Abstract
Phage display is one of the important and effective molecular biology techniques and has remained indispensable for research community since its discovery in the year 1985. As a large number of nucleotide fragments may be cloned into the phage genome, a phage library may harbour millions or sometimes billions of unique and distinctive displayed peptide ligands. The ligand–receptor interactions forming the basis of phage display have been well utilized in epitope mapping and antigen presentation on the surface of bacteriophages for screening novel vaccine candidates by using affinity selection-based strategy called biopanning. This versatile technique has been modified tremendously over last three decades, leading to generation of different platforms for combinatorial peptide display. The translation of new diagnostic tools thus developed has been used in situations arising due to pathogenic microbes, including bacteria and deadly viruses, such as Zika, Ebola, Hendra, Nipah, Hanta, MERS and SARS. In the current situation of pandemic of Coronavirus disease (COVID-19), a search for neutralizing antibodies is motivating the researchers to find therapeutic candidates against novel SARS-CoV-2. As phage display is an important technique for antibody selection, this review presents a concise summary of the very recent applications of phage display technique with a special reference to progress in diagnostics and therapeutics for coronavirus diseases. Hopefully, this technique can complement studies on host–pathogen interactions and assist novel strategies of drug discovery for coronaviruses.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction of Phage Display Technique
Phage display technique is an in vitro method used to select a specific fusion peptide displayed on the surface of bacteriophages [1]. Peptide phage libraries containing billions of variants are used to interact with target molecules and the most specifically binding peptide is enriched after 3–5 rounds of biopanning, i.e. affinity selection of specific peptide from a large repertoire of clones. Phage display technique can be used for targeting different biological entities, such as proteins, viruses, bacteria, cancer cells and non-organic substances, like metals, alloys, semiconductors, toxins and plastic surfaces [2,3,4,5]. The detailed information about filamentous phage genomes has been the key to success for widespread applications of phage display technology. The technique has many applications, such as selection of monoclonal antibody and bioactive peptides, epitope mapping, selection of disease specific antigen mimics, cell-specific or organ-specific peptides, peptidomimetics and development of drug delivery systems [6,7,8,9]. Thus, phage display has been widely used in immunotherapy, regenerative medicine, development of biosensors and synthetic biology [5, 10,11,12,13].
Phage Display Systems
In phage display system, exogenous DNA fragment is inserted into a specific site (phage coat protein encoding gene) in phage genome. Upon host infection by these phage particles, expression of the insert encoding amino acids in the form of fusion peptide takes place displaying them on the exposed surface which is further able to interact with wide variety of external target molecules and this phenotype–genotype association helps researchers isolate target-specific ligands displayed on the phage surface [14, 15]. There are different phage display systems available but the F pilus-specific phage (Ff), encompassing the strains f1, M13 or fd, and T7 lytic bacteriophages have been most extensively used [16, 17].
The M13 filamentous phage particle exhibits a cylindrical shape, about 930 nm in length and 6 nm in diameter (Fig. 1). The M13 genome is single-stranded DNA of 6407 bp consisting of nine genes encoding 11 different proteins [18]. Five of the genome-encoded proteins are coat proteins and the rest of the six proteins are involved in replication and assembly. Although most of the studies have been restricted to the fusion of exogenous DNA fragment with capsid proteins pIII and pVIII, where pVIII is the major coat protein expressed in ~ 2700 copies forming the cylindrical side of the phage and pIII is a minor coat protein (present in 5 copies) at an end of the filamentous phage particle and is responsible for adsorption and extrusion. But in some studies phage display systems involving pVI, pVII and pIX (tail virion proteins involved in coat protein assembly and budding) fusions have also been reported [19]. The protein of interest (POI) fused with phage coat protein may be encoded either in a phage genome or by a phagemid. When a phagemid is used, complementation by a helper phage is required to support virion production and using phagemid is advantageous during subsequent selection process. M13 phage display libraries present diversity in the range of 105 to 1012 and are greatly dependent upon the transformation efficiency of the host E. coli. The M13 phage display system is a suitable tool for display of the appropriately folded proteins carrying disulphide bonds as the proteins are secreted through the periplasmic space. Many functional antibody fragments, enzymes and inhibitors have been displayed and selected using this system [20, 21]. However, it also has the minor limitation of poor display of cytoplasmic proteins on the membrane [22].
On the other hand using T7—a lytic phage originally described as a mutant phage of E. coli––has several advantageous features over other phage display systems [23,24,25,26,27] including display of cytoplasmic proteins—which is a major limitation of the M13 filamentous phage display system. T7 bacteriophage contains linear double-stranded DNA of 39,937 bp packed in an icosahedral capsid [27] and encodes ~ 50 proteins [28] including six major proteins––gp10A, gp10B, gp8, gp11, gp12 and gp17 encoding major capsid protein, minor capsid protein, connector protein, tail proteins and tail fibre proteins, respectively. Fusion proteins are displayed at the C-terminal end of the T7 capsid protein (gene 10)—which includes gp10A (344 amino acids) and gp10B (397 amino acids), thus displaying fusion protein in high number (i.e. 415 copies) on entire capsid shell [17, 27]. The library efficiency depends upon overall efficiency of cloning and packaging. Other advantages include the following: displaying inserts with stop codon, fast growth to form plaques within 3 h and relatively easier construction of large display libraries compared with M13 and the reduced likelihood of survival of the recombinant phage in the environment as the host cell is required to produce T7 gene 10 to express proteins. Also for promoting proper folding of cytoplasmic proteins carrying bisulphide bonds, complementing hosts, such as BLT5615 or BLT5403 E. coli strains can be used [29,30,31]. Furthermore, the purification process of T7 phage for ELISA and DNA sequencing is also simple to perform.
Among other phage display systems, the phage T4 HOC (highly antigenic outer capsid protein of molecular mass 40 kDa) and SOC (small outer capsid protein of molecular mass 9 kDa), bipartite display system for larger proteins to be displayed in high copy number and insert with stop codon is also available [32, 33]. The temperate phage lambda having double-stranded DNA genome size of 48,502 bp is also capable of displaying fusion proteins in association with gpV (tail protein) and gpD (head protein) [34]. It is capable of displaying complicated, high-molecular-mass proteins as fusions with N- or C-terminal of gpD or C-terminal of gpV which are present in 405 and 6 copies, respectively [35, 36]. As translocation of proteins through the host cell membrane is not required using lambda system, it offers a higher immune response against displayed proteins.
Strategies of Phage Display for Development of Diagnostics
Phage display library may be constructed using any of the above-mentioned phage display systems and screened to obtain the most promising binders exhibiting strongest affinity for the target molecule/pathogen for use in diagnostics and immunotherapy. Either randomly generated fragments of targeted pathogen DNA or synthetic random degenerate oligonucleotide inserts are cloned into the phage genome leading to generation of Natural peptide library (NPL) or Random peptide library (RPL), respectively. NPLs are more likely to mount an antibody response that cross-reacts with the native intact pathogen but a vast majority of clones in these libraries are non-functional, whereas RPLs have extended range of displayed epitopes [37, 38]. When the engineered nucleotide fragments of immunoglobulins encoding the antigen-binding region are displayed, it leads to generation of antibody phage display (APD) library. Antibody expression over the phage surface is a powerful platform that allows the selection of specific antibodies when either immunized patients are not available or immunization is not ethically feasible. Due to the vast applications of this important technique, Nobel Prize in Chemistry for the year 2018 was awarded to George P. Smith and Sir Gregory P. Winter for developing and applying the phage display technique for the discovery and isolation of antibodies [1, 39, 40]. Major advantages of antibody phage display include as follows: production of large-sized combinatorial libraries, improved efficiency than conventional hybridoma system, easy manipulation, safety, no involvement of animal cell culture, lower propagative cost and easy screening procedures involved to select target specific binders [41,42,43]. Specifically for viral pathogens, antibody phage display is an alternative tool to hybridoma technology, since it circumvents the limitations of the immune system [44].
The antibody phage display technique was first described using gene III protein of E. coli filamentous phages and was further modified. The display formats include single-chain fragment variable (scFv), antigen-binding fragment (Fabs), single-chain fabs (scFab), human VH domains (dAbs), variable domains of heavy chain (VHHs)—also called as nanobodies, immunoglobulins of sharks (IgNARs), variable domain of IgNAR (VNAR), etc. [45,46,47,48,49,50] (Fig. 2). A review of different types of combinatorial peptide phage libraries and peptide ligands and applications in terms of drug discovery is sufficiently presented elsewhere [51, 52]. These phage display formats have been used extensively for antibody selection against pathogenic bacteria, viruses (including SARS-CoV, Ebola, Hanta virus, Hendra and Nipah virus, Hepatitis A, C, E virus, Herpes Simplex virus, HIV-1, 2, and Influenza A), eukaryotic pathogens and toxins [53]. Additionally very recently researchers demonstrated that incorporating non-canonical amino acids (ncAAs) in phage display expands the range of peptides it can identify [54].
Different antibody formats used for phage display. Conventional IgG structurally comprising two heavy (H) chains (composed of VH and CH domains) and two light (L) chains (composed of VL and CL domains) and carrying two antigen-binding sites constituted by VH and VL domains. Heavy-chain antibodies of family Camelidae lack both constant and variable light chains (CL and VL) and the antigen-binding site is formed only by the heavy-chain variable domain (VHH/nanobody). The new antigen receptor (IgNAR) antibodies from sharks are constituted of two protein chains, each containing one variable (VNAR) and five constant domains (CH). The phage antibody display format may include as follows: Antigen binding fragment (Fab) of conventional immunoglobulins consisting of VH, CH1, VL and CL domains where CH1 and CL domains are joined by a disulphide bond, Single-chain fragment antigen binding (scFab) consisting of VH, CH1, VL and CL domains where CH1 and VL are joined via linker, fragment variable (FV) consisting of variable regions of heavy-chain (VH) and light-chain (VL), single-chain fragment variable (scFv) consisting of variable regions, i.e. VH and VL joined through a linker, variable domain of human antibodies (VH) or VHH
A Brief Introduction to Coronaviruses
Coronaviruses attracted worldwide attention in year 2003 when Severe Acute Respiratory Syndrome coronavirus (SARS-CoV) epidemic occurred followed by the 2012 Middle East Respiratory Syndrome coronavirus (MERS-CoV) outbreak and, most recently, the novel coronavirus pandemic in year 2019–2020. A total of 8098 people worldwide became sick with SARS outbreak, leading to death in 774 cases [55]. MERS-CoV was identified in Saudi Arabia and it caused human infections through direct or indirect contact with infected dromedary camels who acquired it from bats. WHO reported a high mortality rate of 35% for this infection out of 2519 global cases [56]. SARS-CoV2 was reported from Wuhan province in China in December, 2019, which has till January 05, 2021 caused 84,474,195 confirmed human cases, including 18,48,704 deaths [57]. Coronaviruses are pathogenic viruses of humans and animals belonging to family Coronaviridae, subfamily Coronavirinae. Coronaviruses generally cause species-specific illness in mammals and birds [58]. Coronaviruses are enveloped viruses. They carry non-segmented positive-sense RNA genome and have the largest identified viral RNA genomes, with an approximate length of 30 kilobases; however, the recent menace—SARS-CoV-2—is 18 kb RNA in genome size. Among the four known types, i.e. Alphacoronavirus, Betacoronavirus, Gammacoronavirus and Deltacoronavirus, Alphacoronavirus, group 1 (HCoV-229E), was first reported during studies on common cold [59, 60]. The human coronaviruses known till date include HCoV-NL63 (Alpha coronavirus: group1); HCoV-OC43 and HCoV-HKU1 (Beta coronavirus: group2, lineage A); SARS-CoV and SARS-CoV-2 (Beta coronavirus: group2, lineage B) (Fig. 3); and MERS-CoV (Beta coronavirus: group2, lineage C) [61]. The coronaviruses are primarily viewed as respiratory pathogens with only alpha and beta types infecting mammals and usually causing respiratory illness in humans and gastroenteritis in animals. The deltacoronaviruses and gammacoronaviruses majorly infect birds, but some can also infect mammals. The typical symptom involving upper respiratory tract infection includes common cold. On the other hand, SARS-CoV and MERS-CoV are highly pathogenic to humans, cause severe pneumonia and pose fatal outcomes [58]. Since the initiation of pandemic in Wuhan, China, the major thrust of the researchers has been on epidemiology and pharmacological investigations including vaccines and drug moieties to treat and prevent SARS-CoV-2.
Antibody Selection against Coronaviruses Using Phage Display Technology
Phage display technique has been employed as a complementary approach in previous research attempts for development of diagnostics against coronaviruses. It has been used for studying host–pathogen interactions, selection of neutralizing antibodies and epitope mapping for SARS-CoV-2.
Selection of Neutralizing Antibodies against SARS-CoV-2 (COVID-19)
Based on the sequence similarities of the receptor-binding motif (RBM) between SARS-CoV-2 and SARS-CoV, research groups reported that SARS-CoV-2 could enter the cells by binding of surface spike (S) protein with the Angiotensin converting enzyme-2 (ACE-2) receptor in humans, bats, civet cats and pigs [62, 63]. ACE-2 encodes for a type I transmembrane metallocarboxypeptidase which exhibits homology to ACE, an enzyme long known as a target for the treatment of hypertension [64]. It is highly expressed in tissues of cardiovascular, renal and gastrointestinal system, lungs, testis and prostate and moderately expressed in central nervous system and lymphoid tissues [65,66,67,68]. Host cell proteases, such as endosomal cathepsins, trypsin, elastase, furin and transmembrane protease serine 2 (TMPRSS2) are known to play an important role for SARS-CoV-2 entry as both S protein and ACE-2 are proteolytically modified during the process [69].
In order to screen neutralizing antibodies against currently circulating highly pathogenic SARS-CoV-2, Tian et al. [70] reported potent binding of COVID-19 causing coronavirus S protein by anti-SARS-CoV human antibody (CR3022) [71]. CR3022 has been derived from a scFv phage display library and has shown to bind to the RBD outside the RBM [72]. Although it is known that the blockage of RBM within the RBD and the ACE-2 association site, is a major mechanism of SARS-CoV neutralization but as CR3022—actually raised against SARS-CoV bound potently with SARS-CoV-2 RBD, it was emphasized that neutralizing antibodies can also be raised from epitopes outside of RBM. This offers the possibility of partial protection against COVID-19 [73].
As per the document available from Milken Institute, two pharmaceutical firms, viz., PhRMA and AstraZeneca, are into pre-clinical trials for antibodies targeting the S protein from phage display libraries [74]. Also researchers have been successful in developing new phage capsid nanoparticles which could prevent COVID-19 infections [75]. It is based on the approach of developing highly functional multivalent binders which match with the geometry of binding sites of the viral spike proteins which eventually prevent pathogen entry [76]. Walter et al. [77] have also reported selection of 63 unique anti-RBD sybodies (synthetic nanobodies) against SARS-CoV-2 from three large combinatorial libraries produced by phage display technique [77]. Above-mentioned studies underscore the importance of antibody phage display for assisting in searching a probable solution to SARS-CoV-2.
Antibody Selection against Other Coronaviruses
(i) SARS-CoV
In the first attempt to obtain scFv antibodies against SARS-CoV, three scFvs against the E and N proteins of SARS-CoV were isolated by phage display from an scFv antibody library by biopanning against immobilized purified envelope (E) and nucleocapsid (N) proteins [78]. However, majority of later works were focused on spike protein. Using whole irradiated SARS-CoV as target, human monoclonal antibodies (MABs) were selected out from semisynthetic antibody phage display libraries. A total of six human MABs out of eight selected bound to virus and infected cells and these six MABs could be mapped to the nucleocapsid (N) and S proteins of SARS-CoV. Later using epitope mapping, the importance of residue N479 was defined in binding of the most potent neutralizing MAB—CR3014 using recombinant S fragments (residues 318 to 510) containing naturally occurring mutations [79] (Fig. 4). In a similar attempt to produce human SARS-CoV neutralizing antibody, an immune antibody phage display library was constructed from B cells of SARS convalescent patients and scFv, B1-recognizing SARS-CoV pseudovirus in vivo was obtained and its epitope mapping revealed amino acids of 1023–1189 position constituting the epitope [80]. Identification of a dominant epitope in the S2 domain of spike protein of SARS was also carried out using M13 phage display dodecapeptide library [81]. For neutralizing MAB-2C5 (specific to the spike protein), identification of TPEQQFT-mimotope was carried out using epitope mapping [82]. In another study employing antibody phage display library from the B cells of convalescent SARS patients and screening using inactivated SARS-CoV as antigen, a high-affinity binder scFv-H12 was found to neutralize SARS virions in vitro. The fact that it did not bind to monomeric S1 protein suggested its recognition of oligomeric S1 proteins [83]. Key amino acid residues responsible for antibody binding were also identified through epitope mapping using yeast surface display and RBD of SARS-CoV spike (S) protein [84]. Chicken scFv phage display system was employed for selection of high-affinity antibodies against the SARS-CoV spike protein [85]. The discontinuous epitope of 80R—a potent neutralizing human anti-SARS monoclonal antibody against the spike protein––was identified by screening phage display random peptide libraries with the help of unique computer algorithm ‘Mapitope’ [86]. The selection of SARS-CoV-S protein-specific monoclonal antibodies was also reported by Wu et al. [87] by first constructing a SARS-CoV-S protein-specific phage displayed antigen library and biopanning with two anti-S MABs: S-M1 and S-M2 [87].
An illustration of phage display technique for epitope characterization of monoclonal antibodies binding to spike protein of MERS-CoV using phage display technique. The whole RNA can be extracted and cDNA amplified. Spike protein encoding gene can be amplified by PCR. The gene segments can be obtained by fragmentation using restriction enzymes and segments cloned in phagemid vector. Transformation of phagemids in E. coli and infection with helper phage can lead to production of phages displaying protein encoded by cloned genomic segment. The phage library thus formed can be screened using Anti-S-mABs by ELISA and phages binding specifically can be selected by repeated rounds of biopanning (elution, amplification, binding and washing). DNA sequencing of bound phages leads to epitope mapping
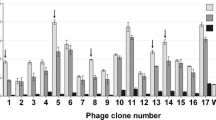
Wang et al. [88] obtained phages with specific binding to SARS-CoV using immobilized RBD target of SARS-CoV spike (S1) protein [88]. The epitope mapping led to identification of displayed peptides HHKTWHPPVMHL and SQWHPRSASYPM, and the two selected phages from 12-mer phage display random peptide library were found to be able to differentiate between SARS-CoV and other coronaviruses using indirect enzyme-linked immunosorbent assays. In another study employing Conformer Libraries’ (phage-displayed RBM linker libraries), Freund et al. [89] reconstituted a functional RBM of SARS coronavirus [89]. Screening the ‘Conformer Libraries’ with specific ligands produced short RBM constructs (ca. 40 amino acids) that could bind both the ACE-2 receptor and the neutralizing MAB 80R (which targeted the RBM and competed with the ACE-2 receptor for binding). Huang et al. [90] identified SARS-CoV ORF6-interacting proteins, including CCNL1, RBMXL2, NPIPB3 and karyopherin alpha 2 which were involved in RNA splicing, nuclear pore complex formation, nuclear export and import of some transcription and splicing factors using the phage-displayed human lung cDNA libraries and examined the association of ORF6-host factor interaction with Type I IFN antagonism [90].
(ii) MERS-CoV
Construction of a phage display library of specific nano-antibodies against MERS-CoV and its application to the screening neutralizing nano-antibodies was reported by He et al. [91]. MERS-CoV RBD recombinant protein was used to immunize alpaca and peripheral blood mononuclear cells (PBMCs) were collected followed by total RNA extraction. The recombinant phages were constructed by introducing VHH gene. A rich phage display library of specific nano-antibodies against MERS-CoV was obtained and used to screen and characterize the nano-antibodies which resulted in identifying neutralizing nano-antibodies against MERS-CoV. Using a Fab phage display library against the S2 subunit of the MERS-CoV spike protein (MERS-S2P), three unique Fabs (S2A3, S2A6 and S2D5) were obtained, two of which (S2A3 and S2D5) bound specifically with MERS-CoV and not other CoVs, indicating them being suitable reagent for developing antibody-based diagnostics in laboratory or hospital settings for point-of-care testing [92]. Construction of nurse shark VNAR phage library and isolation of VNAR binders to viral antigens, including MERS and SARS spike proteins, were recently reported by Feng et al. [93]. A large phage-displayed VNAR library was constructed using VNAR from shark B cells after PCR amplification, cloning in phagemid vectors followed by four rounds of panning, resulting in Type I and II VNARs produced successfully in E. coli. Using next-generation sequencing (NGS) by Illumina MiSeq platform, approximately 1.2 million full-length shark VNAR sequences were found to be displayed in the phage-displayed VNAR antibody library constructed from six naive adult nurse sharks (Ginglymostoma cirratum). This work validated the large diversity of the nurse shark VNAR library produced by Phage display technique as a platform for therapeutic antibody discovery.
(iii) Animal Coronaviruses
A random phage display peptide library was biopanned using purified avian infectious bronchitis virus (IBV) to screen the specific binding phages and a linear peptide with high neutralization titre was identified, which was found to inhibit IBV infection in HeLa cells as well [94]. A similar strategy was also used to identify two linear B-cell epitopes that were recognized by the MABs 6D10 and 4F10 (against the N protein of IBV), which corresponded to the aa sequences (242) FGPRTK (247) and (195) DLIARAAKI (203), respectively, in the IBV-N protein. Alignments of amino acid sequences from a large number of IBV isolates later indicated that the two epitopes, especially (242) FGPRTK (247), were well conserved among IBV coronaviruses causing avian infectious bronchitis [95]. For epitope mapping using a phage display peptide library (12-mer, M13 gene 3-based random peptide phage) for biopanning against spike protein of IBV Sczy3 strain, two linear B-cell epitopes were identified by MABs—1D5 and 6A12, which referred to the amino acid sequences (87)PPQGMAW(93) and (412)IQTRTEP(418), respectively, in the IBV S1 subunit. Further comparisons of sequence revealed that epitope (412) IQTRTEP (418) was conserved among IBVs, while the epitope (87) PPQGMAW (93) was relatively variable among IBVs [96].
Applying epitope mapping against M-protein of porcine transmissible gastroenteritis virus (TGEV) led to development of a phage ELISA showing improved sensitivity and the inhibitory effect of the identified peptide was also confirmed [97]. By using rS-AD (recombinant antigenic site A + D) of TGEV as an immobilized target to identify peptides from a phage display library available commercially, nine phages were selected that specifically bound to it with high affinity and the peptide TLNMHLFPFHTG carrying phage bound with the highest affinity. It was subsequently used to develop a phage-based ELISA for TGEV [98]. To identify antigen epitopes with specificity for porcine epidemic diarrhea virus (PEDV)—causing severe diarrhea and dehydration resulting in substantial morbidity and mortality in newborn piglets, a MAB-5E12 against the immunodominant region of the PEDV spike protein (S1) was used as a target for biopanning a 12-mer phage display, random peptide library, and phage-displayed peptides (3 nos.), viz., L, W and H, were identified that recognized MAB-5E12. The consensus epitope (peptide M) was identified and antiviral properties were established by blocking viral protein expression [99]. Zhang et al. [100] also reported the potential role of scFvs targeted against viral spike protein to inhibit PEDV infectivity by the plaque reduction neutralization assay and in the prevention and treatment of PEDV infection by oral administration [100]. Recently a rapid, low-cost, reliable blocking ELISA (bELISA) was developed using phage display technology employing VHH library and biopanning of nanobodies against PEDV N protein [70]. The sensitivity and specificity of bELISA thus developed were observed to be 100% and 93.18%, respectively. Sun et al. [101] reported construction and evaluation of a murine scFv library against the common receptor porcine aminopeptidase N (pAPN) of porcine coronaviruses TGEV and PEDV using the T7 phage display system [101]. Using recombinant rpAPN-C subunit and applying epitope mapping lead to identification of a peptide sequence with a potential role as a small molecular therapeutic agent against TGEV infection [102]. Identification of Neural Cell Adhesion Molecule (NCAM) as a novel interacting partner of the porcine hemagglutinating encephalomyelitis virus (PHE-CoV) using its S protein for biopanning a T7 phage display cDNA library was performed [103].
Conclusions and Future Perspectives
The phage display technique due to simplicity, high efficiency, in vitro nature, rapidity and low-cost is a powerful tool for selecting target-specific ligands. Efficient selection by biopanning leads to the isolation of ligands with unique, specific and desirable functional characteristics. Its robustness has been amply justified with its use in synthetic biology, immunotherapeutics, diagnostics, development of bioassays and biosensors, epitope mapping, mimotopes and drug discovery. As evidenced in past, phage antibody display has been a subject of many patents in therapeutics and diagnostics. It has remained an important choice for monoclonal antibody selection, diagnosis and treatment of viral infections. Phage antibody display offers a variety of platforms for exploiting antigen binding; however, diversity and stability of displayed library are a concern. Further advancements, such as addition of non-canonical aminoacids, leading to expansion of ligands which can be identified using a library, will help in broadening the possible applications and potentials of phage display technology. The new landmarks may require a close association with nanosciences and chemical engineering to design well-curated semi-synthetic libraries for incorporating more structurally diverse displayed epitopes.
The widespread use of phage display technology to assist in exploration of novel targets through epitope mapping has helped in understanding interactions of coronaviruses with human cell receptors and other molecules. The pandemic of SARS-CoV-2 has recently taken many lives, and although symptomatic treatment is being made available to infected persons, yet there have been no specific options for prevention of nCoV infection. Phage display has offered a deeper insight into protein interactions involved in pathogenesis of SARS-CoV-2. Now it is known that partially neutralizing antibodies derived from scFv library can be obtained outside RBD of spike protein as well. Among the exciting contributions are the pre-clinical trials of two phage display library-derived antibody candidates against S protein of SARS-CoV-2. In order to develop a greater understanding of the infectious biology, pathogenesis and blocking mechanisms of currently circulating, highly pathogenic SARS-CoV-2, phage display may certainly play an effective role by offering a repertoire of blocking peptides and neutralizing antibodies.
Phage display technique has propelled search of potential binders for other human coronaviruses, such as SARS, MERS and animal coronaviruses (IBV, TGEV and PEDV). High-affinity antibodies/small molecular therapeutic agents were identified for viral proteins (E, N and S) of SARS-CoV. Development of phage ELISA for TGEV and point-of-care diagnostics for MERS-CoV using neutralizing antibodies indicates that phage display is indeed a reliable technique. With the escalating advancements in the area of phage display and due to incremental use in multiple applications, it is envisaged that this technology will be improved to overcome its limitations. This progress will foster innovations in an incremental manner to develop effective ways to comprehend the mechanisms involved in infection, cellular interactions, and development of post-exposure therapy for SARS-CoV-2 and others in near future.
References
Smith GP (1985) Filamentous fusion phage: novel expression vectors that display cloned antigens on the virion surface. Science 228:1315–1317. https://doi.org/10.1126/science.4001944
Tan WS, Ho KL (2014) Phage display creates innovative applications to combat Hepatitis B virus. World J Gastroenterol 20:11650–11670. https://doi.org/10.3748/wjg.v20.i33.11650
Bernedo-Navarro RA, Yano T (2016) Phagedisplay and shiga toxin neutralizers. Toxicon 113:60–69. https://doi.org/10.1016/j.toxicon.2016.02.009
Vodnik M, Zager U, Strukelj B, Lunder M (2011) Phage display: selecting straws instead of a needle from a haystack. Molecules 16:790–817. https://doi.org/10.3390/molecules16010790
Wang Y, Gao S, Lv J, Lin Y, Zhou L, Han L (2019) Phage display technology and its applications in cancer immunotherapy. Anticancer Agents Med Chem 19:229–235. https://doi.org/10.2174/1871520618666181029140814
Wu CH, Liu IJ, Lu RM, Wu HC (2016) Advancement and applications of peptide phage display technology in biomedical science. J Biomed Sci 23:8. https://doi.org/10.1186/s12929-016-0223-x
Karimi M, Mirshekari H, Moosavi Basri SM, Bahrami S, Moghoofei M, Hamblin MR (2016) Bacteriophages and phage-inspired nanocarriers for targeted delivery of therapeutic cargos. Adv Drug Deliv Rev 106:45–62. https://doi.org/10.1016/j.addr.2016.03.003
Schiavone M, Fiume G, Caivano A, Laurentiis A, Falcone C, Masci FF et al (2012) Design and characterization of a peptide mimotope of the HIV-1 gp120 bridging sheet. Int J Mol Sci 13:5674–5699. https://doi.org/10.3390/ijms13055674
Lupia A, Mimmi S, Iaccino E, Maisano D, Moraca F, Talarico C et al (2020) Molecular modelling of epitopes recognized by neoplastic B lymphocytes in chronic lymphocytic leukemia. Eur J Med Chem 185:111838. https://doi.org/10.1016/j.ejmech.2019.111838
Hess KL, Jewell CM (2019) Phage display as a tool for vaccine and immunotherapy development. Bioeng Transl Med 5:e10142. https://doi.org/10.1002/btm2.10142
Martins IM, Reis RL, Azevedo HS (2016) Phage display technology in biomaterials engineering: progress and opportunities for applications in regenerative medicine. ACS Chem Biol 11:2962–2980. https://doi.org/10.1021/acschembio.5b00717
Park M (2020) Surface display technology for biosensor applications: a review. Sensors 20:2775. https://doi.org/10.3390/s20102775
Lemire S, Yehl KM, Lu TK (2018) Phage-based applications in synthetic biology. Annu Rev Virol 5(1):453–476. https://doi.org/10.1146/annurev-virology-092917-043544
Parmley SF, Smith GP (1988) Antibody-selectable filamentous fd phage vectors: affinity purification of target genes. Gene 73:305–318. https://doi.org/10.1016/0378-1119(88)90495-7
Scott JK, Smith GP (1990) Searching for peptide ligands with an epitope library. Science 249:386–390. https://doi.org/10.1126/science.1696028
Shim H (2017) Antibody phage display. Adv Exp Med Biol 1053:21–34. https://doi.org/10.1007/978-3-319-72077-7_2
Deng X, Wang L, You X, Dai P, Zeng Y (2018) Advances in the T7 phage display system. Mol Med Rep 17:714–720. https://doi.org/10.3892/mmr.2017.7994
Van Wezenbeek PM, Hulsebos TJ, Schoenmakers JG (1980) Nucleotide sequence of the filamentous bacteriophage M13 DNA genome: comparison with phage fd. Gene 11:129–148. https://doi.org/10.1016/0378-1119(80)90093-1
Løset GA, Sandlie I (2012) Next generation phage display by use of pVII and pIX as display scaffolds. Methods 58:40–46. https://doi.org/10.1016/j.ymeth.2012.07.005
Love KR, Swoboda JG, Noren CJ, Walker S (2006) Enabling glycosyltransferase evolution: a facile substrate-attachment strategy for phage-display enzyme evolution. ChemBioChem 7:753–756. https://doi.org/10.1002/cbic.200600018
Zani ML, Moreau T (2010) Phage display as a powerful tool to engineer protease inhibitors. Biochimie 92:1689–1704. https://doi.org/10.1016/j.biochi.2010.05.003
Takakusagi Y, Ohta K, Kuramochi K, Morohashi K, Kobayashi S, Sakaguchi K et al (2005) Synthesis of a biotinylated camptothecin derivative and determination of the binding sequence by T7 phage display technology. Bioorg Med Chem Lett 15:4846–4849. https://doi.org/10.1016/j.bmcl.2005.07.018
Demerec M, Fano U (1945) Bacteriophage-resistant mutants in Escherichia coli. Genetics 30:119–136
Jespers LS, De Keyser A, Stanssens PE (1996) LambdaZLG6: a phage lambda vector for high-efficiency cloning and surface expression of cDNA libraries on filamentous phage. Gene 173:179–181. https://doi.org/10.1016/0378-1119(96)00217-x
Rosenberg GA, Griffin K, Studier FW, McCormick LM, Berg J, Mierendorf R (1998) T7 select phage display system: a powerful new protein display system based on bacteriophage T7. ScienceOpen. https://www.scienceopen.com/document?vid=d05ae7ab-6b30-48a8-884a-6e451cedbb45. Accessed 1 June 2020
Castagnoli L, Zucconi A, Quondam M, Rossi M, Vaccaro P, Panni S et al (2001) Alternative bacteriophage display systems. Comb Chem High Throughput Screen 4:121–133. https://doi.org/10.2174/1386207013331174
Studier FW (1972) Bacteriophage T7. Science 176:367–376. https://doi.org/10.1126/science.176.4033.367
Studier FW, Dunn JJ (1983) Organization and expression of bacteriophage T7 DNA. Cold Spring Harb Symp Quant Biol 47(Pt 2):999–1007. https://doi.org/10.1101/sqb.1983.047.01.114
Krumpe LRH, Atkinson AJ, Smythers GW, Kandel A, Schumacher KM, McMahon JB et al (2006) T7 lytic phage-displayed peptide libraries exhibit less sequence bias than M13 filamentous phage-displayed peptide libraries. Proteomics 6:4210–4222. https://doi.org/10.1002/pmic.200500606
Du XJ, Wu YN, Zhang WW, Dong F, Wang S (2010) Construction and quality examination of murine naive T7 phage display antibody library. Food Agric Immunol 21:81–90. https://doi.org/10.1080/09540100903414106
Ohtani M, Hikima JI, Jung TS, Kondo H, Hirono I, Aoki T (2013) Construction of an artificially randomized IgNAR phage display library: screening of variable regions that bind to hen egg white lysozyme. Mar Biotechnol 15:56–62. https://doi.org/10.1007/s10126-012-9456-1
Gamkrelidze M, Dąbrowska K (2014) T4 bacteriophage as a phage display platform. Arch Microbiol 196:473–479. https://doi.org/10.1007/s00203-014-0989-8
Wu J, Tu C, Yu X, Zhang M, Zhang N, Zhao M et al (2007) Bacteriophage T4 nanoparticle capsid surface SOC and HOC bipartite display with enhanced classical swine fever virus immunogenicity: a powerful immunological approach. J Virol Methods 139:50–60. https://doi.org/10.1016/j.jviromet.2006.09.017
Beghetto E, Gargano N (2011) Lambda-display: a powerful tool for antigen discovery. Molecules 16:3089–3105. https://doi.org/10.3390/molecules16043089
Cicchini C, Ansuini H, Amicone L, Alonzi T, Nicosia A, Cortese R et al (2002) Searching for DNA-protein interactions by lambda phage display. J Mol Biol 322:697–706. https://doi.org/10.1016/s0022-2836(02)00851-3
Gao J, Wang Y, Liu Z, Wang Z (2010) Phage display and its application in vaccine design. Ann Microbiol 60:13–19. https://doi.org/10.1007/s13213-009-0014-7
Matthews LJ, Davis R, Smith GP (2002) Immunogenically fit subunit vaccine components via epitope discovery from natural peptide libraries. J Immunol 169:837–846. https://doi.org/10.4049/jimmunol.169.2.837
Mullen LM, Nair SP, Ward JM, Rycroft AN, Henderson B (2006) Phage display in the study of infectious diseases. Trends Microbiol 14:141–147. https://doi.org/10.1016/j.tim.2006.01.006
McCafferty J, Griffiths AD, Winter G, Chiswell DJ (1990) Phage antibodies: filamentous phage displaying antibody variable domains. Nature 348:52–554. https://doi.org/10.1038/348552a0
Barderas R, Peña EB (2019) The 2018 Nobel prize in chemistry: phage display of peptides and antibodies. Anal Bioanal Chem 411:2475–2479. https://doi.org/10.1007/s00216-019-01714-4
Bazan J, Całkosiński I, Gamian A (2012) Phage display–a powerful technique for immunotherapy: 1. Introduction and potential of therapeutic applications. Hum Vaccin Immunother 8:1817–1828. https://doi.org/10.4161/hv.21703
Pande J, Szewczyk MM, Grover AK (2010) Phage display: concept, innovations, applications and future. Biotechnol Adv 28:849–858. https://doi.org/10.1016/j.biotechadv.2010.07.004
Ledsgaard L, Kilstrup M, Karatt-Vellatt A, McCafferty J, Laustsen AH (2018) Basics of antibody phage display technology. Toxins 10:236. https://doi.org/10.3390/toxins10060236
Aghebati-Maleki L, Bakhshinejad B, Baradaran B, Motallebnezhad M, Aghebati-Maleki A, Nickho H, Yousefi M, Majidi J (2016) Phage display as a promising approach for vaccine development. J Biomed Sci 23:66. https://doi.org/10.1186/s12929-016-0285-9
Ahmad ZA, Yeap SK, Ali AM, Ho WY, Alitheen NB, Hamid M (2012) ScFvantibody: principles and clinical application. Clin Dev Immunol. https://doi.org/10.1155/2012/980250
Ubah OC, Barelle CJ, Buschhaus MJ, Porter AJ (2016) Phagedisplay derived IgNAR V region binding domains for therapeutic development. Curr Pharm Des 22:6519–6526. https://doi.org/10.2174/1381612822666160907091708
Huang R, Kiss MM, Batonick M, Weiner MP, Kay BK (2016) Generating recombinant antibodies to membrane proteins through phage display. Antibodies 5:11. https://doi.org/10.3390/antib5020011
Hust M, Jostock T, Menzel C, Voedisch B, Mohr A, Brenneis M et al (2007) SinglechainFab (scFab) fragment. BMC Biotechnol 8(7):14. https://doi.org/10.1186/1472-6750-7-14
Nilvebrant J, Tessier PM, Sidhu SS (2016) Engineeredautonomoushumanvariabledomains. Curr Pharm Des 22(43):6527–6537. https://doi.org/10.2174/1381612822666160921143011
Smolarek D, Bertrand O, Czerwinski M (2012) Variable fragments of heavy chain antibodies (VHHs): a new magic bullet molecule of medicine? Postepy Hig Med Dosw 14(66):348–358. https://doi.org/10.5604/17322693.1000334
Kuzmicheva GA, Belyavskaya VA (2016) Peptide phage display in biotechnology and biomedicine. Biomed Khim 62:481–495. https://doi.org/10.18097/PBMC20166205481
Mimmi S, Maisano D, Quinto I, Iaccino E (2019) Phage display: an overview in context to drug discovery. Trends Pharmacol Sci 40:87–91. https://doi.org/10.1016/j.tips.2018.12.005
Kuhn P, Fühner V, Unkauf T, Moreira GMSG, Frenzel A, Miethe S et al (2016) Recombinant antibodies for diagnostics and therapy against pathogens and toxins generated by phage display. Proteomics Clin Appl 10:922–948. https://doi.org/10.1002/prca.201600002
Balfour H (2020) https://www.drugtargetreview.com/news/57698/novel-phage-display-technique-could-lead-to-new-peptide-based-therapeutics/. Accessed June 2020
Centres for Disease Control and Prevention (2017) Severe Acute respiratory syndrome. https://www.cdc.gov/sars/about/fs-sars.html. Accessed Jan 6 2021
World Health Organization (2020). Middle East respiratory syndrome coronavirus (MERS-CoV) – The Kingdom of Saudi Arabia. Disease Outbreak News: Update. https://www.who.int/csr/don/24-february-2020-mers-saudi-arabia/en/. Accessed January 6, 2021.
WHO Coronovirus Disease (COVID-19) Dashboard (2021). WHO health emergency dashboard. https://covid19.who.int/?gclid=CjwKCAiAudD_BRBXEiwAudakXzKCZatLhBuqkduRzYlVRKe_IMy3_LyPywZyY5Q1-YY9rTP6EXmhlhoCX4gQAvD_BwE. Accessed Jan 6 2021
Englund JA, Kim YJ, McIntosh K (2019) Human coronaviruses, including Middle East respiratory syndrome coronavirus. Feigin and Cherry’s textbook of pediatric infectious disease, 8th edn. Philadelphia, pp 1846–54
Tyrrell DA, Bynoe ML (1965) Cultivation of a novel type of common-cold virus in organ cultures. Br Med J 1:1467–1470. https://doi.org/10.1136/bmj.1.5448.1467
Hamre D, Procknow JJ (1966) A new virus isolated from the human respiratory tract. Proc Soc Exp Biol Med 121:190–193. https://doi.org/10.3181/00379727-121-30734
Ye ZW, Yuan S, Yuen KS, Fung SY, Chan CP, Jin DY (2020) Zoonotic origins of human coronaviruses. Int J Biol Sci 16:1686–1697. https://doi.org/10.7150/ijbs.45472
Zhou P, Yang XL, Wang XG, Hu B, Zhang L, Zhang W et al (2020) Discovery of a novel coronavirus associated with the recent pneumonia outbreak in humans and its potential bat origin. BioRxiv. https://doi.org/10.1101/2020.01.22.914952
Hoffmann M, Kleine-Weber H, Schroeder S, Krüger N, Herrler T, Erichsen S et al (2020) SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181:271–80.e8. https://doi.org/10.1016/j.cell.2020.02.052
Riordan JF (2003) Angiotensin-I-converting enzyme and its relatives. Genome Biol 4:225. https://doi.org/10.1186/gb-2003-4-8-225
Kuba K, Imai Y, Ohto-Nakanishi T, Penninger JM (2010) Trilogy of ACE2: a peptidase in the renin-angiotensin system, a SARS receptor, and a partner for amino acid transporters. Pharmacol Ther 128:119–128. https://doi.org/10.1016/j.pharmthera.2010.06.003
Jiang F, Yang J, Zhang Y, Dong M, Wang S, Zhang Q et al (2014) Angiotensin-converting enzyme 2 and angiotensin 1–7: novel therapeutic targets. Nat Rev Cardiol 11:413–426. https://doi.org/10.1038/nrcardio.2014.59
Harmer D, Gilbert M, Borman R, Clark KL (2002) Quantitative mRNA expression profiling of ACE 2, a novel homologue of angiotensin converting enzyme. FEBS Lett 532:107–110. https://doi.org/10.1016/s0014-5793(02)03640-2
Leung WK, To KF, Chan PKS, Chan HLY, Wu AKL, Lee N et al (2003) Enteric involvement of severe acute respiratory syndrome-associated coronavirus infection. Gastroenterology 125:1011–1017. https://doi.org/10.1016/s0016-5085(03)01215-0
Millet JK, Whittaker GR (2015) Host cell proteases: critical determinants of coronavirus tropism and pathogenesis. Virus Res 202:120–134. https://doi.org/10.1016/j.virusres.2014.11.021
Ma Z, Wang T, Li Z, Guo X, Tian Y, Li Y et al (2019) A novel biotinylated nanobody-based blocking ELISA for the rapid and sensitive clinical detection of porcine epidemic diarrhea virus. J Nanobiotechnol 17:96. https://doi.org/10.1186/s12951-019-0531-x
Tian X, Li C, Huang A, Xia S, Lu S, Shi Z, Lu L (2020) Potent binding of 2019 novel coronavirus spike protein by a SARS coronavirus-specific human monoclonal antibody. Emerg Microbes Infect 17(9):382–385. https://doi.org/10.1080/22221751.2020.1729069
Ter Meulen J, van den Brink EN, Poon LLM, Marissen WE, Leung CSW, Cox F et al (2006) Human monoclonal antibody combination against SARS coronavirus: synergy and coverage of escape mutants. PLoS Med 3:e237. https://doi.org/10.1371/journal.pmed.0030237
Chen W-H, Hotez PJ, Bottazzi ME (2020) Potential for developing a SARS-CoV receptor-binding domain (RBD) recombinant protein as a heterologous human vaccine against coronavirus infectious disease (COVID)-19. Hum Vaccin Immunother. https://doi.org/10.1080/21645515.2020.1740560
Mken Institute. Covid-19 treatment and vaccine tracker. https://milkeninstitute.org/sites/default/files/2020-04/Covid19%20Tracker%20NEW4-14-20. Accessed June 2020
Thailand Medical News (2020) Covid-19 Research: Scientists develop new phage capsid nanoparticles that could prevent Covid-19 infections. https://www.thailandmedical.news/news/covid-19-research-scientists-develop-new-phage-capsid-nanoparticles-that-could-prevent-covid-19-infections. Accessed May 1 2020
Lauster D, Klenk S, Ludwig K, Nojoumi S, Behren S, Adam L et al (2020) Phage capsid nanoparticles with defined ligand arrangement block influenza virus entry. Nat Nanotechnol 15:373–379. https://doi.org/10.1038/s41565-020-0660-2
Walter JD, Hutter CAJ, Zimmermann I, Wyss M, Egloff P, Sorgenfrei M et al (2020) Sybodies targeting the SARS-CoV-2 receptor-binding domain. BioRxiv. https://doi.org/10.1101/2020.04.16.045419
Liu H, Ding YL, Han W, Liu MY, Tian RY, Yang SL et al (2004) Recombinant scFv antibodies against E protein and N protein of severe acute respiratory syndrome virus. Acta Biochim Biophys Sin 36:541–547. https://doi.org/10.1093/abbs/36.8.541
Van den Brink EN, Ter Meulen J, Cox F, Jongeneelen MAC, Thijsse A, Throsby M et al (2005) Molecular and biological characterization of human monoclonal antibodies binding to the spike and nucleocapsid proteins of severe acute respiratory syndrome coronavirus. J Virol 79:1635–1644. https://doi.org/10.1128/JVI.79.3.1635-1644.2005
Duan J, Yan X, Guo X, Cao W, Han W, Qi C et al (2005) A human SARS-CoV neutralizing antibody against epitope on S2 protein. Biochem Biophys Res Commun 333:186–193. https://doi.org/10.1016/j.bbrc.2005.05.089
Zhong X, Yang H, Guo ZF, Sin WYF, Chen W, Xu J et al (2005) B-cell responses in patients who have recovered from severe acute respiratory syndrome target a dominant site in the S2 domain of the surface spike glycoprotein. J Virol 79:3401–3408. https://doi.org/10.1128/JVI.79.6.3401-3408.2005
Hua RH, Wu DL, Tong GZ, Wang YF, Tian ZJ, Zhou YJ (2006) Identification of a mimotope peptide bound to the SARS-CoV spike protein specific monoclonal antibody 2C5 with phage-displayed peptide library. Chin J Biotechnol 22:701–706. https://doi.org/10.1016/S1872-2075(06)60051-4
Duan J, Ji X, Feng J, Han W, Zhang P, Cao W et al (2006) A human neutralizing antibody against a conformational epitope shared by oligomeric SARS S1 protein. Antivir Ther 11:117–123
Zhang X, Wang J, Wen K, Mou Z, Zou L, Che X et al (2009) Antibody binding site mapping of SARS-CoV spike protein receptor-binding domain by a combination of yeast surface display and phage peptide library screening. Viral Immunol 22:407–415. https://doi.org/10.1089/vim.2009.0046
Lee Y-C, Leu S-JC, Hu C-J, Shih N-Y, Huang I-J, Wu H-H (2007) Chicken single-chain variable fragments against the SARS-CoV spike protein. J Virol Methods 146:104–111. https://doi.org/10.1016/j.jviromet.2007.06.010
Tarnovitski N, Matthews LJ, Sui J, Gershoni JM, Marasco WA (2006) Mapping a neutralizing epitope on the SARS coronavirus spike protein: computational prediction based on affinity-selected peptides. J Mol Biol 359:190–201. https://doi.org/10.1016/j.jmb.2006.03.008
Wu RP, Meng JZ, He YX (2013) Construction and screening of SARS-CoV S protein-specific phage displayed antigen library. Bing Du Xue Bao 29:280–286
Wang C, Sun X, Suo S, Ren Y, Li X, Herrler G et al (2013) Phages bearing affinity peptides to severe acute respiratory syndromes-associated coronavirus differentiate this virus from other viruses. J Clin Virol 57:305–310. https://doi.org/10.1016/j.jcv.2013.04.002
Freund NT, Roitburd-Berman A, Sui J, Marasco WA, Gershoni JM (2015) Reconstitution of the receptor-binding motif of the SARS coronavirus. Protein Eng Des Sel 28:567–575. https://doi.org/10.1093/protein/gzv052
Huang S-H, Lee T-Y, Lin Y-J, Wan L, Lai C-H, Lin C-W (2017) Phage display technique identifies the interaction of severe acute respiratory syndrome coronavirus open reading frame 6 protein with nuclear pore complex interacting protein NPIPB3 in modulating type I interferon antagonism. J Microbiol Immunol Infect 50:277–285. https://doi.org/10.1016/j.jmii.2015.07.002
He L, Li J, Ren S, Sun S, Guo Y, Qiu H et al (2017) Construction and identification of nanobody phage display library targeting Middle East respiratory syndrome coronavirus. Xi Bao Yu Fen Zi Mian Yi Xue Za Zhi. 33:1662–1668
Kim Y, Lee H, Park K, Park S, Lim J-H, So MK et al (2019) Selection and characterization of monoclonal antibodies targeting middle east respiratory syndrome coronavirus through a human synthetic Fab phage display library panning. Antibodies. https://doi.org/10.3390/antib8030042
Feng M, Bian H, Wu X, Fu T, Fu Y, Hong J et al (2019) Construction and next-generation sequencing analysis of a large phage-displayed VNAR single-domain antibody library from six naïve nurse sharks. Antib Ther 2:1–11. https://doi.org/10.1093/abt/tby011
Peng B, Chen H, Tan Y, Jin M, Chen H, Guo A (2006) Identification of one peptide which inhibited infectivity of avian infectious bronchitis virus in vitro. Sci China C Life Sci 49:158–163. https://doi.org/10.1007/s11427-006-0158-7
Han Z, Zhao F, Shao Y, Liu X, Kong X, Song Y et al (2013) Fine level epitope mapping and conservation analysis of two novel linear B-cell epitopes of the avian infectious bronchitis coronavirus nucleocapsid protein. Virus Res 171:54–64. https://doi.org/10.1016/j.virusres.2012.10.028
Zou N, Xia J, Wang F, Duan Z, Miao D, Yan Q et al (2015) Two novel neutralizing antigenic epitopes of the s1 subunit protein of a QX-like avian infectious bronchitis virus strain Sczy3 as revealed using a phage display peptide library. Vet Immunol Immunopathol 168:49–55. https://doi.org/10.1016/j.vetimm.2015.08.008
Zou H, Zarlenga DS, Sestak K, Suo S, Ren X (2013) Transmissible gastroenteritis virus: identification of M protein-binding peptide ligands with antiviral and diagnostic potential. Antiviral Res 99:383–390. https://doi.org/10.1016/j.antiviral.2013.06.015
Suo S, Wang X, Zarlenga D, Bu R-E, Ren Y, Ren X (2015) Phage display for identifying peptides that bind the spike protein of transmissible gastroenteritis virus and possess diagnostic potential. Virus Genes 51:51–56. https://doi.org/10.1007/s11262-015-1208-7
Cao L, Ge X, Gao Y, Zarlenga DS, Wang K, Li X et al (2015) Putative phage-display epitopes of the porcine epidemic diarrhea virus S1 protein and their anti-viral activity. Virus Genes 51:217–224. https://doi.org/10.1007/s11262-015-1234-5
Zhang F, Chen Y, Ke Y, Zhang L, Zhang B, Yang L et al (2019) Single chain fragment variable (scFv) antibodies targeting the spike protein of porcine epidemic diarrhea virus provide protection against viral infection in piglets. Viruses. https://doi.org/10.3390/v11010058
Sun D, Shi H, Chen J, Shi D, Zhu Q, Zhang H et al (2012) Generation of a mouse scFv library specific for porcine aminopeptidase N using the T7 phage display system. J Virol Methods 182:99–103. https://doi.org/10.1016/j.jviromet.2012.03.021
Guo D, Zhu Q, Feng L, Sun D (2013) Screening and antiviral analysis of phages that display peptides with an affinity to subunit C of porcine aminopeptidase. Monoclon Antib Immunodiagn Immunother 32:326–329. https://doi.org/10.1089/mab.2013.0038
Gao W, He W, Zhao K, Lu H, Ren W, Du C et al (2010) Identification of NCAM that interacts with the PHE-CoV spike protein. Virol J 7:254. https://doi.org/10.1186/1743-422X-7-254
Acknowledgements
The support from Indian Council of Agricultural Research, New-Delhi, India is duly acknowledged.
Funding
This work was supported by Indian Council of Agricultural Research, New Delhi, India.
Author information
Authors and Affiliations
Contributions
TA performed writing original draft, formal analysis, review and editing. NV contributed to conceptualization, methodology and data curation. BCB was involved in conceptualization, formal analysis, review and editing. RKV performed formal analysis, review and editing; MV and PB contributed to visualization and editing. AK contributed to supervision and resources. BNT was involved in conceptualization, methodology and resources.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Anand, T., Virmani, N., Bera, B.C. et al. Phage Display Technique as a Tool for Diagnosis and Antibody Selection for Coronaviruses. Curr Microbiol 78, 1124–1134 (2021). https://doi.org/10.1007/s00284-021-02398-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-021-02398-9