Abstract
One novel ascomycetous yeast strain TF5-16-2 was isolated from water samples of Tuofengling crater lake located in Da Hinggan Ling Mountain, in the Inner Mongolia province of China. Morphological, physiological characteristics, as well as phylogenetic analyses of D1/D2 domains of the large subunit rRNA (LSU), ITS region, small subunit rRNA (SSU), and elongation factor-1α (EF-1α) were performed and finally confirmed the phylogenetic placement of strain TF5-16-2 in the genus Wickerhamomyces. Sequences analysis revealed that strain TF5-16-2 differed from its most closely related phylogenetic neighbors ‘Candida’ silvicultrix CBS 6269T and Wickerhamomyces anomalus CBS 5759T by 8.0% (including 2.3% gaps), 8.5% (including 2.4% gaps) divergences in D1/D2 domains of LSU, and 11% (including 4.3% gaps) and 13% (including 4.4% gaps) divergences in ITS region, respectively. As the considerable sequence divergence and distinguishable physiological characteristics, strain TF5-16-2 was proposed as a new species of the genus Wickerhamomyces, with the name Wickerhamomyces kurtzmanii sp. nov. (holotype = CGMCC 2.5597, Mycobank number is MB829959).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Wickerhamomyces was proposed by Kurtzman et al. [1] based on the phylogenetic analysis of concatenated sequences of LSU, SSU, and EF-1α. It consists of a group of species that were earlier assigned to the genus Pichia [2]. Up to date, the genus Wickerhamomyces accommodates 34 species, embracing the newly published species Wickerhamomyces xylosivorus [3] and Wickerhamomyces menglaensis [4]. Members of the genus Wickerhamomyces occur widely in the natural environments, and have been isolated from different habitats including soil [5,6,7,8], phylloplane [9], tree exudates [10], flowers [11, 12], digestive tract of insects [13, 14], larvae of diptera [15], birds [16], natural fermentation of coffee cherries [17], and brined vegetables [18]. However, freshwater environment including crater lake is a special habitat for Wickerhamomyces members, as they are rarely discovered in aquatic environment. During our survey of the diversity of culturable yeast in these crater lakes, strain TF5-16-2 isolated from Tuofengling crater lake in Greater Khingan Mountain of China was identified as a novel species of Wickerhamomyces based on phylogenetic analyses of LSU, SSU, and EF-1α and morphological and physiological comparison, for which the name Wickerhamomyces kurtzmanii sp. nov. was proposed.
Materials and Methods
Yeasts Isolation
Water samples were obtained from Tuofengling Crater lake (with a height of 1284 m) in Da Hinggan Ling Mountain in September 2015. Samples were taken from five randomly chosen sites on the surface of Tuofengling Lake at the depth of 0.5–1.0 m. The pH value of lake water was 7.5, which was obtained in situ using a portable water quality analyzer (HQ40d, US). Five independent 1 L water samples were collected in sterile bottles, kept at 4 ℃ and transported to the laboratory. Isolation of yeast strains was performed as follows: a tenfold dilution series (10−1–10−4) was prepared for each sample. And 100 μl of each dilution was spread over the YM [0.3% (w/v) yeast extract, 0.3% (w/v) malt extract, 0.5% (w/v) peptone, 1% (w/v) glucose, and 1.5% (w/v) agar] plates supplemented with 0.02% chloramphenicol and then incubated at 20 ℃ for up to 7 days. Three replicated plates were inoculated for each dilution. All emerging yeast colonies on the plates were picked and were purified by repeated streaking on YM agar plates. Subsequently, strains were preserved by lyophilization and in the liquid nitrogen in China General Microbiological Culture Collection Center (CGMCC).
Phenotypic Characterization
Strain TF5-16-2 was characterized morphologically and physiologically using standard methods according to Kurtzman et al. [19]. The assimilation of carbon and nitrogen compounds was examined in liquid medium at 22 ℃. Urease activity was tested with Christensen’s urea agar. Dalmau tubes were prepared using Yeast Carbon Base broth supplemented with 0.01% yeast extract to detect the fermentation ability. Cell morphology was observed by light microscopy and scanning electron microscopy (SEM, HITACHI, SU8180) after 3 days of growth in YM broth. Ascospore formation was investigated by incubating strain TF5-16-2 on 5% malt extract agar, YM agar, corn meal agar (CMA), and potato dextrose agar (PDA, BD), at 22 ℃ for 2 weeks.
Phylogenetic Analysis
The sequences were analyzed corresponded to the small subunit of the ribosomal DNA (SSU), the D1/D2 domain of the large submit (LSU or 26S) rDNA, and the translation elongation factor 1-α (TEF1). DNA extraction was performed by using the commercial Eukaryotic Genomic DNA Extraction Kit (Aidlab Biotech, Beijing, China) according to the manufacturer’s instructions. The primers for amplification of ITS region were ITS1 and ITS4 [20], for the D1/D2 domains of the LSU were NL1 and NL4 [21], for SSU were P1F and U3R [22], and for the EF-α were EF1-983F and EF1-2218R [23]. These sequences were determined by commercial sequencing facilities (Ruibo Biotech Co., Beijing, China) and then deposited on GenBank (with the accession numbers MK573939, MK573961, MK573960, and MK580818).
The comparisons of the sequences were carried out using BLASTN. Phylogenetic analysis was performed following the previously described methods [24]. Sequences of the closely related species were retrieved from Genbank database (https://www.ncbi.nlm.nih.gov/) and aligned iteratively by using the multiple alignment program CLUSTAL X [25]. The Phylogenetic trees were constructed by both the Maximum-Likelihood and Neighbor-Joining methods in MEGA 7.0 [26]. Single sequence of D1/D2 domains of LSU rDNA and concatenated sequences set of SSU-LSU-EF-1α generated along with the sequences of all Wickerhamomyces species and related ‘Candida’ species were analyzed. The jModel test was performed in order to select the most appropriate evolution model [27] for the phylogenetic analysis of D1/D2 domains of LSU and the concatenated sequences of SSU-LSU-EF-1α. The Tamura–Nei model was chosen for the D1/D2 partition and the GTR + G + I model was used for the concatenated sequences. Confidence values were estimated from bootstrap analyses of 1000 replicates [28].
Results and Discussion
Yeast Strains Isolated From Water Sample of Crater Lake
A total of 69 yeast strains were isolated from five water samples of Tuofengling lake, belonging to 17 species of 12 genera. Vishniacozyma victoriae, Martiniozyma abiesophila, and Kuraishia floccosa were present as the most abundant species with the number of 19, 17, and 10, respectively. Some species including Cystobasidium pinicola, Dioszegia butyracea, Vishniacozyma victoriae, and Rhodotorula graminis were present by only one strain.
Species Delineation and Phylogenetic Placement
The results of sequence alignment on NCBI indicated that the D1/D2 domain of strain TF5-16-2 showed the highest similarities to strains ‘Candida’ silvicultrix CBS 6269T and W. anomalus CBS 5759T with the values of 92% (including 53 nt substitutions and 21 gaps out of 926 nt) and 91.5% (including 57 nt substitutions and 22 gaps out of 929 nt), respectively; the divergences in ITS region between strain TF5-16-2 and ‘Candida’ silvicultrix CBS 6269T, W. anomalus CBS 5759T were 11% (including 4.3% gaps) and 13% (including 4.4% gaps), respectively.
Maximum-Likelihood analysis based on the D1/D2 domains of the LSU showed that the strain TF5-16-2 located in a Wickerhamomyces subclade containing twelve species of Wickerhamomyces: W. anomalus, W. sylviae, W. edaphicus, W. ciferrii, W. siamensis, W. subpelliculosus, W. lynferdii, W. arborarius, W. spegazzinii, W. queroliae, W. sydowiorum, and ‘Candida’ silvicultrix (Fig. 1), indicating that strain TF5-16-2 was a member of the genus Wickerhamomyces. Furthermore, the ML-tree based on the concatenated sequences of LSU + SSU + EF-1α showed that strain TF5-16-2 formed a tight clade together with W. anomalus, W. subpelliculosus, ‘Candida’ sivicultrix, W. ciferrii, W. sydowiorum, and W. lynferdii (Fig. 2). The NJ trees showed essentially the same topography as that of the ML trees. The above results indicated that strain TF5-16-2 belonged to the genus Wickerhamomyces and is distinguished from any described species of the genus Wickerhamomyces.
Phylogenetic tree derived from Maximum-likelihood analysis of the D1/D2 domains of LSU, showing the placement of Wickerhamomyces kurtzmanii sp. nov TF5-16-2 and related species in Wickerhamomyces clade including Candida species. Saccharomyces cerevisiae NRRL Y-12632T was used as outgroup. Bootstrap values (%) over 50% based on 1000 replication are given at nodes. The filled circles indicate nodes recovered using the neighbor-joining method. Bar, 0.05 substitutions per nucleotide position
Phylogenetic tree derived from Maximum-likelihood analysis based on the concatenated sequences of SSU, the D1/D2 domains of LSU and EF-1α showing the placement of Wickerhamomyces kurtzmanii sp. nov TF5-16-2 and related species in Wickerhamomyces clade including Candida species. Saccharomyces cerevisiae NRRL Y-12632T was used as outgroup. Bootstrap values (%) above 50% based on 1000 replication are given at nodes. The filled circles indicate nodes recovered using the neighbor-joining method. Bar, 0.05 substitutions per nucleotide position
Phenotypic and Growth Characteristics
The strain TF5-16-2 was examined for sporulation by using CMA, PDA, 5% malt extract, and YM agar at 22℃. Conjugation occurred between two separate cells (Fig. 3b) after 5–7 days growth on PDA or CMA (Fig. 3b). One to four hat-shaped ascospores are formed in each ascus (Fig. 3c, d). Ascospores were released when asci were deliquescent (Fig. 3e). Pseudohyphae were not formed. In sum, strain TF5-16-2 was clearly distinct from the other species of the Wickerhamomyces clade by its inability of growth at higher than 30 ℃ (Table 1).
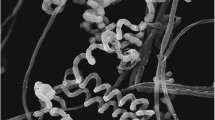
Wickerhamomyces kurtzmanii sp. nov. TF5-16-2. a Scanning electron micrographs of Wickerhamomyces kurtzmanii sp. nov. TF5-16-2 after 3 days of growth at 22 ℃ in YM broth; formation of asci and ascospores; b the conjugated cells; c the intact ascus; d the deliquesced ascus; e the released ascospores. Bar = 5 μm
Unfortunately, we have not yet isolated other strain which might be conspecific species of TF5-16-2 in the last four years. We strived to propose this novel species represented by single strain based on polyphasic approaches including molecular phylogenetic, morphological, and physiological analyses.
Description of Wickerhamomyces kurtzmanii Li, Zhou, and Wang sp. nov.
Wickerhamomyces kurtzmanii (in honor of the prominent yeast taxonomist Cletus P. Kurtzman, who established the genus of Wickerhamomyces).
After growth in YM broth for 3 days, cells are ovoid to ellipsoidal with 2.5–4.0 × 3.5–5.5 μm, and occur singly, in pairs or in small clusters (Fig. 3a). Sediment and ring are formed. Budding is multilateral. Growth is cream-white, butyrous, with a smooth surface and an entire margin. Conjugation occurred between two separate cells (Fig. 3b). Asci form one to four hat-shaped ascospores after 5–7 days growth on PDA or CMA agar at 22 ℃. Asci was deliquescent and then ascospores were released (Fig. 3c–e). Pseudohyphae are not formed.
Growth is positive at 4–25 ℃ (with the optimal range of 20–22 ℃), weakly positive at 28 ℃, but negative at 30 ℃. Growth in vitamin-free medium is negative. Growth occurred in 10% Glucose (w/v), but not in media containing 10% NaCl and 50% Glucose (w/v). Urease activity and diazonium blue B reaction are negative.
Fermentation is not observed. Sucrose, melibiose (weak), glycerol, d-mannitol (weak), salicin, dl-lactic acid, and succinic acid were assimilated as sole carbon resource. Galactose, sorbose, maltose, melezitose, cellobiose, trehalose, lactose, raffinose, soluble starch, d-xylose, l-arabinose, α-methyl glucoside, citric acid, inositol, inulin, ribose, l-rhamnose, erythritol, ribitol, galactitol, and d-glucitol were not assimilated. Sodium nitrite, potassium nitrate, cadaverine, ethylamine, and l-lysine were assimilated as sole nitrogen resource, but creatinine was not. Production of starch-like compounds is negative.
The strain TF5-16-2 isolated from the water of crater lake in Da Hinggan Ling Mountain, North of China. CGMCC 2.5597T as the holotype strain was preserved in a metabolically inactive state at China General Microbiological Culture Collection (CGMCC). The ex-type culture was deposited at the Yeast Collection of the Westerdijk Fungal Biodiversity Institute (CBS), Utrecht, The Netherlands, as CBS 15418, and at Korean Collection for Type Cultures (KCTC) as KCTC 27823. The Mycobank accession number is MB 829959.
Abbreviations
- LSU:
-
Large subunit rRNA
- ITS:
-
Internal transcribed spacer;
- SSU:
-
Small subunit rRNA
- EF-1α:
-
Elongation factor-1α
References
Kurtzman CP, Robnett CJ, Basehoar-Powers E (2008) Phylogenetic relationships among species of Pichia, Issatchenkia and Williopsis determined from multigene sequence analysis, and the proposal of Barnettozyma gen. nov., Lindnera gen. nov. and Wickerhamomyces gen. nov. FEMS Yeast Res 8:939–954
Kurtzman CP, Robnett CJ (1998) Identification and phylogeny of ascomycetous yeasts from analysis of nuclear large subunit (26S) ribosomal DNA partial sequences. Antonie Van Leeuwenhoek 73:331–371
Kobayashi R, Kanti A, Kawasaki H (2017) Three novel species of d-xylose-assimilating yeasts, Barnettozyma xylosiphila sp. nov., Barnettozyma xylosica sp. nov. and Wickerhamomyces xylosivorus fa, sp. nov. Int J Syst Evol Microbiol 67:3971–3976
Chai CY, Huang LN, Cheng H, Liu WJ, Hui FL (2019) Wichkerhamomyces menglaensis fa, sp. nov., a yeast species isolated from rotten wood. Int J Syst Evol Microbiol. https://doi.org/10.1099/ijsem.0.003350
Limtong S, Yongmanitchai W, Kawasaki H, Fujiyama K (2009) Wickerhamomyces edaphicus sp. nov. and Pichia jaroonii sp. nov., two ascomycetous yeast species isolated from forest soil in Thailand. FEMS Yeast Res 9:504–510
Kurtzman CP (2011) Wickerhamomyces Kurtzman, Robnett & Basehoar-Powers. In: Kurtzman CP, Fell JW, Boekhout T (eds) The Yeasts, a Taxonomic Study. Elsevier, Oxford, pp 87–110
Shin KS, Bae KS, Lee KH, Park DS, Kwon GS (2011) Wickerhamomyces ochangensis sp. nov., an ascomycetous yeast isolated from the soil of a potato field. Int J Syst Evol Microbiol 61:2543–2546
Limtong S, Nitiyon S, Kaewwichian R, Jindamorakot S, Am-In S, Yongmanitchai W (2012) Wickerhamomyces xylosica sp. nov. and Candida phayaonensis sp. nov.: two xylose-assimilating yeast species from soil. Int J Syst Evol Microbiol 6:2786–2792
Kaewwichian R, Kawasaki H, Limtong S (2013) Wickerhamomyces siamensis sp. nov., a novel yeast species isolated from the phylloplane in Thailand. Int J Syst Evol Microbiol 63:1568–1573
de García V, Brizzio S, Libkind D, Rosa CA, van Broock M (2010) Wickerhamomyces patagonicus sp. nov., an ascomycetous yeast species from Patagonia, Argentina. Int J Syst Evol Microbiol 60:1693–1696
Groenewald M, Robert V, Smith MT (2011) Five novel Wickerhamomyces and Metschnikowia related yeast species, Wickerhamomyces chaumierensis sp. nov., Candida pseudoflosculorum sp. nov., Candida danieliae sp. nov., and Candida eppingiae sp. nov., isolated from plants. Int J Syst Evol Microbiol 61:2015–2022
Nakase T, Jindamorakot S, Am-In S, Ninomiya S, Kawasaki H (2012) Wickerhamomyces tratensis sp. nov. and Candida namnaoensis sp. nov., two novel ascomycetous yeast species in the Wickerhamomyces clade found in Thailand. J Gen Appl Microbiol 58:145–152
James SA, Barriga EJ, Barahona PP, Harrington TC, Lee CF, Bond CJ, Roberts IN (2014) Wickerhamomyces arborarius fa sp. nov., an ascomycetous yeast species found in arboreal habitats on three different continents. Int J Syst Evol Microbiol 64:1057–1061
Hui FL, Chen L, Chu XY, Niu QH, Ke T (2013) Wickerhamomyces mori sp. nov., an anamorphic yeast species found in the guts of wood-boring insect larvae. Int J Syst Evol Microbiol 63:1174–1178
Rosa CA, Morais PB, Lachance MA, Santos RO, Melo WGP, Viana RHO, Braganc MAL, Pimenta RS (2009) Wickerhamomyces queroliae sp. nov. and Candida jalapaonensis sp. nov., two yeast species isolated from Cerrado ecosystem in North Brazil. Int J Syst Evol Microbiol 59:1232–1236
Francesca N, Carvalho C, Almeida PM, Sannino C, Settani L, Sampaio JP, Moschetti G (2013) Wickerhamomyces sylviae fa, sp. nov., an ascomycetous yeast species isolated from migratory birds. Int J Syst Evol Microbiol 63:4824–4830
Silva CF, Schwan RF, Sousa Dias Ë, Wheals AE (2000) Microbial diversity during maturation and natural processing of coffee cherries of Coffea arabica in Brazil. Int J Food Microbiol 60:251–260
Etchells JL, Bell TA (1950) Film yeasts on commercial cucumber brines. Food Technol 4:77–83
Kurtzman CP, Fell JW, Boekhout T, Robert V (2011) Methods for isolation, phenotypic characterization and maintenance of yeasts. In: Kurtzman CP, ed. The yeasts, a taxonomic study, 5th ed, pp 88–110.
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide for methods and applications. Academic Press, New York, pp 315–322
O'Donnell K (1993) Fusarium and its near relatives. In: Reynolds DR, Taylor JW (eds) The fungal holomorph: mitotic, meiotic and pleomorphic speciation in fungal systematic. CAB International, Wallingford, pp 225–233
Wang QM, Groenewald M, Takashima M, Theelen B, Han PJ et al (2016) Phylogeny of yeasts and related filamentous fungi within Pucciniomycotina determined from multigene sequence analyses. Stud Mycol 81:27–53
Rehner SA, Buckley E (2005) A Beauveria phylogeny inferred from nuclear ITS and EF 1-αsequences: evidence for cryptic diversification and links to Cordyceps teleomorphs. Mycologia 97:84–98
Pratibha J, Nguyen HD, Melnik VA, Bhat DJ, White GP, Seifert KA (2015) Lectotypification, epitypification, and molecular phylogeny of the synnematous hyphomycete Pseudogliophragma indicum, the second genus in the Wiesneriomycetaceae. Mycoscience 56:387–395
Thompson JD, Gibson TJ, Plewniak F (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Posada D (2008) JModelTest: phylogenetic model averaging. Mol Biol Evol 25:1253–1256
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Masiulionis VE, Pagnocca FC (2016) Wickerhamomyces spegazzinii sp. nov., an ascomycetous yeast isolated from the fungus garden of Acromyrmex lundii nest (Hymenoptera: Formicidae). Int J Syst Evol Microbiol 66:2141–2145
Barahona PP, Harrington TC, Lee CF, Bond CJ, Roberts IN (2014) Wickerhamomyces arborarius f.a sp. nov., an ascomycetous yeast species found in arboreal habitats on three different continents. Int J Syst Evol Microbiol 64:1057–1061
Braganca MA, Pimenta RS (2009) Wickerhamomyces queroliae sp. nov. and Candida jalapaonensis sp. nov., two yeast species isolated from Cerrado ecosystem in North Brazil. Int J Syst Evol Microbiol 59:1232–1236
Acknowledgement
The authors gratefully acknowledge financial support from the National Natural Science Funds of China (31400106) and the National Science and Technology Foundation Project (2014FY210400). We thank Dr. Chun-Li Li for his kind help in the use of scanning electron microscopes.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhou, Y., Jia, BS., Han, PJ. et al. Wickerhamomyces kurtzmanii sp. nov. An Ascomycetous Yeast Isolated From Crater Lake Water, Da Hinggan Ling Mountain, China. Curr Microbiol 76, 1537–1544 (2019). https://doi.org/10.1007/s00284-019-01773-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-019-01773-x