Abstract
Diols are versatile chemicals used for multiple manufacturing products. In some previous studies, Escherichia coli has been engineered to produce 1,2-propanediol (1,2-PDO) and 1,3-propanediol (1,3-PDO) from glucose. However, there are no reports on the direct production of these diols from starch instead of glucose as a substrate. In this study, we directly produced 1,2-PDO and 1,3-PDO from starch using E. coli engineered for expressing a heterologous α-amylase, along with the expression of 1,2-PDO and 1,3-PDO synthetic genes. For this, the recombinant plasmids, pVUB3-SBA harboring amyA gene for α-amylase production, pSR5 harboring pct, pduP, and yahK genes for 1,2-PDO production, and pSR8 harboring gpd1-gpp2, dhaB123, gdrAB, and dhaT genes for 1,3-PDO production, were constructed. Subsequently, E. coli BW25113 (ΔpflA) and BW25113 strains were transformed with pVUB3-SBA, pSR5, and/or pSR8. Using these transformants, direct production of 1,2-PDO and 1,3-PDO from starch was demonstrated under microaerobic condition. As a result, the maximum production titers of 1,2-PDO and 1,3-PDO from 1% glucose as a sole carbon source were 13 mg/L and 150 mg/L, respectively. The maximum production titers from 1% starch were similar levels (30 mg/L 1,2-PDO and 120 mg/L 1,3-PDO). These data indicate that starch can be an alternative carbon source for the production of 1,2-PDO and 1,3-PDO in engineered E. coli. This technology could simplify the upstream process of diol bioproduction.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biorefining is a technology that produces biobased fuels and chemicals from primarily starch-based crops such as corn [1, 2]. Since diols are versatile compounds harboring two hydroxyl groups, they are used as platform chemicals for many industrial applications. Biobased diols are promising targets produced from biomass through the biorefinery process [3]. Among biobased diols, 1,3-propanediol (1,3-PDO) has enabled synthesis of multiple synthetic polymers such as new biobased polyester, poly(trimethylene terephthalate) [4, 5].
Both 1,2-PDO and 1,3-PDO are C3 diols that can be produced by engineered and non-engineered microbes, using glucose, glycerol, or xylose as the carbon source [6,7,8]. Clostridium spp. and Corynebacterium glutamicum produce 1,2-PDO from glucose or xylose [9,10,11,12]. Clostridium acetobutylicum and ` produce 1,3-PDO from glycerol [13,14,15]. Escherichia coli is a suitable host for many biobased chemical production processes since it grows rapidly and can easily be genetically manipulated. There are reports of 1,2-PDO production from glucose in recombinant E. coli strains harboring 1,2-PDO synthetic genes pct, pduP, and yahK [6, 8]. In another study, gpd1 and gpp2 from Saccharomyces cerevisiae and the dha operon (dhaB123, gdrAB, and dhaT) from K. pneumoniae were introduced into E. coli DH5α and BL21 to produce 1,3-PDO from glycerol or glucose [8, 16]. However, there is no report of direct production of these diols from starch instead of glucose because E. coli unassimilated oligo- and polysaccharides [17].
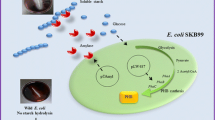
The cell surface display is a protein expression technology for expression of desired proteins on the cell membrane, using membrane anchors such as glycosylphosphatidylinositol [18, 19]. The cell surface display enables the beneficial production of desired products from starch in multiple microbial strains, including S. cerevisiae, Lactobacillus casei, and C. glutamicum [20,21,22,23]. Recently, an E. coli recombinant strain producing itaconic acid has been engineered to display the starch-digesting enzyme, α-amylase from Streptococcus bovis NRIC1535, fused with an OprI’ lipoprotein anchor on its cell membrane, so that the recombinant strain produces itaconic acid directly from starch [24, 25]. These findings suggest that α-amylase expression on the cell surface could allow E. coli to produce 1,2-PDO and 1,3-PDO directly from starch. Direct production of 1,2-PDO and 1,3-PDO in E. coli from starch simplifies the upstream process of the bioproduction of these diols, leading to lower production costs.
In this study, we demonstrate the production of 1,2-PDO and 1,3-PDO directly from starch in E. coli by expressing α-amylase on the cell surface, along with the expression of diol synthetic genes.
Materials and Methods
Bacterial Strains and Media
The bacterial strains and plasmids used in this study are listed in Table 1. Spectinomycin and carbenicillin were added to cultures at final concentrations of 100 µg/mL and 50 µg/mL, respectively. For diol production, recombinant E. coli were grown at 30 °C and 150 rpm under microaerobic condition in M9 medium (15.14 g/L, Na2HPO4, 3 g/L KH2PO4, 0.5 g/L NaCl), supplemented with 2 g/L yeast extract [16] and 10 g/L glucose or starch as a carbon source. Microaerobic cultivation was performed in 200-mL flasks filled with 133 mL medium and sealed with rubber stoppers. The culture medium was further supplemented with 0.12 g/L MgSO4, 0.028 g/L CaCl2, and 1 mg/L thiamin hydrochloride for 1,2-PDO production. Vitamin B12 (10 µM) was added into the medium for 1,3-PDO production. Isopropyl-β-d-1-thiogalactopyranoside (IPTG) was added to a final concentration of 0.5 mM to induce gene expression when the optical density of cultures at 600 nm (OD600) reached 0.4 to 0.6.
Plasmid Construction
The primer sets for PCR amplification used in this study are listed in S. Table 1. The vector pGV3, the pct gene from Megasphaera elsdenii JCM 1772 (accession No. M26493), the pduP gene from Salmonella enterica subsp. enterica NBRC 13245 (accession No. AB68030), and the yahK gene from E. coli BW25113 (accession No. U00096) were amplified by PCR using Q5 High-Fidelity DNA Polymerase (New England Biolabs, MA, USA). The amplified genes were ligated with the amplified pGV3 vector fragment using the Gibson assembly system [26] with the Gibson Assembly Master mix (New England Biolabs), resulting in pSR5.
The gpd1-gpp2 fusion gene [16] was synthesized by GenScript Japan Co. (Tokyo, Japan). The vectors pTV118N and pGV3, the gpd1-gpp2 fusion gene, and dha operon (dhaB123, gdrAB, and dhaT) from K. pneumoniae NCIMB 418 (accession no. U30903) were amplified by PCR. The amplified gpd1-gpp2 fusion gene was ligated with the amplified pTV118N vector fragment by Gibson assembly, resulting in pSR3. The amplified dha operon was ligated with the amplified pGV3 vector fragment by Gibson assembly, resulting in pSR4. The regions of lac promoter (Plac) and gpd1-gpp2 fusion gene and pSR4 were amplified by PCR. The amplified Plac-gpd1-gpp2 fragment was ligated with the amplified pSR4 plasmid fragment by Gibson assembly, resulting in pSR8. The amyA gene from S. bovis NRIC 1535 (accession no. AB000829) was amplified by PCR using KOD plus DNA Polymerase (TOYOBO, Osaka, Japan). The amplified amyA gene was ligated at BglII and PstI sites of pVUB3 plasmid vector, resulting in pVUB3-SBA.
Analytical Methods
The recombinant strains cultivated in LB medium at 30 °C for 18 h were inoculated in fresh LB medium, and cultured to OD600 of 0.1. After the cultures were subsequently grown at 30 °C until they reached an OD600 to 0.5, IPTG was added to the cultures to a final concentration of 0.5 mM and incubated at 30 °C for 18 h for amyA expression. Cultures were centrifuged at 15,000 rpm for 5 min, and then the culture supernatant was removed and the bacterial pellet was resuspended in 20 mM phosphate buffer to bring the OD600 to 1.0. The α-amylase activity was tested using a commercial α-Amylase activity assay kit (Kikkoman Corp., Chiba, Japan). The culture supernatant was cultivated overnight and then tested by iodine starch reaction. Glucose concentration was determined using a commercial Glucose CII-test kit (Fujifilm Wako Co., Osaka, Japan). Starch concentration was measured by the Glucose CII-test kit after digesting starch to glucose by treatment of the culture supernatant with 1% sulfuric acid at 120 °C for 2 h.
Concentrations of lactate, 1,2-PDO, glycerol, and 1,3-PDO in culture supernatants were quantified using a Prominence HPLC system (Shimadzu, Kyoto Japan) equipped with an Aminex HPX-87H column (Bio-Rad, CA, USA) and a refractive index detector (GL Science, Tokyo, Japan). A mobile phase (5 mM H2SO4) was eluted at a flow rate of 0.5 mL/min at 65 °C.
Results
Engineering E. coli
The plasmids pSR5, pSR8, and pVUB3-SBA for 1,2-PDO and 1,3-PDO production, and α-amylase expression were constructed, respectively (Fig. 1). E. coli BW25113 (ΔpflA) and E. coli BW25113 strains were transformed with pVUB3-SBA, pSR5, and/or pSR8, resulting in E. coli strains EC_3, EC_4, EC_5, and EC_6. The α-amylase activity was measured in E. coli strains, showing that the activities of E. coli BW25113, EC_1, EC_2, EC_4, and EC_6 were 0.06 ± 0.04 U/mL, 0.01 ± 0.00 U/mL, 4.6 ± 0.0 U/mL, 2.70 ± 0.00 U/mL, and 2.53 ± 0.51 U/mL, respectively (Fig. 2a). There was no significant difference (P > 0.05) between their activities. The culture supernatants of E. coli BW25113 and EC_2 showed negligible amylolytic activity (BW25113, 0.03 ± 0.02 U/mL; EC_2, 0.31 ± 0.07 U/mL). These results show that α-amylase expressed is not secreted into the cultures but located on the cell surface. To test for amylolytic activity in the cultures of E. coli harboring pVUB3-SBA, iodine tests were performed. The E. coli BW25113 culture showed amylolytic activity but the E. coli EC_6 culture did not (Fig. 2b). This result shows that α-amylase digests starch in EC_6.
Specific α-amylase activities and iodine starch reaction of E. coli cultures. a α-Amylase activities of E. coli cultures grown at 30 °C for 18 h. A, E. coli BW25113; B, E. coli EC_1; C, E. coli EC_2; D, E. coli EC_4 E, E. coli EC_6. White and black bars correspond to specific α-amylase activities of the cells in phosphate buffer and the culture supernatants, respectively. b Iodine starch reaction with E. coli cultures. E. coli BW25113 and EC_6 were cultivated at 30 °C for 72 h and then tested
Direct Production of 1,2-PDO from Glucose or Starch
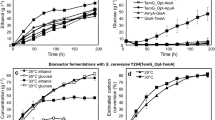
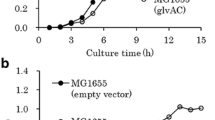
We performed 1,2-PDO production in engineered E. coli in M9 medium supplemented with 1% glucose or 1% starch as the sole carbon source under microaerobic condition. Glucose added in the culture was completely consumed by 96 h (Fig. 3a). The OD600 after 96 h cultivation was 2.60 ± 0.12 when 1% glucose was used. E. coli EC_4 produced 6.11 ± 0.11 g/L lactate and 13.0 ± 0.0 mg/L 1,2-PDO from 1% glucose after 96 h cultivation. The lactate and 1,2-PDO yields from glucose were 60.7% and 0.31% (mol/mol), respectively, and 1,2-PDO yield from lactate was 0.25% (mol/mol). The specific growth rate was 0.11 ± 0.01 h−1 when 1,2-PDO was produced from 1% glucose.
The same strain was cultivated in M9 medium supplemented with 1% starch as the sole carbon source under microaerobic condition. The concentration of residual sugar detected was 1.97 ± 0.60 g/L when starch was used after 96 h cultivation (Fig. 3b). The OD600 after 96 h cultivation was 2.18 ± 0.12 when 1% starch was used. The specific growth rate was 0.08 ± 0.00 h−1 when 1,2-PDO was produced from starch. There was no significant difference between the specific growth rates during cultivation from glucose and starch (P > 0.05). After 96 h cultivation with 1% starch, production of 3.08 ± 0.00 g/L lactate and 28.0 ± 0.0 mg/L 1,2-PDO was observed. The lactate and 1,2-PDO yields from starch were 38.3 and 0.8% (mol/mol), respectively, and 1,2-PDO yield from lactate was 1.1% (mol/mol). These results show that starch assimilation and lactate production are rate-limiting reactions for 1,2-PDO production. However, the production level of 1,2-PDO from starch was similar to that from glucose.
Direct Production of 1,3-PDO from Glucose or Starch
We demonstrated 1,3-PDO production in engineered E. coli in M9 medium supplemented with 1% glucose or 1% starch as the sole carbon source under microaerobic condition. Glucose added in the culture was completely consumed by 72 h (Fig. 4a). E. coli EC_6 produced 350.0 ± 0.0 mg/L glycerol and 140.0 ± 0.0 mg/L 1,3-PDO from 1% glucose after 72 h cultivation (OD600 = 1.13 ± 0.03). The specific growth rate was 0.05 ± 0.00 h−1 when 1% glucose was used. The glycerol and 1,3-PDO yields from glucose were 7.1 and 3.4% (mol/mol), respectively, and 1,3-PDO yield from glycerol was 48.4% (mol/mol).
The same strain was cultivated in M9 medium supplemented with 1% starch as the sole carbon source under microaerobic condition. The concentration of residual sugar detected was 0.49 ± 0.06 g/L when starch was used after 72 h cultivation (Fig. 4b). The OD600 and specific growth rate were 1.27 ± 0.00 and 0.05 ± 0.00 h−1, respectively, when 1% starch was used. There was no significant difference between the specific growth rates during cultivation from glucose and starch (P > 0.05). E. coli EC_6 produced 410.0 ± 0.0 mg/L glycerol and 100.0 ± 0.0 mg/L 1,3-PDO from 1% starch after 72 h cultivation (OD600 = 0.83 ± 0.06). The glycerol and 1,3-PDO yields from starch were 8.4 and 2.5% (mol/mol), respectively, and the 1,3-PDO yield from glycerol was 29.5% (mol/mol). These results show that growth and production levels of glycerol and 1,3-PDO are similar when 1% glucose or 1% starch is used (P > 0.05). E. coli cells expressing α-amylase had similar growth rates when grown in the presence of 1% glucose or 1% starch.
Discussion
It has been reported that engineered E. coli strains produce 1,2-PDO and 1,3-PDO from glucose [6,7,8, 16]. However, the direct production of these diols from starch has not been reported so far. In this study, we demonstrated 1,2-PDO and 1,3-PDO production from starch as the sole carbon source in engineered E. coli.
Prior to the demonstration of 1,2-PDO and 1,3-PDO production, we constructed an amylolytic plasmid pVUB3-SBA and evaluated its function. The α-amylase activities of E. coli EC_4 and EC_6 were somewhat lower than that of EC_2 (Fig. 2a). This may be due to competition with RNA polymerase, since the plasmids pVUB3-SBA, pSR5, and pSR8 possess the Plac or its derived promoter (Fig. 1). Starch digestion by E. coli harboring pVUB3-SBA was tested with the iodine starch reaction (Fig. 2b). This showed that starch added to the medium was almost completely digested, but the residual sugar concentration was found to be 1.6 g/L at 96 h (Fig. 3b). The reason starch was not completely consumed is that E. coli unassimilated oligosaccharide [17].
We demonstrated the production of 1,2-PDO from glucose or starch by engineered E. coli. Production titers of 1,2-PDO from glucose in this study were lower than those reported in previous studies [6, 8]. However, lactate production in this study was no more significant than that reported by Niu et al. (3.8 g/L lactate and 0.5 g/L 1,2-PDO from 1% glucose in engineered E. coli MG1655) [8]. The reactions involved in the conversion of lactoyl-CoA to 1,2-PDO require NADH and NADPH, which are partially derived from glucose [8]. Glucose added in the culture was completely consumed by 96 h in this study. Consequently, quantities of NADH and NADPH were less than required for the synthesis of 1,2-PDO from lactoyl-CoA. This suggests that additional glucose improves 1,2-PDO production after glucose is consumed. There was no significant difference between the specific growth rates when 1% glucose and starch were used (P > 0.05). The quantities of lactate produced in E. coli EC_4 from glucose and starch at 96 h cultivation were 6 and 3 g/L, respectively (Fig. 3). This is because glucose was consumed rapidly and completely compared to starch.
We have demonstrated direct production of 1,3-PDO from starch in engineered E. coli. The production level in this study was lower than that reported by Liang et al. (1.15 g/L glycerol and 0.48 g/L 1,3-PDO from 1% glucose in engineered E. coli DH5α) [16]. They reported that 1,3-PDO production was improved by using a stress-induced rpoS promoter instead of the Plac. Because the induction level of rpoS promoter is stronger than that of Plac. In addition, this promoter has been reported to be inducible under stress conditions including cold shock and pH stress [27, 28]. Lee et al. also reported that deletion of the glycerol oxidation pathway in E. coli improves glycerol and 1,3-PDO production [7]. These reports suggest that further engineering of our constructed strain can improve 1,3-PDO production efficiency.
In this study, the glycerol yield from starch was higher than that from glucose, while the 1,3-PDO yield from glucose was higher than that from starch. However, glycerol and 1,3-PDO yields from glucose were 7.1 and 3.4% (mol/mol), respectively. On the other hand, the glycerol and 1,3-PDO yields from starch were 8.4 and 2.5% (mol/mol), respectively. These results showed that there is no significant difference in glycerol yields from glucose and starch, but the 1,3-PDO yield from glucose was higher. This may be because glucose was consumed more slowly than starch. The reactions from dihydroxyacetone phosphate to 1,3-PDO require NADH [29]. Glucose in the culture was almost completely consumed by 24 h in this study. Consequently, the quantity of NADH was less than required for the synthesis of 1,3-PDO from dihydroxyacetone phosphate, causing the 1,3-PDO production level to be lower than when starch was used.
We have demonstrated direct production of 1,2-PDO and 1,3-PDO from starch in engineered E. coli based on cell surface display technology. This technology would play an important role in skipping the saccharification step for digestion of starch to lower-molecular-weight sugars upstream of diol production, leading to accelerated diol production by bioprocess.
Conclusion
In conclusion, 1,2-PDO and 1,3-PDO, the raw materials for several multiple manufacturing products, were produced directly from starch. This technology simplifies the process of saccharification for digestion of sugar in the microbial production process of these diols, leading to cost savings.
References
Fernando S, Adhilari S, Chandrapal C, Murali N (2006) Biorefineries: current status, challenges, and future direction. Energy Fuel 20:1727–1737
Saggy KS, Dey P (2016) An overview of simultaneous saccharification and fermentation of starchy and lignocellulosic biomass for bio-ethanol production. Biofuels 10:287–299
Tomishige K, Nakagawa Y, Tamura M (2017) Production of diols from biomass. Prod Platf Chem Sustain Resour Chapter 11:343–373
Jong ED, Higson A, Walsh P, Wellish M (2011) Bio-based chemicals: value added products from biorefineries. In IEA Bioenergy. pp 1–33
Motel E, Phillips H, Smith A (2015) 1,3-propanediol from crude glycerol. In Senior design reports. 74. http://repository.upenn.edu/cbe_sdr/74
Altras EN, Cameron CD (1999) Metabolic of a 1,2-propanediol pathway in Escherichia coli. App Environ Microbiol 65:1180–1185
Lee HJ, Lama S, Kim RJ, Park HS (2018) Production of 1,3-propanediol from glucose by recombinant Escherichia coli BL21 (DE3). Biotechnol Bioprocess Eng 23:250–258
Niu W, Kramer L, Mueller J, Liu K, Guo J (2018) Metabolic engineering of Escherichia coli for the de novo stereospecific biosynthesis of 1,2-propanediol through lactic acid. Metab Eng Commun 7:e00082
Cameron DC, Cooney LC (1986) A novel fermentation: the production of (R)-1,2-propanediol and acetol by Clostridium thermosaccharolyticum. Bio/Technology 4:651–654
Cameron DC, Altras EN, Holfman LM, Shaw JA (1998) Metabolic engineering of propanediol pathways. Biotechnol Prog 14:116–125
Tran-Din K, Gottschalk G (1985) Formation of D(–)-1,2-propanediol and D(–)-lactate from glucose by Clostridium sphenoides under phosphate limitation. Arch Microbiol 142:87–92
Siebert D, Wendisch FV (2015) Metabolic pathway engineering for production of 1,2-propanediol and 1-propanol by Corynebacterium glutamicum. Bitotechnol Biofuels 8:91–104
Liu H, Xu Y, Zheng Z, Liu D (2010) 1,3-Propanediol and its copolymers: research, development and industrialization. Biotechnol J 5:1137–1148
Maeroet VE, De Mey J, Beauprez S, Maeseneire De, Soetaert WKS (2010) Enhancing the microbial conversion of glycerol to 1,3-propanediol using metabolic engineering. Org Proc Res Dev 15:189–202
Ainala KS, Ashok S, Ko Y, Park S (2013) Glycerol assimilation and production of 1,3-propanediol by Citrobacter amalonaticus Y19. Appl Microbiol Biotechnol 97:5001–5011
Liang Q, Zhang H, Li S, Qi Q (2011) Construction of stress-induced metabolic pathway from glucose to 1,3-propanediol in Escherichia coli. App Microbiol Biotechnol 89:57–62
Yazawa K, Imai K, Tamura Z (1978) Oligosaccharides and Polysaccharides specifically utilizable by bifidobacteria. Chem Pharm Bull 26(1):3306–3311
Lipike NP, Wojcechowicz D, Kurjan J (1989) AGα1 is the structural gene for the Saccharomyces cerevisiae α-agglutinin, a cell surface protein involved in cell-cell interactions during mating. Mol Cell Biol 9:3155–3165
Wojcechowicz D, Lu CF, Kurjan J, Lipke NP (1993) Cell surface anchorage and ligand-binding domains of saccharomyces cerevisiae cell adhesion protein α-agglutinin, a member of the immunoglobulin superfamily. Mol Cell Biol 13:2554–2563
Murai T, Ueda M, Yamamura M, Atomi H, Shibasaki Y, Kamasawa N, Osumi M, Amachi T, Tanaka A (1997) Construction starch-utilizing yeast by cell surface engineering. App Environ Microbiol 63:1362–1366
Yao W, Chu C, Deng X, Zhang Y, Liu M, Zheng P, Sun Z (2009) Display α-amylase on the surface of Corynebacterium glutamicum cells by using NCgl221 as the anchoring protein and production of glutamate from starch. Arch Microbiol 191:751–759
Narita J, Okano K, Kitao T, Isida S, Sewaki T, Sung MH, Fukuda H, Kondo A (2005) Display of α-Amylase on the surface of Lactobacillus casei cells by use of the PgsA anchor protein, and production of lactic acid from starch. Appl Environ Microbiol 72:269–275
Tateno T, Hatada K, Tanaka T, Fukuda H, Kondo A (2009) Development of novel cell surface display in Corynebacterium glutamicum using porin. App Environ Microbiol 84:733–739
Okamoto S, Chin T, Nagata K, Takahashi T, Ohara H, Aso Y (2014) Production of itaconic in Escherichia coli expressing recombinant α-amylase using starch as substrate. J Biosci Bioeng 119(5):548–553
Aso Y, Tsubaki M, Long DHB, Murakami R, Nagata K, Okano H, Dung PTN, Ohara H (2018) Continuous production of D-lactic acid from cellobiose in cell recycle fermentation using β-glucosidase-displaying Escherichia coli. J Biosci Bioeng 127(4):441–446
Gibson GD, Young L, Chuang YR, Venter CJ, Clyde AH III, Hamilton OS (2009) Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat Methods 6:343–345
Kang Z, Wang Q, Zhang H, Qi Q (2008) Construction of a stress-induced system in Escherichia coli for efficient polyhydroxyalkanoates production. Appl Microbiol Biotechnol 79:203–208
Hengge-Aronis R (2002) Signal transduction and regulatory mechanisms involved in control of the sigma (S) (RpoS) subunit of RNA polymerase. Microbiol Mol Biol Rev 66:373–395
Cláudio JRF, Débora T, Hélène SB, Adilia D, Christopher MT, Thomas W, Jean MF (2019) Construction of a synthetic pathway for the production of 1,3-propanediol from glucose. Sci Rep 9:11576
Acknowledgements
This work was supported by Japan Science and Technology Agency-Mirai R&D Program Grant Number JPMJMI17EC. We thank the National BioResource Project (National Institute of Genetics, Japan), the Biological Resource Center (NITE, Japan), the National Collection of Industrial, Food and Marine Bacteria (NCIMB Ltd., United Kingdom), the Japan Collection of Microorganisms (RIKEN BioResource Research Center, Japan), and the NODAI Culture Collection Center (Tokyo University of Agriculture, Japan) for providing us with E. coli BW25113, S. enterica subsp. enterica NBRC 13245 and pVUB3, K. pneumoniae NCIMB 418, M. elsdenii JCM 1772, and S. bovis NRIC 1535, respectively.
Author information
Authors and Affiliations
Contributions
Substantial contribution to conception and design or the acquisition and analysis of data: RS, YA. Drafting or critically revising the manuscript: TT, HO, YA. Approval of the final submitted version: RS, TT, HO, YA.
Corresponding author
Ethics declarations
Conflict of interest
The authors declared that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sato, R., Tanaka, T., Ohara, H. et al. Engineering Escherichia coli for Direct Production of 1,2-Propanediol and 1,3-Propanediol from Starch. Curr Microbiol 77, 3704–3710 (2020). https://doi.org/10.1007/s00284-020-02189-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-020-02189-8