a fully functional RecF pathway, supporting the view that the RecBCD pathways do not act as a backup in the absence of RecF (as RecF does in the absence of RecB), but is genuinely involved in SSG repair (Fig. 2). These findings raise the question of what dictates RecBCD versus RecF pathway choice for SSG. Could the extent of ssDNA at the SSG determine the pathway choice? It will be interesting to further explore this role of RecB. Are additional co-factors required to help RecB in SSG repair, or does RecB act alone in promoting the loading of RecA directly on SSB-coated ssDNA generated at the gap? Previous work revealed that in addition to the helicase and nuclease activity of RecBCD, the complex also facilitates the loading of RecA (Churchill et al. 1999) and that this activity is essential for HR (Arnold and Kowalczykowski 2000). While this activity was shown to be coupled to DNA unwinding in a substrate mimicking a DSB, it will be interesting to see if such RecA loading activity could also be observed on a gapped DNA.
In a normal situation, RecBCD pathway (blue arrow) is in charge of 99 % of DSB repair. In the absence of RecBCD, RecF pathway (green arrow) can substitute to RecBCD and process DSB with the same efficiency. While the RecF pathway is responsible for a majority of SSG repair, recent data show that RecB also contributes to SSG repair
Finally, our data show that in the absence of both RecF and RecB the frequency of SSG repair is as low as in a recA mutant. It appears therefore that the action of RecA is very limited in the absence of its key co-factors RecF and RecB. Interestingly, even in the absence of RecA, cells are still able to achieve some exchange of genetic information between the damaged and non-damaged strands. In the future, we will exploit our genetic system to further characterize the actors involved in RecA-independent SSG repair.
References
Anderson DG, Kowalczykowski SC (1997) The translocating RecBCD enzyme stimulates recombination by directing RecA protein onto ssDNA in a chi-regulated manner. Cell 90:77–86
Arnold DA, Kowalczykowski SC (2000) Facilitated loading of RecA protein is essential to recombination by RecBCD enzyme. J Biol Chem 275:12261–12265
Bentchikou E, Servant P, Coste G, Sommer S (2010) A major role of the RecFOR pathway in DNA double-strand-break repair through ESDSA in Deinococcus radiodurans. PLoS Genet 6:e1000774. doi:10.1371/journal.pgen.1000774
Churchill JJ, Anderson DG, Kowalczykowski SC (1999) The RecBC enzyme loads RecA protein onto ssDNA asymmetrically and independently of chi, resulting in constitutive recombination activation. Genes Dev 13:901–911
Clark AJ (1973) Recombination deficient mutants of E. coli and other bacteria. Annu Rev Genet 7:67–86. doi:10.1146/annurev.ge.07.120173.000435
Cockram CA, Filatenkova M, Danos V et al (2015) Quantitative genomic analysis of RecA protein binding during DNA double-strand break repair reveals RecBCD action in vivo. Proc Natl Acad Sci USA 112:E4735–E4742. doi:10.1073/pnas.1424269112
Dillingham MS, Kowalczykowski SC (2008) RecBCD enzyme and the repair of double-stranded DNA breaks. Microbiol Mol Biol Rev 72:642–71– Table of Contents. doi:10.1128/MMBR.00020-08
Handa N, Morimatsu K, Lovett ST, Kowalczykowski SC (2009) Reconstitution of initial steps of dsDNA break repair by the RecF pathway of E. coli. Genes Dev 23:1234–1245. doi:10.1101/gad.1780709
Horii Z, Clark AJ (1973) Genetic analysis of the recF pathway to genetic recombination in Escherichia coli K12: isolation and characterization of mutants. J Mol Biol 80:327–344
Kuzminov A (1999) Recombinational repair of DNA damage in Escherichia coli and bacteriophage lambda. Microbiol Mol Biol Rev 63:751–813
Laureti L, Demol J, Fuchs RP, Pagès V (2015) Bacterial proliferation: keep dividing and don’t mind the gap. PLoS Genet 11:e1005757. doi:10.1371/journal.pgen.1005757
Lesterlin C, Ball G, Schermelleh L, Sherratt DJ (2013) RecA bundles mediate homology pairing between distant sisters during DNA break repair. Nature. doi:10.1038/nature12868
Lloyd RG, Evans NP, Buckman C (1987) Formation of recombinant lacZ + DNA in conjugational crosses with a recB mutant of Escherichia coli K12 depends on recF, recJ, and recO. Mol Gen Genet 209:135–141
Morimatsu K, Kowalczykowski SC (2003) RecFOR proteins load RecA protein onto gapped DNA to accelerate DNA strand exchange: a universal step of recombinational repair. Mol Cell 11:1337–1347
Naiman K, Philippin G, Fuchs RP, Pagès V (2014) Chronology in lesion tolerance gives priority to genetic variability. Proc Natl Acad Sci USA 111:5526–5531. doi:10.1073/pnas.1321008111
Pagès V, Mazon G, Naiman K et al (2012) Monitoring bypass of single replication-blocking lesions by damage avoidance in the Escherichia coli chromosome. Nucleic Acids Res 40:9036–9043. doi:10.1093/nar/gks675
Roman LJ, Eggleston AK, Kowalczykowski SC (1992) Processivity of the DNA helicase activity of Escherichia coli recBCD enzyme. J Biol Chem 267:4207–4214
Spies M, Kowalczykowski SC (2005) Homologous Recombination by the RecBCD and RecF Pathways. Bact Chromosome. doi:10.1128/9781555817640.ch21
Wang TC, Smith KC (1983) Mechanisms for recF-dependent and recB-dependent pathways of postreplication repair in UV-irradiated Escherichia coli uvrB. J Bacteriol 156:1093–1098
Acknowledgments
The author is extremely grateful to his colleagues Luisa Laureti and Mauro Modesti for their many helpful comments and discussion. The work in the author’s laboratory is supported by Agence National de la Recherche grant GENOBLOCK (ANR-14-CE09-0010-01).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Rights and permissions
About this article
Cite this article
Pagès, V. Single-strand gap repair involves both RecF and RecBCD pathways. Curr Genet 62, 519–521 (2016). https://doi.org/10.1007/s00294-016-0575-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-016-0575-5