Abstract
We previously demonstrated that the nop-1 gene encodes a putative green-light opsin photoreceptor that is highly expressed in cultures that support asexual sporulation (conidiation) in Neurospora crassa. In this study, we demonstrate that nop-1 is a late-stage conidiation gene, through analysis of nop-1 transcript levels in wild-type strains and mutants blocked at various stages of conidiation. nop-1 message amounts are similar with constant illumination or darkness during conidiation, consistent with developmental, but not light, regulation of nop-1 expression. Furthermore, photoinduction experiments using wild type and mutants defective in components of the blue light sensing pathway (wc-1 and wc-2) indicate that nop-1 mRNA levels are not appreciably affected by brief light exposure during conidiation. Surprisingly, nop-1 message amounts are greatly elevated in wc-2 mutants in light or dark, suggesting that the wc-2 gene product regulates nop-1 expression in a light-independent manner. Analysis of expression patterns for al-2, con-10 and con-13, genes regulated by conidiation and/or blue light, showed that nop-1 has significant and reproducible effects on all three genes during various stages of conidiation. The results suggest that NOP-1 directly or indirectly modulates carotenogenesis and repression of conidiation-specific gene expression in N. crassa.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In Neurospora crassa, macroconidiation (conidiation) is the process of producing multinucleate asexual spores (macroconidia or conidia). This developmental program is initiated by exposure to an air–water interface in cultures with adequate nutrition (rev. in Springer 1993; Ebbole 1998; Borkovich et al. 2004). However, conidiation can be induced in submerged cultures in the presence of limiting nitrogen or carbon or heat stress (That and Turian 1978; Plesofsky-Vig et al. 1983). In the early stages of conidiation, newly-formed aerial hyphae grow away from the surface and change their growth mode from hyphal tip elongation to repeated apical budding, forming hyphal segments with minor constrictions (Springer and Yanofsky 1989). These proconidial aerial chains further mature to form a conidiophore structure late in conidiation that allows for the release of free conidia. Macroconidiation is entrained by the circadian rhythm in N. crassa (rev. in Dunlap and Loros 2006), presumably to synchronize spore production and release at optimal times of the day to ensure successful germination (Bell-Pedersen et al. 1996).
Mutants have been identified that are blocked at several different stages of conidiation (Springer and Yanofsky 1989). Early-stage mutants include fluffyoid (fld), aconidiate-2 (acon-2), fluffy (fl), and aconidiate-3 (acon-3). These mutants cannot develop past formation of major constrictions. The FL protein is a Gal4p-type C6 zinc cluster transcription factor that is necessary and sufficient to initiate macroconidiation in N. crassa (Bailey and Ebbole 1998; Bailey-Shrode and Ebbole 2004; Rerngsamran et al. 2005). Late-stage mutants are conidial-separation-1 (csp-1), conidial-separation-2 (csp-2), and easily wettable (eas). The csp-1 and csp-2 mutants are defective in the release of free conidia. eas mutants cannot form the hydrophobic layer found on mature conidia, as they contain a defective hydrophobin gene (Bell-Pedersen et al. 1992; Lauter et al. 1992). The granular (gran) and frost (fr) mutants form abnormal conidia. fr is homologous to Saccharomyces cerevisiae CDC1, and fr and CDC1 have been linked to Mn2+ homeostasis in N. crassa and S. cerevisiae, respectively (Paidhungat and Garrett 1997; Sone and Griffiths 1999).
Conidiation can be synchronized in N. crassa mycelial mats freshly-harvested from liquid culture, and this process has been utilized to identify genes whose expression is specific to conidiation (con genes: Berlin and Yanofsky 1985a, b). The expression patterns of con genes have been monitored throughout the conidial development and individual con genes were classified as early or late, depending on the timing of their expression. For example, con-1, con-2, con-3, con-4 and con-5 are expressed early in conidiation, while con-10, con-11, and con-13 are late-stage con genes (Berlin and Yanofsky 1985a).
Blue-light has been shown to enhance conidiation and other asexual developmental processes, including the induction of mycelial carotenoid biosynthesis, entrainment of the conidiation circadian rhythm, increased yield of conidia, hyperpolarization of hyphal membranes, and conidiophore phototropism (rev. in Linden et al. 1997; He and Liu 2005). The N. crassa wc-1 and wc-2 mutants are defective in all blue-light responses (rev. in Linden et al. 1997; He and Liu 2005) and the respective genes encode zinc-finger transcription factors with PAS domains (Ballario et al. 1996; Linden and Macino 1997). WC-1 and WC-2 together form a white collar complex (WCC) that regulates expression of blue-light inducible genes (Ballario et al. 1996). The WC-1 protein binds a flavin adenine nucleotide cofactor and functions as the actual blue light receptor (Froehlich et al. 2002; He et al. 2002).
The wc-1 and wc-2 mutants have been used to characterize the influence of blue light on the expression of conidiation-specific genes. The al-1 and al-2 genes, encoding enzymes required for carotenoid synthesis, are developmentally regulated, peaking during the formation of major constrictions and at the separation of free conidia (Li and Schmidhauser 1995). Both al-1 and al-2 can be photoinduced in wild type, but not in wc-1 and wc-2 strains (Li and Schmidhauser 1995). In contrast, another carotenoid biosynthetic gene, al-3, shows WC-dependent photoinducibility only during the early stages of conidiation (Li et al. 1997). The conidiation-specific genes con-5, con-10, con-6, and con-8 are also subject to photoinduction by blue light during conidiation, but response times vary among the different genes (Lauter and Russo 1991; Lauter and Yanofsky 1993; Carattoli et al. 1995).
We have previously shown that the N. crassa nop-1 gene encodes a member of the microbial opsin family (Bieszke et al. 1999a; Sharma et al. 2006). Microbial opsins had been well-studied in Archaea (rev. in Brown and Jung 2006; Spudich 2006), but NOP-1 was the first such protein identified in eukaryotic microbes. nop-1 expression was only detected in cultures grown under conditions that support conidial development (Bieszke et al. 1999a). Analysis of Δnop-1 mutant strains revealed subtle defects in the pattern of conidiation in the presence of the mitochondrial ATPase inhibitor oligomycin (Bieszke et al. 1999a). In addition, heterologously-expressed NOP-1 protein bound all-trans retinal to form a green-light absorbing pigment with the characteristics of archaeal sensory rhodopsins, suggesting that NOP-1 function may be important for sensory perception in N. crassa (Bieszke et al. 1999b). Other work has demonstrated that NOP-1 does not function as a proton pump, in spite of conservation of acidic residues (Asp-131 and Glu-142) at two sites required for proton translocation in bacteriorhodopsin (Brown et al. 2001; Bergo et al. 2002). This contrasts with the observations that an opsin from the filamentous fungus Leptosphaeria maculans has proton translocation activity (Waschuk et al. 2005) and mutation of the residue corresponding to Asp-131 in the L. maculans opsin leads to a rhodopsin with the characteristics of NOP-1 (Furutani et al. 2006; Fan et al. 2007). The in vivo significance of such differences is currently unknown, as a mutant lacking the L. maculans rhodopsin has not yet been reported (Idnurm and Howlett 2001).
In this study, we further define the expression pattern and the conditions that regulate nop-1 expression during conidiation and introduce a possible role for NOP-1 in impacting transcriptional regulation in N. crassa. We monitor nop-1 transcript levels throughout conidiation in wild-type strains. We analyze nop-1 mRNA amount in several mutants with defects in conidiation and assess the effects of brief light pulses and the wc genetic background on nop-1 expression. Finally, we compare transcript levels for several conidiation and/or light-regulated genes in wild-type and Δnop-1 strains. Our results suggest that nop-1 is a non-photoinducible late-stage conidiation gene that is negatively regulated by the wc-2 gene product. Our data also support a direct or indirect role for NOP-1 in transcriptional regulation during conidial development and carotenogenesis.
Materials and methods
Strains and growth conditions
The strains used in this study are listed in Table 1. Fluorescent light fixtures with 2-15 Watt F15T8/Natural Sunshine lamps (Philips, USA) were the source of white light for experiments. All tissue samples were immediately quick-frozen in liquid nitrogen after collection. Conidia were propagated on solid Vogel’s minimal medium (VM; high nitrogen, with sucrose as carbon source) as described (Davis and Serres 1970) and used to inoculate cultures or directly used for isolation of RNA (see below). Vegetative plate cultures, consisting of hyphae, aerial hyphae and conidiophores, were prepared by inoculation of cellophane-overlaid VM plates in the center with conidia, followed by incubation at room temperature for three days under fluorescent lights. For analysis of gene expression in sexually-differentiated cultures, synthetic crossing medium (SCM; Westergaard and Mitchell 1947) plates (overlaid with cellophane) were inoculated in the center with conidia and incubated under fluorescent lights at room temperature for 6 days. For characterization of the conidiation mutants, cellophane-overlaid VM plates were inoculated in the center using conidia from the various strains. The plates were incubated at room temperature under fluorescent lights for 3 days.
In order to evaluate gene expression during a time course of conidiation, strains were grown as described (Berlin and Yanofsky 1985b) with the following modifications. Conidia from various strains were inoculated to a final density of 1 × 106 conidia/ml into silanized 500 ml flasks containing 100 ml liquid VM. The flasks were covered with aluminum foil and shaken at 200 RPM for 20 h at 30°C. The cultures were then filtered under a red safe light onto 5.5 cm Whatman #1 filters and the resulting cell pads placed on top of a monolayer of 0.45 mm glass beads in a 5.5 cm petri dish containing 5 ml of liquid VM. For induction of conidiation under constant conditions, the plates were either incubated under fluorescent white light (fluence rate = 5.6 W/m2) or in the dark, in a humid atmosphere. For photoinduction experiments, the plates were incubated in the dark for the indicated times, and then transferred to fluorescent lights (fluence rate = 5.6 W/m2) for 30 min prior to collection.
Northern analysis
All frozen samples were ground to a fine powder, and 200 mg of tissue was used for RNA extraction as described in Sachs and Yanofsky (1991). Northern analysis was performed essentially as described (Tsui et al. 1994), except with Nytran Plus membranes (Schleicher and Schuell). Radiolabeled probes were generated by random priming (Prime A Gene, Promega, Madison, WI, USA) from the following DNA fragments: a 600 bp KpnI-BglII fragment from pJAB13, a cDNA containing the 5′end of the nop-1 open reading frame (ORF), a 5.2 Kb EcoRV fragment of pTJS542 containing the al-2 gene (Schmidhauser et al. 1994), and a 500 bp BamHI-EcoRI fragment from pBW100 (obtained from D. Ebbole) containing con-10. Expression of a rRNA gene was detected by labeling a cosmid containing the gene (obtained from D. Ebbole) or by direct photography of Northern blots. In order to assess con-13 mRNA levels, a fragment containing most of the coding region (830 bp) in cosmid X11D11 (Orbach et al. 1990) was amplified using PCR and the following primers. The sense primer anneals upstream of the start codon and has the sequence 5′-GCTGGTGTGTGTCAACGCTTAGTG-3′, while the antisense primer anneals after the first intron in the genomic clone and has the sequence 5′-GGAATAGTGGTCTGCGACGACGCAAG-3′. The identity of the PCR product was verified by sequencing, and the fragment was labeled as described above for use as a probe.
mRNA levels were quantitated by performing densitometric analysis of bands on films after exposure of Northern blots using the Alpha Imager TM 2200 Documentation and Analysis System (Alpha Innotech, San Leandro, CA, USA). All the values were standardized to rRNA.
Results
nop-1 is a late-stage conidiation gene
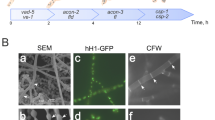
The macroconidiation pathway has been extensively analyzed, and morphological changes have been monitored with respect to time (Springer and Yanofsky 1989). Conidiation begins with the formation of aerial hyphae (1–2 h) that go on to develop minor constrictions (2–4 h). Conidial spores begin to form between each minor constriction, and major constrictions subsequently appear (6–8 h). Septation then occurs between each conidial bud at the major constriction sites (8–10 h), but the conidia remain attached by connectives that form between each spore (10–12 h). These terminally differentiated structures are referred to as conidiophores, and, at the end of their development, mature conidia are released (≥ 16 h).
We previously noted that expression of the N. crassa opsin, nop-1, is specific to conditions that induce both asexual and sexual spore production (Bieszke et al. 1999a). To further characterize the expression pattern of nop-1 during conidial development, total RNA was isolated from wild-type cultures incubated to allow synchronized production of conidia in constant light (Berlin and Yanofsky 1985b). RNA was also extracted from purified mature conidia, as well as conidiating vegetative (VM) and sexually differentiated (SCM) plate cultures producing conidia. Northern analysis of these RNA samples revealed that nop-1 expression is initiated early in conidiation, at 2–4 h, with elevated levels at the later stages, 12–24 h (Fig. 1). The temporal regulation of nop-1 expression during conidiation is similar in cultures exposed to constant darkness (data not shown). nop-1 mRNA is also abundant in mature conidia and in conidiating VM and SCM plate cultures (Fig. 1).
Expression of nop-1 and conidiation-specific (con) genes during a time course of conidiation. Total RNA was isolated from cultures collected at various times during conidiation (0, 2, 4, 8, 12, 16, 20, and 24 h after transfer to an air–liquid interface), as well as from mature conidia (conidia) and vegetative (VM plates) and sexually differentiated plate cultures (SCM plates) of the wild-type strain 74A. Samples containing 10 μg of total RNA were subjected to Northern analysis, using the nop-1, con-10, con-13 and rRNA genes (loading control) as probes. The blot shown is representative of two independent experiments
Genes whose expression is specific to conidiation (con genes) have been identified (Berlin and Yanofsky 1985a). One such con gene, con-10, is well-characterized in terms of its expression pattern during conidiation. con-10 is abundantly expressed during the later stages of conidiation (Berlin and Yanofsky 1985a). The expression pattern of con-10 is remarkably similar to nop-1 (Fig. 1), including the high transcript levels in purified conidia, and VM and SCM plate cultures. Directly upstream of con-10 is another con gene, con-13, whose transcription is independent of con-10 (Hager and Yanofsky 1990). The con-13 message is of low abundance and therefore difficult to detect, but this gene is consistently expressed midway through conidiation (Fig. 1). Interestingly, con-13 message cannot be detected after conidiophore production or in mature conidia (Fig. 1). The message is also weakly expressed in VM and SCM plate cultures, in contrast to nop-1 and con-10. Thus, con-10 and nop-1 appear to have similar expression patterns, while that for con-13 differs.
Mutants that are blocked at different stages of conidiation have been isolated (rev. in Springer and Yanofsky 1989). In order to further characterize the timing of nop-1 expression during conidiation, we analyzed nop-1 mRNA levels in vegetative plate cultures from the early conidiation-defective mutants fluffyoid (fld), fluffy (fl) and acondiate-3 (acon-3), as well as in the late-stage specific conidial-separation-2 (csp-2) mutant. Previous genetic analysis showed that acon-2 and fld act at the stage of formation of minor constrictions and are upstream of fl and acon-3, which are both required for development of major constrictions (Springer and Yanofsky 1989). The csp-2 gene acts later, at the stage of separation of free conidia (Springer and Yanofsky 1989). The frost (fr) and granular (gran) mutants were also tested, as they exhibit altered colony morphologies and produce aberrant conidia.
Northern analysis revealed that nop-1 mRNA can not be detected in the vegetative mutants fr and gran, or in the early-stage developmental mutants fld and fl (Fig. 2). In contrast, nop-1 is weakly-expressed in the acon-3 and csp-2 mutants (Fig. 2). The difference in nop-1 expression noted in the fl and acon-3 mutants, which appear to be blocked at the same stage of conidiation, suggests a difference in temporal regulation of gene expression by fl or acon-3 that is not apparent at the gross morphological level. The lack of nop-1 mRNA in the fr and gran mutants, which develop aberrant conidia, suggests that the defects in these mutants may occur early in the conidiation timeline. The absence of nop-1 mRNA in mutants blocked at early stages of conidiation and the weak transcription of nop-1 observed in the csp-2 mutant blocked in the later stages of conidiation are consistent with the observation that nop-1 expression is specific to late conidial development.
nop-1 expression in developmental mutants. Samples containing 10 μg of total RNA from the indicated strains (Table 1) were used to prepare Northern blots. Membranes were probed with the nop-1 and rRNA genes, as shown. Wildtype is strain 74A. Blot shown is representative of two independent experiments
Expression of con-10 is induced by heat shock temperatures in wild-type conidiating cultures (Lee and Ebbole 1998). To determine if elevated temperature can enhance nop-1 expression, levels of nop-1 mRNA in conidiating cultures at room temperature were compared to those in identical cultures shifted to 45°C (heat shock conditions) for the last hour of incubation. The results showed that nop-1 levels were not elevated by heat (data not shown). Thus, despite their similar expression pattern during conidiation, con-10 and nop-1 do not share heat shock-inducibility.
nop-1 is not photoinducible and is negatively regulated by WC-2
Blue light has been shown to enhance production of conidia (Lauter et al. 1997) and to induce expression of a number of genes associated with conidiation and asexual development (rev. in Linden et al. 1997). WC-1 and WC-2 are transcription factors required for the blue-light signaling pathway in N. crassa (Ballario et al. 1996; Linden and Macino 1997). Increased expression of blue light-regulated genes is blocked in the wc-1 or wc-2 genetic background (rev. in Linden et al. 1997), He and Liu (2005)]. Examples are the carotenoid biosynthetic genes, al-1 and al-2 (Li and Schmidhauser 1995) and the con genes, con-5, con-6, con-8 and con-10 (Lauter and Russo 1991; Lauter and Yanofsky 1993; Carattoli et al. 1995).
Based on the proposed role of NOP-1 as a photoreceptor, we were interested to determine whether nop-1 is regulated by the blue-light sensing pathway. We first evaluated nop-1 mRNA levels in the wc-1 and wc-2 mutant backgrounds (alleles ER53 and ER33, respectively; see Table 1) under constitutive light and darkness. The pattern of nop-1 expression in the white-collar mutants was similar to that of wild type under both conditions (data not shown). However, since N. crassa possesses a strong photoadaptation pathway in response to long-term light exposure (rev. in He and Liu 2005), we also tested the effect of short (30 min) light pulses on nop-1 expression. We included the photoinducible con-10 gene as a control. For these experiments, we also took advantage of strains carrying deletions of the wc-1 and wc-2 ORFs (Δwc-1 and Δwc-2; see Table 1). Northern analysis revealed that nop-1 could not be photoinduced in any genetic background (Fig. 3). In contrast, con-10 mRNA was abundant in light-exposed wild-type cultures and was absent from Δwc-1 and Δwc-2 mutants (Fig. 3). Surprisingly, nop-1 levels were significantly elevated in the wc-2 background with and without photoinduction (Fig. 3). We obtained the same result with another Δwc-2 strain (UCR#473; data not shown). Although not as robust of a response, we also often observed slightly higher amounts of nop-1 message in Δwc-1 mutants (Fig. 3; data not shown). These results suggest that nop-1 is not photoinduced but instead is negatively regulated in a constitutive manner by components of the major blue-light sensing pathway in N. crassa, most notably by the PAS-domain transcription factor WC-2.
Photoinduction of nop-1 transcript levels in wild type and the blue-light sensing mutants wc-1 and wc-2. Cultures of wild-type (74A), Δwc-1 (FGSC#11712) and Δwc-2 (FGSC#11124) strains were allowed to conidiate for 8 h in constant darkness, then either kept in the dark or exposed to light for 30 min prior to collection and RNA extraction. Northern blots were prepared and probed using nop-1 or con-10. The rRNAs were detected by photography of the ethidium bromide-stained blots. Blot shown is representative of three independent experiments
nop-1 influences the levels of several genes that are expressed during conidiation
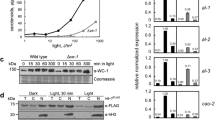
nop-1 expression is increased during the later stages of conidiation. This pattern is consistent with NOP-1 functioning during the formation of conidiophores and/or the release of mature conidia. Thus, it is possible that a NOP-1 pathway may ultimately regulate the expression of other genes specific to conidiation. NOP-1 binds retinal and forms a light-absorbing pigment characteristic of archaeal sensory rhodopsins in vitro (Bieszke et al. 1999a, b) and has been speculated to have similar light-absorbing capabilities in vivo. Therefore, a preliminary analysis of nop-1 as a potential regulator of light and/or conidiation-specific gene expression was initiated by evaluating mRNA levels of al-2, con-10 and con-13 in wild-type and Δnop-1 strains. al-2 is a blue-light regulated gene that encodes phytoene synthase, an important enzyme for carotenoid biosynthesis in N. crassa (Schmidhauser et al. 1994). As mentioned above, con-10 expression is also blue-light regulated, but the function of its gene product is unknown (Corrochano et al. 1995). con-13 is another con gene of unknown function whose expression is not blue-light regulated (Hager and Yanofsky 1990; Lauter and Russo 1991). Therefore, comparison of levels of the al-2, con-10 and con-13 transcripts in the Δnop-1 and wild-type background would reveal whether NOP-1 function was specific to light-regulation (depicted by photoinduced changes in al-2 and con-10 expression) or directed to conidiation (differences in con-10 and con-13 expression levels). mRNA levels were assessed in three independent experiments and a representative Northern blot is shown in Fig. 4. Densitometry was used to quantitate mRNA levels for the three genes at 8, 12, 16, and 24 h into conidiation (Table 2; some values not available due to low mRNA levels).
Photoinduced gene expression during conidiation in Δnop-1 and wild-type strains. Cultures were grown in the dark for the indicated times and then either kept in the dark (D) or transferred to light for 30 min (L) before collection and subsequent RNA extraction. Samples containing 10 μg of total RNA from wild-type strain 74A (W) and Δnop-1 strain 39-1 (triangle) were used to prepare Northern blots. Membranes were probed with the al-2, con-13, con-10 and rRNA genes, as indicated. Blot shown is representative of three independent experiments
The results show that nop-1 exerts a subtle, but reproducible effect on levels of all three transcripts during at least two time points during conidiation (Fig. 4; Table 2). nop-1 affects al-2 expression at two time points in light-exposed cultures; levels of al-2 mRNA are reduced twofold at 12 h, but are twice those of wild type at 16 h (Fig. 4; Table 2). Levels of con-13 message are elevated in Δnop-1 mutants relative to wild type at 8 h into conidiation (light or dark; Fig. 4; Table 2) and at 12 h in dark-grown cultures (Table 2; Δnop-1 lane in Fig. 4 is underloaded). We also noted minor apparent photosuppression of con-13 transcript in both Δnop-1 and wild-type strains at 8 h into conidiation (Fig. 4). The con-10 transcript is present at higher levels in Δnop-1 strains relative to wild type at 12 and 16 h in dark-grown cultures; photoinduced levels are also elevated in Δnop-1 mutants at 16 h (Fig. 4; Table 2). Thus, NOP-1 appears to play a minor role in influencing both light and conidiation-specific gene expression in N. crassa.
Discussion
Conidiation is subject to complex regulation in N. crassa, as it is influenced by light, desiccation, CO2 levels, nutrient-availability, and can be entrained by the circadian rhythm (rev. in Davis 2000). Light and developmental regulation of nop-1 mRNA levels during conidiation was investigated to further probe possible functions for NOP-1. nop-1 is highly expressed in the later stages of conidiation independent of light, consistent with a developmentally-regulated gene. Furthermore, the pattern of weak or absent nop-1 transcription in the developmental mutants supports conidiation-specific expression of nop-1. Interestingly, nop-1 expression parallels the formation of conidiophores and release of mature conidia, suggesting that NOP-1 may function during the later stages of conidiation.
The pattern of nop-1 expression is similar to that of con-10 during conidiation. The function of con-10 is unknown, but the encoded protein is similar to a Bacillus subtilis protein that is expressed under conditions of starvation and stress (Mueller et al. 1992; Ebbole 1996). Although nop-1 expression is not enhanced by elevated temperature, the NOP-1 protein may play a role during heat-shock or other stress responses. NOP-1 is related to a fungal opsin-related protein (ORP) group that includes N. crassa ORP-1 (Bieszke et al. 1999a) and several members that are putative heat shock/stress response proteins in yeast (Seymour and Piper 1999; Hahn et al. 2004). One ORP, Saccharomyces cerevisiae HSP30, negatively regulates the plasma membrane H+-ATPase during heat shock (Piper et al. 1997). Δnop-1 strains were tested in the presence of plasma, vacuolar, and mitochondrial H+-ATPase inhibitors (Bieszke et al. 1999a). Δnop-1 mutants exhibited defects in vegetative growth in the presence of the mitochondrial H+-ATPase inhibitor, oligomycin (Bieszke et al. 1999a), suggesting a possible function for NOP-1 in the response to stress. The observed links between NOP-1, CON-10, heat shock proteins and stress-inducing factors suggest that the late-stage conidial gene products NOP-1 and CON-10 may function in the same regulatory pathway under certain stress conditions.
Both early and late responses to blue-light exposure have been documented in N. crassa (rev. in Linden et al. 1997). Blue-light initiates early responses that are important for entraining the circadian rhythm of conidiation, controlling membrane potential during hyphal elongation, and inducing carotenogenesis in mycelial cultures. The late blue-light effects include the phototrophic responses and enhancement of spore production during both asexual and sexual reproduction. Both early and late responses are regulated by two blue-light dependent transcription factors, WC-1 and WC-2. We did not observe a significant increase in nop-1 mRNA levels after a brief light pulse at any point during conidiation in wild type or the wc mutants. However, nop-1 expression levels are elevated in wc-2 mutants in light or dark conditions. These results suggest that nop-1 expression is negatively regulated by wc-2 in a light-independent manner. These observations also support WC-1-independent regulation of gene expression by WC-2; this is of interest, as WC-2 is a PAS-domain containing transcription factor that, in contrast to WC-1, does not bind flavin or serve directly as the blue-light photoreceptor (Froehlich et al. 2002; He et al. 2002).
NOP-1 may have a sensory role in the later-stages of conidiation, as NOP-1 was shown to form a green-light absorbing pigment upon binding the all-trans retinal chromophore and to possess spectral properties similar to those of the archaeal sensory rhodopsins (Bieszke et al. 1999b). Green light effects are not well characterized in fungi, but green light has been shown to affect spore production in the plant-pathogens Trichometasphaeria turcica and Alternaria solani, either alone or in a synergistic interaction with blue light (rev. in Klein 1992). In N. crassa, conidiophore phototropism and production of mature conidia, both late-stage conidiation events, are subject to regulation by blue light (Siegel et al. 1968; Lauter 1996). Thus, NOP-1 acting as a green-light receptor could potentially regulate these or other conidial developmental processes as an adjunct to the blue-light signaling pathway.
Carotenogenesis is induced by blue-light in basal hyphae, but constitutive during conidiation in N. crassa (Harding and Turner 1981). Carotenoids are thought to play a protective function in fungi, serving as scavengers for reactive oxygen species (Schroeder and Johnson 1995; Michan et al. 2003; Iigusa et al. 2005). The al-2 gene is required for carotenogenesis and serves as a reporter gene for this process. The al-2 transcript is photoinduced during conidiation, in a WC-1- and WC-2-dependent manner (Li and Schmidhauser 1995). Relative to wild type, al-2 transcript levels are reduced at 12 h, but elevated at 16 h in light-exposed Δnop-1 conidiating cultures. We cannot determine whether the differences seen in the Δnop-1 photoinduced samples are light-dependent, as the al-2 transcript could not be detected in 12-h wild-type and Δnop-1 dark-grown cultures. However, this result may point to a sensory role for NOP-1, possibly as a green-light receptor serving as an adjunct to the blue-light pathway. A link between opsins/ORPs and carotenogenesis has also been revealed in the filamentous fungus Fusarium fujikurori (Prado et al. 2004). The ORP-encoding gene carO is found in a carotenoid gene cluster. Expression of carO is deregulated by overproduction of carotenoids and is induced by light and by heat shock. However, similar to other fungal opsin and orp genes, targeted deletion of carO did not reveal any phenotypes.
The effect of nop-1 on expression of the conidiation-specific genes con-10 and con-13 is light-independent, as their expression is significantly elevated in the Δnop-1 mutants compared to wild type with and without light exposure. This suggests a role for NOP-1 in negative regulation of these two con genes during development. Interestingly, nop-1 expression is highest throughout the formative stages of conidiation and remains elevated even into the time points after conidial release. Since Δnop-1 strains do not exhibit obvious defects in conidiation (Bieszke et al. 1999a), any contribution of NOP-1 to a negative-regulatory process during conidiation must be functionally redundant to another pathway and/or operate under different environmental conditions.
Taken together, our results suggest that NOP-1 may play an auxiliary role during late-stage conidial events and/or stress responses. It is possible that NOP-1 may act in concert or share overlapping functions with other proteins, namely N. crassa ORP-1. Future studies will focus on genome-wide transcriptional profiling to identify genes regulated by green light and nop-1, and on genetic approaches to help discern the role of nop-1 and orp-1 in the diverse photobiology of N. crassa.
References
Bailey LA, Ebbole DJ (1998) The fluffy gene of Neurospora crassa encodes a Gal4p-type C6 zinc cluster protein required for conidial development. Genetics 148:1813–1820
Bailey-Shrode L, Ebbole DJ (2004) The fluffy gene of Neurospora crassa is necessary and sufficient to induce conidiophore development. Genetics 166:1741–1749
Ballario P, Vittorioso P, Magrelli A, Talora C, Cabibbo A, Macino G (1996) White collar-1, a central regulator of blue light responses in Neurospora, is a zinc finger protein. EMBO J 15:1650–1657
Bell-Pedersen D, Dunlap JC, Loros JJ (1992) The Neurospora circadian clock-controlled gene, ccg-2, is allelic to eas and encodes a fungal hydrophobin required for formation of the conidial rodlet layer. Genes Dev 6:2382–2394
Bell-Pedersen D, Shinohara ML, Loros JJ, Dunlap JC (1996) Circadian clock-controlled genes isolated from Neurospora crassa are late night- to early morning-specific. Proc Natl Acad Sci USA 93:13096–13101
Bergo V, Spudich EN, Spudich JL, Rothschild KJ (2002) A Fourier transform infrared study of Neurospora rhodopsin: similarities with archaeal rhodopsins. Photochem Photobiol 76:341–349
Berlin V, Yanofsky C (1985a) Isolation and characterization of genes differentially expressed during conidiation of Neurospora crassa. Mol Cell Biol 5:849–855
Berlin V, Yanofsky C (1985b) Protein changes during the asexual cycle of Neurospora crassa. Mol Cell Biol 5:839–848
Bieszke JA, Braun EL, Bean LE, Kang S, Natvig DO, Borkovich KA (1999a) The nop-1 gene of Neurospora crassa encodes a seven transmembrane helix retinal-binding protein homologous to archaeal rhodopsins. Proc Natl Acad Sci USA 96:8034–8039
Bieszke JA, Spudich EN, Scott KL, Borkovich KA, Spudich JL (1999b) A eukaryotic protein, NOP-1, binds retinal to form an archaeal rhodopsin-like photochemically reactive pigment. Biochemistry 38:14138–14145
Borkovich KA, et al. (2004) Lessons from the genome sequence of Neurospora crassa: tracing the path from genomic blueprint to multicellular organism. Microbiol Mol Biol Rev 68:1–108
Brown LS, Dioumaev AK, Lanyi JK, Spudich EN, Spudich JL (2001) Photochemical reaction cycle and proton transfers in Neurospora rhodopsin. J Biol Chem 276:32495–32505
Brown LS, Jung KH (2006) Bacteriorhodopsin-like proteins of eubacteria and fungi: the extent of conservation of the haloarchaeal proton-pumping mechanism. Photochem Photobiol Sci 5:538–546
Carattoli A, Kato E, Rodriguez-Franco M, Stuart WD, Macino G (1995) A chimeric light-regulated amino acid transport system allows the isolation of blue light regulator (blr) mutants of Neurospora crassa. Proc Natl Acad Sci USA 92:6612–6616
Corrochano LM, Lauter FR, Ebbole DJ, Yanofsky C (1995) Light and developmental regulation of the gene con-10 of Neurospora crassa. Dev Biol 167:190–200
Davis RH (2000) Neurospora: contributions of a model organism. Oxford University Press, New York
Davis RH, Serres FJd (1970) Genetic and microbiological research techniques in Neurospora crassa. Methods Enzymol 71A:79–143
Dunlap J, Loros J (2006) How fungi keep time: circadian system in Neurospora and other fungi. Curr Opin Microbiol 9:579–587
Dunlap JC et al (2007) Enabling a community to dissect an organism: overview of the Neurospora functional genomics project. Adv Genet 57:49–96
Ebbole DJ (1996) Morphogenesis and vegetative differentiation in filamentous fungi. J Genet 75:361–374
Ebbole DJ (1998) Carbon catabolite repression of gene expression and conidiation in Neurospora crassa. Fungal Genet Biol 25:15–21
Fan Y, Shi L, Brown LS (2007) Structural basis of diversification of fungal retinal proteins probed by site-directed mutagenesis of Leptosphaeria rhodopsin. FEBS Lett 581:2557–2561
Froehlich AC, Liu Y, Loros JJ, Dunlap JC (2002) White Collar-1, a circadian blue light photoreceptor, binding to the frequency promoter. Science 297:815–819
Furutani Y, et al. (2006) Conformational coupling between the cytoplasmic carboxylic acid and the retinal in a fungal light-driven proton pump. Biochemistry 45:15349–15358
Hager KM, Yanofsky C (1990) Genes expressed during conidiation in Neurospora crassa: molecular characterization of con-13. Gene 96:153–159
Hahn JS, Hu Z, Thiele DJ, Iyer VR (2004) Genome-wide analysis of the biology of stress responses through heat shock transcription factor. Mol Cell Biol 24:5249–5256
Harding RW, Turner RV (1981) Photoregulation of the carotenoid biosynthetic pathway in albino and white collar mutants of Neurospora crassa. Plant Physiol 68:745–748
He Q, Cheng P, Yang Y, Wang L, Gardner KH, Liu Y (2002) White collar-1, a DNA binding transcription factor and a light sensor. Science 297:840–843
He Q, Liu Y (2005) Molecular mechanism of light responses in Neurospora: from light-induced transcription to photoadaptation. Genes Dev 19:2888–2899
Idnurm A, Howlett BJ (2001) Characterization of an opsin gene from the ascomycete Leptosphaeria maculans. Genome 44:167–171
Iigusa H, Yoshida Y, Hasunuma K (2005) Oxygen and hydrogen peroxide enhance light-induced carotenoid synthesis in Neurospora crassa. FEBS Lett 579:4012–4016
Klein RM (1992) Effects of green light on biological systems. Biol Rev Camb Philos Soc 67:199–284
Lauter F-R, Yamashiro CT, Yanofsky C (1997) Light stimulation of conidiation in Neurospora crassa: studies with the wild-type strain and mutants wc-1, wc-2 and acon-2. J Photochem Photobiol B 37:203–211
Lauter FR (1996) Molecular genetics of fungal photobiology. J Genet 75:375–386
Lauter FR, Russo VE (1991) Blue light induction of conidiation-specific genes in Neurospora crassa. Nucleic Acids Res 19:6883–6886
Lauter FR, Russo VE, Yanofsky C (1992) Developmental and light regulation of eas, the structural gene for the rodlet protein of Neurospora. Genes Dev 6:2373–2381
Lauter FR, Yanofsky C (1993) Day/night and circadian rhythm control of con gene expression in Neurospora. Proc Natl Acad Sci USA 90:8249–8253
Lee K, Ebbole DJ (1998) Tissue-specific repression of starvation and stress responses of the Neurospora crassa con-10 gene is mediated by RCO1. Fungal Genet Biol 23:269–278
Li C, Sachs MS, Schmidhauser TJ (1997) Developmental and photoregulation of three Neurospora crassa carotenogenic genes during conidiation induced by desiccation. Fungal Genet Biol 21:101–108
Li C, Schmidhauser TJ (1995) Developmental and photoregulation of al-1 and al-2, structural genes for two enzymes essential for carotenoid biosynthesis in Neurospora. Dev Biol 169:90–95
Linden H, Ballario P, Macino G (1997) Blue light regulation in Neurospora crassa. Fungal Genet Biol 22:141–150
Linden H, Macino G (1997) White collar 2, a partner in blue-light signal transduction, controlling expression of light-regulated genes in Neurospora crassa. Embo J 16:98–109
Michan S, Lledias F, Hansberg W (2003) Asexual development is increased in Neurospora crassa cat-3-null mutant strains. Eukaryot Cell 2:798–808
Mueller JP, Bukusoglu G, Sonenshein AL (1992) Transcriptional regulation of Bacillus subtilis glucose starvation-inducible genes: control of gsiA by the ComP–ComA signal transduction system. J Bacteriol 174:4361–4373
Orbach MJ, Sachs MS, Yanofsky C (1990) The Neurospora crassa arg-2 locus. Structure and expression of the gene encoding the small subunit of arginine-specific carbamoyl phosphate synthetase. J Biol Chem 265:10981–10987
Paidhungat M, Garrett S (1997) A homolog of mammalian, voltage-gated calcium channels mediates yeast pheromone-stimulated Ca2+ uptake and exacerbates the cdc1(Ts) growth defect. Mol Cell Biol 17:6339–6347
Piper PW, Ortiz-Calderon C, Holyoak C, Coot P, Cole M (1997) Hsp30, the integral plasma membrane heat-shock-protein of Saccharomyces cerevisiae, is a stress-inducible regulator of plasma membrane H+-ATPase. Cell Stress Chaperones 2:12–24
Plesofsky-Vig N, Light D, Brambl R (1983) Paedogenetic conidiation in Neurospora crassa. Exp Mycol 7:283–286
Prado MM, Prado-Cabrero A, Fernandez-Martin R, Avalos J (2004) A gene of the opsin family in the carotenoid gene cluster of Fusarium fujikuroi. Curr Genet 46:47–58
Rerngsamran P, Murphy MB, Doyle SA, Ebbole DJ (2005) Fluffy, the major regulator of conidiation in Neurospora crassa, directly activates a developmentally regulated hydrophobin gene. Mol Microbiol 56:282–297
Sachs MS, Yanofsky C (1991) Developmental expression of genes involved in conidiation and amino acid biosynthesis in Neurospora crassa. Dev Biol 148:117–128
Schmidhauser TJ, Lauter FR, Schumacher M, Zhou W, Russo VE, Yanofsky C (1994) Characterization of al-2, the phytoene synthase gene of Neurospora crassa. Cloning, sequence analysis, and photoregulation. J Biol Chem 269:12060–12066
Schroeder WA, Johnson EA (1995) Singlet oxygen and peroxyl radicals regulate carotenoid biosynthesis in Phaffia rhodozyma. J Biol Chem 270:18374–18379
Seymour IJ, Piper PW (1999) Stress induction of HSP30, the plasma membrane heat shock protein gene of Saccharomyces cerevisiae, appears not to use known stress-regulated transcription factors. Microbiology 145(Pt 1):231–239
Sharma AK, Spudich JL, Doolittle WF (2006) Microbial rhodopsins: functional versatility and genetic mobility. Trends Microbiol 14:463–469
Siegel RW, Matsuyama SS, Urey JC (1968) Induced macroconidia formation in Neurospora crassa. Experientia 24:1179–1181
Sone T, Griffiths AJ (1999) The frost gene of Neurospora crassa is a homolog of yeast cdc1 and affects hyphal branching via manganese homeostasis. Fungal Genet Biol 28:227–237
Springer ML (1993) Genetic control of fungal differentiation: the three sporulation pathways of Neurospora crassa. Bioessays 15:365–374
Springer ML, Yanofsky C (1989) A morphological and genetic analysis of conidiophore development in Neurospora crassa. Genes Dev 3:559–571
Spudich JL (2006) The multitalented microbial sensory rhodopsins. Trends Microbiol 14:480–487
That TC, Turian G (1978) Ultrastructural study of microcyclic macroconidiation in Neurospora crassa. Arch Microbiol 116:279–288
Tsui H-CT, Pease AJ, Koehler TM, Winkler ME (1994) Detection and quantitation of RNA transcribed from bacterial chromosomes and plasmids. In: Adolph KW (ed) Methods in molecular genetics. Academic, San Diego, pp 197–200
Waschuk SA, Bezerra AG Jr, Shi L, Brown LS (2005) Leptosphaeria rhodopsin: bacteriorhodopsin-like proton pump from a eukaryote. Proc Natl Acad Sci USA 102:6879–6883
Westergaard M, Mitchell HK (1947) Neurospora V. A synthetic medium favoring sexual reproduction. Am J Bot 34:573–577
Acknowledgments
We acknowledge Marek Nemcovic, Donald Natvig, Jennifer Loros, Ann Kays and Svetlana Krystofova for many helpful discussions, Daniel Ebbole for plasmids and the John Spudich laboratory for use of their Alpha Imager TM 2200 Documentation and Analysis System. This work was supported by National Science Foundation Grant MCB-0296055 (to K.A.B.).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by G. Braus.
Rights and permissions
About this article
Cite this article
Bieszke, J.A., Li, L. & Borkovich, K.A. The fungal opsin gene nop-1 is negatively-regulated by a component of the blue light sensing pathway and influences conidiation-specific gene expression in Neurospora crassa . Curr Genet 52, 149–157 (2007). https://doi.org/10.1007/s00294-007-0148-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-007-0148-8