Abstract
The complete mitochondrial genome sequences of Brassica species have provided insight into inter- and intraspecific variation of plant mitochondrial genomes. However, the size of mitochondrial genome sequenced for Brassica oleracea hitherto does not match to its physical mapping data. This fact led us to investigate B. oleracea mitochondrial genome in detail. Here we report novel B. oleracea mitochondrial genome, derived from var. capitata, a cabbage cultivar ‘‘Fujiwase’’. The genome was assembled into a 219,952-bp circular sequence that is comparable to the mitochondrial genomes of other Brassica species (ca. 220–232 kb). This genome contained 34 protein-coding genes, 3 rRNA genes and 17 tRNA genes. Due to absence of a large repeat (140 kb), the mitochondrial genome of ‘‘Fujiwase’’ is clearly smaller than the previously reported mitochondrial genome of B. oleracea accession ‘‘08C717’’ (360 kb). In both mitotypes, all genes were identical, except cox2-2, which was present only in the Fujiwase type. At least two rearrangement events via large and small repeat sequences have contributed to the structural differences between the two mitotypes. PCR-based marker analysis revealed that the Fujiwase type is predominant, whereas the 08C717 type coexists at low frequency in all B. oleracea cultivars examined. Intraspecific variations in the mitochondrial genome in B. oleracea may occur because of heteroplasmy, coexistence of different mitotypes within an individual, and substoichiometric shifting. Our data indicate that the Fujiwase-type genome should be used as the representative genome of B. oleracea.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Unlike animal mitochondrial genomes, which are compact and relatively uniform in size, plant mitochondrial genomes are large and variable in size, ranging from 208 kb (Brassica hirta) to 11.3 Mb (Silene conica) (Palmer and Herbon 1987; Sloan et al. 2012a). Mitochondrial genomes consist mostly of noncoding sequences and duplications of large segments. Plant mitochondrial genomes also contain multiple repeats (Alverson et al. 2011), and highly frequent recombination via these repeats leads to extensive structural differences between closely related species (Davila et al. 2011; Tanaka et al. 2012). Frequent recombination is also involved in heteroplasmy, defined as the coexistence of various mitotypes within an individual (Woloszynska 2010). Plants usually have the main mitochondrial genome accompanied by minor genome molecules, called sublimons. Sublimons are generated by recombinations via short repeats (50–1,000 bp) present in the main genome. Plant mitochondrial DNA (mtDNA) also contains sequences of plastid and nuclear origin (reviewed by Kubo and Newton 2008), and sequences originated from foreign genomes by horizontal transfer (Bergthorsson et al. 2003). Frequent recombination and incorporation of foreign DNA generates dynamic structural variations of mtDNA within a species or genus. Comparison between several mitotypes within one genus or species provides insight into the mechanisms of evolution of plant mitochondrial genomes. Such comparative analysis showed intraspecific variations in mitochondrial genome size and structure (Allen et al. 2007; Darracq et al. 2010; Fujii et al. 2010; Sloan et al. 2012b).
Brassica includes six domesticated species, which are important as vegetable and oilseed crops. The relationships between the nuclear genomes of these six species are known as U’s triangle (U1935). Except B. nigra (BB), complete mitochondrial genome sequences from five Brassica species [B. rapa (AA), B. oleracea (CC), B. juncea (AABB), B. napus (AACC) and B. carinata (BBCC)] among the six cultivated species have been reported (Chang et al. 2011). While, the entire sequences of two mitotypes (nap and pol) have been revealed in B. napus and used to study the mechanism of cytoplasmic male sterility (Chen et al. 2011; Handa 2003). Comparisons of the sequences of the Brassica species show that the mitotype of B. juncea (jun) was derived from B. rapa (cam), whereas the B. napus pol and nap mitotypes were derived from a primary mitotype very similar to cam. Because of a large repeat (141 kb), the mitochondrial genome of B. oleracea accession ‘‘08C717’’ (360 kb) is clearly larger than those of other Brassica species (ca. 220–232 kb). It is also much larger than that previously reported from physical mapping (219.7 kb) (Chétritl et al. 1984; Palmer 1988), suggesting variation of mitochondrial genomes within B. oleracea.
We determined the complete sequence of another mitotype from B. oleracea and compared the two B. oleracea sequences to investigate mitotype variation within the species. We conclude that the mitotype of cv. ‘‘Fujiwase’’ is the major mitochondrial genome of B. oleracea and that of ‘‘08C717’’ reported by Chang et al. (2011) is a sublimon.
Materials and methods
Plant material
We determined the complete mitochondrial sequence of cabbage (B. oleracea var. capitata) cv. ‘‘Fujiwase’’. Including it, we tested 22 B. oleracea cultivars of seven kinds of vegetables, whose names are shown in Table 2, for the mitotypes.
Mitochondrial DNA extraction
MtDNA was isolated from ‘‘Fujiwase’’ leaves according to the method of Bonen and Gray (1980) using a discontinuous density gradient (1.15, 1.30, and 1.45 M sucrose) with minor modifications. DNase I-treated mitochondria were collected from the interface between 1.30 and 1.45 M sucrose, and mtDNA was purified by CsCl–ethidium bromide centrifugation and precipitated with ethanol.
Sequencing and sequence assembly
Pyrosequencing on the GS-FLX system (Roche) and de novo assembly were conducted at Hokkaido System Science (Sapporo, Japan). The total length of sequenced DNA was 29,344,724 bp. Sequences were assembled into 61 contigs (average length of 5,798 bp) and connected to develop a master circular genome (shown in Supplementary Fig. 1 and Supplementary Table 1). Contigs similar to the plastid genome and those with low depth values (read depth <50) were ignored. All linkages between contigs were confirmed by genomic PCR followed by sequencing. The sequences were assembled in Sequencher software ver. 4.9 (Gene Codes, Ann Arbor, Michigan, USA).
Sequence analysis
We identified the genes for known mitochondrial proteins and rRNAs and repeated sequences by BLAST search (http://blast.ncbi.nlm.nih.gov/Blast.cgi). A tRNA gene search was conducted with the tRNAscan-SE program. (http://lowelab.ucsc.edu/tRNAscan-SE/) (Lowe and Eddy 1997). Genome map was generated using CGviewer (Stothard and Wishart 2005).
Detection of the 08C717 mitotype
PCR primers were designed from the Fujiwase-type and 08C717-type sequences. The accession 08C717 mitotype was sequenced by Chang et al. (2011). Specific primers P1 (Fujiwase-type R: ATTGAATCAAAGCGTCGGCTAA) or P2 (08C717-type R: GTCCGTCGACAGCTGAATGTT) were used with the common primer P3 (common F: GCAATTGAGGATGGAAGCAATC). P1 and P3 amplify a 873-bp fragment of the Fujiwase mitotype. P2 and P3 amplify a 535-bp fragment of the 08C717-type mitotype. Multiplex PCR with P1, P2, and P3 was conducted to determine whether the mitotype of ‘‘Fujiwase’’ or ‘‘08C717’’ is predominant.
Total DNA was isolated from 0.1 g of young leaves using the DNeasy Plant Mini Kit (Qiagen, Valencia, CA, USA). PCR reaction mixtures contained 12.5 μL Go-Taq polymerase (Promega, Tokyo, Japan) and 1.0 μL each primer (10 μM), and were adjusted to 25 μL with ultra-pure water. Approximately 20-ng genomic DNA was used as a template. The genomic PCR procedure was as follows: 2 min at 94 °C; 30 cycles of 30 s at 94 °C, 30 s at 55 °C, and 1 min at 72 °C; and a final extension of 4 min at 72 °C. PCR products were separated in 1 % agarose gel, stained with ethidium bromide, and visualized under a UV transilluminator.
Determination of the copy number ratio of the two mitotypes by real-time PCR
The copy number (CN) was determined for each mitotype by absolute quantification against standard curves generated using serial 1:10 dilutions of P1–P3 or P2–P3 PCR products. The ratio was calculated as CNFujiwase/CN08C717. PCR was performed in a 7500 Real-Time PCR system (Applied Biosystems, Foster City, CA, USA) using SYBR Green Real-Time PCR Master Mix-Plus (TOYOBO, Osaka, Japan). Reactions were carried out in a total volume of 25 μL with 0.2 μM each primer (final concentration). The PCR protocol was as follows: 1 min at 95 °C to activate the polymerase; and 40 cycles of 15 s at 95 °C, 15 s at 60 °C, and 45 s at 72 °C. Specific primers P4 (Fujiwase-type R for RT: TGAGTAAGTTGTTGCATGAATGGTC) or P5 (08C717-type R for RT: GCCCAGTTCACCAAACATAACC) were used with the common primer P6 (common F for RT: GGGGATCGAAGCCTAAATCAAG). P4 is located on Fujiwase-type specific region, while P5 and P6 correspond to R1 and syntenic region 2, respectively. The sizes of PCR product with P4–P6 and P5–P6 are expected to be 201 and 278 bp. Each sample was analyzed three times.
Results
Brassica oleracea cv. ‘‘Fujiwase’’ mitochondrial genome
The mitochondrial genome of cabbage cv. ‘‘Fujiwase’’ (AP012988) was assembled into a 219,952-bp circular molecule (coverage value 106.2× in average) (Fig. 1). The overall GC content of this mtDNA, 45.2 %, was comparable to those of mtDNAs of other Brassica species (Chang et al. 2011).
Gene organization of two B. oleracea mitochondrial genomes
The Fujiwase-type mitochondrial genome contains 54 genes, including 34 protein-coding genes, 3 rRNA genes, and 17 tRNA genes (Table 1). Among protein-coding genes, 19 are components of the electron transport chain and ATP synthase: nine subunits of complex I (NAD1, 2, 3, 4, 4L, 5, 6, 7, and 9), apocytochrome b (COB) of complex III, four subunits of complex IV (COX1, 2-1, 2-2, and 3), and five subunits of complex V (ATP1, 4, 6, 8, and 9). Five proteins (ccmB, C, FN1, FN2 and FC) are involved in cytochrome c biogenesis. Eight genes encode ribosomal proteins (rpL2, 5, and 16, and rps3, 4, 7, 12, and 14), and the two remaining genes encode maturase and orfX. The three rRNAs are rrn5, 18, 26.
Reorganization of the Fujiwase and 08C717 mitotypes
The two B. oleracea mtDNA sequences contain three syntenic regions (Fig. 2) with very high identity (no SNP and total 12-bp indel in total 214,927-bp syntenic regions). In addition, the Fujiwase type has a specific region (5,025 bp) between syntenic regions 1 and 3.
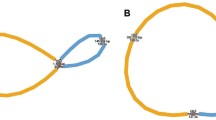
Possible rearrangements that had occurred between the Fujiwase- and 08C717-type genomes via short repeats. Large blank arrows indicate syntenic regions. Arrows of the same color show the position and orientation of repeats. The picture in brackets explains the process of rearrangement between Fujiwase- and 08C717-type genomes. The rearrangement could occur through homologous recombination via RB and two short repeats (146 and 312 bp), which are present within R1
We investigated large repeats (>1 kb) in the two mitochondrial genomes. We defined a pair of sequences as repeats if sequence identity between them was >90 %. A pair of large repeats (2,428 bp), called RB (Chang et al. 2011), was identified in both mitotypes. In the Fujiwase type, RB repeats are located in syntenic region 2 and a specific region, whereas in the 08C717 type, both RB repeats are located in the two copies of syntenic region 2. In several Brassica mitochondrial genomes, RB repeats were reported as a large direct repeat, which includes the cox2 region (Chang et al. 2011; Handa 2003). Such large repeats are involved in formation of the multipartite structure (Maréchal and Brisson 2010). The Brassica mitochondrial genome could be recombined into two subgenomic circles via RB. The sizes of the two subgenomic circles in the Fujiwase type are 170,039 and 49,913 bp. The RB repeats are also related to the reorganization of syntenic regions 1 and 2 and the Fujiwase-type-specific region (Fig. 2). In the 08C717-type mtDNA, one additional pair of large repeats (R1, 3,605 bp) is present, whereas there is only one R1 copy in the Fujiwase-type mtDNA.
We also identified short repeats (<1 kb), which may be involved in reorganization of the Fujiwase- and 08C717-type genomes (Fig. 2). In the Fujiwase-type genome, the 146-bp repeats are present in syntenic region 3 and, in an inverted orientation, at the border of the specific region. The 312-bp repeats are present within syntenic region 3 and, in an inverted orientation, at the border of syntenic region 2. These repeats may be associated with the generation of a linkage between syntenic regions 2 and 3 in the 08C717-type genome.
Fujiwase and 08C717 mitotypes coexist in B. oleracea cultivars
Recently it was reported that two mitotypes coexist in B. napus, and substoichiometric shifting can account for the occurrence of cytoplasmic male sterility (Chen et al. 2011). We intended to investigate whether the two mitotypes, Fujiwase-type and 08C717-type, coexist similarly in B. oleracea. To examine the possibility of the coexistence of the two mitotypes, we designed two pairs of mitotype-specific PCR primers (Fig. 3). Forward primer P3 corresponded to syntenic region 3. The reverse primer specific to the Fujiwase type corresponded to the Fujiwase-type-specific region, whereas the other reverse primer, specific to the 08C717 type, corresponded to the large repeat R1. Both primer pairs amplified corresponding fragments from Fujiwase mtDNA (Fig. 4a, b, lane 1). These results indicate that 08C717-type molecules exist in ‘‘Fujiwase’’ at low frequency. We also checked whether these mitotypes coexist in other B. oleracea cultivars. As B. oleracea includes a wide range of cultivar groups, such as cabbage, broccoli, cauliflower, kohlrabi, and kale, we used a broad range of B. oleracea cultivars for this purpose. Fragments of both mitotypes were amplified in all 22 cultivars analyzed (Fig. 4a, b), but the Fujiwase type was amplified predominantly in multiplex PCR analysis in all cultivars (Fig. 4c). Therefore, the Fujiwase-type mitochondrial genome can be considered as the major genome in a broad range of B. oleracea cultivars, whereas the 08C717 type coexists at low frequency. The ratios between the abundances of the Fujiwase-type and 08C717-type genomes, estimated by using real-time PCR, varied between approximately 250 (“Neo Ruby”) and 550 (“Aojiru-yo”) in different cultivars (Table 2).
Location of primers specific for the two B. oleracea mitotypes. P1 (Fujiwase-type R) and P2 (08C717-type R) are located in a Fujiwase-type-specific region (line) and a repeated sequence (red arrow), respectively. P3 (common F) is located in syntenic region 3. P1 and P3 amplify a 873-bp fragment of the Fujiwase mitotype. P2 and P3 amplify a 535-bp fragment of the 08C717 mitotype
Genomic PCR analysis to distinguish the two B. oleracea mitotypes. Genomic PCR amplified the molecules of Fujiwase-type (a primers P1 and P3), 08C717-type (b primers P2 and P3), and both types (c multiplex PCR with primers P1, P2, and P3). B. oleracea cultivars: 1, ‘‘Fujiwase’’ (cabbage); 2, ‘‘Neo Ruby’’ (cabbage); 3, ‘‘Grand Duke’’ (kohl rabi); 4, ‘‘Aojiru-yo’’ (kale); 5, ‘‘Delikatess White’’ (kohl rabi); 6, ‘‘Dwarf Green Curled’’ (kale); 7, ‘‘Curly Scarlet’’ (kale); 8, ‘‘Sicilia Violetto’’ (cauliflower); 9, ‘‘Romanesco Natalino’’ (cauliflower); 10, ‘‘Andes’’ (cauliflower); 11, ‘‘All The Year Round’’ (cauliflower); 12, ‘‘Savoy Ormskirk’’ (cabbage); 13, ‘‘Langedijk 4’’ (cabbage); 14, ‘‘Greyhound’’ (cabbage); 15, ‘‘Bloemendaalse Gele’’ (cabbage); 16, ‘‘Evesham Special’’ (Brussels sprout); 17, ‘‘Red Arrow’’ (broccoli); 18, ‘‘Rabe 60 Days’’ (broccoli); 19, ‘‘Early White Sprouting’’ (broccoli); 20, ‘‘Early Purple Sprouting’’ (broccoli); 21, ‘‘Purple Vienna’’ (kohl rabi); 22, ‘‘Kailan’’ (Chinese kale); 23, Negative control; M, φx174/HaeIII marker
Discussion
In this study, we determined a novel complete mitochondrial genome sequence of B. oleracea as Fujiwase-type. The size of the Fujiwase mitotype agreed with the previously reported data (219 kb) from physical mapping (Palmer 1988), and was close to 216.8 kb reported by Chétritl et al. (1984) for the mitochondrial genome of cauliflower of B. oleracea.
The 08C717-type mtDNA genome reported by Chang et al. (2011) (360,271 bp) is larger than the Fujiwase-type genome because of a large repeat of 140 kb. Approximately 97 % of the Fujiwase-type sequence shows high similarity to the 08C717-type sequence, whereas the remaining 3 % is specific to the Fujiwase type.
Most of the 08C717-type genes are present in two copies because of a large repeat region (Table 1), and are identical to those of the Fujiwase type. The only exception is cox2-2, which is absent in the 08C717 type because of a DNA rearrangement near this gene. The cox2-2 gene is present in cam, jun, pol, and nap mitotypes, but is absent in car of B. carinata (Chang et al. 2011).
The comparison between two mitotypes revealed that the two types of mitochondrial genome in B. oleracea consist of three syntenic regions, and Fujiwase-type has a specific region (5,025 bp) between syntenic region 3 and 1 (Fig. 2). We also identified large repeats and short repeats (<1 kb), which may be involved in reorganization of the Fujiwase- and 08C717-type genomes. These repeats can explain rearrangement between Fujiwase- and 08C717-type genomes via homologous recombination (Fig. 2). RB repeats (2,428 bp) are present in syntenic region 2, and between specific region of Fujiwase-type and syntenic region 1. These RB repeats can account for the occurrence of a linkage between syntenic region 2 and 1 in the 08C717-type genome. As for short repeat, the 146-bp repeats are present in syntenic region 3 and, in an inverted orientation, at the border of the specific region in the Fujiwase-type genome. The 312-bp repeats are present within syntenic region 3 and, in an inverted orientation, at the border of syntenic region 2. These repeats may be associated with the generation of a linkage between syntenic regions 2 and 3 in the 08C717-type genome. Thus, 08C717-type genome could arise from Fujiwase-type genome via these two recombination events.
Interestingly, both of short repeats are located within the large repeat R1. Recombination events via short repeats generate new pair of large repeated sequence (R1) in 08C717-type genome (Fig. 2). It has been reported that plant mitochondrial genomes are rich in repeated sequences (Alverson et al. 2011). Mitochondrial repeated sequence could be originated via recombination of the shorter repeats in plants.
The ratios between the abundances of the Fujiwase-type and 08C717-type genomes varied between ~250 and 550 in different cultivars (Table 2). This variation is lower than the variation in the CN ratios of the nap and pol mitotypes, which coexist in B. napus cultivars (Chen et al. 2011). Chen et al. (2011) determined the CN ratios of the nap and pol mitotypes by TaqMan qPCR method, and the CN ratios of orf222 and orf224 (genes specific for the nap and pol mitotypes, respectively) showed a two to six orders of magnitude variation among cultivars. Therefore, the Fujiwase-type/08C717-type ratio in B. oleracea is more restricted than the nap-type/pol-type ratio in B. napus.
The 08C717 type was found to be the main genome in accession ‘‘08C717’’ by Chang et al. (2011), although the authors neither described the cultivar name and characteristics nor determined the cultivar group. Sublimons are generated by recombination via short repeats and are sometimes amplified rapidly, replacing the main genome in a process called substoichiometric shifting, while the former main genome is suppressed to a low-frequency level (Feng et al. 2009; Janska et al. 1998; Small et al. 1989; Woloszynska and Trojanowski 2009). This phenomenon might explain the prevalence of the 08C717-type genome in ‘‘08C717’’. Substoichiometric shifting is possibly caused by environmental conditions or mutations in the nuclear recombination surveillance machinery, including MsH1, RecA3, and OSB1 (Shedge et al. 2007; Zaegel et al. 2006; reviewed by Maréchal and Brisson 2010). In future studies, these recombination surveillance genes should be investigated in accession ‘‘08C717’’.
In conclusion, Fujiwase-type mitochondrial genome sequenced here is predominant in B. oleracea, whereas the 08C717-type genome usually coexists with the Fujiwase mitotype at low frequency. The 08C717 molecules are two to three orders of magnitude less abundant than the Fujiwase type. Our data suggest that the Fujiwase-type genome should be used for the phylogenic analysis of plant mitochondria as the representative of B. oleracea, because of the size comparable to the mitochondrial genomes of other Brassica species and predominance in B. oleracea.
References
Allen JO, Fauron CM, Minx P, Roark L, Oddiraju S, Guan NL, Meyer L, Sun H, Kim K, Wang C et al (2007) Comparisons among two fertile and three male-sterile mitochondrial genomes of maize. Genetics 177:1173–1192
Alverson AJ, Zhuo S, Rice DW, Sloan DB, Palmer JD (2011) The mitochondrial genome of the legume Vigna radiata and the analysis of recombination across short mitochondrial repeats. PLoS One 6:e16404
Bergthorsson U, Adams KL, Thomason B, Palmer JD (2003) Widespread horizontal transfer of mitochondrial genes in flowering plants. Nature 424:197–201
Bonen L, Gray MW (1980) Organization and expression of the mitochondrial genome of plants. I. The genes for wheat mitochondrial ribosomal and transfer RNA: evidence for an unusual arrangement. Nucleic Acids Res 8:319–335
Chang S, Yang T, Du T, Huang Y, Chen J, Yan J, He J, Guan R (2011) Mitochondrial genome sequencing helps show the evolutionary mechanism of mitochondrial genome formation in Brassica. BMC Genomics 12:497–508
Chen J, Guan R, Chang S, Du T, Zhang H, Xing H (2011) Substoichiometrically different mitotypes coexist in mitochondrial genomes of Brassica napus L. PLoS One 6:e17662
Chétritl P, Mathieu C, Muller JP, Vedel F (1984) Physical and gene mapping of cauliflower (Brassica oleracea) mitochondrial DNA. Curr Genet 8:413–421
Darracq A, Varré JS, Touzet P (2010) A scenario of mitochondrial genome evolution in maize based on rearrangement events. BMC Genomics 11:233
Davila J, Arrieta-Montiel M, Wamboldt Y, Cao J, Hagmann J, Shedge V, Xu YZ, Weigel D, Mackenzie S (2011) Double-strand break repair processes drive evolution of the mitochondrial genome in Arabidopsis. BMC Biol 9:64
Feng X, Kaur AP, MacKenzie SA, Dweikat IM (2009) Substoichiometric shifting in the fertility reversion of cytoplasmic male sterile pearl millet. Theor Appl Genet 118:1361–1370
Fujii S, Kazama T, Yamada M, Toriyama K (2010) Discovery of global genomic re-organization based on comparison of two newly sequenced rice mitochondrial genomes with cytoplasmic male sterility-related genes. BMC Genomics 11:209
Handa H (2003) The complete nucleotide sequence and RNA editing content of the mitochondrial genome of rapeseed (Brassica napus L.): comparative analysis of the mitochondrial genomes of rapeseed and Arabidopsis thaliana. Nucleic Acids Res 31:5907–5916
Janska H, Sarria R, Woloszynska M, Arrieta-Montiel M, Mackenzie SA (1998) Stoichiometric shifts in the common bean mitochondrial genome leading to male sterility and spontaneous reversion to fertility. Plant Cell 10:1163–1180
Kubo T, Newton KJ (2008) Angiosperm mitochondrial genomes and mutations. Mitochondrion 8:5–14
Lowe TM, Eddy SR (1997) tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 25:955–964
Maréchal A, Brisson N (2010) Recombination and the maintenance of plant organelle genome stability. New Phytol 186:299–317
U N (1935) Genome analysis in Brassica with special reference to the experimental formation of B. napus and peculiar mode of fertilization. Jpn J Bot 7:389–452
Palmer JD (1988) Intraspecific variation and multicircularity in Brassica mitochondrial DNAs. Genetics 118:341–351
Palmer JD, Herbon LA (1987) Unicircular structure of the Brassica hirta mitochondrial genome. Curr Genet 11:565–570
Shedge V, Arrieta-Montiel M, Christensen AC, Mackenzie SA (2007) Plant mitochondrial recombination surveillance requires unusual RecA and MutS homologs. Plant Cell 19:1251–1264
Sloan DB, Alverson AJ, Chuckalovcak JP, Wu M, McCauley DE, Palmer JD, Taylor DR (2012a) Rapid evolution of enormous, multichromosomal genomes in flowering plant mitochondria with exceptionally high mutation rates. PLoS Biol 10:e1001214
Sloan DB, Müller K, McCauley DE, Taylor DR, Štorchová H (2012b) Intraspecific variation in mitochondrial genome sequence, structure, and gene content in Silene vulgaris, an angiosperm with pervasive cytoplasmic male sterility. New Phytol 196:1228–1239
Small I, Suffolk R, Leaver CJ (1989) Evolution of plant mitochondrial genomes via substoichiometric intermediates. Cell 58:69–76
Stothard P, Wishart DS (2005) Circular genome visualization and exploration using CGView. Bioinformatics 21:537–539
Tanaka Y, Tsuda M, Yasumoto K, Yamagishi H, Terachi T (2012) A complete mitochondrial genome sequence of Ogura-type male-sterile cytoplasm and its comparative analysis with that of normal cytoplasm in radish (Raphanus sativus L.). BMC Genomics 13:352–363
Woloszynska M (2010) Heteroplasmy and stoichiometric complexity of plant mitochondrial genomes—though this be madness, yet there’s method in’t. J Exp Bot 61:657–671
Woloszynska M, Trojanowski D (2009) Counting mtDNA molecules in Phaseolus vulgaris: sublimons are constantly produced by recombination via short repeats and undergo rigorous selection during substoichiometric shifting. Plant Mol Biol 70:511–521
Zaegel V, Guermann B, Le Ret M, Andres C, Meyer D, Erhardt M, Canaday J, Gualberto JM, Imbault P (2006) The plant-specific ssDNA binding protein OSB1 is involved in the stoichiometric transmission of mitochondrial DNA in Arabidopsis. Plant Cell 18:3548–3563
Acknowledgments
This study was supported in part by the Private University Strategic Research Foundation Support Program, Grants-in-Aid for Scientific Research, Scientific Research (B) (No. 22380008), and the Program for Promotion of Basic and Applied Researches for Innovations in Bio-oriented Industry (BRAIN), Japan.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. Hohmann.
Electronic supplementary material
Below is the link to the electronic supplementary material.
294_2014_433_MOESM1_ESM.pdf
Validation of contig linkage by PCR analysis. (A) Map of linkages between contigs. (B) PCR analysis to confirm the linkages. The primer information used for this PCR analysis is described in Supplementary Table 1 (PDF 221 kb)
Rights and permissions
About this article
Cite this article
Tanaka, Y., Tsuda, M., Yasumoto, K. et al. The complete mitochondrial genome sequence of Brassica oleracea and analysis of coexisting mitotypes. Curr Genet 60, 277–284 (2014). https://doi.org/10.1007/s00294-014-0433-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-014-0433-2