Abstract
The study provides phenotypic and molecular analyses of the antibiotic resistance in 20 Lactobacillus strains including 11 strains newly isolated from fermented plant material. According to the results of disc diffusion method, 90% of tested lactobacilli demonstrated sensitivity to clindamycin and 95% of strains were susceptible to tetracycline, erythromycin, and rifampicin. Ampicillin and chloramphenicol were found to inhibit all bacteria used in this study. The vast majority of tested strains revealed phenotypic resistance to vancomycin, ciprofloxacin, and aminoglycosides. Most of Lactobacillus strains showed high minimum inhibitory concentrations (MICs) of cefotaxime, ceftriaxone, and cefazolin and therefore were considered resistant to cephalosporins. All the strains exhibited multidrug resistance. The occurrence of resistance genes was associated with phenotypic resistance, with the exception of phenotypically susceptible strains that contained genes for tetracycline (tetK, tetL) and erythromycin (ermB, mefA) resistance. The vanX gene for vancomycin resistance was among the most frequently identified among the lactobacilli (75% of strains), but the occurrence of the parC gene for ciprofloxacin resistance was sporadic (20% of strains). Our results mainly evidence the intrinsic nature of the resistance to aminoglycosides in lactobacilli, though genes for enzymatic modification of streptomycin aadA and aadE were found in 20% of tested strains. The occurrence of extended spectrum beta-lactamases (ESBL) was unknown in Lactobacillus, but our results revealed the blaTEM gene in 80% of strains, whereas blaSHV and blaOXA-1 genes were less frequent (20% and 15% of strains, respectively).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lactobacilli are Gram-positive bacteria of high biotechnological and natural significance. They populate nutrient-rich habitats associated with food, feed, plants, animals, and humans [15]. In latter, Lactobacillus spp. belong to the resident gut microbiota being responsible for many of its beneficial effects on human health [28]. As a result, lactobacilli are widely used as probiotics to maintain or to replenish the gut microbiota after antibiotic treatment [23].
With regard to antibiotic resistance of the lactobacilli, vancomycin and ciprofloxacin have low inhibitory effect among the majority of Lactobacillus species [23, 54]. Lactobacilli are generally susceptible to the inhibitors of protein synthesis, such as chloramphenicol, macrolides, lincosamides, and tetracycline, but their resistance to aminoglycosides is often high [1, 23, 27, 32]. Moreover, lactobacilli are usually sensitive to the cell wall-targeting β-lactams such as penicillin, but are more resistant to cephalosporins [1, 23]. Resistance to other antibiotics varies greatly among lactobacilli.
Antibiotic resistance of probiotic Lactobacillus strains is an essential property for their application to reinforce the concomitant antibiotic therapy. On the other hand, intake of bacteria with acquired antibiotic resistance poses the risk of its dissemination in the gastrointestinal microbiota and totally in the environment. Studying of antibiotic susceptibility pattern and resistance genes in Lactobacillus spp. is a comparably recent approach [30]. Up to now, the presence of several antibiotic resistance genes, both intrinsic and acquired, has been reported in Lactobacillus spp. For example, chloramphenicol-resistance cat gene has been found in L. reuteri [16] and L. plantarum [55]. Different erythromycin-resistance genes (ermB, ermA, ermC, ermT) and at least 11 tetracycline resistance genes (tetW, tetM, tetS, tetO, tetQ, tet36, tetZ, tetO/W/32/O/W/O, tetW/O, tetK, and tetL) have been detected to date in lactobacilli [38, 42, 57], among which tetM and ermB were suggested to be the most widely-distributed among Lactobacillus spp.[3, 4, 6, 14]. Some of them were found to be located on plasmids and in transposons and thus were considered acquired [14, 18, 42, 44, 47]. These findings evidence the view on the food and probiotic bacteria as reservoirs of antibiotic resistance genes and facilitate the revision of GRAS (generally recognized as safe) status of lactic acid bacteria (LAB).
However, there is still much to be explored about the problem of antibiotic resistance of lactobacilli. Assessment of antibiotic resistances among lactobacilli is confounded by the lack of standards for susceptibility testing, as well as susceptibility breakpoints for most antibiotics. Determination of antibiotic susceptibility patterns of a representative number of different strains from each species is necessary for working out of reliable criteria for the differentiation between the intrinsic and the acquired antibiotic resistance in probiotic bacteria. Although some effort has been made to this end, work has only been carried out for some antibiotics and particular Lactobacillus species, such as L. casei, L. acidophilus, L. reuteri, L. rhamnosus, L. delbrueckii, L. brevis, L. fermentum [2, 4, 20, 37, 42]. Besides, antibiotic resistance profile of Lactobacillus strains for practical application should be studied by both phenotypic and genotypic methods, because a susceptible phenotype may carry silent genes, which are detected by PCR-based molecular methods [54].
In the present study, antibiotic resistance pattern of 20 Lactobacillus strains was investigated through the disc diffusion method and microdilution method as well as molecular methods to check the presence of different antibiotic resistance genes.
Materials and Methods
Isolation of Bacteria and Growth Conditions
Lactobacilli were isolated from probiotics, commercial dairy products and fermented plant material and identified in our previous studies [5, 7]. L. brevis DSM 20,054, L. buchneri DSM 20,057, L. brevis ssp. gravesensis LMG 7934, and L. rhamnosus B-8238, obtained from Culture Collections, were used as reference strains (Table 1). The cultures preserved as glycerol stocks were activated by growing in sterile de Man, Rogosa, Sharpe (MRS) broth [12] under microaerophilic conditions at 37 °C for 24 h. The cultures thus activated were stored at 5 °C with fortnightly subculture in MRS broth. Working cultures were prepared in sterilized MRS broth using 1% inocula followed by incubation under microaerophilic conditions at 37 °C for 16–18 h.
Antibiotic Susceptibility Testing
Antibiotic susceptibility was assessed by the disk diffusion method, as described earlier [5, 7]. In brief, all strains were diluted in 0.85% saline to obtain turbidity equivalent to McFarland scale 0.5 and aliquots were pour-plated on MRS agar plates. Antibiotic discs (Scientific Research Centre of Pharmacotherapy, Russia) were placed on the surface of inoculated plates. After 48 h incubation in anaerobic conditions (Anaerogas GasPak, NIKI MLT, Russia) at 37 °C, inhibition halos were measured in mm (means ± SD of 3 trials) and interpreted as susceptible (S), moderately susceptible (MS), or resistant (R) according to [9, 43], as indicated in Table S1.
The MIC values of cephalosporins were determined by the broth microdilution method in MRS broth in 96-well nontreated cell culture plates (Eppendorf). Cefazolin, ceftriaxone, and cefotaxime (all Sigma-Aldrich) were tested in concentration range of 0.5–1024 μg/ml obtained after a series of two-fold dilutions in MRS broth. Wells were inoculated with 200 μl of the bacterial culture (3 × 107 CFU/ml) and incubated at 37 °C. The MIC was read after 24 h of incubation as the lowest concentration of an antibiotic at which visible growth was inhibited.
Detection of Antibiotic Resistance Genes
Genomic DNA was extracted from lactobacilli cells and purified as described earlier [4]. Antibiotic resistance genes tested by PCR are indicated in Table S2. The PCR reaction was carried out in a total volume of 25 μl containing DNA template, 10 pmol of each primer (Table S2) [10, 16, 17, 21, 25, 27, 31, 40, 48, 50, 52, 59, 60], 1U Taq DNA polymerase, each of four dNTPs at a concentration of 200 μM, and PCR buffer containing Tris–HCl, KCl (NH4)2SO4, MgSO4, and Triton X-100. The amplification program was as following: initial denaturation step of 94 °C for 5 min, and then 35 cycles of 94 °C for 30 s, annealing temperature (Table S2) for 30 s, and a final extension step at 72 °C for 7 min. The obtained PCR fragments were analyzed by electrophoreses on a 1% agarose gel, stained with Midori Green DNA Stain (Nippon Genetics Europe, Germany) and visualized with UV light. The positive amplicons obtained in these PCRs were confirmed by sequencing. PCR products were purified with a GenJET Plasmid Miniprep Kit (Thermo Scientific, Lithuania) according to the instructions of the manufacturer. The purified products were sequenced with the forward and reverse primer (Evrogen JSC, Russian). The obtained primary nucleotide sequences were analyzed using the NCBI-BLAST algorithm and GenBank database.
Results
Phenotypic Resistance of Lactobacilli
In this study, 20 Lactobacillus strains were analyzed for the antibiotic susceptibility by disc diffusion method and were classified either as resistant (R), moderately susceptible (MS), or sensitive (S) based on zones of growth inhibition. All tested Lactobacillus strains were susceptible to ampicillin and chloramphenicol. Sensitivity to rifampicin and to the inhibitors of protein synthesis erythromycin, tetracycline, and clindamycin was also widely distributed among lactobacilli (Fig. 1b). Most of Lactobacillus strains were resistant to vancomycin (95% of strains), ciprofloxacin (95%) and aminoglycosides used in this work: amikacin (95%), kanamycin (100%), gentamicin (90%), streptomycin (85%). Noteworthily, that well-known probiotic strain L. planrarum 8PA3 possessed an antibiotic resistance profile not typical for Lactobacillus. For example, it revealed sensibility to vancomycin, while the vast majority of other tested lactobacilli were resistant to this antibiotic. Besides, unlike most lactobacilli, L. plantarum AG15 revealed rifampicin resistance.
The broth microdilution method was used to characterize the resistance of lactobacilli to cephalosporins. Microbiological breakpoints categorizing Lactobacillus species as resistant to cefotaxime, ceftriaxone and cefazolin are not determined yet. Therefore, we assumed that strains with high MIC values (64 and 128 µg/ml) were more likely to be resistant while two L. plantarum isolates (strains FCa1L and S6) with MIC values 0.5 μg/mL for three tested cephalosporins were considered as sensitive to them.
Genotypic Resistance of Lactobacilli
The Lactobacillus strains were tested for the presence of antibiotic resistance genes by PCR. The results are presented in Table 2 and fig. S1. Four silage Lactobacillus isolates were positive for genes encoding erythromycin-resistance ermB and mefA. The tetK gene was detected in L. buchneri DSM 20057 and tetL was found in L. plantarum FCa1L and in two L. plantarum silage isolates S1 and AG15 (Fig. S1, b). Detection of tetK gene was confirmed by the results of NCBI BLAST search for the homologs of tetracycline-resistance protein of Staphylococcus aureus PM1 (YP_006958133.1) in the genome sequence of L. buchneri DSM 20057. As a result, we identified nine secondary transporters which belong to the major facilitator superfamily (MFS) proteins and facilitate the transport across bacterial membranes. Among them, the EmrB QacA subfamily drug resistance transporter (KRK67099.1) gave the highest query cover (99%) and shared 41% similarity and 24% identity with the typical tetracycline efflux protein TetK.
Other erythromycin and tetracycline-resistance determinants (ermA, mefE, tetM) were not detected in any strain.
The gene parC associated with resistance to ciprofloxacin was detected in L. plantarum FCa3L, L. brevis DSM 20,054, L. brevis ssp. gravesensis LMG 7934, and L. buchneri DSM 20,057. No PCR products were obtained for another ciprofloxacin resistance gene gyrA. In addition, the PCR analysis showed that none of tested Lactobacillus strains possessed aac(6′)-aph(2″), ant(6), aph(3′)-III, and ant(2″)-I genes, which encode enzymatic inactivation of aminoglycosides. Yet, streptomycin resistance genes aadA and aadE were found in L. rhamnosus B-8238, L. plantarum FCa3L, L. plantarum AG1, and L. plantarum AG10.
The vancomycin resistance gene vanX was detected in 15 Lactobacillus strains (Table 2, Fig. S1, c), while other genes of this cluster vanA and vanE were not revealed in any strain.
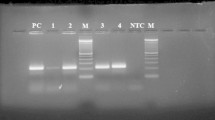
Sixteen tested Lactobacillus strains gave a 0.5 kbp band, presumably corresponding to blaTEM gene of cephalosporins resistance (Fig S1, a). The amplicons were sequenced and a 99% similarity with the blaTEM gene of Acinetobacter baumannii (GenBank Accession No. MK764360.1) and Escherichia sp. (GenBank Accession No. NG050218.1) was revealed by NCBI BLAST algorithm. The genes blaOXA-1 and blaSHV also related to cephalosporins resistance were less frequent in tested Lactobacillus strains and were detected in 3 and 4 strains, correspondingly.
The chloramphenicol resistance gene cat as well as the mecA gene were not detected in any strain.
Discussion
In the present study, we characterized phenotypic and genotypic antibiotic resistance profiles of 20 Lactobacillus strains, including 11 strains newly isolated from fermented plant material. In addition, four reference strains and five probiotic and dairy isolates were included in this study (Table 1). Irrespective of the origin, all the strains exhibited phenotypic resistance to a number of antibiotics revealing multidrug resistance pattern (Fig. 1, a). Moreover, all the strains carried at least one gene for antibiotic resistance (Tables 2, 3).
It is well known that lactobacilli are generally susceptible to antibiotics inhibiting nucleic acid synthesis (except for ciprofloxacin) and protein synthesis (except for aminoglycosides), and to cell wall synthesis inhibitors (except for vancomycin) [1, 11, 27, 34]. Indeed, in our study, 95% of tested Lactobacillus strains were susceptible to tetracycline, erythromycin, and rifampicin, and 90% of strains demonstrated sensitivity to clindamycin (Fig. 1b). Ampicillin and chloramphenicol were found to inhibit all bacteria used in this study (Fig. 1b). The latter notion coincided with the genomic program of tested lactobacilli. The chloramphenicol acetyl transferase gene cat which is associated with resistance to chloramphenicol was not detected in any of the strains. Gene mecA which encodes penicillin-binding protein 2A and confers resistance to penicillin-like antibiotics also was not found in tested Lactobacillus strains.
Tetracycline or erythromycin-resistant lactobacilli have been encountered in previous studies [4, 11, 21, 23, 42, 56]. Genes encoding these two resistances are often located on mobile genetic elements such as conjugative plasmids and transposons [3, 14, 18, 44, 47]. Therefore, detection of resistances to tetracycline and erythromycin in bacterial strains for food and agricultural applications always constitutes the risk of spread of antibiotic resistance genes in the environment. The most common determinants for resistance to tetracycline found in lactobacilli are genes tet (K, M, O, Q, S, W, 36), sometimes also present in combination [3, 23]. Among erythromycin-resistance genes, the ermB gene is the most frequently found among Lactobacillus spp.[42, 57].
In the present study, using PCR, we detected tetracycline efflux genes tetK (in L. buchneri DSM 20,057) and tetL (in L. plantarum FCa1L, L. plantarum S1, and L. plantarum AG15). The ermB gene, encoding 23S ribosomal rRNA methyltransferase, was found in three silage isolates, and macrolide efflux gene mefA was detected in another silage isolate L. plantarum AG15. All the strains which contained erythromycin and tetracycline-resistance genes displayed phenotypic susceptibility to these antibiotics. Similarly, in our previous investigation L. fermentum 5–1 sensitive to tetracycline was discovered to carry silent genes tetK and tetM, and L. fermentum 3–4 sensitive to erythromycin was positive for ermC [4]. These discrepancies between the resistance phenotype and genotype may be due to defective expression of resistance genes and have been described earlier by [41].
It is well documented that lactobacilli are usually resistant to aminoglycosides [23, 60]. This resistance is considered intrinsic and originates from the low impermeability of lactobacillar cell surface for aminoglycosides [9, 32, 34]. Yet, resistance genes for enzymatic modification of aminoglycosides such as aac(6′)-aph(2″), ant(6) and aph(3′)-III have been reported in several Lactobacillus spp.[50]. The vast majority of tested strains revealed phenotypic resistance to streptomycin, kanamycin, gentamycin, and amikacin (Table 2). The aminoglycoside adenyltransferase genes aadA and aadE which confer resistance to streptomycin were identified in four strains (Table 2, Fig S1, b), but other aminoglycoside-resistance genes (aac(6′)-aph(2″), aph(3′)-III, ant(6), ant(2″)-I) were not detected by PCR analysis in any of the tested lactobacilli. Thus, according to our results, among Lactobacillus spp. widely distributed resistance to aminoglycosides is likely to be intrinsic, though enzymatic inactivation of streptomycin is possible in some strains.
Lactobacilli have high natural resistance to vancomycin and ciprofloxacin. However, susceptibility to these antibiotics was shown to be species-dependent and varied several folds between species [11]. Hence, some resistant strains may harbor spontaneous mutations or acquired genes. In this work, we studied resistances to vancomycin and ciprofloxacin to find out their genetic determinants and assess their potential transferability.
We demonstrated that all tested Lactobacillus strains except for L. plantarum 8PA3 were resistant to vancomycin (Table 2). The vancomycin resistance has been reported to be intrinsic, chromosomally encoded and not inducible or transferable in lactobacilli [35, 51]. The most studied mechanism of vancomycin resistance in Lactobacillus spp. includes the replacement of the terminal D-alanine residue by d-lactate or d-serine in muramyl-pentapeptide molecule [13, 54]. Vancomycin has low-affinity binding to such altered peptidoglycan termini, and thus lactobacilli are generally resistant to vancomycin. The d-alanyl-d-alanine dipeptidase, a product of vanX gene, is critical for vancomycin resistance, because it prevents synthesis of the usual D-alanyl-D-alanine termini of the peptidoglycan precursor side chain. Homologs of vanX were found in genomes of sequenced Lactobacillus spp. (e.g. KRK67761.1 in L. buchneri DSM 20,057, ERK42887.1 in L. brevis DSM 20,054, ARW34669.1 in L. plantarum SRCM102022, according to NCBI GenBank database). Besides, in two strains of L. plantarum (LP1, LP2), the vanX gene was detected after sequencing and alignment [40]. In this study, PCR analysis revealed vanX gene in 15 tested Lactobacillus strains (Table 2, fig. S1, c). Guo et al. [24] also showed that vanX gene was frequent in lactobacilli. Among the genes of the vancomycin resistance cluster, only vanA gene is considered transferable via conjugation within the plasmid DNA [56, 58] or the conjugative transposon [26, 56]. In our work, no PCR products were amplified with vanA and vanE primer sets. Therefore. we conclude that tested Lactobacillus strains did not carry these resistance genes.
According to our results, all tested Lactobacillus strains showed resistance to ciprofloxacin, except for L. rhamnosus B-8238 which was moderately susceptible. Frequently encountered within the genus Lactobacillus resistance to ciprofloxacin has been earlier described by [11, 32, 39]. It is considered to arise from intrinsic characteristics, such as cell wall impermeability or efflux mechanism [27]. However, other mechanisms can be involved in the development of resistance to fluoroquinolones in Gram-positive bacteria. Some are the consequence of mutations involving genes encoding DNA gyrase and topoisomerase IV, essential type II topoisomerases necessary for DNA replication, chromosome segregation and DNA compaction in the cell [36, 49]. Here, we tested mutations in the quinolone resistance-determining region (QRDR) of the parC (topoisomerase IV) and gyrA (DNA gyrase) genes using PCR and consequent DNA sequencing of the obtained amplicons. The PCR fragments for the parC gene were obtained in four Lactobacillus strains and none of the strains possessed gyrA gene. Our data partly corroborate the results of [20, 27], which also demonstrated absence of typical mutations in the QRDR of gyrA or parC genes for ciprofloxacin resistance in Lactobacillus spp.
Regarding beta-lactams, lactobacilli are generally susceptible to penicillins, but more resistant or variable to cephalosporins [1, 53]. With few exceptions, Lactobacillus strains showed high MIC values of cefotaxime, ceftriaxone, and cefazolin (Table 3), as previously reported by [1, 22, 29]. Notably, among tested Lactobacillus strain sensitivity to cefotaxime was more frequent rather than to two other cephalosporins (Table 3). Indeed, cefotaxime has been shown to be the most active type I β-lactamase inhibitor, in comparison to the other β-lactam antibiotics [19]. The understanding of the mechanisms underlying resistance to cephalosporins is still very limited in lactobacilli. Although multiple beta-lactamases can be identified in the available genomic sequences of Lactobacillus spp., the presence of extended spectrum beta-lactamase (ESBL) in Gram-positive lactic acid bacteria remains obscure [33]. The major ESBL enzymes are TEM, SHV, CTX-M KPC, VIM, IMP, NDM-1, and OXA [8, 46]. Using PCR amplification and subsequent sequencing we identified the blaTEM gene in 80% of tested Lactobacillus trains, whereas blaSHV and blaOXA-1 genes were less frequent. To our knowledge, this is the first data on the detection of ESBL in lactobacilli. Although it is believed, that SHV enzymes confer much higher resistance than do TEM enzymes [45], according to our results TEM enzyme was the most frequent lactamase responsible for resistance to cephalosporins in lactobacilli.
Conclusions
Phenotypic multidrug resistance was revealed in all tested Lactobacillus strains. Studying of corresponding resistance genes showed that blaTEM (80% of strains) and vanX (75%) were the most frequently identified, and the occurrence of ermB (15%), mefA (5%), tetK (5%), tetL (15%), parC (20%), aadA (5%), aadE (15%), blaSHV (20%), and blaOXA-1 (15%) was less frequent. To our knowledge, genes for ESBLs were found in lactobacilli for the first time. Consideration should be given to the potentially transferable resistance determinants such as ermB, tetK, and tetL which were found in this work, fueling the debate about the safe use of lactobacilli in food and their potential to spread resistance in the environment. Future studies should be focused on horizontal transfer of detected resistance genes to other species.
Data Availability
The data used to support the findings of this study are included within the article.
References
Ammor MS, Flórez AB, Mayo B (2007) Antibiotic resistance in non-enterococcal lactic acid bacteria and bifidobacterial. Food Microbiol 24:559–570. https://doi.org/10.1016/j.fm.2006.11.001
Ammor MS, Flórez AB, Van Hoek AHAM, de los Reyes-Gavilán CG, Aarts HJM, Margolles A, et al (2008) Molecular characterization of intrinsic and acquired antibiotic resistance in lactic acid bacteria and bifidobacteria. J Mol Microbiol Biotechnol 14:6–15. https://doi.org/10.1159/000106077
Ammor MS, Gueimonde M, Danielsen M, Zagorec M, van Hoek AH, de Los Reyes-Gavilán CG, Mayo B, Margolles A (2008) Two different tetracycline resistance mechanisms, plasmid-carried tet(L) and chromosomally located transposon-associated tet(M), coexist in Lactobacillus sakei Rits 9. Appl Environ Microbiol 74(5):1394–1401. https://doi.org/10.1128/AEM.01463-07
Anisimova E, Yarullina D (2018) Characterization of erythromycin and tetracycline resistance in Lactobacillus fermentum strains. Int J Microbiol. https://doi.org/10.1155/2018/3912326
Anisimova EA, Yarullina DR, Ilinskaya ON (2017) Antagonistic activity of lactobacilli isolated from natural ecotopes. Microbiology 86(6):708–713. https://doi.org/10.1134/S0026261717060054
Bernardeau M, Vernoux JP, Henri-Dubernet S, Guéguen M (2008) Safety assessment of dairy microorganisms: the Lactobacillus genus. Int J Food Microbiol 126:278–285. https://doi.org/10.1016/j.ijfoodmicro.2007.08.015
Bruslik NL, Akhatova DR, Toimentseva AA, Abdulkhakov SR, Yarullina DR (2015) Estimation of probiotic lactobacilli drug resistance. J Antibiotics and Chemotherapy 60(3–4):6–13
Canton R, Coque TM (2006) The CTX-M beta-lactamase pandemic. Curr Opin Microbiol 9:466–475. https://doi.org/10.1016/j.mib.2006.08.011
Charteris WP, Kelly PM, Morelli L, Collins JK (1998) Antibiotic susceptibility of potentially probiotic Lactobacillus species. J Food Protection 61(12):1636–1643
Colom K, Perez J, Alonson R, Fernandez-Aranguiz A, Larino E, Cisterna R (2003) Simple and reliable multiplex PCR assay for detection of blaTEM, bla SHV and bla OXA-1 genes in Enterobacteriaceae. FEMS Microbiol Lett 223:147–151. https://doi.org/10.1016/S0378-1097(03)00306-9
Danielsen M, Wind A (2003) Susceptibility of Lactobacillus spp. to antimicrobial agents. Int J Food Microbiol 82:1–11
De Man JC, Rogosa M, Sharpe MT (1960) A medium for the cultivation of lactobacilli. J Appl Bacteriol 23:130–135
Delcour J, Ferain T, Deghorain M, Palumbo E, Hols P (1999) The biosynthesis and functionality of the cell-wall of lactic acid bacteria. Antonie Van Leeuwenhoek 76:159–184
Devirgiliis C, Coppola D, Barile S, Colonna B, Perozzi G (2009) Characterization of the Tn916 conjugative transposon in a food-borne strain of Lactobacillus paracasei. Appl Environ Microbiol 75(12):3866–71. https://doi.org/doi:10.1128/AEM.00589-09
Duar RM, Lin XB, Zheng JZ, Martino ME, Grenier T, Pérez-Muñoz ME, Leulier F, Gänzle M, Walter J (2017) Lifestyles in transition: evolution and natural history of the genus Lactobacillus. FEMS Microbiol Rev 41:S27–S48. https://doi.org/10.1093/femsre/fux030
Egervärn M, Roos S, Lindmark H (2009) Identification and characterization of antibiotic resistance genes in Lactobacillus reuteri and Lactobacillus plantarum. J Appl Microbiol 107(5):1658–1668. https://doi.org/10.1111/j.1365-2672.2009.04352.x
Faridi A, Kareshk AT, Fatahi-Bafghi M, Ziasistani M, Ghahraman MRK, Seyyed-Yousefi SZ, Shakeri N, Kalantar-Neyestanaki D (2018) Detection of methicillin-resistant Staphylococcus aureus (MRSA) in clinical samples of patients with external ocular infection. Iranian J Microbiol 10(4):215–219
Feld L, Schjorring S, Hammer K, Licht TR, Danielsen M, Krogfelt K, Wilcks A (2008) Selective pressure affects transfer and establishment of a Lactobacillus plantarum resistance plasmid in the gastrointestinal environment. J Antimicrob Chemother 61(4):845–852. https://doi.org/10.1093/jac/dkn033
Fu KP, Neu HC (1978) Beta-lactamase stability of HR756, a novel cephalosporin compared to that of cefuroxime and cefoxitin. Antimicrob Agents Chemother 14(3):322–326. https://doi.org/10.1128/aac.14.3.322
Fukao M, Tomita H, Yakabe T, Nomura T, Ike Y, Yajima N (2009) Assessment of antibiotic resistance in probiotic strain Lactobacillus brevis KB290. J Food Prot 72(9):1923–1929
Gevers D, Danielsen M, Huys G, Swings J (2003) Molecular characterization of tet(M) genes in Lactobacillus isolates from different types of fermented dry sausage. Appl Environ Microbiol 69(2):1270–1275. https://doi.org/10.1128/aem.69.2.1270-1275.2003
Gharajalar SN, Firouzamandi M (2017) Molecular detection of antibiotic resistance determinants in Lactobacillus bacteria isolated from human dental plaques. J Med Microbiol Infect Dis 5(3–4):51–55
Gueimonde M, Sánchez B, de Los Reyes-Gavilán CG, Margolles A (2013) Antibiotic resistance in probiotic bacteria. Front Microbiol 4:202. https://doi.org/10.3389/fmicb.2013.00202
Guo H, Pan L, Li L, Lu J, Kwok L, Menghe B, Zhang H, Zhang W (2017) Characterization of antibiotic resistance genes from Lactobacillus isolated from traditional dairy products. J Food Sci 82(3):724–730. https://doi.org/10.1111/1750-3841.13645
Han JH, Li XF, Gao WH, Walczak P, Zhang BL, Jia YM (2013) Susceptibility of Lactobacillus pentosus strains isolated from fermented products to streptomycin and kanamycin. International Food Research Journal 20(4):1927–1931
Handwerger S, Skoble J (1995) Identification of chromosomal mobile element conferring high-level vancomycin resistance in Enterococcus faecium. Antimicrob Agents Chemother 39(2446):2453
Hummel AS, Hertel C, Holzapfel WH, Franz CM (2007) Antibiotic resistances of starter and probiotic strains of lactic acid bacteria. Appl Environ Microbiol 73(3):730–739
Ilinskaya ON, Ulyanova VV, Yarullina DR, Gataullin IG (2017) Secretome of intestinal bacilli: a natural guard against pathologies. Front Microbiol 8:1666. https://doi.org/10.3389/fmicb.2017.01666
James L, Beena AK, Anupa A, Sreeshma N (2016) Antibiogram of lactobacilli isolated from four different niches. J Microbiol Microb Technol 1(1):4
Jorgensen JH, Hindler JF, Reller LB, Weinstein MP (2007) New consensus guidelines from the Clinical and Laboratory Standards Institute for antimicrobial susceptibility testing of infrequently isolated or fastidious bacteria. Clin Infect Dis 44:280–286. https://doi.org/10.1086/510431
Kastner S, Perreten V, Bleuler H, Hugenschmidt G, Lacroix C, Meile L (2006) Antibiotic susceptibility patterns and resistance genes of starter cultures and probiotic bacteria used in food. J Syst Appl Microbiol 29(2):145–155. https://doi.org/10.1016/j.syapm.2005.07.009
Katla AK, Kruse H, Johnsen G, Herikstad H (2001) Antimicrobial susceptibility of starter culture bacteria used in Norwegian dairy products. Int J Food Microbiol 67(1–2):147–152
Khan U, Afsana S, Kibtia M, Hossain M, Choudhury N, Ahsan CR (2019) Presence of blaCTX-M antibiotic resistance gene in Lactobacillus spp. isolated from Hirschsprung diseased infants with stoma. J Infect Dev Ctries 13(5):426–433. https://doi.org/10.3855/jidc.10968
Kirtzalidou E, Pramateftaki P, Kotsou M, Kyriacou A (2011) Screening for lactobacilli with probiotic properties in the infant gut microbiota. Anaerobe 17(6):440–443. https://doi.org/10.1016/j.anaerobe.2011.05.007
Klein G, Hallman C, Casas IA, Abad J, Lowers J, Reuter G (2000) Exclusion of vanA, vanB and vanC type glycopeptide resistance in strains of Lactobacillus reuteri and Lactobacillus rhamnosus used as probiotics by polymerase chain reaction and hybridization methods. J Appl Microbiol 89(5):815–824
Klein G, Pack A, Reuter G (1998) Antibiotic resistance patterns of enterococci and occurrence of vancomycin-resistant enterococci in raw minced beef and pork in Germany. Appl Environ Microbiol 64(5):1825–1830
Korhonen JM, Danielsen M, Mayo B, Egervarn H, Axelsson L, Huys G et al (2008) Antimicrobial susceptibility and proposed microbiological cut-off values of lactobacilli by phenotypic determination. Int J Probiotics Prebiotics 3:257–268
Lahtinen SJ, Boyle RJ, Margolles A, Frías R, Gueimonde M (2009) Safety assessment of probiotics. In: Charalampopoulos D, Rastall RA (eds) Prebiotics and Probiotics Science and Technology. Springer-Verlag, Berlin, pp 1193–1225
Li S, Li Z, Wei W, Ma C, Song X, He W, Tian J, Huo X (2015) Association of mutation patterns in GyrA and ParC genes with quinolone resistance levels in lactic acid bacteria. J Antibiot (Tokyo) 68(2):81–87. https://doi.org/10.1038/ja.2014.113
Liu C, Zhang ZY, Dong K, Yuan JP, Guo XK (2009) Antibiotic resistance of probiotic strains of lactic acid bacteria isolated from marketed foods and drugs. Biomed Environ Sci 22(5):401–412. https://doi.org/10.1016/S0895-3988(10)60018-9
Martel A, Meulenaere V, Devriese LA, Decostere A, Haesebrouck F (2003) Macrolide and lincosamide resistance in the gram-positive nasal and tonsillar flora of pigs. Microb Drug Resist 9(3):293–297. https://doi.org/10.1089/107662903322286508
Mayrhofer S, van Hoeck AHAM, Mair C, Huys G, Arts HJM, Kneifel W et al (2010) Antibiotic susceptibility of members of the Lactobacillus acidophilus group using broth microdilution and molecular identification of their resistance determinants. Int J Food Microbiol 144(1):81–87. https://doi.org/10.1016/j.ijfoodmicro.2010.08.024
Melo TA, Dos Santos TF, Pereira LR, Passos HM, Rezende RP, Romano CC (2017) Functional profile evaluation of Lactobacillus fermentum TCUESC01: A New Potential Probiotic Strain Isolated during Cocoa Fermentation. Biomed Res Int. https://doi.org/10.1155/2017/5165916
Mendonça AA, de Lucena BT., de Morais MM., de Morais MA Jr (2016) First identification of Tn916-like element in industrial strains of Lactobacillus vini that spread the tet-M resistance gene. FEMS Microbiol Lett https://doi.org/10.1093/femsle/fnv240
Moosdeen F (1997) The evoluation of resistance to cephalosporins. Clin Infect Dis 24(3):487–493. https://doi.org/10.1093/clinids/24.3.487
Naas T, Cuzon G, Bogaerts P, Glupczynski Y, Nordmann P (2011) Evaluation of a DNA microarray (Check-MDR CT102) for the rapid detection of TEM, SHV and CTX-M extended-spectrum ß-lactamases (ESBLs), and of KPC, OXA-48, VIM, IMP, and NDM-1 carbapenemases. J Clin Microbiol 49(4):1608–1613. https://doi.org/doi:10.1128/JCM.02607-10
Nicoloff H, Bringel F (2003) ISLpl1 is a functional IS30-related insertion element in Lactobacillus plantarum that is also found in other lactic acid bacteria. Appl Environ Microbiol 69(10):6032–6040. https://doi.org/10.1128/AEM.69.10.6032-6040.2003
Ouoba LI, Lei V, Jensen LB (2008) Resistance of potential probiotic lactic acid bacteria and bifidobacteria of African and European origin to antimicrobials: determination and transferability of the resistance genes to other bacteria. Int J Food Microbiol 121(2):217–224. https://doi.org/10.1016/j.ijfoodmicro.2007.11.018
Petersen A, Jensen LB (2004) Analysis of gyrA and parC mutations in enterococci from environmental samples with reduced susceptibility to ciprofloxacin. FEMS Microbiol Lett 231(1):73–76. https://doi.org/10.1016/S0378-1097(03)00929-7
Rojo-Bezares B, Saenz Y, Poeta P, Zarazaga M, Ruiz-Larrea F, Torres C (2006) Assessment of antibiotic susceptibility within lactic acid bacteria strains isolated from wine. Int J Food Microbiol 111(3):234–240. https://doi.org/10.1016/j.ijfoodmicro.2006.06.007
Saarela M, Mättö J, Mattila-Sandholm T (2002) Safety aspects of Lactobacillus and Bifidobacterium species originating from human oro-gastrointestinal tract or from probiotic products. Microbial Ecology in Health and Disease 14(4):233–240. https://doi.org/10.1080/08910600310002127
Sabouni F, Movahedi Z, Mahmoudi S, Pourakbari B, Valian SK, Mamishi S (2016) High frequency of vancomycin resistant Enterococcus faecalis in children: an alarming concern. Journal of preventive medicine and hygiene 57(4):E201–E204
Salminen MK, Rautelin H, Tynkkynen S, Poussa T, Saxelin M, Valtonen V, Järvinen A (2006) Lactobacillus bacteremia, species identification, and antimicrobial susceptibility of 85 blood isolates. A Clin Infect Dis 42(5):e35–44. https://doi.org/10.1086/500214
Sharma P, Tomar SK, Goswami P, Sangwan V, Singh R (2014) Antibiotic resistance among commercially available probiotics. Food Res Int 57:176–195. https://doi.org/10.1016/j.foodres.2014.01.025
Sukmarini L, Mustopa AZ, Normawati M, Muzdalifah I (2014) Identification of antibiotic-resistance genes from lactic acid bacteria in indonesian fermented foods. HAYATI Journal of Biosciences 21:3:144–150 EISSN: 2086–4094. https://doi.org/10.4308/hjb.21.3.144
Thal L, Donabedian S, Robinson-Dunn B, Chow JW, Dembry L, Clewell DB, Alshab D, Zervos MJ (1998) Molecular analysis of glycopeptide-resistant Enterococcus faecium isolates collected from Michigan hospitals over a 6-year period. J Clinical Microbiol 36(11):3303–3308
van Hoek AHAM, Margolles A, Damig KJ, Korhonen JM, Zycka-Krzesinka J, Bardowsky JK et al (2008) Molecular assessment of erythromycin and tetracycline resistance genes in lactic acid bacteria and bifidobacteria and their relation to the phenotypic resistance. International Journal of Probiotics and Prebiotics 3(4):271–280
Werner G, Klare I, Witte W (1999) Large conjugative vanA plasmids in vancomycin-resistant Enterococcus faecium. J Clinical Microbiol 37(7):2383–2384
Werner G, Willems RJL, Hildebrandt B, Klare IW (2003) Witte Influence of transferable genetic determinants on the outcome of typing methods commonly used for Enterococcus faecium. J Clinical Microbiol 41(4):1499–1506. https://doi.org/10.1128/JCM.41.4.1499-1506.2003
Zhou JS, Pillidge CJ, Gopal PK, Gill HS (2005) Antibiotic susceptibility profiles of new probiotic Lactobacillus and Bifidobacterium strains. Int J Food Microbiology 98(2):211–217. https://doi.org/10.1016/j.ijfoodmicro.2004.05.011
Acknowledgements
This work was supported by the program of competitive growth of Kazan Federal University, RFBR Grants No 18–34-00268 and 17–00-00456.
Author information
Authors and Affiliations
Contributions
EA designed the experiments, carried out the study, interpreted the data and drafted the manuscript. DY designed the study, supervised all experiments and reviewed the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interests
The authors declare that there is no conflict of interests regarding the publication of this paper.
Ethical Approval
Ethical approval was not required.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Anisimova, E.A., Yarullina, D.R. Antibiotic Resistance of LACTOBACILLUS Strains. Curr Microbiol 76, 1407–1416 (2019). https://doi.org/10.1007/s00284-019-01769-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-019-01769-7