Abstract
Mitochondrial DNA (mtDNA) repair occurs in all eukaryotic organisms and is essential for the maintenance of mitochondrial function. Evidence from both humans and yeast suggests that mismatch repair is one of the pathways that functions in overall mtDNA stability. In the mitochondria of the yeast Saccharomyces cerevisiae, the presence of a homologue to the bacterial MutS mismatch repair protein, MSH1, has long been known to be essential for mitochondrial function. The mechanisms for which it is essential are unclear, however. Here, we analyze the effects of two point mutations, msh1-F105A and msh1-G776D, both predicted to be defective in mismatch repair; and we show that they are both able to maintain partial mitochondrial function. Moreover, there are significant differences in the severity of mitochondrial disruption between the two mutants that suggest multiple roles for Msh1p in addition to mismatch repair. Our overall findings suggest that these additional predicted functions of Msh1p, including recombination surveillance and heteroduplex rejection, may be primarily responsible for its essential role in mtDNA stability.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Maintenance and repair of the mitochondrial genome is of central importance to all respiring organisms. Mitochondria carry out the terminal steps of respiration, generating ATP and O2 via oxidative phosphorylation. Although the majority of mitochondrial proteins are encoded in the nucleus, many of those involved in electron transport and ATP synthesis are mitochondrially encoded. For this reason, mutations to mtDNA usually result in disruption of oxidative phosphorylation, strongly affecting tissues with high energy requirements, such as muscle and neural tissue (Shoubridge 2001). Mitochondrial mutation and malfunction are also associated with the progression of Huntington’s and Alzheimer’s neurodegenerative diseases (Graeber and Muller 1998). We are interested in identifying factors that contribute to mitochondrial (mt)DNA maintenance and fidelity in the budding yeast Saccharomyces cerevisiae, focusing in this study on the mismatch repair homologue Msh1p.
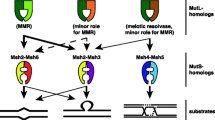
Msh1p, encoded by the nuclear MSH1 gene, plays an essential role in mtDNA maintenance. It is the only one of six yeast MutS homologues that is known to function in the mitochondria. MutS is the initiating protein of the well studied methyl-directed mismatch repair pathway in Escherichia coli. The primary role of MutS is the recognition of mispaired bases in newly synthesized DNA, to which it binds and subsequently activates downstream steps in mismatch repair (Modrich 1991). Other roles, including suppression of homeologous recombination, have also been defined for MutS (Chambers et al. 1996; Evans and Alani 2002; Evans et al. 2000; Junop et al. 2003). The yeast nuclear homologues MSH2, MSH3, and MSH6 are involved in these functions, while the remaining two, MSH4 and MSH5, are not required for mismatch repair but instead function in meiotic crossing-over (Hollingsworth et al. 1995; Pochart et al. 1997; Ross-Macdonald and Roeder 1994).
In E. coli, a MutS dimer initiates mismatch repair by recognizing and binding to mismatched DNA (Modrich and Lahue 1996; Su and Modrich 1986). Recent findings suggest that tetrameric MutS may also be involved in the initiation step (Bjornson et al. 2003). The MutS–DNA complex recruits the MutL protein dimer in an ATP-dependent manner, forming a ternary complex that contacts the MutH endonuclease. Activation by MutS–MutL causes MutH to cleave the nascent strand containing the mismatch. Cleavage is followed by exonuclease digestion of the DNA past the mismatched site, which is then resynthesized and ligated to the growing nascent strand (Harfe and Jinks-Robertson 2000; Modrich 1991).
In yeast, cells lacking Msh1p were shown to lose respiration competence within 8–10 generations, marked by rearrangements and deletions of the mitochondrial genome and a higher rate of spontaneous point mutation relative to the wild type (Reenan and Kolodner 1992a). Further analysis of Msh1p revealed intrinsic ATPase and mismatch-binding activities and showed that ATP-binding modulated the specificity of mismatch recognition in vitro (Chi and Kolodner 1994a, b). Despite the high degree of similarity between Msh1p and MutS, a much more extensively characterized protein, the precise biological function of Msh1p in the mitochondria is still unclear (Culligan et al. 2000; Eisen 1998).
All proteins in the MutS family possess DNA-binding and ATPase activities. The mechanism of mismatch surveillance, recognition, and binding is mediated by a conserved Phe residue in the N-terminal region of MutS (Bowers et al. 1999; Drotschmann et al. 2001; Lamers et al. 2000; Malkov et al. 1997; Schofield et al. 2001; Yamamoto et al. 2000). Studies of this N-terminal domain have provided a model whereby MutS “scans” for mismatches by deforming the DNA backbone, allowing the Phe residue to attempt to bind a mismatched base pair. Scanning continues until a mismatch is recognized and bound, inducing a conformational change in the protein that enables the recruitment of MutL and activation of downstream events. Studies in yeast and E. coli have confirmed that mutations made to this Phe result in a mutator phenotype consistent with a loss of mismatch repair (Bowers et al. 1999; Drotschmann et al. 2001).
ATP-binding and hydrolysis play an equally important role in mismatch repair (Alani et al. 1997; Bowers et al. 1999; Drotschmann et al. 2002; Gradia et al. 1997; Studamire et al. 1998). The exact contribution of ATPase activity to mismatch repair remains unresolved, however, although a large body of genetic and biochemical studies has attempted to explore this question (Alani et al. 2003; Junop et al. 2001, 2003; Acharya et al. 2003; Gradia et al. 1997, 1999). While differing mechanistically, all three models agree that ATPase activity and DNA-binding modulate each other and that the activity of both functional domains is required for mismatch repair.
Mutational analyses of the yeast nuclear genes MSH2, MSH3, and MSH6 have shown that mutating these conserved functional domains leads to a loss of nuclear mismatch repair (Alani et al. 1997; Studamire et al. 1998). One well studied mutation is a Gly→Asp conversion of the terminal glycine in the “p-loop” (a six-amino-acid motif for phosphate-binding, part of the Walker A-binding motif) in the ATPase domain of the gene. The yeast nuclear alleles msh2-G693D and msh6-G987D show significant increases in point mutations and microsatellite alterations due to a mismatch repair defect (Alani et al. 1997; Sia et al. 1997; Studamire et al. 1998). These alleles also display reduced ATPase activity (Alani et al. 1997; Studamire et al. 1998).
Recent studies show that the relevance of mitochondrial mismatch repair extends beyond yeast. Evidence for mismatch repair activity in rat and human mitochondrial lysates suggests that, although there is no identified Msh1p homologue in these systems, mitochondrial mismatch repair can occur, possibly using an alternate MutS homologue (Mason et al. 2003). In this paper, we investigate the effects of two mutations in the S. cerevisiae MSH1 gene that each displays only a partial defect in mtDNA maintenance, despite the prediction that they are mismatch repair-deficient. Additionally, we discuss evidence that physiological changes in the mitochondria are associated with MSH1 disruption and propose that multiple functions, including mismatch repair, account for the essential nature of Msh1p in the mitochondria.
Materials and methods
Yeast strains, plasmids, and growth conditions
The yeast strains used in this study were isogenic with DFS188 (MATa ura3-52 leu2-3,112 lys2 his3 arg8::hisG), which was derived from D273-10B. EAS504 was generated by deletion of the TRP1 gene from DFS188, using the plasmid pNKY1009 (Alani et al. 1987). To construct plasmids expressing the wild-type MSH1 gene, the MSH1 gene from −298 to the stop codon were amplified from DFS188 genomic DNA, using the primers 5′-CTTGAACTGCAGATGATACATTGA-3′ and 5-GGCTGCAGGTTATTCCCAATATTTCGCGGG-3′. The amplified fragment was cloned into pCR2.1 (Invitrogen, Carlsbad, Calif.) to generate pSM2. The XhoI–BamHI fragment containing MSH1 from this plasmid was ligated to XhoI- and BamHI-digested pRS415 and pRS416 to generate plasmids pSM4 and pSM3, respectively. The MSH1 gene was deleted in EAS504 (DFS188-Δtrp1) to generate YSM2 by one-step transplacement of MSH1 with the kanMX gene, using the primers 5′-CAATAATATAGATGGTACATAACATATGCGCAAGAAAACGTAAAGGCCACCGTACGCTGCAGGTCGAC-3′ and 5′-CAAATTATATACAATTAATAGTTGTATTCAAAAGTTATCCCAATATTTCGCATCGATGAATTCGAGCTCG-3′ (Sia et al. 2000). These strains failed to respire due to loss of Msh1p. To reintroduce functional mitochondria into strain YSM2, we mated it to isogenic strain EAS462 (DFS188α) and selected on glycerol medium with 200 μg/ml geneticin, to generate diploid strain YSM17. We transformed YSM17 with the pSM3 plasmid, selecting on synthetic medium lacking uracil to generate the diploid strain YSM18. Tetrads were dissected by standard methods onto glycerol medium lacking uracil, then replica-plated to dextrose media with 200 μg/ml geneticin to confirm antibiotic resistance. This selection identified YSM19 (YSM2+pRS416-MSH1, ρ+), a respiring, msh1Δ::kanMX haploid strain carrying the wild-type MSH1 gene on plasmid pSM3. The mutant alleles pLAJ12 (msh1-G776D), pSM105 (msh1-F105A), and pSM106 (msh1-F105A/G776D) were generated in pSM4 by site-directed mutagenesis, using the Quikchange XL kit (Stratagene, Cedar Creek, Tex.) and the primer pairs 5′-GTTTTACACAAATGGGGTTCAGCTTATGAACTTTACTTTGAACAA-3′ and 5′-TTGTTCAAAGTAAAGTTCATAAGCTGACCCCATTGGTGTTAAAAC-3′ for msh1-F105A and 5′-GTGTCTTAAGAATGTAGATTTATCACCCATATTCGGTCCGGTAAT-3′ and 5′-ATTACGGACCGAATATGGGTGATAAATCTACATTCTTAAGACAG-3′ for msh1-G776D.
YP rich medium contained 1% yeast extract, 2% Bacto peptone, and 2% dextrose or 2% glycerol as specified in the text. Synthetic medium contained 0.17% yeast nitrogen base without amino acids, 0.5% ammonium sulfate, and 2% dextrose or 2% glycerol, and was supplemented as described by Sherman (1991). YG-Ery medium contained 2% glycerol, 2% yeast extract, and 4 mg/ml erythromycin. Media descriptions are abbreviated as follows: YPD for YP rich medium with dextrose, YPG for YP medium with glycerol, SD for synthetic medium with dextrose, −Ura for media lacking uracil supplementation, −Leu for media lacking leucine supplementation, −Lys for media lacking lysine supplementation. SD −Leu−Ura, for example, is synthetic dextrose medium lacking both leucine and uracil.
Plasmid shuffle
Strains containing MSH1 alleles were constructed via plasmid-shuffling. Plasmids pSM4 (MSH1), pLAJ12 (msh1-G776D), pSM105 (msh1-F105A), and pSM106 (msh1-F105A/G776D) were each transformed into YSM19 and selected on SD −Leu−Ura to generate YSM39, YSM41, YSM40, and YSM42, respectively. Colonies were replica-plated onto SD −Leu supplemented with 5-fluoro-orotic acid (5-FOA; 1 g/l) to select for loss of pSM4. After 2 days incubation at room temperature, colonies were replicated from 5-FOA medium to SD −Ura, to YPG, and then to SD −Leu to confirm the loss of wild-type MSH1 and retention of the mutant allele, and to test the respiratory capacity of cells sustained only by the mutant.
Fluctuation analysis of mitochondrial point mutation accumulation
For each strain, yeast were streaked for independent colonies on YPG and 20 independent colonies were then picked into 5 ml YPG liquid cultures and grown for 2 days at 30°C. Appropriate dilutions were made and plated onto YPG and YG media supplemented with 50 mM sodium phosphate (pH 6.5) and 4 g/l erythromycin. These plates were incubated at 30°C for 7 days. We calculated the rate of spontaneous conversion to erythromycin resistance by the median method (Lea and Coulson 1949). Each rate calculated represents the mean value of two independent fluctuation tests with overlapping 95% confidence intervals.
Determination of respiration loss and growth curves
Strains assayed were grown in YPG liquid medium to saturation at 30°C. Then, at T0, they were diluted 1:50 into 10 ml SD −Leu. At the time points given, 10 μl were removed, appropriately diluted, and plated onto YPG medium supplemented with 0.1% dextrose. Respiring colonies grow normally on these plates, while non-respiring cells are limited by the availability of dextrose and form very small colonies. Plates were scored after 5 days incubation at 30°C. For growth curves, strains were cultured overnight in SD−Leu and then diluted 1:50 into 10 ml SD −Leu at T0. Growth was measured by reading the optical density at 600 nm (OD600) of each culture at the time points given.
Fluorescence microscopy for mitochondrial morphology
Yeast strains were grown overnight at 30°C in either YPD or YPG. A total quantity of 1 ml from each mid-log phase culture was supplemented with Mitotracker Red (Molecular Probes, Eugene, Ore.) to a final concentration of 100 pM and incubated for 1 h at 30°C. Images of 4′,6-diamidino-2-phenylindole (DAPI) staining were obtained by additionally supplementing cells with 10 nM DAPI during the last 5 minutes of incubation. Cells were harvested, washed two times in appropriate medium, and spotted onto poly-L-lysine-coated slides. Fluorescence microscopy was performed on a Zeiss Axioplan 2 microscope.
Determination of H2O2 sensitivity
Cells were assayed essentially as described by Meeusen et al. (1999). Briefly, 1 ml cells were grown overnight in YPG, washed once with YPD, and resuspended in 1 ml YPD containing 0–10 mM H2O2. Cells were incubated at 30°C for 1 h, washed twice in YPD, and plated to YPG containing 0.1% dextrose to support growth of both respiring and non-respiring cells (enhancing the petite phenotype of the latter, which cease to grow after consuming the available dextrose). Plates were incubated at 30°C for 4 days and scored for percent respiring colonies.
Msh1p antibody production and immunofluorescence
MSH1 was cloned into pETBlue1 using primers 5′-ATGCATCATCATCATCATCACAAGCATTTCTTTAGGCTACCG-3′ and 5′-TTATCCCAATATTTCGCGGGC-3′ and generating an insertion of 6×His at the N-terminus of Msh1p. This vector was transformed into E. coli and cells were grown to mid-log phase in LB with 100 μg/ml ampicillin. Isopropyl-β-D-thiogalactopyranoside was added to a final concentration of 1 mM and cells were induced for 3 h while shaking at 37°C. Cells were harvested and protein lysates made, using the Novagen HisBind protocol (Novagen, Madison, Wis.). Msh1p was then purified using the Novagen HisBind kit. Purified Msh1p eluate was run on a sodium dodecyl sulfate (SDS)-polyacrylamide gel and stained with Coomassie blue dye to resolve protein bands, which were excised and Msh1p identity confirmed by mass spectroscopy. Protein for antibody production was sent in acrylamide slices to Pocono Rabbit Farm and Laboratory (Canadensis, Pa.). Recovered serum aliquots were monitored for antibody content by probing Western blots of 6×His-purified Msh1p and by the immunofluorescence of DFS188 wild-type cells.
DFS188 wild-type yeast cells were grown in YPD and harvested at mid-log phase. Formaldehyde was added directly to 1 ml cell culture to a final concentration of 3.7% and incubated at 30°C for 5 min. Cells were harvested, washed twice with PBS, pH 7.4, and twice again with PBS–1 M sorbitol, resuspending to a final volume of 1 ml. A total quantity of 5 μl β-mercaptoethanol (14.2 M) and 30 μl zymolyase (10 mg/ml) were added and cells were then incubated at 30°C for 20 min with shaking. Cells were harvested, washed in PBS, and adhered to poly-L-lysine-coated slides. Slides were plunged into −20°C methanol for 2 min and then into −20°C acetone for 5 s and allowed to dry. Primary antibody (α-Cox3p or α-Msh1p) was applied at an appropriate dilution (1:100 or 1:250, respectively) and incubated overnight at 37°C. Cells were washed for 3 h in Tris-buffered saline with 0.05% Tween-20 (TBST), drained, and secondary antibody was applied at 1:500 dilution. For Cox3p, anti-mouse Alexa 488 was used, and for Msh1p, anti-rabbit 568 was used (Molecular Probes, Eugene, Ore.). Cells were washed for 1 h in TBST, mounted in Gelvitol medium, and imaged with a Leica Microsystems confocal microscope.
Protein lysates and Western blotting
Protein lysates were prepared by the method of cell disruption using glass beads, as described in Ausubel et al. (1994) with the following changes: cells were grown in 200 ml synthetic dextrose medium to an OD600 of about 0.2. Cells were harvested, resuspended in 0.3 ml lysis buffer, and prepared as described. Lysate was cleared with spin at 3,000 rpm for 5 min at 4°C. Quantification of protein concentration was determined using the Bradford reagent protein assay (Bio-Rad Laboratories, Hercules, Calif.). A sample (approximately 400 μg total protein) from each strain was run on a denaturing SDS 8% polyacrylamide gel and blotted using standard methods. Antibody against Msh1p was prepared from rabbit serum as follows: 1 ml serum was applied to a DEAE Affi-Gel Blue column (5 ml; Bio-Rad Laboratories, Hercules, Calif.), washed with 10 ml PBS and eluted with 10 ml elution buffer (1.4 M NaCl, 0.020 M Tris-HCl, pH 8.0). Eluate was collected in 0.6-ml fractions and protein concentration was quantified by Bradford assay. Blots were incubated with a 1:10 dilution of DEAE-purified antibody in TBST overnight at room temperature, washed 4× 10 min in TBST, incubated with 1:1500 horseradish peroxidase-conjugated anti-rabbit antibody (Zymed, San Francisco, Calif.), and washed as before. Reaction of Msh1p with the antibody was detected using the ECL Western kit (Amersham Pharmacia Biotech, Piscataway, N.J.).
Results
Mutation studies of the MutS family proteins confirmed the importance of its conserved domains for overall protein function. In particular, analysis of the nuclear mismatch repair genes MSH2, MSH3, and MSH6 showed that point mutations resulting in single amino acid substitutions within these domains lead to loss of mismatch repair.
Previous studies of the effects of MSH1 disruption in vivo were all carried out in strains retaining wild-type gene expression. Chi and Kolodner (1994a, b) demonstrated an increase in point mutation in a MSH1/msh1 diploid, while Koprowski et al. (2002) showed that overexpression of a plasmid-borne msh1-G776D mutant resulted in the same (Chi and Kolodner 1994b; Koprowski et al. 2002). In this study, we constructed strains lacking wild-type Msh1p expression, allowing us to isolate the msh1 mutant phenotypes. This approach may clarify the functions that are required for Msh1p-dependent mtDNA maintenance.
Msh1p mismatch repair mutants retain partial respiratory function
Figure 1 shows a diagrammatic representation of Msh1p and partial alignments of its ATPase (upper) and mismatch recognition (lower) domains with MutS, Msh2p, and Msh6p. The ATPase alignment shows the six-amino-acid motif known as the p-loop, used for phosphate-binding, and the terminal Gly which was mutated to Asp for this study. This Gly→Asp mutation reduces ATP-binding and eliminates hydrolysis, disrupting mismatch repair in MSH2, MSH3, MSH6, and MutS (Studamire et al. 1998; Wu and Marinus 1994). Figure 1 (lower) gives an alignment of the mismatch recognition domain, showing the conservation of residues that participate in DNA-binding and mismatch specificity. In particular, the importance of the Phe residue conserved across Msh1p, Msh6p, and MutS has been shown by substituting Phe with Ala and demonstrating a loss of mismatch repair in the resulting mutants (Bowers et al. 1999; Drotschmann et al. 2001; Dufner et al. 2000).
Schematic representation of Msh1p showing conserved regions as shaded rectangles. From N-terminus to C-terminus are the mismatch recognition (dark grey), ATPase (light grey), and helix-turn-helix (black) functional domains. The two alignments show portions of these domains containing the amino acid substitutions used in this study. The upper panel shows the six-amino-acid p-loop motif (shaded box) within the ATPase domain, containing the Gly→Asp mutation in bold. The lower paner shows the alignment of the N-terminal mismatch recognition/binding domain containing the Phe→Ala mutation in bold. Numbers refer to the position of the first amino acid in each alignment. The diagram is not to scale
Since the Gly→Asp and Phe→Ala mutants behave consistently in all MutS homologues examined, we constructed the msh1-G776D and msh1-F105A alleles via site-directed mutagenesis to assess their effects in MSH1. Previous studies of the msh1-G776D mutant utilized a plasmid-borne mutant construct that was expressed in the presence of the wild-type MSH1 gene (Koprowski et al. 2002). We introduced our mutant constructs via plasmid shuffle into a strain lacking chromosomal MSH1 expression to examine the resulting phenotypes in the absence of wild-type protein. Intact mitochondrial DNA is required for respiration to occur, enabling us to use it as an indirect marker for mtDNA maintenance.
The multi-step plasmid shuffle technique is necessary to construct these strains without irreversible loss of mitochondrial function prior to assaying the mutants (Fig. 2a). We deleted the chromosomal copy of the MSH1 allele in a strain that carries a second MSH1 copy on a plasmid bearing the URA3 auxotrophic marker. The ability of these strains to respire is dependent on the plasmid-borne allele. Following the introduction of a second, msh1-bearing plasmid carrying the LEU2 auxotrophic marker, we selected against the presence of the URA3 plasmid by growth in medium containing 5-FOA, a drug that is toxic to cells expressing Ura3p. The resulting strain, shown at the right of Fig. 2a, has as its only source of Msh1p the mutant allele carried on the LEU2 plasmid and is maintained under selection for plasmid retention.
Growth phenotypes of msh1 alleles before and after plasmid shuffle. a Model of plasmid shuffle scheme showing cells undergoing loss of the URA3-containing plasmid. Yeast cells are represented by thick black lines and nuclei are represented by thin lines. Ovals represent plasmid vectors, double lines represent the nuclear chromosome, and shaded boxes represent alleles of MSH1. The black bar indicates the knockout substitution of MSH1 with the kanMX antibiotic resistance gene. The black star indicates a point mutation resulting in a single amino acid substitution. A functional mitochondrion is surrounded by short grey speckles (left). Question marks indicate unknown mitochondrial function (right). Arrows indicate growth on 5-FOA-containing medium. b Cells on SD − Leu− Ura (upper panel) and YPG (lower panel) prior to 5-FOA selection. All cells contain pSM4 (containing URA3) carrying MSH1 and variants of pSM3 (containing LEU2) carrying msh1 alleles. Labels refer to the allele carried by the LEU2 plasmid. c Cells on SD-Leu (upper panel) and YPG (lower panel) following 5-FOA selection. Cells contain only pSM3 carrying msh1 allelic mutants
Using this strategy, we introduced MSH1, msh1-F105A, msh1-G776D, msh1-F105A/G776D, and an empty vector control into cells containing wild-type MSH1 on the URA3 plasmid. These strains are shown on selective medium (SD − Leu− Ura) and glycerol medium (YPG) in Fig. 2b. All strains grow comparably on both media, demonstrating that they are respiration-competent when harboring both plasmids. After 5-FOA selection, these phenotypes change as shown in Fig. 2c. There is robust growth on medium lacking leucine, indicating that the plasmids bearing msh1 alleles are still present in the cells, but this growth is reduced in the msh1 mutant strains when grown on glycerol medium. Notably, however, cells expressing msh1-F105A and msh1-G776D are still partially able to maintain respiring colonies. In contrast, the double mutant msh1-F105A/G776D is severely impaired and shows growth on glycerol medium that is comparable with the vector alone, indicating complete loss of respiration.
We performed a quantitative analysis of these phenotypes by examining the growth profiles and respiration loss of each mutant strain. Figure 3a,b show the respiration loss profiles of the strains carrying plasmid-borne MSH1 alleles before and after the plasmid shuffle, respectively. For this assay, strains were grown in glycerol to saturation. Selection for respiration was released at T0 by shifting cells to synthetic dextrose medium lacking uracil and leucine, allowing non-respiring cells to accumulate while maintaining selection for the two plasmid-borne msh1 alleles. Samples were removed and plated onto glycerol medium plus limiting dextrose to score the percent respiring cells, as described in the Materials and methods.
Growth and respiration loss phenotypes of MSH1 and msh1 strains. Each panel shows MSH1 (closed circles), msh1-F105A (open squares), msh1-G776D (closed triangles), and msh1-F105A/G776D (crosses) in synthetic dextrose medium lacking either leucine and uracil (a,d) or leucine (b,e). a Respiration loss of cells prior to shuffle, carrying wild-type MSH1 and mutant msh1 alleles, over a 24-h period. The assay was conducted as described in the Materials and methods. Each point in the respiration loss charts represents the mean value of 3–5 samplings (n is ca. 200 colonies/sample). b Respiration loss of post-shuffle cells carrying only msh1 mutant alleles (see abbreviations above). c Respiration loss of wild-type yeast bearing the pRS415 vector (closed squares) or carrying MSH1 (open diamonds) and mip1-D347A (open circles). d Typical growth profile of cells prior to plasmid shuffle. e Typical growth profile of cells following plasmid shuffle
The MSH1/MSH1 strain retained the highest percent respiring cells over a 24-h period, ranging from 98% to 60% (Fig. 3a). The MSH1/msh1-G776D strain showed only a modest average decrease, with a range of 84–67%, suggesting that the detrimental effects of the ATPase mutation do not strongly affect mitochondrial function in the presence of wild-type protein. This is especially striking considering that, in MutS and its yeast nuclear homologues, this G→D mutation is phenotypically dominant, due to the requirement of both ATPase domains in the protein dimer for full ATPase function (Lamers et al. 2000; Studamire et al. 1998). By contrast, the F105A mutation gives rise to a significant loss of respiring cells over the time-course, dropping from 80% to 24%. Respiration loss is enhanced by the additional presence of the G776D mutation, as shown by the double mutant in the MSH1/msh1-F105A/G776D strain, which begins with only 42% respiring and ends with 2%.
We then examined the respiration loss of the strains carrying only one msh1 allele and found their phenotypes to be consistent with those observed in the presence of wild-type MSH1 (Fig. 3b). The msh1-F105A and msh1-G776D mutant strains show reduced respiration maintenance relative to MSH1, although this reduction is modest in cells expressing msh1-G776D. Cells bearing wild-type MSH1 range from 80% to 60%, while msh1-G776D cells range from 65% to 47%. The msh1-F105A strain retains almost no respiration capability in the absence of selection when MSH1 is lost, maintaining only 0.5–2.0% respiring cells. The double mutant cannot be assayed by this method as it is completely respiration-deficient.
A comparison of these data with the respiration loss profiles of strains concurrently expressing endogenous MSH1 reveals that plasmid-borne MSH1 does not fully complement loss of the chromosomal gene copy. As shown in Fig. 3c, wild-type DFS188 strains carrying either the empty vector pRS415 or pRS415-MSH1 both maintain near 100% levels of respiration capability. Moreover, expression of Msh1p from both the plasmid-borne and chromosomal gene copies does not appear to affect maintenance of respiring cells. Figure 3c also shows that the mip-D347A polymerase mutant, which displays a 475-fold higher rate of point-mutation accumulation, maintains high levels of respiring cells in a profile comparable with that of msh1-G776D (Phadnis and Sia 2004). This mutation is a substitution in the Exo3 motif of the 3′–5′ exonuclease domain, which decreases proofreading exonuclease activity approximately 500-fold during in vitro assays (Foury and Vanderstraeten 1992).
Growth rates of the msh1 mutant strains reveal additional differences between the two mutations. In Fig. 3d, strains expressing both alleles display differences in growth rate that are inverse to their respiration capability when grown in synthetic dextrose medium lacking leucine and uracil. The MSH1/MSH1 and MSH1/msh1-G776D strains both grow slowly, with a doubling time of 6 h, relative to the MSH1/msh1-F105A and MSH1/msh1-F105A/G776D strains, with doubling times of 3.2 h and 3.6 h, respectively. By contrast, growth rates of the strains expressing a single MSH1, grown in synthetic dextrose medium lacking leucine, vary much less (Fig. 3e). All strains, with the exception of msh1-F105A, grow with doubling times of 2.3 h (MSH1) and 2.5 h (msh1-G776D, msh1-F105A/G776D), while msh1-F105A lags behind, doubling every 3.6 h. A control ρ− strain bearing plasmid-borne MSH1 grew similarly to the two ATPase-deficient strains, doubling every 2.5 h (data not shown).
Confirmation of Msh1p localization in the mitochondria
Previous localization of Msh1p to the mitochondria was demonstrated using indirect immunofluorescence against a Msh1p-12CA5 epitope fusion protein overexpressed from the GAL10 promoter (Chi and Kolodner 1994b). We used purified protein to raise antibody against wild-type Msh1p and subsequently examine the localization of untagged, endogenously expressed Msh1p (Fig. 4a). Consistent with previous results, we found that Msh1p localizes to the mitochondria in the absence of overexpression. Immunofluorescence of wild-type DFS188 cells with anti-Msh1p produced a punctate staining pattern that was coincident with antibody staining against Cox3p, an electron-transport protein synthesized in the mitochondria and localized in the mitochondrial membrane (Fig. 4b). In the msh1-Δ strain, which is non-respiring and therefore contains aberrant mitochondria, we observed only non-specific staining with anti-Msh1p that did not correlate with anti-Cox3p staining (data not shown). To ensure that all of the plasmid-borne alleles express protein similarly, Western blot analysis was performed (Fig. 4c). There are comparable levels of Msh1p expressed from the chromosomal and each plasmid-borne allele and an absence of protein at the same position in the msh1-Δ strain.
Msh1p localization to mitochondria in vivo. a Differential interference contrast (DIC) image of wild-type DFS188 cells. b Confocal image of anti-Msh1p localization showing punctate staining. c Confocal image of the same cells showing anti-Cox3p, a mitochondrial marker. d Overlay of anti-Msh1p and anti-Cox3p images showing co-localization of staining in the mitochondria. All cells were grown in glycerol medium, fixed, stained, and imaged by confocal microscopy as described in the Materials and methods. e Signal intensity of anti-Msh1p (red) and anti-Cox3p (green) across a section of the image overlay marked by a red bar in d. f Western blot of: (from left to right) wild-type DFS188, msh1-Δ, and the strains containing plasmid-borne MSH1, msh1-F105A, msh1-G776D, and msh1-F105A/G776D, probed with anti-Msh1p. Blotting, staining, and detection were carried out as described in the Materials and methods. Each lane shows approximately 400 μg total protein. All lanes show equivalent expression of Msh1p by each allele, except msh1-Δ, which shows no expression
msh1-F105A and msh1-G776D show increases in point mutation rate of mtDNA
Loss of Msh1p severely disrupts mtDNA stability after very few generations, a profile at odds with a predicted role primarily in mismatch repair (Chi and Kolodner 1994b; Reenan and Kolodner 1992a). To determine whether Msh1p does play a role in single-base mismatch repair in mtDNA, we measured the rate of mitochondrial point mutation accumulation by fluctuation test. A total of 15–20 individual colonies from each strain were grown in liquid glycerol medium, diluted, and plated onto glycerol medium containing erythromycin. Resistance to erythromycin arises from single nucleotide changes in the gene encoding the 28S mitochondrial ribosomal subunit. Thus, erythromycin resistance serves as a reporter of the accumulation of point mutations.
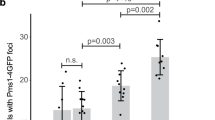
As shown in Fig. 5, strains expressing wild-type Msh1p gave rise to point mutations at a rate of 4.0×10−8 mutations/cell division. Both of the msh1 mutant strains showed significantly higher rates of mutation accumulation. Msh1-F105A displayed a rate of 1.4×10−6 mutations/cell division, 35-fold over the wild-type rate, while msh1-G776D showed a slightly lower rate of 7.6×10−7 mutations/cell division, 19-fold over the wild-type rate. Overall, however, these rates are surprisingly modest compared with the Polγ polymerase mutant mip1-D347A, which displays a rate of 1.9×10−5 mutations/cell division. The comparison is especially striking, given that mip-D347A maintains a respiration capability similar to msh1-G776D Fig. 3c).
Rates of point mutation accumulation/cell division in mitochondrial DNA of strains carrying mutant alleles of MSH1 (msh1-F105A, msh1-G776D) or MIP1 (mip1-D347A). The frequency of EryR was measured by fluctuation analysis; and rates were calculated by the median method, as described in the Materials and methods. Each bar represents the mean value of two separate assays, each consisting of 20 independently isolated colonies of each strain. Rates for mip1-D347A were published by Phadnis and Sia (2004)
Sensitivity to H2O2-induced oxidative stress is not Msh1p-dependent
To determine whether Msh1p has a role in protecting the cell from oxidative stress, we assayed the wild-type and mutant MSH1 strains for respiration competence following exposure to H2O2. Saturated cultures in glycerol medium were washed and transferred to liquid dextrose medium containing H2O2 at concentrations of 0–10 mM. After 1 h incubation, the cells were diluted and plated onto limiting dextrose medium to allow measurement of both viability (measured by total colony count) and respiration competence (measured by the percent respiring cells). The viability of cells decreased similarly with exposure to higher concentrations of H2O2 in all three strains, and at 10 mM H2O2 showed an approximately 2-fold reduction in colony-forming units relative to cells plated directly from dextrose media alone (data not shown). Figure 6 shows a concurrent reduction in respiration capability as H2O2 exposure increases, but there is no significant difference between the wild-type and mutant-bearing strains, suggesting that the effects of Msh1p disruption do not affect cell sensitivity to oxidative damage. This is clearest when comparing the MSH1 and msh1-G776D strains, as the msh1-F105A cells do not retain enough respiring cells to show a clear trend. Dzerzbicki et al. (2004) recently suggested that in the absence of mitochondrial base excision repair (BER), which usually repairs oxidative damage, mitochondrial mismatch repair (mediated by Msh1p) acts as a “back-up” pathway to reduce the accumulation of mutations. However, in the absence of mutations that disrupt BER, we see no apparent effects of Msh1p disruption on cell sensitivity to oxidative damage.
Mitochondrial morphology is altered in msh1-F105A and msh1-G776D cells
It was shown that changes in mtDNA stability are correlated with alterations in mitochondrial morphology (Boldogh et al. 2003; Hanekamp et al. 2002; MacAlpine et al. 2000). To determine whether the partial function alleles of Msh1p affect mitochondrial morphology, we stained cells with Mitotracker Red, a cationic fluorescent dye that passively diffuses across the mitochondrial membrane and remains sequestered in the mitochondria. Because sequestration is membrane potential-dependent, it can only stain active mitochondria. Figure 7 shows stained mitochondria in cells taken from glycerol and synthetic dextrose media, demonstrating the differences in structure between wild-type and mutant mitochondria grown both with and without respiration selection.
H2O2 sensitivity of msh1 strains. Cells were assayed as described in the Materials and methods. Each point represents the mean of five samplings (n is ca. 200 cells/sample). Mid-log phase cells were harvested and resuspended in YPD medium containing H2O2 at 0, 1, 5, or 10 mM, incubated for 1 h with agitation, washed, diluted, and plated. After 4 days growth at 30°C, plates were scored for growth and percent respiration
All cells grown in glycerol medium displayed highly reticular mitochondrial structures (Fig. 7, column 2). There was a slight tendency for the msh1-F105A and msh1-G776D mitochondria to appear more knobby or granular, rather than being smooth. In addition, the networks seemed less extensively distributed. However, when the same strains were grown in dextrose medium, we observed large differences in mitochondrial morphology. Cells expressing wild-type Msh1p still showed a reticular network, although glucose repression of mitochondrial biogenesis prevented the same extensive network as seen in glycerol-grown cells (Pon and Schatz 1991). All mutants, however, showed marked disruptions ranging from irregularity of the mitochondrial tubules (msh1-G776D) to complete condensation of mitochondrial material into brightly staining spots (msh1-F105A, msh1-F105A/G776D).
To confirm that we were observing mitochondrial structures and to assess whether Msh1p disruption affects mtDNA, we co-stained cells with Mitotracker Red and DAPI. Figure 8 shows MSH1 cells (top row), msh1-F105A cells (middle row), and msh1-G776D cells. Clearly visible in the MSH1 cells were many nucleoid bodies distributed throughout the same regions stained with Mitotracker Red, confirming that these structures were mitochondria. Relative to the wild type, both the F105A and G776D mutations led to an apparent reduction in nucleoid number, although they fluoresced more intensely. Distinct from nucleoids, the nucleus in each cell could be seen as a larger, more dimly stained spot.
Morphological changes associated with Msh1p disruption. Mid-log-phase cells (OD600 ca. 0.3) were vitally stained with Mitotracker Red and imaged as described in the Materials and methods. Cells in columns 1 and 2 were grown in YPG prior to staining, while columns 3 and 4 show cells grown in SD−Leu. Each image shows a typical representative of the majority of cells in each field
Additionally, we recorded a 24-h progression of mitochondrial dynamics in cells following release from respiration selection (Fig. 9). Identical to Fig. 3, saturated cultures in glycerol medium were diluted at T0 into synthetic dextrose medium, removed at the time points shown, stained live with Mitotracker Red and DAPI, and photographed. Strains bearing wild-type MSH1 are shown on the left of each column, demonstrating the normal progression of mitochondrial dynamics as a culture grows to saturation. At T0, cells taken from glycerol display highly reticular and extensively networked mitochondria, seen also in Fig. 7. As growth in dextrose progresses, the mitochondria fuse into thicker tubular structures that gradually thin and fragment until, near saturation at T24, they form smaller punctate spots that cluster at the cell periphery.
Distribution of nucleoids within MSH1 and msh1 strains grown in dextrose medium. Mid-log phase cells (OD600 ca. 0.3) were vitally stained with Mitotracker Red (MtR) and DAPI and imaged as described in the Materials and methods. The arrow indicates the position of the nucleus as seen by nuclear DNA staining
Progression of mitochondrial disruption in msh1 cells in dextrose medium. Actively growing cells were removed at time-points (T0–T24, hours) indicated, vitally stained with Mitotracker Red and DAPI, and visualized by fluorescence microscopy. Note that Mitotracker Red does not stain mitochondrial material without an active membrane potential. Lack of staining indicates loss of membrane potential, not loss of mitochondrial material
Immediately adjacent are cells bearing msh1-F105A. Note that the similarity to wild-type cells at T0 becomes increasingly divergent over the course of culture growth, until the membrane potential is nearly eliminated by T24, indicating the loss of mitochondrial membrane potential and function. In conjunction, the nucleoid number decreases, though mtDNA is still present in cells that lack a mitochondrial membrane potential. Msh1-G776D cells, to the right of each column, display a mild disruption that is intermediate between the MSH1 and msh1-F105A cells in both mitochondrial structural aberration and nucleoid number reduction.
Discussion
The dependence of mitochondrial genome stability on the function of Msh1p is well documented (Chi and Kolodner 1994a, b; Koprowski et al. 2002; Reenan and Kolodner 1992a, b). Characterizing Msh1p as a mitochondrial mismatch repair protein is logical, based on both its homology to MutS and the mutator phenotype produced by Msh1p disruption, but the significance of its mismatch repair capability regarding mitochondrial function is unknown. Studies of MutS family proteins from several model systems support the possibility that Msh1p may serve multiple functions in the maintenance of mtDNA (for a review, see Schofield and Hsieh 2003). We propose that mismatch repair is a relatively minor element of mitochondrial genome maintenance due to the phenotypes we observe in the msh1 mutant strains and the factors affecting mtDNA stability which differentiate it from nuclear DNA. These include a highly repetitive and A/T-rich composition and multiple genome copies per cell, all of which may make ectopic recombination a greater threat to overall genome stability than point mutation. Additionally, it is known that the Polγ mitochondrial polymerase has a very low rate of misincorporation, further reducing a strong requirement for post-replicative mismatch repair (Roberts and Kunkel 1996). Msh1p does appear to inhibit both polymerase slippage and mismatch accumulation, but mtDNA stability may also depend on other potential functions of the protein, such as Msh1p-mediated suppression of ectopic recombination (this study; Koprowski et al. 2002).
The msh1-F105A and msh1-G776D mutants display differences in phenotypic severity and penetrance
As shown by respiration loss assay, the effects of msh1-F105A and msh1-G776D on mitochondrial function are markedly dissimilar. This is partially mitigated by the presence of wild-type protein, but is still quite large (Fig. 3a,b). The data reveal differences between the two mutants, both in the degree of mitochondrial disruption and in the penetrance of each genotype. The F105A mismatch recognition mutation confers almost total loss of respiration, but shows a high degree of rescue in the presence of wild-type protein, suggesting that it may be co-dominant. Conversely, the G776D mutation does not greatly disrupt function, but this disruption is little changed in the presence of wild-type protein, suggesting that it is a dominant mutation, as was suggested by Koprowski et al. (2002). One potential explanation for these behaviors is that Gly 776 participates in ATP-binding and hydrolysis, which requires the ATPase domains of both subunits of the protein dimer, while Phe 105 is only required in one subunit of each dimer. It is interesting to note that the presence of the double mutant Msh1p-F105A/G776D confers the greatest reduction in respiration capability of the strains containing two Msh1p species. This suggests that the mutant protein, although unable to support mitochondrial function, is still able to interact with other Msh1p monomers, possibly forming non-functional multimers. One important consideration of these interpretations, however, is that the plasmid-borne alleles do not fully complement loss of endogenous MSH1, so the affects of these mutant msh1 constructs on respiration are likely to be more severe than if the mutations were present in endogenous MSH1.
Although wild-type Msh1p does provide partial complementation of respiration loss, it appears to adversely affect growth rate. When cells containing two MSH1 alleles are grown without a respiratory requirement, the growth rate is actually hindered in the two strains which have (presumably) higher levels of activity, Msh1p/Msh1p and Msh1p/Msh1p-G776D, while the remaining two grow at a higher rate. By contrast, the single-copy strains show much greater similarity in growth rate. We observed previously that overexpression of Msh1p is toxic to the cell, which could explain the reduced growth ability of the double-copy Msh1p strains (Sia and Sia 2002; unpublished data). Each MSH1 allele is borne on an ARS-CEN vector plasmid, which is normally present at low copy (Sikorski and Hieter 1989). Since the plasmid-borne MSH1 is expressed from its endogenous promoter, the levels of Msh1p expression we see from the plasmid should be similar to expression from the chromosome. However, with multiple plasmid species in the cell, both maintained by auxotrophic selection, it is possible that expression of Msh1p could rise to toxic levels. Figure 3c shows that the ability to respire is not affected by Msh1p expression from both a plasmid-borne and chromosomal copy of MSH1.
In addition to phenotypic differences regarding respiration and growth rate, we also find a moderate, ca. 2-fold difference in the mutator phenotypes conferred by each mutation. However, it is relatively minor compared with the high rate of mutation conferred by the polymerase mutant mip1-D347A. The comparison is especially striking, given that the mip-D347A strain maintains a respiration capability similar to msh1-G776D throughout the time-course (Fig. 3c). Previous findings reveal that polymerase mutants with mutation rates that are several orders of magnitude higher than the wild type are capable of respiration on a non-fermentable carbon source (Hu et al. 1995; Vanderstraeten et al. 1998). We believe that the phenotype exhibited by the mip1-D347A mutant is an example of point mutation accumulation alone. This suggests that mtDNA changes more profound than point mutation (or at much higher rates) are required for complete loss of respiration. In light of this, neither the msh1-F105A nor the msh1-G776D mutator phenotypes are sufficient to explain the rapid destabilization of mtDNA following Msh1p loss.
Two mutations, different disruptions: the importance of mismatch recognition in mtDNA
Since the G776D mutation results in an ATPase-deficient protein that is presumably incapable of mismatch repair, msh1-G776D would produce a null, msh1-Δ phenotype, were its sole function mismatch repair. We clearly showed, however, that more severe phenotypes can be induced by disrupting the DNA-binding and mismatch recognition activities of Msh1p through the F105A mutation, suggesting there are additional functions of Msh1p that are mismatch repair-independent. It should be noted that the analogous G→D mutation in Msh2p, Msh3p, Msh6p, and MutS confers a phenotype consistent with complete loss of mismatch repair (Alani et al. 1997; Haber and Walker 1991; Kijas et al. 2003; Studamire et al. 1999; Studamire et al. 1998). This loss is not concurrent with a complete loss of ATPase activity in vitro, which is reduced 3- to 7-fold in mutant/wild-type heterodimers and about 10-fold in mutant homodimers (Studamire et al. 1998). However, even one mutant monomer is sufficient to prevent two crucial steps in mismatch repair: the ATP-binding-dependent conformational change that causes dissociation from the bound mismatch and the ATP hydrolysis-dependent recruitment of MutL homologues and formation of the ternary complex that activates excision of the mismatch-containing strand (Kijas et al. 2003).
These findings suggest that efficiency of repair is not dependent on the rate of ATPase activity per se, but rather the ability of ATP to bind and alter the DNA-binding specificity and mismatch affinity of MutS homologues at the appropriate moment in the pathway and also recruit downstream MutL homologue partners. Additionally, disruption of ATP-binding and hydrolysis appear to primarily affect the mismatch repair pathway.
By contrast, mismatch recognition is a crucial component of both mismatch repair and recombination surveillance by the MutS family (Bowers et al. 1999; Chambers et al. 1996; Datta et al. 1997; Dufner et al. 2000; Junop et al. 2003). In the yeast nucleus, heterodimers Msh2p/3p and Msh2p/6p have two identified roles in double-strand break repair (DSBR) pathways. The first involves regulation of homologous recombination at the point of strand invasion, known as heteroduplex rejection or antirecombination. If the invading strand contains heterologies that produce mismatches when paired with the prospective template, Msh2p/3p binds to the mismatches and blocks further processing. The attempted pairing is predicted to be unwound by the helicase activity of Sgs1p (Jinks-Robertson, personal communication; Myung et al. 2001; Sugawara et al. 2004). Loss of heteroduplex rejection can lead to genomic instability as large deletions and rearrangements arise between incorrectly paired substrates. An increase in recombination between different species may also occur, compromising the interspecies barrier (Young and Ornston 2001).
The second DSBR-related function of Msh2p/3p is non-homologous tail removal during single-strand annealing (SSA), an in cis repair process involving the pairing of homologous regions flanking a DSB (Sugawara et al. 2000). The role of the Msh2p/3p complex appears to be stabilization of the newly paired single strands and recruitment of the Rad1/10 exonuclease to cleave the non-homologous tail of the SSA intermediate, a function which becomes increasingly unnecessary with larger and more stable homologous regions (Sugawara et al. 1997).
MSH1 is essential for mitochondrial function as a factor of mitochondrial genome stability, in marked contrast to MSH2, MSH3, and MSH6, which play analogous roles in nuclear genome maintenance but are not essential for cellular viability. This demonstrates a difference in the importance of MutS homologue-mediated functions between these two subcellular compartments. In addition to its previously described differences from nuclear DNA, mtDNA is also highly recombinogenic, such that genetic linkage is not detectable between markers that are ≥1,000 base pairs apart (Dujon 1981). By contrast, the extremely low error rate of DNA Polγ imposes an accordingly low requirement for mitochondrial mismatch repair (Roberts and Kunkel 1996). We propose a model consistent with available data in which the ATPase-dependent functions of Msh1p include mismatch repair, but showing that the mismatch recognition-dependent functions include both mismatch repair and heteroduplex recognition and rejection. This may explain why the F105A mutation produces a greater disruption of mitochondrial function than the G776D mutation. To test this prediction, the effects of Msh1p on recombination in mtDNA is being addressed.
Physiological effects of Msh1p disruption
In order to investigate the effects of Msh1p disruption on a physiological level, we examined two features that are often associated with mitochondrial disruption: increased sensitivity to oxidative damage and aberrant mitochondrial morphology. Oxidative damage is a major contributor to mitochondrial malfunction and there are many mechanisms that repair this damage. To address the possibility that Msh1p functions in such a pathway, we assessed the sensitivity of cells carrying either wild-type or mutant alleles of MSH1 to H2O2 exposure. As shown in Fig. 6, there is no detectable difference in sensitivity conferred by disruption of Msh1p function. This is consistent with the findings of Meeusen et al. (1999), who showed that, when exposed to UV, gamma radiation, and H2O2, wild-type yeast are most resistant to H2O2. This suggests that the pathways that act in the repair of oxidative damage are very efficient, relative to other types of DNA-damage repair (Meeusen et al. 1999). Both Msh2p/6p and Msh1p have been shown to adopt some activities of the BER pathway under certain circumstances, demonstrating that some mismatch repair proteins are capable of participating in oxidative damage repair (Ni et al. 1999), Dzerzbicki et al. 2004). When BER is active, however, as it is in this study, there is no apparent effect of Msh1p disruption on sensitivity to oxidative damage.
Conversely, we do see a dramatic effect on both mitochondrial morphology and mtDNA organization when Msh1p activity is disrupted. As seen in the top row in Fig. 7, functional mitochondria display a highly networked structure, which in dextrose medium is reduced due to glucose repression of mitochondrial biogenesis (Pon and Schatz 1991). The msh1 mutants maintained in glycerol show both a slight thickening and irregular knobbiness of the mitochondrial tubules. In dextrose, these phenotypes are manifest as a pronounced condensation into either a few irregular tubules, as in msh1-G776D, or one large spot, as shown by both msh1-F105A and msh1-F105A/G776D. Note that these phenotypes are visible with Mitotracker Red staining, indicating that morphological changes occur before the loss of membrane potential that signals a breakdown in mitochondrial function.
MtDNA is organized into nucleoprotein complexes known as nucleoids. Figure 8 shows cells stained with DAPI to reveal the nucleoid locations and with MitoTracker Red to show mitochondrial structure. All mtDNA is located in the same regions as those stained by Mitotracker Red, confirming that the stained structures are mitochondria. Moreover, closer examination shows that wherever mitochondrial staining is visible, there is at least one nucleoid present, suggesting that mitochondrial fission does not occur without each new fragment containing at least some mtDNA.
The time-course of mitochondrial changes shown in Fig. 9 offers additional insights into the intermediate phases of progressive mitochondrial malfunction. Compared with the normal progression of MSH1 cells, the flexibility of both msh1-F105A and msh1-G776D mutants in adopting new conformations specific to the metabolic needs of the cell appears hindered. This becomes readily apparent by about 8 h after release from respiration-restrictive growth. The size and density of nucleoids at this time are also clearly different from the wild type. These trends continue as the cultures approach saturation, resulting in almost total loss of mitochondrial function by 24 h in msh1-F105A. Msh1-G776D fares better, retaining a phenotype closer to that of MSH1.
Many proteins involved in mtDNA stability and transmission are membrane or membrane-associated factors whose disruption results in aberrant mitochondrial morphology (for a review, see Hermann and Shaw 1998). Meeusen and Nunnari (2003) recently demonstrated that a protein complex known as the two-membrane-spanning (TMS) structure serves to anchor mtDNA to the membrane. These anchored nucleoids are proposed to be the exclusive sites of mtDNA replication (Meeusen and Nunnari 2003). MMM1, MDM10, and MDM12 are nuclear genes that encode proteins in the outer membrane complex of the TMS and are thought to connect the membrane/nucleoid structure to a cytosolic actin network that drives transmission of mitochondrial material to daughter cells (Boldogh et al. 2003). Deletion of any of these genes results in an aberrant mitochondrial morphology similar to the msh1 mutants (Boldogh et al. 2003; Kondo-Okamoto et al. 2003; Meeusen and Nunnari 2003). Another class of proteins encoded by the YME genes is also important in maintaining mitochondrial morphology. These were isolated in screens for increased rates of mtDNA transfer to the nucleus, a phenomenon termed yeast mtDNA escape. Deletion of YME1, in addition to mtDNA instability, also produces a condensation of mitochondria into single punctate spots (Campbell and Thorsness 1998). These findings demonstrate that maintenance of mtDNA and maintenance of mitochondrial morphology are linked processes, giving credence to a potential role for Msh1p in the maintenance of mitochondrial structure via its role in the maintenance of mtDNA.
Conclusion
Emerging discoveries about the behavior of mtDNA, in both yeast and humans, suggest that both mismatch repair and recombination surveillance are important for respiration and overall cellular function. However, our data suggest that mismatch repair is not the primary function of the mitochondrial MutS homologue Msh1p. Instead, its ability to maintain mitochondrial genome stability through identifying and preventing ectopic recombination may be of more profound importance. Additionally, we predict that Msh1p fits into the larger picture of mitochondrial organization in conjunction with the nucleoid complex, carrying out recombination surveillance and post-replication mismatch repair within the actively replicating nucleoid.
References
Acharya S, Foster PL, Brooks P, Fishel R (2003) The coordinated functions of the E. coli MutS and MutL proteins in mismatch repair. Mol Cell 12:233–246
Alani EA, Lo C, Kleckner N (1987) A method for gene disruption that allows repeated use of URA3 selection in the construction of multiply disrupted yeast strains. Genetics 116:541–545
Alani E, Sokolsky T, Studamire B, Miret JJ, Lahue RS (1997) Genetic and biochemical analysis of Msh2p-Msh6p: role of ATP hydrolysis and Msh2p-Msh6p subunit interactions in mismatch base pair recognition. Mol Cell Biol 17:2436–2447
Alani EA, Lee JY, Schofield MJ, Kijas AW, Hsieh P, et al (2003) Crystal structure and biochemical analysis of the MutS:ADP:Beryllium fluoride complex suggests a conserved mechanism for ATP interactions in mismatch repair. J Biol Chem 278:16088–16094
Allen DJ, Mahkov A, Grilley M, Taylor J, Thresher R, et al (1997) MutS mediated heteroduplex loop formation by a translocation mechanism. EMBO J 16:4467–4476
Ausubel FM, Brent R, Kingson RE, Moore DD, Seidman JG, et al (1994) Current protocols in molecular biology. Wiley, New York
Bjornson KP, Blackwell LJ, Sage H, Baitinger C, Allen D, et al (2003) Assembly and molecular activities of the MutS tetramer. J Biol Chem 278:34667–34673
Boldogh IR, Nowakowski DW, Yang HC, Chung H, Karmon S, et al (2003) A protein complex containing Mdm10p, Mdm12p, and Mmm1p links mitochondrial membranes and DNA to the cytoskeleton-based segregation machinery. Mol Biol Cell 14:4618–4627
Bowers J, Sokolsky T, Quach T, Alani E (1999) A mutation in the MSH6 Subunit of the Saccharomyces cerevisiae MSH2-MSH6 complex disrupts mismatch recognition. J Biol Chem 274:16115–16125
Campbell CL, Thorsness MK (1998) Escape of mitochondrial DNA to the nucleus in yme1 yeast is mediated by vacuolar-dependent turnover of abnormal mitochondrial compartments. J Cell Sci 111:2455–2464
Chambers SR, Hunter N, Louis EJ, Borts RH (1996) The mismatch repair system reduces meiotic homeologous recombination and stimulates recombination-dependent chromosome loss. Mol Cell Biol 16:6110–6120
Chi N, Kolodner RD (1994a) The effect of DNA mismatches on the ATPase Activity of MSH1, a protein in yeast mitochondria that recognizes DNA mismatches. J Biol Chem 269:29993–29997
Chi N, Kolodner RD (1994b) Purification and characterization of MSH1, a yeast mitochondrial protein that binds to DNA mismatches. J Biol Chem 269:29984–29992
Culligan KM, Meyer-Gauen G, Lyons-Weiler J, Hays JB (2000) Evolutionary origin, diversification and specialization of eukaryotic MutS homolog mismatch repair proteins. Nucleic Acids Res 28:463–471
Datta A, Hendrix M, Lipsitch M, Jinks-Robertson S (1997) Dual roles for DNA sequence identity and the mismatch repair system in the regulation of mitotic crossing over in yeast. Proc Natl Acad Sci USA 94:9757–9762
Drotschmann K, Yang W, Brownewell FE, Kool ET, Kunkel TA (2001) Asymmetric recognition of DNA local distortion: structure-based functional studies of eukaryotic Msh2-Msh6. J Biol Chem 276:46225–46229
Drotschmann K, Yang W, Kunkel TA (2002) Evidence for sequential action of two ATPase active sites in yeast Msh2-Msh6. DNA Repair 1:743–753
Dufner P, Marra G, Raschle M, Jiricny J (2000) Mismatch recognition and DNA-dependent stimulation of the ATPase activity of hMutSalpha is abolished by a single mutation in the hMSH6 subunit. J Biol Chem 275:36550–36555
Dujon B (1981) Mitochondral genetics and functions. In: Strathern JN, Jones EW, Broach JR (eds) Molecular biology of the yeast Saccharomyces: life cycle and inheritance. Cold Spring Harbor Laboratory, Plainview, pp 505–635
Dzerzbicki P, Koprowski P, Fikus MU, Malc E, Ciesla Z (2004) Repair of oxidative damage in mitochondrial DNA of Saccharomyces cerevisiae: involvement of the MSH1-dependent pathway. DNA Repair 3:403–411
Eisen JA (1998) A phylogenomic study of the MutS family of proteins. Nucleic Acids Res 26:4291–4300
Evans E, Alani E (2002) Roles for mismatch repair factors in regulating genetic recombination. Mol Cell Biol 20:7839–7844
Evans E, Sugawara N, Haber JE, Alani E (2000) The Saccharomyces cerevisiae Msh2 mismatch repair protein localizes to recombination intermediates in vivo. Mol Cell 5:789–799
Foury F, Vanderstraeten S (1992) Yeast mitochondrial DNA mutators with deficient proofreading exonuclease activity. EMBO J 11:2717–2726
Gradia S, Acharya S, Fishel R (1997) The human mismatch recognition complex hMSH2-hMSH6 functions as a novel moelcular switch. Cell 91:995–1005
Gradia S, Do S, Wilson T, Acharya S, Mahkov A, et al (1999) hMSH2-hMSH6 forms a hydrolysis-independent sliding clamp on mismatched DNA. Mol Cell 3:255–261
Graeber MB, Muller U (1998) Recent developments in the molecular genetics of mitochondrial disorders. J Neurol Sci 153:251–263
Haber LT, Walker GC (1991) Altering the conserved nucleotide binding motif in the Salmonella typhimurium MutS mismatch repair protein affects both its ATPase and mismatch binding activities. EMBO J 10:2707–2715
Hanekamp T, Thorsness MK, Rebbapragada I, Fisher EM, Seebart C, et al (2002) Maintenance of mitochondrial morphology is linked to maintenance of the mitohcondrial genome in Saccharomyces cerevisiae. Genetics 162:1147–1156
Harfe BD, Jinks-Robertson S (2000) DNA mismatch repair and genetic instability. Annu Rev Genet 34:359–399
Hermann GJ, Shaw JM (1998) Mitochondrial dynamics in yeast. Annu Rev Cell Dev Biol 14:265–303
Hollingsworth NM, Ponte L, Halsey C (1995) MSH5, a novel MutS homolog, facilitates meiotic reciprocal recombination between homologs in Saccharomyces cerevisiae but not mismatch repair. Genes Dev 9:1728–1739
Hu JP, Vanderstraeten S, Foury F (1995) Isolation and characterization of ten mutator alleles of the mitochondrial DNA polymerase-encoding MIP1 gene from Saccharomyces cerevisiae. Gene 160:105–110
Junop MS, Obmolova G, Rausch K, Hsieh P, Yang W (2001) Composite active site of an ABC ATPase: MutS uses ATP to verify mismatch recognition and authorize DNA repair. Mol Cell 7:1–12
Junop MS, Yang W, Funchain P, Clendenin W, Miller JH (2003) In vitro and in vivo studies of MutS, MutL and MutH mutants: correlation of mismatch repair and DNA recombination. DNA Repair 2:387–405
Kijas AW, Studamire B, Alani EA (2003) Msh2 separation of function mutations confer defects in the initiation steps of mismatch repair. J Mol Biol 331:128–138
Kondo-Okamoto N, Shaw JM, Okamoto K (2003) Mmm1p spans both the outer and inner mitochondrial membranes and contains distinct domains for targeting and foci formation. J Biol Chem 278:48997–49005
Koprowski P, Fikus M, Mieckowski P, Ciesla Z (2002) A dominant mitochondrial mutator phenotype of Saccharomyces cerevisiae conferred by msh1 alleles altered in the sequence encoding the ATP-binding domain. Mol Genet Genomics 266:988–994
Lamers MH, Perrakis A, Enzlin JH, Winterwerp HHK, Wind ND, et al (2000) The crystal structure of DNA mismatch repair protein MutS binding to a G.T mismatch. Nature 407:711–717
Lea DE, Coulson CA (1949) The distribution of the number of mutants in bacterial populations. J Genet 49:264–285
MacAlpine DM, Perlman PS, Butow RA (2000) The numbers of individual mitochondrial DNA molecules and mitochondrial DNA nucleoids in yeast are co-regulated by the general amino acid control pathway. EMBO J 19:767–775
Malkov VA, Biswas I, Camerini-Otero RD, Hsieh P (1997) Photocross-linking of the NH2-terminal region of Taq MutS protein to the major groove of a heteroduplex DNA. J Biol Chem 272:23811–23817
Mason P, Matheson EC, Hall AG, Lightowlers RN (2003) Mismatch repair activity in mammaliam mitochondria. Nucleic Acids Res 31:1052–1058
Meeusen S, Nunnari J (2003) Evidence for a two membrane-spanning autonomous mitochondrial DNA replisome. J Cell Biol 163:503–510
Meeusen S, Tieu Q, Wong E, Weiss E, Schieltz D, et al (1999) MGM101p is a novel component of the mitochondrial nucleoid that binds DNA and is required for the repair of oxidatively damaged mitochondrial DNA. J Cell Biol 145:291–304
Modrich P (1991) Mechanisms and biological effects of mismatch repair. Annu Rev Genet 25:229–253
Modrich P, Lahue RS (1996) Mismatch repair in replication fidelity, genetic recombination, and cancer biology. Annu Rev Genet 25:229–253
Myung K, Datta A, Chen C, Kolodner RD (2001) SGS1, the Saccharomyces cerevisiae homologue of BLM and WRN, suppress genome instability and homeologous recombination. Nat Genet 27:113–116
Ni TT, Marschisky GT, Kolodner RD (1999) MSH2 and MSH6 are required for removal of adenine misincorporation opposite 8-oxo-guanine in S. cerevisiae. Mol Cell 4:439–444
Obmolova G, Ban C, Hsieh P, Yang W (2000) Crystal structures of mismatch repair protein MutS and its complex with a substrate DNA. Nature 407:703–710
Phadnis N, Sia EA (2004) Role of the putative structural protein sed1p in mitochondrial genome maintenance. J Mol Biol 342:1115–1129
Pochart P, Woltering D, Hollingsworth NM (1997) Conserved properties between functionally distinct MutS homologs in yeast. J Biol Chem 272:30345–30349
Pon L, Schatz G (1991) Biogenesis of yeast mitochondria. (The molecular and cellular biology of yeast mitochondria: genome dynamics, protein synthesis, and energetics) Cold Spring Harbor Laboratory, Plainview, pp 333–406
Reenan R, Kolodner RK (1992a) Characterization of insertion mutations in the Saccharomyces cerevisiae MSH1 and MSH2 genes: evidence for separate mitochondrial and nuclear functions. Genetics 132:975–985
Reenan RA, Kolodner RD (1992b) Isolation and characterization of two Saccharomyces cerevisiae genes encoding homologs of the bacterial MutS mismatch repair proteins. Genetics 132:963–973
Roberts J, Kunkel T (1996) Fidelity of DNA replication. In: Moll D (ed) DNA replication in eukaryotic cells. Cold Spring Harbor Laboratory, Plainview, pp 217–248
Ross-Macdonald P, Roeder GS (1994) Mutation of a meiosos-specific MutS homolog decreases crossing over but not mismatch correction. Cell 79:1069–1080
Schofield MJ, Hsieh P (2003) DNA mismatch repair: molecular mechanisms and biological function. Annu Rev Microbiol 57:579–608
Schofield MJ, Brownewell FJ, Nayak S, Du C, Kool ET, et al (2001) The Phe-X-Glu DNA binding motif of MutS. J Biol Chem 276:45505–45508
Sherman F (1991) Getting started with yeast. Methods Enzymol 194:3–21
Shoubridge EA (2001) Nuclear genetic defects of oxidative phosphorylation. Hum Mol Genet 10:2277–2284
Sia EA, Kokoska RJ, Dominska M, Greenwell P, Petes TD (1997) Microsatellite instability in yeast: dependence on repeat unit size and DNA mismatch repair genes. Mol Cell Biol 17:2851–2858
Sia EA, Butler CA, Dominska M, Greenwell P, Fox TD, et al (2000) Analysis of microsatellite mutations in the mitochondrial DNA of Saccharomyces cerevisiae. Proc Natl Acad Sci USA 97:250–255
Sikorski RS, Hieter P (1989) A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122:19–27
Studamire B, Quach T, Alani E (1998) Saccharomyces cerevisiae Msh2p and Msh6p ATPase activities are both required during mismatch repair. Mol Cell Biol 18:7590–7601
Studamire B, Price G, Sugawara N, Haber JE, Alani EA (1999) Separation-of-function mutations in Saccharomyces cerevisiae that confer miamatch repair defects but do not affect non-homologous tail removal during recombination. Mol Cell Biol 19:7558–7567
Su S, Modrich P (1986) Escherichia coli mutS-encoded protein binds to mismatched DNA base pairs. Proc Natl Acad Sci USA 83:5057–5061
Sugawara N, Paques F, Colaiacovo M, Haber JM (1997) Role of Saccharomyces cerevisiae Msh2 and Msh3 repair proteins in double-strand break-induced recombination. Proc Natl Acad Sci USA 94:9214–9219
Sugawara N, Ira G, Haber JE (2000) DNA length dependence of the single-strand annealing pathway and the role of Saccharomyces cerevisiae RAD59 in double-strand break repair. Mol Cell Biol 20:5300–5309
Sugawara N, Goldfarb T, Studamire B, Alani EA, Haber JE (2004) Roles for MSH mismatch repair proteins and SGS1 in heteroduplex rejection during single strand annealing between mismatched DNA sequences (in press)
Vanderstraeten S, Van der Brule S, Hu JP, Foury F (1998) The role of 3′-5′ exonucleolytic proofreading and mismatch repair in yeast mitochondrial DNA error avoidance. J Biol Chem 273:23690–23697
Wu TH, Marinus MG (1994) Dominant negative mutations in the mutS gene of Escherichia coli. J Bacteriol 176:5393–5400
Yamamoto A, Schofield MJ, Biswas I, Hsieh P (2000) Requirement for Phe36 for DNA binding and mismatch repair by Escherichia coli MutS protein. Nucleic Acids Res 28:3564–3569
Young DM, Ornston LN (2001) Functions of the mismatch repair gene mutS from Acinobacter sp. strain ADP1. J Bacteriol 183:6822–6831
Acknowledgements
This work was supported by the National Institutes of Health grant GM63626-01. E.A.S. is the recipient of a Burroughs–Wellcome Fund Career Award. Purchase of the Leica TCS SP spectral confocal microscope was supported by shared instrumentation awards from the National Science Foundation (9512886) and the National Institute of Health (S10RR11358) and by matching funds from the University of Rochester. We would like to thank Dr. Rita Miller for use of the Zeiss Axioplan 2 microscope. We would also like to thank Leah Jablonski for construction of the msh1-G776D mutation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Brunner
Rights and permissions
About this article
Cite this article
Mookerjee, S.A., Lyon, H.D. & Sia, E.A. Analysis of the functional domains of the mismatch repair homologue Msh1p and its role in mitochondrial genome maintenance. Curr Genet 47, 84–99 (2005). https://doi.org/10.1007/s00294-004-0537-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-004-0537-1